Abstract
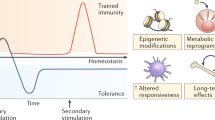
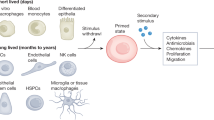
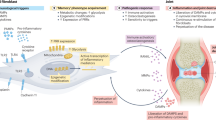
Innate immune memory is a vital mechanism of myeloid cell plasticity that occurs in response to environmental stimuli and alters subsequent immune responses. Two types of immunological imprinting can be distinguished—training and tolerance. These are epigenetically mediated and enhance or suppress subsequent inflammation, respectively. Whether immune memory occurs in tissue-resident macrophages in vivo and how it may affect pathology remains largely unknown. Here we demonstrate that peripherally applied inflammatory stimuli induce acute immune training and tolerance in the brain and lead to differential epigenetic reprogramming of brain-resident macrophages (microglia) that persists for at least six months. Strikingly, in a mouse model of Alzheimer’s pathology, immune training exacerbates cerebral β-amyloidosis and immune tolerance alleviates it; similarly, peripheral immune stimulation modifies pathological features after stroke. Our results identify immune memory in the brain as an important modifier of neuropathology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Netea, M. G., Latz, E., Mills, K. H. G. & O’Neill, L. A. J. Innate immune memory: a paradigm shift in understanding host defense. Nat. Immunol. 16, 675–679 (2015).
Netea, M. G. et al. Trained immunity: A program of innate immune memory in health and disease. Science 352, aaf1098 (2016).
Saeed, S. et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science 345, 1251086 (2014).
Cheng, S. C. et al. mTOR- and HIF-1α-mediated aerobic glycolysis as metabolic basis for trained immunity. Science 345, 1250684 (2014).
Biswas, S. K. & Lopez-Collazo, E. Endotoxin tolerance: new mechanisms, molecules and clinical significance. Trends Immunol. 30, 475–487 (2009).
Novakovic, B. et al. β-Glucan reverses the epigenetic state of LPS-induced immunological tolerance. Cell 167, 1354–1368 (2016).
Kleinnijenhuis, J. et al. Bacille Calmette-Guerin induces NOD2-dependent nonspecific protection from reinfection via epigenetic reprogramming of monocytes. Proc. Natl Acad. Sci. USA 109, 17537–17542 (2012).
Tay, T. L. et al. A new fate mapping system reveals context-dependent random or clonal expansion of microglia. Nat. Neurosci. 20, 793–803 (2017).
Füger, P. et al. Microglia turnover with aging and in an Alzheimer’s model via long-term in vivo single-cell imaging. Nat. Neurosci. 20, 1371–1376 (2017).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Heneka, M. T., Kummer, M. P. & Latz, E. Innate immune activation in neurodegenerative disease. Nat. Rev. Immunol. 14, 463–477 (2014).
Iadecola, C. & Anrather, J. The immunology of stroke: from mechanisms to translation. Nat. Med 17, 796–808 (2011).
Perry, V. H., Cunningham, C. & Holmes, C. Systemic infections and inflammation affect chronic neurodegeneration. Nat. Rev. Immunol. 7, 161–167 (2007).
Goldmann, T. et al. A new type of microglia gene targeting shows TAK1 to be pivotal in CNS autoimmune inflammation. Nat. Neurosci. 16, 1618–1626 (2013).
Datta, M. et al. Histone deacetylases 1 and 2 regulate microglia function during development, homeostasis, and neurodegeneration in a context-dependent manner. Immunity http://www.cell.com/immunity/fulltext/S1074-7613(18)30075-X (2018).
Banks, W. A. & Robinson, S. M. Minimal penetration of lipopolysaccharide across the murine blood-brain barrier. Brain Behav. Immun. 24, 102–109 (2010).
Saederup, N. et al. Selective chemokine receptor usage by central nervous system myeloid cells in CCR2-red fluorescent protein knock-in mice. PLoS One 5, e13693 (2010).
Sturchler-Pierrat, C. et al. Two amyloid precursor protein transgenic mouse models with Alzheimer disease-like pathology. Proc. Natl Acad. Sci. USA 94, 13287–13292 (1997).
Ostuni, R. et al. Latent enhancers activated by stimulation in differentiated cells. Cell 152, 157–171 (2013).
Kaikkonen, M. U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Cramer, T. et al. HIF-1α is essential for myeloid cell-mediated inflammation. Cell 112, 645–657 (2003).
Caron, E., Self, A. J. & Hall, A. The GTPase Rap1 controls functional activation of macrophage integrin αMβ2 by LPS and other inflammatory mediators. Curr. Biol. 10, 974–978 (2000).
Li, Y. et al. Rap1a null mice have altered myeloid cell functions suggesting distinct roles for the closely related Rap1a and 1b proteins. J. Immunol. 179, 8322–8331 (2007).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Mills, E. L. et al. Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell 167, 457–470 (2016).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 (2017).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581 (2017).
Shi, Y. et al. ApoE4 markedly exacerbates tau-mediated neurodegeneration in a mouse model of tauopathy. Nature 549, 523–527 (2017).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 1617, eaal3222 (2017).
Wang, C. et al. Targeting the mTOR signaling network for Alzheimer’s disease therapy. Mol. Neurobiol. 49, 120–135 (2014).
Ulland, T. K. et al. TREM2 maintains microglial metabolic fitness in Alzheimer’s disease. Cell 170, 649–663 (2017).
Kaufmann, E. et al. BCG educates hematopoietic stem cells to generate protective innate immunity against tuberculosis. Cell 172, 176–190 (2018).
Arts, R. J. W. et al. BCG vaccination protects against experimental viral infection in humans through the induction of cytokines associated with trained immunity. Cell Host Microbe 23, 89–100 (2018).
Heneka, M. T., Golenbock, D. T. & Latz, E. Innate immunity in Alzheimer’s disease. Nat. Immunol. 16, 229–236 (2015).
O’Banion, M. K. Does peripheral inflammation contribute to Alzheimer disease? Evidence from animal models. Neurology 83, 480–481 (2014).
Naik, S. et al. Inflammatory memory sensitizes skin epithelial stem cells to tissue damage. Nature 550, 475–480 (2017).
Liu, F., Yuan, R., Benashski, S. E. & McCullough, L. D. Changes in experimental stroke outcome across the life span. J. Cereb. Blood Flow Metab. 29, 792–802 (2009).
Sturchler-Pierrat, C. & Staufenbiel, M. Pathogenic mechanisms of Alzheimer’s disease analyzed in the APP23 transgenic mouse model. Ann. NY Acad. Sci 920, 134–139 (2000).
Neher, J. J. et al. Phagocytosis executes delayed neuronal death after focal brain ischemia. Proc. Natl Acad. Sci. USA 110, E4098–E4107 (2013).
Hefendehl, J. K. et al. Repeatable target localization for long-term in vivo imaging of mice with 2-photon microscopy. J. Neurosci. Methods 205, 357–363 (2012).
Eisele, Y. S. et al. Peripherally applied Aβ-containing inoculates induce cerebral β-amyloidosis. Science 330, 980–982 (2010).
Varvel, N. H. et al. Replacement of brain-resident myeloid cells does not alter cerebral amyloid-β deposition in mouse models of Alzheimer’s disease. J. Exp. Med. 212, 1803–1809 (2015).
Rottenberg, H. & Wu, S. Quantitative assay by flow cytometry of the mitochondrial membrane potential in intact cells. Biochim. Biophys. Acta 1404, 393–404 (1998).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Halder, R. et al. DNA methylation changes in plasticity genes accompany the formation and maintenance of memory. Nat. Neurosci. 19, 102–110 (2016).
Quinlan, A. R. BEDTools: The Swiss-army tool for genome feature analysis. Curr. Protoc. Bioinformatics 47, 11.12.1–11.12.34 (2014).
Acknowledgements
We thank P. Rizzu for experimental advice, L. Walker for manuscript comments and D. Bryce for statistical advice. This study was supported by a PhD fellowship from the Studienstiftung des Deutschen Volkes (A.-C.W.), a Roman Herzog Fellowship from the Hertie Foundation (J.J.N.), and grants from the network ‘Neuroinflammation in Neurodegeneration’ (State of Baden-Wuerttemberg, Germany; M.J. and M.P.), the Sobek-Stiftung (M.P.), the DFG (SFB992, Reinhart-Koselleck-Grant to M.P., SFB704 to J.L.S.), the European Research Council (A.F.) the Fortüne Program (Med. Faculty, Univ. Tuebingen; 2075-1-0; J.J.N.), the Fritz Thyssen Foundation (Cologne, Germany; J.J.N.) and the Paul G. Allen Family Foundation (Seattle, USA; J.J.N.). M.B. and J.L.S are members of the Excellence Cluster ImmunoSensation.
Author information
Authors and Affiliations
Contributions
K.D., A.-C.W., J.W., L.M.H., K.W., A.S., T.B, O.S., M.D. and J.J.N. performed microglial isolation, in vivo and ex vivo experiments and histological/biochemical analyses. M.G., L.K., G.J., T.P.C., V.C., R.I., C.K., A.F., M.B., T.U., J.L.S. and J.J.N. performed ChIP–seq and RNA-seq analyses. J.J.N conceived the study and coordinated the experiments together with M.J., A.F., M.P., M.B., J.L.S. and M.S. J.J.N. wrote the manuscript, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Acute responses to LPS injections.
a, Weight changes after injection of LPS (wild-type mice: n = 11, 11, 11, 11, 4 from left to right for PBS, n = 9, 9, 9, 8, 7 for 1 × LPS, n = 10, 10, 10, 10, 7 for 4 × LPS; APP animals: n = 14, 14, 14, 14, 7 for PBS, n = 8, 8, 8, 5, 5 for 1 × LPS; n = 10, 10, 10, 10, 10 for 4 × LPS; Cre mice: n = 5, 5, 4). b, c, Morphological changes in microglia (n = 6, 6, 6, 6, 6 mice). Scale bar, 50 µm. d, Numbers of microglia and activated (GFAP+) astrocytes (microglia: n = 6, 7, 8, 6, 6 mice, astrocytes: n = 6, 8, 9, 7, 5 mice). e, Blood and brain levels of LPS after daily injections with 500 µg per kg bodyweight (n = 4, 3, 3, 3, 3 animals). f, Assessment of iron entry from the blood (detected by Prussian blue staining) shows positive staining in an aged (>25 months) APP transgenic mouse, but not after repeated intraperitoneal LPS injections (n = 3 mice analysed). g, In heterozygous mice expressing red fluorescent protein (RFP) under the type 2 CC chemokine receptor (Ccr2) promoter, no entry of CCR2-expressing blood monocytes was detected after repeated LPS injection (staining for RFP; insert shows RFP-positive monocytes in the choroid plexus; n = 3 mice analysed). Scale bar, 100 µm. Data are means ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001 for one-way ANOVA with Tukey correction.
Extended Data Fig. 2 Cytokine response after acute LPS injections.
a, Additional cytokines (Fig. 1) analysed in the serum (top) and brain (bottom) 3 h after each daily intraperitoneal LPS injection on four consecutive days in 3-month-old mice (control mice received PBS injections; n = 16, 11, 12, 9, 9, 7, 7 and 5, 13, 4, 6, 9, 4, 5 mice for groups from left to right). b, c, Cytokine response in the blood only in wild-type (b, n = 6, 7, 8, 5, 5, 3, 3 mice) or APP23 mice (c, n = 10, 3, 3, 3, 4, 3, 3 mice). d, e, Cytokine response in the brain only in wild-type (d, n = 6, 7, 8, 5, 5, 3, 3 mice) or APP23 mice (e, n = 10, 4, 4, 4, 4, 4, 4 mice). Data are means ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001 for independent-samples median test with correction for multiple comparisons.
Extended Data Fig. 3 APP levels and processing, neuritic dystrophy and astrocyte activation in 9-month-old APP23 animals.
a, b, Micrograph of fluorescent staining for amyloid plaque (Methoxy-X04; green) and amyloid precursor protein (APP; red) (a) shows neuritic dystrophy surrounding the amyloid deposit, which is unchanged by LPS treatments (b; n = 5, 5, 5 animals). c, Overall Pearson’s correlation of plaque size with neuritic dystrophy (APP area; n = 49, 39, 42 plaques for PBS, 1 × LPS, 4 × LPS groups). d, Western blotting analysis (for gel source data, see Supplementary Fig. 1) of brain homogenates for APP and C-terminal fragment-β (CTFβ; n = 7, 4, 7 mice), and soluble APPβ ELISA (n = 6, 6, 6 mice). e, Micrograph of activated astrocytes (glial fibrillar acidic protein: GFAP) surrounding an amyloid plaque (Congo red) and quantification of the number of plaque-associated GFAP-positive astrocytes (n = 6, 6, 5 mice). Scale bar, 10 µm (a), 20 µm (e). Data are means ± s.e.m. *P < 0.05 for one-way ANOVA with Tukey correction.
Extended Data Fig. 4 Cytokine levels in 9-month-old animals.
a, Cytokine measurements in brain homogenates of 9-month-old wild-type (n = 8, 8, 7 mice) and APP23 mice (n = 14, 10, 10 mice) treated i.p. with 1 × LPS or 4 × LPS at 3 months of age. b, Cytokine measurements in the serum of 9-month-old wild-type (WT; n = 14, 9, 13 mice) and APP23 mice (APP; n = 18, 12, 14 mice) after i.p. stimulation with 1 × LPS or 4 × LPS at 3 months of age. c, Cytokine measurements in the serum of wild-type mice stimulated i.p. with 1 × LPS or 4 × LPS at 3 months of age and re-stimulated with an additional LPS injection (500 µg kg–1) at 9 months of age (n = 10, 7, 10 animals). Data are means ± s.e.m. *P < 0.05, **P < 0.01 for two-way ANOVA with Tukey correction. In b a significant main effect for genotype is indicated by bars spanning all conditions of the same genotype.
Extended Data Fig. 5 Cytokine levels after brain ischaemia and in blood of 4-month-old mice.
Three-month-old animals were injected i.p. with 1 × LPS or 4 × LPS and incubated for 4 weeks before receiving a stroke. a, Cytokine measurements in brain homogenates 24 h after stroke (n = 5, 7, 5, 5 animals). b, Cytokine measurements in the serum (n = 6, 6, 6 animals). Data are means ± s.e.m. ***P < 0.001 for one-way ANOVA with Tukey correction.
Extended Data Fig. 6 Microglial sorting strategy.
Microglia were sorted as CD11bhigh and CD45low cells (population P4) from 9-month-old APP23 mice or wild-type littermates following i.p. injections of 1 × LPS or 4 × LPS at 3 months of age.
Extended Data Fig. 7 Analysis of microglial enhancers.
Microglial enhancers were analysed in 9-month-old wild-type and APP23 (APP) mice treated intraperitoneally with 1 × LPS or 4 × LPS at 3 months of age. a, Exemplary UCSC browser images of genomic region around the Hif1a gene (normalized to input and library dimension). b, Numbers of regions with differentially regulated H3K4me1 levels. c, Heatmaps of H3K4me1 regions (centred on H3K27ac peaks). d, Pairwise correlations between the two replicates of H3K4me1 read densities in differentially regulated regions. e–g, Analyses of H3K27ac levels analogous to b–d for H3K4me1. n = 2 replicates (8–10 mice per replicate); differential enhancers showed a cumulative Poisson P < 0.0001.
Extended Data Fig. 8 Transcription factor motif analysis of active enhancer regions.
Motif analysis was performed for selected conditions to identify transcription factors involved in the differential activation of enhancers (using putative enhancer regions present in both replicates within 500 bp around enhancer peaks). a, For all active enhancers, motif analysis was performed using the union H3K27ac peak file and standard background (random genomic sequence). b, Pairwise comparisons between conditions, using the first condition’s H3K27ac peak file as input and the second condition’s peak file as background. As motif enrichment was often relatively low, the analysis was focused on transcription factor (families), whose motifs occurred at least twice in ‘known’ (black) and ‘de novo’ motifs (blue). Motifs are identified by HOMER software using hypergeometric testing (no adjustment for multiple comparisons was made).
Extended Data Fig. 9 Peripherally applied cytokines induce immune memory in the brain.
a, Experimental design. b, Cytokine responses in the brain, four weeks after peripheral cytokine application (n = 17, 5, 5, 21, 8, 8, 15 mice from left to right). Note that TNF dose-dependently enhances (low dose) or decreases (high dose) certain cytokines. Similar to high dose TNF, certain cytokines are also reduced by peripheral application of IL-10 four weeks earlier. c, Cytokine responses in the periphery are unaffected (n = 8, 21, 9, 5,10 mice). Data are means ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001 for one-way ANOVA with Tukey correction.
Supplementary information
Supplementary Figure
Gel source data for western blot in Extended Data Fig. 3
Supplementary Table 1
Differential microglial H3K4me1 levels
Supplementary Table 2
Differential microglial H3K27ac levels
Supplementary Table 3
Microglial gene expression matrix
Supplementary Table 4
WGCNA analysis
Video 1: Effect of microglia-specific Tak1 knockout on sickness behaviour
Microglia-specific (CX3CR1-CreER induced) knockout of transforming growth factor-β-activated kinase 1 (Tak1) alleviates sickness behaviour in mice 3 h after injection of the second dose of LPS. CX3CR1-Cre+ mice at the end of the movie are the first and third mice from the left
Rights and permissions
About this article
Cite this article
Wendeln, AC., Degenhardt, K., Kaurani, L. et al. Innate immune memory in the brain shapes neurological disease hallmarks. Nature 556, 332–338 (2018). https://doi.org/10.1038/s41586-018-0023-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0023-4
This article is cited by
-
Hypoxia inducible factor-1α regulates microglial innate immune memory and the pathology of Parkinson’s disease
Journal of Neuroinflammation (2024)
-
The endotoxin hypothesis of Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
Astrocyte cells in the brain have immune memory
Nature (2024)
-
Nuclear GAPDH in cortical microglia mediates cellular stress-induced cognitive inflexibility
Molecular Psychiatry (2024)
-
Cortical microinfarcts potentiate recurrent ischemic injury through NLRP3-dependent trained immunity
Cell Death & Disease (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.