Abstract
Ubiquitination is initiated by transfer of ubiquitin (Ub) from a ubiquitin-activating enzyme (E1) to a ubiquitin-conjugating enzyme (E2), producing a covalently linked intermediate (E2–Ub)1. Ubiquitin ligases (E3s) of the ‘really interesting new gene’ (RING) class recruit E2–Ub via their RING domain and then mediate direct transfer of ubiquitin to substrates2. By contrast, ‘homologous to E6-AP carboxy terminus’ (HECT) E3 ligases undergo a catalytic cysteine-dependent transthiolation reaction with E2–Ub, forming a covalent E3–Ub intermediate3,4. Additionally, RING-between-RING (RBR) E3 ligases have a canonical RING domain that is linked to an ancillary domain. This ancillary domain contains a catalytic cysteine that enables a hybrid RING–HECT mechanism5. Ubiquitination is typically considered a post-translational modification of lysine residues, as there are no known human E3 ligases with non-lysine activity. Here we perform activity-based protein profiling of HECT or RBR-like E3 ligases and identify the neuron-associated E3 ligase MYCBP2 (also known as PHR1) as the apparent single member of a class of RING-linked E3 ligase with esterification activity and intrinsic selectivity for threonine over serine. MYCBP2 contains two essential catalytic cysteine residues that relay ubiquitin to its substrate via thioester intermediates. Crystallographic characterization of this class of E3 ligase, which we designate RING-Cys-relay (RCR), provides insights into its mechanism and threonine selectivity. These findings implicate non-lysine ubiquitination in cellular regulation of higher eukaryotes and suggest that E3 enzymes have an unappreciated mechanistic diversity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Deshaies, R. J. & Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 (2009).
Huibregtse, J. M., Scheffner, M., Beaudenon, S. & Howley, P. M. A family of proteins structurally and functionally related to the E6-AP ubiquitin-protein ligase. Proc. Natl Acad. Sci. USA 92, 2563–2567 (1995).
Scheffner, M., Nuber, U. & Huibregtse, J. M. Protein ubiquitination involving an E1–E2–E3 enzyme ubiquitin thioester cascade. Nature 373, 81–83 (1995).
Wenzel, D. M., Lissounov, A., Brzovic, P. S. & Klevit, R. E. UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids. Nature 474, 105–108 (2011).
Hewings, D. S., Flygare, J. A., Bogyo, M. & Wertz, I. E. Activity-based probes for the ubiquitin conjugation–deconjugation machinery: new chemistries, new tools, and new insights. FEBS J. 284, 1555–1576 (2017).
Pao, K. C. et al. Probes of ubiquitin E3 ligases enable systematic dissection of parkin activation. Nat. Chem. Biol. 12, 324–331 (2016).
Niphakis, M. J. & Cravatt, B. F. Enzyme inhibitor discovery by activity-based protein profiling. Annu. Rev. Biochem. 83, 341–377 (2014).
Borodovsky, A. et al. Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme family. Chem. Biol. 9, 1149–1159 (2002).
Zhen, M., Huang, X., Bamber, B. & Jin, Y. Regulation of presynaptic terminal organization by C. elegans RPM-1, a putative guanine nucleotide exchanger with a RING-H2 finger domain. Neuron 26, 331–343 (2000).
Wan, H. I. et al. Highwire regulates synaptic growth in Drosophila. Neuron 26, 313–329 (2000).
Grill, B., Murphey, R. K. & Borgen, M. A. The PHR proteins: intracellular signaling hubs in neuronal development and axon degeneration. Neural Dev. 11, 8 (2016).
Byrne, R., Mund, T. & Licchesi, J. D. F. Activity-based probes for HECT E3 ubiquitin ligases. ChemBioChem 18, 1415–1427 (2017).
Stieglitz, B., Morris-Davies, A. C., Koliopoulos, M. G., Christodoulou, E. & Rittinger, K. LUBAC synthesizes linear ubiquitin chains via a thioester intermediate. EMBO Rep. 13, 840–846 (2012).
You, J. & Pickart, C. M. A HECT domain E3 enzyme assembles novel polyubiquitin chains. J. Biol. Chem. 276, 19871–19878 (2001).
Plechanovová, A., Jaffray, E. G., Tatham, M. H., Naismith, J. H. & Hay, R. T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012).
Dou, H., Buetow, L., Sibbet, G. J., Cameron, K. & Huang, D. T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Lechtenberg, B. C. et al. Structure of a HOIP/E2–ubiquitin complex reveals RBR E3 ligase mechanism and regulation. Nature 529, 546–550 (2016).
Dove, K. K. et al. Structural studies of HHARI/UbcH7–Ub reveal unique E2–Ub conformational restriction by RBR RING1. Structure 25, 890–900 (2017).
Xiong, X. et al. The Highwire ubiquitin ligase promotes axonal degeneration by tuning levels of Nmnat protein. PLoS Biol. 10, e1001440 (2012).
Babetto, E., Beirowski, B., Russler, E. V., Milbrandt, J. & DiAntonio, A. The Phr1 ubiquitin ligase promotes injury-induced axon self-destruction. Cell Rep. 3, 1422–1429 (2013).
Milde, S., Gilley, J. & Coleman, M. P. Subcellular localization determines the stability and axon protective capacity of axon survival factor Nmnat2. PLoS Biol. 11, e1001539 (2013).
Zheng, N., Wang, P., Jeffrey, P. D. & Pavletich, N. P. Structure of a c-Cbl–UbcH7 complex: RING domain function in ubiquitin-protein ligases. Cell 102, (533–539 (2000).
Olsen, S. K. & Lima, C. D. Structure of a ubiquitin E1–E2 complex: insights to E1–E2 thioester transfer. Mol. Cell 49, 884–896 (2013).
Cadwell, K. & Coscoy, L. Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 309, 127–130 (2005).
Shimizu, Y., Okuda-Shimizu, Y. & Hendershot, L. M. Ubiquitylation of an ERAD substrate occurs on multiple types of amino acids. Mol. Cell 40, 917–926 (2010).
Wang, X., Herr, R. A. & Hansen, T. H. Ubiquitination of substrates by esterification. Traffic 13, 19–24 (2012).
Alpi, A. F., Chaugule, V. & Walden, H. Mechanism and disease association of E2-conjugating enzymes: lessons from UBE2T and UBE2L3. Biochem. J. 473, 3401–3419 (2016).
Conforti, L., Gilley, J. & Coleman, M. P. Wallerian degeneration: an emerging axon death pathway linking injury and disease. Nat. Rev. Neurosci. 15, 394–409 (2014).
Ali, Y. O., Bradley, G. & Lu, H. C. Screening with an NMNAT2-MSD platform identifies small molecules that modulate NMNAT2 levels in cortical neurons. Sci. Rep. 7, 43846 (2017).
Stieglitz, B. et al. Structural basis for ligase-specific conjugation of linear ubiquitin chains by HOIP. Nature 503, 422–426 (2013).
Stanley, M. et al. Orthogonal thiol functionalization at a single atomic center for profiling transthiolation activity of E1 activating enzymes. ACS Chem. Biol. 10, 1542–1554 (2015).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Pruneda, J. N. et al. Structure of an E3:E2–Ub complex reveals an allosteric mechanism shared among RING/U-box ligases. Mol. Cell 47, 933–942 (2012).
Wu, P. Y. et al. A conserved catalytic residue in the ubiquitin-conjugating enzyme family. EMBO J. 22, 5241–5250 (2003).
Yunus, A. A. & Lima, C. D. Lysine activation and functional analysis of E2-mediated conjugation in the SUMO pathway. Nat. Struct. Mol. Biol. 13, 491–499 (2006).
Brownell, J. E. et al. Substrate-assisted inhibition of ubiquitin-like protein-activating enzymes: the NEDD8 E1 inhibitor MLN4924 forms a NEDD8–AMP mimetic in situ. Mol. Cell 37, 102–111 (2010).
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Lovell, S. C. et al. Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins 50, 437–450 (2003).
Acknowledgements
We thank D. Campbell and J. Varghese of the MRC PPU Proteomics Facility, the MRC PPU DNA Sequencing Facility, the European Synchrotron Radiation Facility (ESRF); J. Hastie, H. McLauchlan and F. Brown from MRC PPU Reagents and Services for MYCBP2 antibody production; F. Zuccotto and I. Gilbert for software access; and H. Walden and R. T. Hay for reading of the manuscript and suggestions. This work was funded by the Scottish Funding Council, the UK Medical Research Council (MC_UU_12016/8), BBSRC (BB/P003982/1), and pharmaceutical companies supporting the Division of Signal Transduction Therapy (Boehringer-Ingelheim, GlaxoSmithKline and Merck KGaA). D.M.F.v.A. is funded by a Wellcome Trust Investigator Award (110061).
Author information
Authors and Affiliations
Contributions
S.V. and K.-C.P. designed research. K.-C.P. carried out experiments with assistance from S.V. N.T.W. performed cloning. A.K. prepared E1, E2 panel and recombinant NMNAT2 protein. K.R. mounted crystals and collected synchrotron diffraction data. M.S. contributed to project conception and carried out molecular synthesis. P.D.M. performed structural modelling, R.S. carried out SEC–MALS. K.H. performed bioinformatics analysis. D.M.F.v.A. solved the MYCBP2(cat) crystal structure. S.V. coded PERL scripts, processed mass spectrometry data and wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
S.V., K.-C.P. and M.S. are authors on patents relating to work presented in this article.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 LC–MS characterization of biotinylated-ABP intermediates and biotinylated ABPs.
a–f, E3s can have distinct E2 preferences5, so to obtain broad coverage we prepared biotinylated ABPs based on the promiscuous E2 UBE2D2 (ABP1), and the HECT/RBR-specific E2 UBE2L3 (ABP2). As controls we prepared ABPs containing a point mutation in the E2-recognition component to disrupt or impair E3 ligase binding, UBE2D2(F62A) (ABP3) and UBE2L3(F63A) (ABP4)7. Ubiquitin in ABP1 and ABP3 was extended by a single residue relative to that previously reported7 (Ub1–74 rather than Ub1–73), as this improved labelling efficiency of the RBR E3 HOIP. a, Characterization of biotin-labelled, truncated ubiquitin-thioester intermediate, biotin–Ub1–73-SR used for ABP2 and ABP4. HPLC chromatogram monitoring UV absorbance at 214 nm (as for all subsequent intermediates and ABPs). ESI mass spectrum (inset left) and deconvoluted mass spectrum (inset right). Expected mass = 9,093 Da (−Met); found mass = 9,091 Da. b, Characterization of UBE2L3 ABP2. ESI mass spectrum (inset left) and deconvoluted mass spectrum (inset right). Expected mass = 29,286.9 Da (−Met); found mass = 29,282 Da. c, Characterization of UBE2L3(F63A) ABP4. ESI mass spectrum (inset left) and deconvoluted mass spectrum (inset right). Expected mass = 29,210.8 Da (−Met); found mass = 29,206 Da. d, Characterization of probe intermediate with extended ubiquitin C terminus, biotin-Ub1–74-SR used to make ABP1 and ABP3. ESI mass spectrum (inset left) and deconvoluted mass spectrum (inset right). Expected mass = 9,249.2 Da (−Met); found mass = 9,247 Da. e, Characterization of UBE2D2 ABP1. ESI mass spectrum (inset left) and deconvoluted mass spectrum (inset right). Expected mass = 29,268.8 Da (−Met); found mass = 29,264 Da. f, Characterization of UBE2D2(F62A) ABP2. ESI mass spectrum (inset left) and deconvoluted mass spectrum (inset right). Expected mass = 29,192.7 Da (−Met); found mass = 29,186 Da. Intermediates and probes have been prepared and characterized more than three times with similar results.
Extended Data Fig. 2 Activity-based proteomic profiling of neuroblastoma SH-SY5Y cells.
a–d, Parallel profiling of neuroblastoma SH-SY5Y cell extracts was carried out with ABPs 1–4. As an additional control, cells were left untreated or treated with inhibitors of oxidative phosphorylation, oligomycin and antimycin A, which enables activity-dependent labelling of the RBR E3 Parkin7. ABP-labelled proteins were enriched against streptavidin resin followed by on-resin tryptic digestion. Obtained peptides were analysed by data-dependent LC–MS/MS. Recovered proteins were filtered against E3-associated PFAM domain terms (RING, HECT, IBR, zf-UBR) and proteins with fewer than three spectral counts were excluded. E3s that did not demonstrate more than 14-fold spectral count enrichment compared to control purifications, in which ABP was withheld, were also excluded. E1s yielded a strong signal, because they undergo transthiolation and are highly enriched by our ABPs. The number of spectral counts for the majority of HECT/RBR E3s was reduced by > 50% relative to their parental counterpart when the binding-defective control probes ABP3 and ABP4 were used (Fig. 1c). The aggregate number of recovered HECT/RBR E3s from ABP1 and ABP2 was 33 (22 HECT and 11 RBR), representing around 80% of the currently annotated HECT/RBR E3s (Fig. 1d). A subset of E3s remain permissive to control probes ABP3 and ABP4; we cannot establish whether this is because the respective E3s are labelled in an activity-independent manner or whether they are permissive to the F62A or F63A mutation. Furthermore, ABP-dependent spectral count signals are not normalized against protein abundance. Therefore, we cannot deconvolute the effects of E3 activation stoichiometry from E3 abundance (that is, highly abundant E3s that are in a low-activation state could yield disproportionately high signals in our data). a, The number of spectral counts for the recovered E1 and E3 proteins plotted against protein ID for UBE2D2 ABP1 and its respective control probe UBE2D2(F62A) ABP3. MYCBP2 yields a high signal with ABP1, which is reduced by more than 50% with ABP3. A number of RING E3s that bind the ABP are also labelled, and for the majority of cases this is presumed to be mechanistically off-target labelling that is exacerbated by the high sensitivity of mass spectrometry-based detection. Another possibility is that hitherto undiscovered RING-linked E3s are being labelled. b, The number of spectral counts for the recovered E1 and E3 proteins plotted against protein ID for UBE2L3 ABP2 versus the respective control probe UBE2L3(F63A) ABP4. MYCBP2 is not detected with UBE2L3 ABP2 and ABP4. c, The number of spectral counts for E1 and E3 proteins obtained with ABP1 for untreated versus oligomycin and antimycin A-treated cells. d, The number of spectral counts for E1 and E3 proteins obtained with ABP2 for untreated versus oligomycin and antimycin A-treated cells. Parkin peptides were only recovered from cells treated with oligomycin and antimycin A, consistent with activity-dependent Parkin labelling. Thus, for at least a subset of detected E3s, spectral counts correlate with E3 activity.
Extended Data Fig. 3 ABPs label MYCBP2(cat) C4520 with high selectivity.
a, Recombinant MYCBP2(cat) was profiled with His-tagged ABPs based on the E2s UBE2D2 (ABP5) and UBE2D3 (ABP6). The experiment was repeated twice with similar results. b, Putative active-site cysteines in MYCBP2 were determined by ABP profiling of a panel of cysteine-to-serine mutants. MYCBP2(cat) mutant (3 μM) was incubated with ABP6 (12 μM) at 30 °C for 4 h. ABP-treated samples were resolved by SDS–PAGE and visualized by Coomassie staining and immunoblotting against the hexahistidine reporter tag on the ABP. Mutation of three cysteine residues (C4506, C4520 and C4537) abolished ABP labelling. The asterisk corresponds to inadvertent cleavage of the hexahistidine tag from the ABP due to trace protease contamination of the E3 preparations. The experiment was repeated twice with similar results. c, Using the pLink software, 38 spectral matches corresponding to cysteine-labelling sites in wild-type MYCBP2cat were identified. Thirty-six of these corresponded to C4520. One of the two remaining matches corresponded to C4440, a predicted Zn-coordinating residue in the RING domain. The other remaining match corresponded to C4600, which did not significantly affect ABP labelling when mutated. The table lists the predicted and found fragment ions for the representative spectrum depicted in Fig. 2b. The spectrum is for a 5+ precursor ion (expected m/z = 614.5094; observed m/z = 614.5088). A mass tolerance of 20 p.p.m. was applied for fragment ion assignment. Experiment was carried out once.
Extended Data Fig. 4 Esterification activity of MYCBP2(cat) and further data in support of a dual cysteine mechanism that operates in cis.
a, Mass spectrum of condensation products between ubiquitin and glycerol, and ubiquitin and Tris. Expected mass for ubiquitin condensation with glycerol = 8,639 Da; found mass = 8,637 Da. Expected mass for ubiquitin condensation with Tris = 8,668 Da; found mass = 8,666 Da. This experiment was repeated twice with similar results. b, Chemical structures of Tris and glycerol. c, Discharge activity towards Tris and glycerol for all of the tested MYCBP2(cat) cysteine-to-serine mutants (selected mutants shown in Fig. 2c). The C4506S mutation abolishes discharge activity, but because C4506S resides within a Cys-X-X-Cys Zn-binding motif, this was assumed to be a structural defect rather than a catalytic defect. The C4561S mutation undergoes aberrant thioester adduct formation; this may be because the C4561S mutation (also in a structurally important Cys-X-X-Cys Zn-binding motif) unfolds the protein and liberates Cys residues, which would otherwise be occupied as Zn ligands. These experiments were repeated twice with similar results. d, Coomassie stain of the thioester–ester trapping assay with GST–MYCBP2(cat). After the in-gel fluorescence scan, as shown in Fig. 2e, the gel was Coomassie stained. The experiment was repeated at least three times with similar results. e, The RCR E3 ligase activity is dependent on both C4520 and C4572. The combination of an inactive C4520S mutant with an inactive C4572A mutant did not restore activity; hence there appears to be cis-cooperation between these two residues (*elevated concentrations of E3 mutants). This experiment was repeated twice with similar results. f, Furthermore, consistent with cis-cooperation, SEC-MALS data for untagged MYCBP2(cat) were consistent with a monodisperse species with a calculated molecular weight of 30.06 ± 6.00 kDa (theoretical molecular weight of MYCBP2(cat) monomer = 30.08 kDa). The experiment was repeated twice with similar results.
Extended Data Fig. 5 MYCBP2 has serine/threonine ubiquitin esterification activity with a preference for threonine.
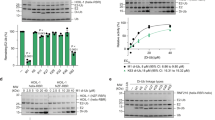
a, HPLC chromatogram of discharge reaction of wild-type MYCBP2(cat) onto threonine (50 mM). Note that esterified threonine with a free amino terminus can undergo O–N acyl transfer, forming a peptide-linked species. b, Integrated single-quadrupole electrospray-ionization mass spectrum of the entire peak highlighted in the above chromatogram. Inset shows the deconvoluted mass spectrum (as shown in Fig. 3b). Expected mass of Thr–Ub = 8,666 Da; found mass = 8,664 Da. c, HPLC chromatogram of MYCBP2(cat)(C4520S) discharge reaction in the presence of threonine (50 mM). d, Integrated single-quadrupole electrospray-ionization mass spectrum of the entire peak highlighted in the above chromatogram. Inset shows the deconvoluted mass spectrum (as shown in Fig. 3b). Expected mass of unmodified ubiquitin = 8,565 Da; found mass = 8,563 Da. All of the above experiments were repeated three times with similar results. e, Deconvoluted mass spectra for ubiquitin species in the presence of amino acid (50 mM). The intensities of the ubiquitin reactant and product are reflective of their relative abundance. Observed molecular weight of ubiquitinated serine (Ub–Ser) = 8,650 Da; theoretical molecular weight = 8,652 Da. The observed mass at 8,591 Da corresponds to a side product that is only observed after extended incubation. Ubiquitinated lysine (Ub–Lys) observed molecular weight = 8,691 Da; theoretical molecular weight = 8,693 Da. Assuming exponential ubiquitin consumption, t1/2 is around 5 min for threonine. For serine, t1/2 is tenfold slower. Lysine ubiquitination is E3-independent as a similar degree of modification is observed in the absence of E3. The experiment was repeated twice with similar results. f, Coomassie stain of threonine gel presented in Fig. 3c. Also shown is the deconvoluted mass spectrum representative of all ubiquitin species at the 60 min time point. Observed mass of Cy3B–Ub modified threonine peptide = 10,033 Da; theoretical mass = 10,036 Da. g, Coomassie stain of the serine gel presented in Fig. 3c. Also shown is the deconvoluted mass spectrum representative of all ubiquitin species at the 60-min time point. Observed mass of Cy3B–Ub = 9,534 Da; theoretical mass = 9,537 Da. Observed mass of Cy3B-Ub modified serine peptide = 10,019 Da; theoretical mass = 10,022 Da. h, Coomassie stain of lysine gels, in the presence and absence of E3, presented in Fig. 3c. Inefficient modification of the lysine peptide is observed, which is moderately enhanced in the absence of E3. Experiments shown in f–h were repeated more than three times. i, Top, observed rate constant (0.024 min−1) for MYCBP2(cat)-threonine-mediated single-turnover E2–Ub discharge, determined by in-gel fluorescence of Cy3B-labelled ubiquitin. The E2 was UBE2D3 and the substrate was threonine (50 mM) (n = 3). Bottom, representative replicate gel used for quantification. j, Top, observed rate constant for UBE3C-lysine mediated single-turnover E2–Ub discharge was too slow to measure. The E2 was UBE2L3 and the substrate was lysine (50 mM) (n = 2). Bottom, representative replicate gel used for quantification. k, Top, observed rate constant (0.52 min−1) for HHARI-lysine mediated single-turnover E2–Ub discharge. The E2 was UBE2L3 and the substrate was lysine (50 mM) (n = 2). The major component of this rate is attributable to autoubiquitination of lysine residues within HHARI because when lysine is withheld, kobs HHARI-mediated E2–Ub discharge is 0.39 min−1 and this is only partially outcompeted by the addition of lysine (n corresponds to the number of biologically independent experiments).
Extended Data Fig. 6 E2 requirements of MYCBP2.
a, To establish whether RCR E3 ligase activity occurs exclusively via the proposed E3–Ub thioester intermediates, or alternatively, mediates direct transfer of ubiquitin from E2–Ub (characteristic of RING E3s), we tested RCR E3 activity with a number of UBE2D3 mutants (N77S, D87A, I88A, L97A, L104A, S108A and D117A) that can be diagnostic for these two scenarios5. Single-turnover E2–Ub discharge assays employing Cy3B-labelled ubiquitin demonstrate that MYCBP2 has E2 requirements that are consistent with neither a HECT/RBR nor a RING mechanism. The N77 and D117 mutations alter E2 amino acids involved in pKa suppression of the acceptor nucleophile and are required for RING activity35,36. Additionally, a characteristic of RING activity is the adoption of a ‘closed’ E2–Ub conformation that involves E2 residues D87, I88, L97, L104 and S108. Unlike the RING requirements, UBE2D3, S108 and D117 were dispensable for E2–E3 transthiolation activity. The I88A mutant had reduced activity, the N77S mutant had strongly impaired activity, whereas the D87A, L97A and L104A mutants had negligible activity. Furthermore, RBR E3 activity is permissive to the L104A mutant18. Thus, based on our current understanding of the E2 requirements of these E3 classes, MYCBP2 has E2 requirements that are consistent with neither a HECT/RBR-like mechanism nor a RING-like mechanism. However, we cannot formally exclude the possibility that MYCBP2cat induces a closed E2–Ub conformation, characteristic of RING E3s, as it does not contain a prohibitive RING-domain loop insertion19. b, Quantification of the different E2 mutant activities. Mean of percentage E2–Ub discharge. n = 2 biologically independent experiments. c, Seventeen E2s were tested for threonine discharge activity with GST–MYCBP2(cat). UBE2D1, UBE2D3 and UBE2E1 were the only E2s that demonstrated detectable activity. -, position of unmodified E2; *ubiquitin-charged E2. Unexpectedly, the HECT/RBR-specific E2 UBE2L3 shows negligible activity with MYCBP2. Certain E2s undergo E3-independent polyubiquitin chain formation and/or autoubiquitination. In the presence of UBE2Q2, GST–MYCBP2 undergoes minor degradation resulting in the appearance of two lower-molecular-weight species. Consequently, we also carried out the assay with untagged MYCBP2 that did not undergo degradation; this produced similar results, showing that UBE2Q2 does not support MYCBP2 activity. The experiment was repeated twice with similar results.
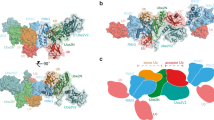
Extended Data Fig. 7 Structural comparison and representative stereo views of the crystallographic model of MYCBP2(cat).
a, Wide-field view. Regions are distinguished by colour in the stick representation: RING domain (blue), linker helix (purple), helix–turn–helix motif (green) and tandem-cysteine domain (orange). The mesh represents the experimental 2|Fobs|−|Fcalc| electron density map contoured at 1.5σ. C4572 is the downstream catalytic cysteine residue in the esterification site. The mediator loop region is formed between A4518 and G4527 and is disordered in the structure. b, Close up of the mediator loop region. c, Close up of the esterification site. T4380 motif from the symmetry-related molecule (T4380(sym)) is shown and represented in grey ball and stick. d, Superposition of MYCBP2 RING domain with the RING domain from the canonical RING E3 ligase RNF416, and from the RBR E3 ligase HOIP18. e, The linker helix and helix–turn–helix motif that connect the RING domain to the tandem cysteine domain. f, Diagram depicting the Zn coordination network for the tandem cysteine domain. Catalytic residues (numbered) are distributed throughout the tandem cysteine polypeptide. g, The tandem cysteine domain that confers threonine specificity is present in all MYCBP2 orthologues. All residues shown to be required for threonine esterification activity are conserved. Asterisks correspond to Zn-binding residues, grey arrows correspond to β-strands, gold rectangles correspond to 310-helices, and the red cylinder corresponds to an α-helix.
Extended Data Fig. 8 Modelling of E2-MYCBP2(cat) complex.
a, Ubiquitin adduct formation for catalytic mutants of GST-tagged MYCBP2(cat). The H4583N mutant undergoes near-quantitative ubiquitin-adduct formation. The adduct is largely removed after thiol treatment, indicating that ubiquitin is linked to the E3 via a thioester. The diffuse nature of the upper band might be due to the presence of a trapped thioester-linked ubiquitin on C4520, and C4572, as the H4583N mutation prevents substrate deprotonation. The C4572S/H4583N double mutant forms only a single ubiquitin adduct that is thioester-linked, presumably to the C4520 residue. This indicates that formation of the engineered ester-linked adduct on a mutated S4572 residue is dependent on the presence of a general base. C4520 does not appear to have a base in its proximity, hence its activity could be due to its intrinsic pKa which results in it being nucleophilic at physiological pH in the absence of a general base. This could explain why we failed to produce an engineered ester adduct on a mutant S4520 residue, as serine is fully protonated at physiological pH. The experiment was repeated twice with similar results. b, Superposition of the RING domain from the RBR E3 ligase HOIP in complex with E2 (PDB ID: 5EDV; ubiquitin linked to E2 has been omitted owing to a direct clash with the tandem cysteine domain18) allows modelling of the E2 into our structure (grey cartoon representation). The catalytic C85 residue in E2 (mutated in silico from Lys to Cys18) is proximal to C4520, which undergoes transthiolation with E2–Ub. Right, a top-down close up of the mediator loop region. The eight missing residues that form the mediator loop are shown schematically in brown text. c, Model of the proposed ubiquitin relay intermediate as shown in Fig. 5e but from an alternative perspective. In the experimental structure, tandem-cysteine-domain residues are shown in orange and mediator-loop residues are in dark brown. In the model, tandem-cysteine-domain residues are in light orange and mediator-loop residues are in mauve. The modelled E2, based on the superposition in d, is in grey cartoon. Essential cysteines C4520 and C4572 are in yellow and coloured by atom type. Ubiquitin residues G75–G76 are in blue ball-and-stick representation and are coloured by atom type. Gly residues in the mediator loop that are likely to be important for loop mobility are displayed in mauve ball and stick and coloured by atom type. N4570 and H4583 side chains have been rotated by the specified angles to relieve steric clash. d, As in c, but amino acid side chains that have been flipped to relieve steric clash with the modelled mediator loop are labelled in blue. e, All phi and psi angles in the modelled structure fall within accepted values as determined by Ramachandran analysis with the RAMPAGE server41.
Supplementary information
Supplementary Figure 1
This file contains the gel source data
Supplementary Data
Raw MS data for untreated cells profiled with ABP1
Supplementary Data
Raw MS data for untreated cells profiled with ABP2
Supplementary Data
No probe control for OA-treated cells
Supplementary Data
No probe control for untreated cells
Supplementary Data
Raw MS data for OA-treated cells profiled with ABP1
Supplementary Data
Raw MS data for OA-treated cells profiled with ABP2
Supplementary Data
Raw MS data for OA-treated cells profiled with ABP3
Supplementary Data
Raw MS data for OA-treated cells profiled with ABP4
Rights and permissions
About this article
Cite this article
Pao, KC., Wood, N.T., Knebel, A. et al. Activity-based E3 ligase profiling uncovers an E3 ligase with esterification activity. Nature 556, 381–385 (2018). https://doi.org/10.1038/s41586-018-0026-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0026-1
This article is cited by
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
Just how big is the ubiquitin system?
Nature Structural & Molecular Biology (2024)
-
UBE2A and UBE2B are recruited by an atypical E3 ligase module in UBR4
Nature Structural & Molecular Biology (2024)
-
Noncanonical assembly, neddylation and chimeric cullin–RING/RBR ubiquitylation by the 1.8 MDa CUL9 E3 ligase complex
Nature Structural & Molecular Biology (2024)
-
Breaking the K48-chain: linking ubiquitin beyond protein degradation
Nature Structural & Molecular Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.