Abstract
Hepatocytes are replenished gradually during homeostasis and robustly after liver injury1, 2. In adults, new hepatocytes originate from the existing hepatocyte pool3,4,5,6,7,8, but the cellular source of renewing hepatocytes remains unclear. Telomerase is expressed in many stem cell populations, and mutations in telomerase pathway genes have been linked to liver diseases9,10,11. Here we identify a subset of hepatocytes that expresses high levels of telomerase and show that this hepatocyte subset repopulates the liver during homeostasis and injury. Using lineage tracing from the telomerase reverse transcriptase (Tert) locus in mice, we demonstrate that rare hepatocytes with high telomerase expression (TERTHigh hepatocytes) are distributed throughout the liver lobule. During homeostasis, these cells regenerate hepatocytes in all lobular zones, and both self-renew and differentiate to yield expanding hepatocyte clones that eventually dominate the liver. In response to injury, the repopulating activity of TERTHigh hepatocytes is accelerated and their progeny cross zonal boundaries. RNA sequencing shows that metabolic genes are downregulated in TERTHigh hepatocytes, indicating that metabolic activity and repopulating activity may be segregated within the hepatocyte lineage. Genetic ablation of TERTHigh hepatocytes combined with chemical injury causes a marked increase in stellate cell activation and fibrosis. These results provide support for a ‘distributed model’ of hepatocyte renewal in which a subset of hepatocytes dispersed throughout the lobule clonally expands to maintain liver mass.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stanger, B. Z. Cellular homeostasis and repair in the mammalian liver. Annu. Rev. Physiol 77, 179–200 (2015).
Zaret, K. S. & Grompe, M. Generation and regeneration of cells of the liver and pancreas. Science 322, 1490–1494 (2008).
Malato, Y. et al. Fate tracing of mature hepatocytes in mouse liver homeostasis and regeneration. J. Clin. Invest. 121, 4850–4860 (2011).
Schaub, J. R., Malato, Y., Gormond, C. & Willenbring, H. Evidence against a stem cell origin of new hepatocytes in a common mouse model of chronic liver injury. Cell Reports 8, 933–939 (2014).
Yanger, K. et al. Adult hepatocytes are generated by self-duplication rather than stem cell differentiation. Cell Stem Cell 15, 340–349 (2014).
Tarlow, B. D., Finegold, M. J. & Grompe, M. Clonal tracing of Sox9+ liver progenitors in mouse oval cell injury. Hepatology 60, 278–289 (2014).
Tarlow, B. D. et al. Bipotential adult liver progenitors are derived from chronically injured mature hepatocytes. Cell Stem Cell 15, 605–618 (2014).
Raven, A. et al. Cholangiocytes act as facultative liver stem cells during impaired hepatocyte regeneration. Nature 547, 350–354 (2017).
Calado, R. T. et al. Constitutional telomerase mutations are genetic risk factors for cirrhosis. Hepatology 53, 1600–1607 (2011).
Hartmann, D. et al. Telomerase gene mutations are associated with cirrhosis formation. Hepatology 53, 1608–1617 (2011).
Rudolph, K. L., Chang, S., Millard, M., Schreiber-Agus, N. & DePinho, R. A. Inhibition of experimental liver cirrhosis in mice by telomerase gene delivery. Science 287, 1253–1258 (2000).
Jungermann, K. & Kietzmann, T. Zonation of parenchymal and nonparenchymal metabolism in liver. Annu. Rev. Nutr 16, 179–203 (1996).
Wang, B., Zhao, L., Fish, M., Logan, C. Y. & Nusse, R. Self-renewing diploid Axin2+ cells fuel homeostatic renewal of the liver. Nature 524, 180–185 (2015).
Font-Burgada, J. et al. Hybrid periportal hepatocytes regenerate the injured liver without giving rise to cancer. Cell 162, 766–779 (2015).
Planas-Paz, L. et al. The RSPO-LGR4/5-ZNRF3/RNF43 module controls liver zonation and size. Nat. Cell Biol. 18, 467–479 (2016).
Tanami, S. et al. Dynamic zonation of liver polyploidy. Cell Tissue Res 368, 405–410 (2017).
Batista, L. F. & Artandi, S. E. Understanding telomere diseases through analysis of patient-derived iPS cells. Curr. Opin. Genet. Dev 23, 526–533 (2013).
Nault, J. C. et al. High frequency of telomerase reverse-transcriptase promoter somatic mutations in hepatocellular carcinoma and preneoplastic lesions. Nat. Commun. 4, 2218 (2013).
Pech, M. F. et al. High telomerase is a hallmark of undifferentiated spermatogonia and is required for maintenance of male germline stem cells. Genes Dev. 29, 2420–2434 (2015).
Montgomery, R. K. et al. Mouse telomerase reverse transcriptase (mTert) expression marks slowly cycling intestinal stem cells. Proc. Natl Acad. Sci. USA 108, 179–184 (2011).
Schepers, A. G., Vries, R., van den Born, M., van de Wetering, M. & Clevers, H. Lgr5 intestinal stem cells have high telomerase activity and randomly segregate their chromosomes. EMBO J 30, 1104–1109 (2011).
Itzkovitz, S. et al. Single-molecule transcript counting of stem-cell markers in the mouse intestine. Nat. Cell Biol. 14, 106–114 (2011).
Gao, G.-P. et al. Novel adeno-associated viruses from rhesus monkeys as vectors for human gene therapy. Proc. Natl Acad. Sci. USA 99, 11854–11859 (2002).
Elias, H. A re-examination of the structure of the mammalian liver; the hepatic lobule and its relation to the vascular and biliary systems. Am. J. Anat. 85, 379–456 (1949).
Recknagel, R. O., Glende, E. A. Jr, Dolak, J. A. & Waller, R. L. Mechanisms of carbon tetrachloride toxicity. Pharmacol. Ther. 43, 139–154 (1989).
Kopp, J. L., Grompe, M. & Sander, M. Stem cells versus plasticity in liver and pancreas regeneration. Nat. Cell Biol. 18, 238–245 (2016).
Davis, R. Cre recombinase-expressing mice generated for the NIH Neuroscience Blueprint Cre Driver Network. MGI Direct Data Submission http://www.credrivermice.org (2009).
Wu, S., Ying, G., Wu, Q. & Capecchi, M. R. A protocol for constructing gene targeting vectors: generating knockout mice for the cadherin family and beyond. Nat. Protocols 3, 1056–1076 (2008).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev 15, 3243–3248 (2001).
Wu, Z., Autry, A. E., Bergan, J. F., Watabe-Uchida, M. & Dulac, C. G. Galanin neurons in the medial preoptic area govern parental behaviour. Nature 509, 325–330 (2014).
Zolotukhin, S. et al. Recombinant adeno-associated virus purification using novel methods improves infectious titer and yield. Gene Ther 6, 973–985 (1999).
Aurnhammer, C. et al. Universal real-time PCR for the detection and quantification of adeno-associated virus serotype 2-derived inverted terminal repeat sequences. Hum. Gene Ther. Methods 23, 18–28 (2012).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, New York, 2009).
Snippert, H. J., Schepers, A. G., Delconte, G., Siersema, P. D. & Clevers, H. Slide preparation for single-cell-resolution imaging of fluorescent proteins in their three-dimensional near-native environment. Nat. Protocols 6, 1221–1228 (2011).
Cui, W., Taub, D. D. & Gardner, K. qPrimerDepot: a primer database for quantitative real time PCR. Nucleic Acids Res 35, D805–D809 (2007).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Kim, N. W. & Wu, F. Advances in quantification and characterization of telomerase activity by the telomeric repeat amplification protocol (TRAP). Nucleic Acids Res 25, 2595–2597 (1997).
Acknowledgements
This work was supported by grants from the NIH (NCI CA197563 and NIA AG056575; S.E.A.), the Emerson Foundation (S.E.A.), the DFG (C.R.G), and California TRDRP (P.N.). We thank members of the Artandi laboratory, R. Nusse, P. Beachy, M. Kay and M. Krasnow for critical comments.
Reviewer information
Nature thanks S. Forbes, K. Zaret and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.L. and S.E.A. conceived the study. S.L., E.M.N. and S.E.A. designed the experiments. S.L. and C.R.G. created the Tert knock-in line. S.L. and E.M.N. performed the lineage tracing and EdU incorporation experiments. L.C. performed the TRAP assay. S.L. and P.N. performed histological analysis. S.L. and S.W. performed the AAV experiments. S.L. and A.G. performed RNA-seq analyses. S.L. and S.E.A. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Generation and characterization of the TertCreERT2/+ knock-in line.
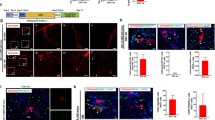
a, TertCreERT2 targeting strategy and Southern blot strategy. b–c, Southern-blots using a 5′ probe (b), a NeoR probe (c), and a 3′ probe (d). KI, knock-in cells; WT, wild-type cells. For gel source data, see Supplementary Fig. 1. e–g, TertCreERT2/+Rosa26mTmG/+ mouse ES cells, which respond to Cre-mediated recombination by switching from membrane Tomato to membrane EGFP expression (e), showed either membrane Tomato (f, overlaid on bright-field image) or membrane EGFP (g, overlaid on bright-field image) in response to 500 nM 4-hydroxy tamoxifen (4-HT). h, Hepatocytes from TertCreERT2/+Rosa26LSL-Tomato/+ livers before and after FACS enrichment. i, Tamoxifen dose–response curve for TertCreERT2/+Rosa26LSL-Tomato/+ livers (n = 3 mice for each group; horizontal bar shows mean). j, Quantification of the TRAP assay shown in Fig. 1e by densitometry. k–n, Co-immunofluorescence for Tomato (red) and CD45 (k, blood cells, 202 cells examined), CD68 (l, Kupffer cells, 179 cells examined), GFAP (m, stellate cells, 158 cells examined) and PECAM (n, endothelial cells, 167 cells examined) in TertCreERT2/+Rosa26LSL-Tomato/+ livers after 3-day trace with DAPI (blue) staining. Experiments repeated twice for b–d, f, g, k–n. Scale bars, 100 µm in g, h, 50 µm in k–n.
Extended Data Fig. 2 Ploidy and nuclear profiles of the TERTLow and TERTHigh lineages.
a–c, Ploidy analysis by Hoechst incorporation and FACS in TERTLow (a) and TERTHigh (b) hepatocytes. c, Quantification showed no significant difference between TERTLow and TERTHigh cells regarding ploidy (n = 5 mice, each represented by unique dot shapes). d–f, Nucleus count by Tomato (red), phalloidin (green) and DAPI (blue) in livers traced for 3 days (d) and 6 months (e). f, Quantification showed no significant difference between TERTLow and TERTHigh cells in binucleus fractions (n = 4 mice for each group, each represented by unique dot shapes). Experiments repeated twice. Scale bar, 50 µm.
Extended Data Fig. 3 Characterization of the lineage expansion of TERTHigh hepatocytes.
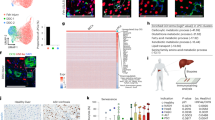
a–e, Immunofluorescence performed on TertCreERT2/+Rosa26LSL-Tomato/+ livers after one-year trace showed that only TERTHigh hepatocytes gave rise to hepatocytes. f–i, Repeated injections (f) showed that TERTHigh cells formed a constant proportion of the liver. Lineage expansion over one injection (g) and three injections (h) was quantified (i, n = 3 mice for each group; horizontal bars show mean). j, Heat map showing differentially regulated genes among all TERTLow and TERTHigh samples. Class 1 and class 2 refer to genes significantly downregulated and upregulated in TERTHigh samples, respectively. Genes assigned to DAVID-generated annotation clusters shown on the right. Experiments repeated twice. Scale bar, 200 μm.
Extended Data Fig. 4 Zonal pattern of TERTHigh lineage hepatocytes.
a–d, Stitched images of immunofluorescence for Tomato protein (red) and GS (green) in liver sections from TertCreERT2/+Rosa26LSL-Tomato/+ mice treated with tamoxifen and traced for three days (a), three months (b), six months (c) or one year (d). e–g, FACS-isolated and cytospun hepatocytes from TertCreERT2/+Rosa26LSL-Tomato/+ mice treated with tamoxifen and traced for three days were stained for CPS1 (red) and GS (green) in TERTLow (e) and TERTHigh hepatocytes (f), and quantified for the GS+ fraction of all cells (g, n = 3 mice; horizontal bars show mean). Experiments repeated three times. Scale bars, 200 μm.
Extended Data Fig. 5 Distribution of proliferating hepatocytes in Tert+/+ and TertCreERT2/+ livers in homeostasis and after injury.
a–f, Livers were stained with anti-Ki-67 antibody by standard immunohistochemistry. a–d, Ki-67+ nuclei are indicated by brown colours in uninjured livers (a, b), and CCl4 (10 μl per 10 g weight) injured livers (c, d), with haematoxylin counterstain in light blue. e, f, Green chromogen was used to indicate Ki-67+ nuclei in DDC (0.1%) treated livers. Hepatocyte nuclei were distinguished by size and morphology. Examples of Ki-67+ hepatocyte nuclei are shown in insets. g, Quantification of Ki-67+ hepatocytes and their distribution along the central–portal axis. The position index (P.I.) was determined by the distance to the most adjacent central vein (CV) (x), the distance to the most adjacent portal vein (PV) (y), and the distance between the central and portal veins (z), following the law of cosines. h–j, Two-sided Kolmogorov–Smirnov tests were performed to analyse the distributions of Ki-67+ hepatocytes along the central–portal axis. Histograms (bin-width = 0.1) and shaded curves of the kernel density estimation with Gaussian approximation are shown (mean ± s.e.m.). No significant differences were found between Tert+/+ and TertCreERT2/+ livers in uninjured livers (h, n = 4 mice for each group; each mouse represented by unique dot shapes; P = 0.58), in CCl4-injured livers (i, n = 3 mice for each group; each mouse represented by unique dot shapes, P = 0.32), or in DDC injured livers (j, n = 3 mice for each group; each mouse represented by unique dot shapes; P = 0.98). Experiments repeated twice. Scale bar, 200 μm.
Extended Data Figure 6 EdU incorporation assays.
a, Scheme of experiments. b–g, EdU incorporation in livers of TertCreERT2/+Rosa26LSL-Tomato/+ mice treated with tamoxifen, traced for three days, then treated with EdU in drinking water for 7 days (1 mg ml−1); overlay image (b), HNF4A (c), DAPI (d), EdU (e) and Tomato (f). Dashed boxes, EdU+HNF4A+Tomato+ cells. g, Quantification of EdU incorporation into hepatocytes (n = 5 mice, each represented by unique dot colours). h–k, EdU incorporation into livers of Tert+/+ (h) and TertCreERT2/+ (i) mice were compared. Co-immunofluorescence for GS (red) and CK19 (white) was overlaid with EdU (green) and DAPI (blue). j, Quantification of the distribution of EdU+ hepatocytes (pericentral, in GS+ zones; periportal, 0–2 cell layers adjacent to the portal vein space or CK19+ bile ducts; mid-lobular, neither pericentral nor periportal). Dot colours represent individual mice. k, Total EdU+ hepatocytes in Tert+/+ and TertCreERT2/+ livers (n = 5 mice for Tert+/+ livers; n = 4 mice for TertCreERT2/+ livers). Experiments repeated twice. Scale bars, 50 μm in d, 200 μm in i.
Extended Data Figure 7 Single-molecule RNA FISH on wild-type hepatocytes.
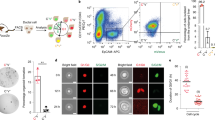
a, Experiment performed on wild-type hepatocytes isolated by FACS and cytospun. Red foci show individual Tert mRNA molecules. Control experiment by omitting the detection probe for Tert. c, Quantification by focus counts (n = 3 mice, each represented by unique dot shapes; mean + s.e.m.). Experiments repeated three times. Scale bar, 50 μm.
Extended Data Figure 8 Responses of Tert+/+ and TertCreERT2/+ livers to injuries.
a, b, Haematoxylin and eosin (H&E) staining of uninjured livers. c-d, H&E staining of livers 3 days after CCl4 injection. White dotted lines encircle the damaged pericentral area. e, f, H&E staining of livers 7 days after CCl4 injection. g, h, H&E staining of livers 1 month after DDC treatment. Experiments repeated five times. Scale bar, 200 μm.
Extended Data Figure 9 Progeny of TERTHigh hepatocytes can adopt ductal fate after DDC injury.
a–d, Immunofluorescence analysis of TertCreERT2/+Rosa26LSL-Tomato/+ livers treated with tamoxifen and DDC, and traced for 1 month (a, overlay image; b, Tomato; c, CK19. d, DAPI). e, Quantification of the percentage of CK19+Tomato+ cells among all Tomato+ cells (n = 5 mice, mean ± s.e.m. 10.0 ± 1.2%) f, Quantification of the percentage of CK19+Tomato+ cells among all CK19+ cells (n = 5 mice, mean ± s.e.m. 6.1 ± 1.0%). Bars show mean. Experiments repeated three times. Scale bar, 50 μm.
Extended Data Figure 10 Characterization of AAV-lsl-DTA and AAV-flex-DTA.
a–d, Epifluorescence of EGFP and DAPI staining of livers 4 days after injection with AAV-GFP (a), AAV-lsl-DTA (b), AAV-flex-DTA (c), and uninjected control (d). e, Diagram of AAV-flex-DTA and recombination events that lead to DTA expression. f, Survival effects of AAV-TBG viruses. Combined injection of AAV-lsl-DTA and AAV-Cre (red line) or AAV-flex-DTA and AAV-Cre (green line) lead to a narrow window of complete mortality between 4.5 and 6 days; by contrast, injection of AAV-GFP and AAV-Cre, AAV-lsl-DTA, or AAV-flex-DTA did not result in mortality. Between 4 and 6 mice were used for each regimen. Surviving mice were monitored for up to 2 months. g–j, H&E staining of liver sections from mice injected with AAV-lsl-DTA alone (g), AAV-flex-DTA alone (h), AAV-lsl-DTA and AAV-Cre (i), or AAV-flex-DTA and AAV-Cre (j). k–r, Livers injected with AAV-flex-DTA and tamoxifen showed a reduction in TERTHigh cells (k, l), as well as increases in collagen deposition (m, n), activated stellate cells (o, p) and ductal cells (q, r). Experiments repeated three times for a–d, and twice for g–r. Scale bars, 200 µm.
Supplementary information
Supplementary Information
This file contains the source data for the gel images (Supplementary Figure 1) and the Gating Strategy
Supplementary Table
This file contains the RNA-seq results
Source data
Rights and permissions
About this article
Cite this article
Lin, S., Nascimento, E.M., Gajera, C.R. et al. Distributed hepatocytes expressing telomerase repopulate the liver in homeostasis and injury. Nature 556, 244–248 (2018). https://doi.org/10.1038/s41586-018-0004-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0004-7
This article is cited by
-
Discovery of a selective TRF2 inhibitor FKB04 induced telomere shortening and senescence in liver cancer cells
Acta Pharmacologica Sinica (2024)
-
Heterogeneity of hepatocyte dynamics restores liver architecture after chemical, physical or viral damage
Nature Communications (2024)
-
The association between telomere length and non-alcoholic fatty liver disease: a prospective study
BMC Medicine (2023)
-
The MRN complex maintains the biliary-derived hepatocytes in liver regeneration through ATR-Chk1 pathway
npj Regenerative Medicine (2023)
-
Genetic recording of in vivo cell proliferation by ProTracer
Nature Protocols (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.