Abstract
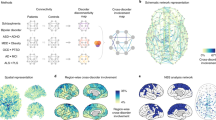
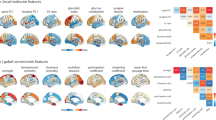
Many human brain disorders are associated with characteristic alterations in the structural and functional connectivity of the brain. In this article, we explore how commonalities and differences in connectome alterations can reveal relationships across disorders. We survey recent literature on connectivity changes in neurological and psychiatric disorders in the context of key organizational principles of the human connectome and observe that several disturbances to network properties of the human brain have a common role in a wide range of brain disorders and point towards potentially shared network mechanisms underpinning disorders. We hypothesize that the distinct dimensions along which connectome networks are organized (for example, ‘modularity’ and ‘integration’) provide a general coordinate system that allows description and categorization of relationships between seemingly disparate disorders. We outline a cross-disorder ‘connectome landscape of dysconnectivity’ along these principal dimensions of network organization that may place shared connectome alterations between brain disorders in a common framework.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Sporns, O., Tononi, G. & Kotter, R. The human connectome: a structural description of the human brain. PLOS Comput. Biol. 1, e42 (2005).
Sporns, O. Discovering the Human Connectome (MIT Press, Cambridge, 2012).
van den Heuvel, M. P. & Fornito, A. Brain networks in schizophrenia. Neuropsychol. Rev. 24, 32–48 (2014).
Fornito, A., Zalesky, A. & Breakspear, M. The connectomics of brain disorders. Nat. Rev. Neurosci. 16, 159–172 (2015).
van den Heuvel, M. P., Bullmore, E. T. & Sporns, O. Comparative connectomics. Trends Cogn. Sci. 20, 345–361 (2016).
Bullmore, E. & Sporns, O. The economy of brain network organization. Nat. Rev. Neurosci. 13, 336–349 (2012).
Avena-Koenigsberger, A., Misic, B. & Sporns, O. Communication dynamics in complex brain networks. Nat. Rev. Neurosci. 19, 17–33 (2017).
Horn, A. et al. The structural-functional connectome and the default mode network of the human brain. Neuroimage 102, 142–151 (2014).
Hagmann, P. et al. Mapping the structural core of human cerebral cortex. PLOS Biol. 6, e159 (2008).
van den Heuvel, M. P. et al. Functionally linked resting-state networks reflect the underlying structural connectivity architecture of the human brain. Hum. Brain Mapp. 30, 3127–3141 (2009).
Zhang, K. & Sejnowski, T. J. A universal scaling law between gray matter and white matter of cerebral cortex. Proc. Natl Acad. Sci. USA 97, 5621–5626 (2000).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Birnbaum, R. & Weinberger, D. R. Genetic insights into the neurodevelopmental origins of schizophrenia. Nat. Rev. Neurosci. 18, 727–740 (2017).
Silbersweig, D. A. & Rauch, S. L. Neuroimaging in psychiatry: a quarter century of progress. Harv. Rev. Psychiatry 25, 195–197 (2017).
Bassett, D. S. & Sporns, O. Network neuroscience. Nat. Neurosci. 20, 353–364 (2017).
Kaiser, M. The potential of the human connectome as a biomarker of brain disease. Front. Hum. Neurosci. 7, 484 (2013).
Petrella, J. R. Use of graph theory to evaluate brain networks: a clinical tool for a small world? Radiology 259, 317–320 (2011).
Pasquini, L. et al. Individual correspondence of amyloid-beta and intrinsic connectivity in the posterior default mode network across stages of Alzheimer’s disease. J. Alzheimers Dis. 58, 763–773 (2017).
Jung, M. et al. Default mode network in young male adults with autism spectrum disorder: relationship with autism spectrum traits. Int. J. Psychophysiol. 94, 212–212 (2014).
Whitfield-Gabrieli, S. et al. Hyperactivity and hyperconnectivity of the default network in schizophrenia and in first-degree relatives of persons with schizophrenia. Proc. Natl Acad. Sci. USA 106, 1279–1284 (2009).
Wise, T. et al. Instability of default mode network connectivity in major depression: a two-sample confirmation study. Transl Psychiatry 7, e1105 (2017).
Chenji, S. et al. Investigating default mode and sensorimotor network connectivity in amyotrophic lateral sclerosis. PLOS ONE 11, e0157443 (2016).
Lee, K. et al. Disruption, emergence and lateralization of brain network hubs in mesial temporal lobe epilepsy. Neuroimage Clin. 20, 71–84 (2018).
Rudie, J. D. et al. Altered functional and structural brain network organization in autism. Neuroimage Clin. 2, 79–94 (2012).
Lord, A. et al. Changes in community structure of resting state functional connectivity in unipolar depression. PLOS ONE 7, e41282 (2012).
Vaessen, M. J. et al. Abnormal modular organization of functional networks in cognitively impaired children with frontal lobe epilepsy. Cereb. Cortex 23, 1997–2006 (2013).
Alexander-Bloch, A. F. et al. Disrupted modularity and local connectivity of brain functional networks in childhood-onset schizophrenia. Front. Syst. Neurosci. 4, 147 (2010).
Zhan, L. et al. Baseline connectome modular abnormalities in the childhood phase of a longitudinal study on individuals with chromosome 22q11.2 deletion syndrome. Hum. Brain Mapp. 39, 232–248 (2018).
Waller, L. et al. Evaluating the replicability, specificity, and generalizability of connectome fingerprints. Neuroimage 158, 371–377 (2017).
Grisanzio, K. A. et al. Transdiagnostic symptom clusters and associations with brain, behavior, and daily function in mood, anxiety, and trauma disorders. JAMA Psychiatry 75, 201–209 (2018).
Insel, T. et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am. J. Psychiatry 167, 748–751 (2010).
Kraemer, H. C., Noda, A. & O’Hara, R. Categorical versus dimensional approaches to diagnosis: methodological challenges. J. Psychiatr. Res. 38, 17–25 (2004).
Goodkind, M. et al. Identification of a common neurobiological substrate for mental illness. JAMA Psychiatry 72, 305–315 (2015).
Foss-Feig, J. H. et al. Searching for cross-diagnostic convergence: neural mechanisms governing excitation and inhibition balance in schizophrenia and autism spectrum disorders. Biol. Psychiatry 81, 848–861 (2017).
Coleman, K. & Pierre, P. J. Assessing anxiety in nonhuman primates. ILAR J. 55, 333–346 (2014).
McTeague, L. M. et al. Identification of common neural circuit disruptions in cognitive control across psychiatric disorders. Am. J. Psychiatry 174, 676–685 (2017).
Bota, M., Dong, H. W. & Swanson, L. W. Combining collation and annotation efforts toward completion of the rat and mouse connectomes in BAMS. Front. Neuroinform. 6, 2 (2012).
Chiang, A. S. et al. Three-dimensional reconstruction of brain-wide wiring networks in Drosophila at single-cell resolution. Curr. Biol. 21, 1–11 (2010).
Smith, S. M. et al. Correspondence of the brain’s functional architecture during activation and rest. Proc. Natl Acad. Sci. USA 106, 13040–13045 (2009).
Hilger, K. et al. Intelligence is associated with the modular structure of intrinsic brain networks. Sci. Rep. 7, 16088 (2017).
Stevens, A. A. et al. Functional brain network modularity captures inter- and intra-individual variation in working memory capacity. PLOS ONE 7, e30468 (2012).
Gao, Q. et al. Extraversion and neuroticism relate to topological properties of resting-state brain networks. Front. Hum. Neurosci. 7, 257 (2013).
Adelstein, J. S. et al. Personality is reflected in the brain’s intrinsic functional architecture. PLOS ONE 6, e27633 (2011).
van den Heuvel, M. P. & Sporns, O. Network hubs in the human brain. Trends Cogn. Sci. 17, 683–696 (2013).
van den Heuvel, M. P. & Sporns, O. An anatomical substrate for integration among functional networks in human cortex. J. Neurosci. 33, 14489–14500 (2013).
Gomez-Gardenes, J. et al. From modular to centralized organization of synchronization in functional areas of the cat cerebral cortex. PLOS ONE 5, e12313 (2010).
van den Heuvel, M. P. & Sporns, O. Rich-club organization of the human connectome. J. Neurosci. 31, 11 (2011).
Collin, G. et al. Structural and functional aspects relating to cost and benefit of rich club organization in the human cerebral cortex. Cereb. Cortex 24, 2258–2267 (2014).
Zamora-Lopez, G., Zhou, C. & Kurths, J. Cortical hubs form a module for multisensory integration on top of the hierarchy of cortical networks. Front. Neuroinform. 4, 1 (2010).
van den Heuvel, M. P. et al. High-cost, high-capacity backbone for global brain communication. Proc. Natl Acad. Sci. USA 109, 11372–11377 (2012).
Senden, M. et al. Rich club organization supports a diverse set of functional network configurations. Neuroimage 96, 174–182 (2014).
Baggio, H. C. et al. Rich club organization and cognitive performance in healthy older participants. J. Cogn. Neurosci. 27, 1801–1810 (2015).
Seidlitz, J. et al. Morphometric similarity networks detect microscale cortical organization and predict inter-individual cognitive variation. Neuron 97, 231–247 (2018).
Daianu, M. et al. Disrupted rich club network in behavioral variant frontotemporal dementia and early-onset Alzheimer’s disease. Hum. Brain Mapp. 37, 868–883 (2016).
Ray, S. et al. Structural and functional connectivity of the human brain in autism spectrum disorders and attention-deficit/hyperactivity disorder: a rich club-organization study. Hum. Brain Mapp. 35, 6032–6048 (2014).
Rubinov, M. Constraints and spandrels of interareal connectomes. Nat. Commun. 7, 13812 (2016).
Chen, Y. et al. Trade-off between multiple constraints enables simultaneous formation of modules and hubs in neural systems. PLOS Comput. Biol. 9, e1002937 (2013).
Sporns, O. & Betzel, R. F. Modular brain networks. Annu. Rev. Psychol. 67, 613–640 (2016).
Kaiser, M. & Varier, S. Evolution and development of brain networks: from Caenorhabditis elegans to Homo sapiens. Network 22, 143–147 (2011).
Chen, Y. et al. Features of spatial and functional segregation and integration of the primate connectome revealed by trade-off between wiring cost and efficiency. PLOS Comput. Biol. 13, e1005776 (2017).
de Haan, W. et al. Activity dependent degeneration explains hub vulnerability in Alzheimer’s disease. PLOS Comput. Biol. 8, e1002582 (2012).
Buckner, R. L. et al. Cortical hubs revealed by intrinsic functional connectivity: mapping, assessment of stability, and relation to Alzheimer’s disease. J. Neurosci. 29, 1860–1873 (2009).
Verstraete, E. et al. Structural brain network imaging shows expanding disconnection of the motor system in amyotrophic lateral sclerosis. Hum. Brain Mapp. 35, 1351–1361 (2014).
Harrington, D. L. et al. Network topology and functional connectivity disturbances precede the onset of Huntington’s disease. Brain 138, 2332–2346 (2015).
Pedersen, M. et al. The dynamics of functional connectivity in neocortical focal epilepsy. Neuroimage Clin. 15, 209–214 (2017).
Uddin, L. Q., Supekar, K. & Menon, V. Reconceptualizing functional brain connectivity in autism from a developmental perspective. Front. Hum. Neurosci. 7, 458 (2013).
Courchesne, E. & Pierce, K. Why the frontal cortex in autism might be talking only to itself: local over-connectivity but long-distance disconnection. Curr. Opin. Neurobiol. 15, 225–230 (2005).
Griffa, A. et al. Characterizing the connectome in schizophrenia with diffusion spectrum imaging. Hum. Brain Mapp. 36, 354–366 (2015).
Lynall, M. E. et al. Functional connectivity and brain networks in schizophrenia. J. Neurosci. 30, 9477–9487 (2010).
van den Heuvel, M. P. et al. Abnormal rich club organization and functional brain dynamics in schizophrenia. JAMA Psychiatry 70, 783–792 (2013).
Wang, F. et al. Abnormal corpus callosum integrity in bipolar disorder: a diffusion tensor imaging study. Biol. Psychiatry 64, 730–733 (2008).
Roberts, G. et al. Structural dysconnectivity of key cognitive and emotional hubs in young people at high genetic risk for bipolar disorder. Mol. Psychiatry 23, 413–421 (2018).
Filippi, M. et al. Assessment of system dysfunction in the brain through MRI-based connectomics. Lancet Neurol. 12, 1189–1199 (2013).
Griffa, A. et al. Structural connectomics in brain diseases. Neuroimage 80, 515–526 (2013).
Stam, C. J. Modern network science of neurological disorders. Nat. Rev. Neurosci. 15, 683–695 (2014).
Crossley, N. A. et al. The hubs of the human connectome are generally implicated in the anatomy of brain disorders. Brain 137, 2382–2395 (2014).
Martin, G. Network analysis and the connectopathies: current research and future approaches. Nonlinear Dynam. Psychol. Life Sci. 16, 79–90 (2012).
Toga, A. W. & Thompson, P. M. Connectopathy in ageing and dementia. Brain 137, 3104–3106 (2014).
Seeley, W. W. et al. Neurodegenerative diseases target large-scale human brain networks. Neuron 62, 42–52 (2009).
Verstraete, E. et al. Impaired structural motor connectome in amyotrophic lateral sclerosis. PLOS ONE 6, e24239 (2011).
Zhou, J. et al. Predicting regional neurodegeneration from the healthy brain functional connectome. Neuron 73, 1216–1227 (2012).
Cope, T. E. et al. Tau burden and the functional connectome in Alzheimer’s disease and progressive supranuclear palsy. Brain 141, 550–567 (2018).
Raj, A., Kuceyeski, A. & Weiner, M. A network diffusion model of disease progression in dementia. Neuron 73, 1204–1215 (2012).
Zeighami, Y. et al. Network structure of brain atrophy in de novo Parkinson’s disease. eLife 4, e08440 (2015).
Schulthess, I. et al. Functional connectivity changes resemble patterns of pTDP-43 pathology in amyotrophic lateral sclerosis. Sci. Rep. 6, 38391 (2016).
Visanji, N. P. et al. Iron deficiency in parkinsonism: region-specific iron dysregulation in Parkinson’s disease and multiple system atrophy. J. Parkinsons Dis. 3, 523–537 (2013).
Brettschneider, J. et al. Spreading of pathology in neurodegenerative diseases: a focus on human studies. Nat. Rev. Neurosci. 16, 109–120 (2015).
Schmidt, R. et al. Simulating disease propagation across white matter connectome reveals anatomical substrate for neuropathology staging in amyotrophic lateral sclerosis. Neuroimage 124, 762–769 (2016).
Yau, Y. et al. Network connectivity determines cortical thinning in early Parkinson’s disease progression. Nat. Commun. 9, 12 (2018).
Vidal, C. N. et al. Dynamically spreading frontal and cingulate deficits mapped in adolescents with schizophrenia. Arch. Gen. Psychiatry 63, 25–34 (2006).
Cauda, F. et al. The morphometric co-atrophy networking of schizophrenia, autistic and obsessive spectrum disorders. Hum. Brain Mapp. 39, 1898–1928 (2018).
Raj, A. & Powell, F. Models of network spread and network degeneration in brain disorders. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 3, 788–797 (2018).
Sala-Llonch, R., Bartres-Faz, D. & Junque, C. Reorganization of brain networks in aging: a review of functional connectivity studies. Front. Psychol. 6, 663 (2015).
Hilgetag, C. C. et al. The primate connectome in context: principles of connections of the cortical visual system. Neuroimage 134, 685–702 (2016).
Beul, S. F., Grant, S. & Hilgetag, C. C. A predictive model of the cat cortical connectome based on cytoarchitecture and distance. Brain Struct. Funct. 220, 3167–3184 (2015).
van den Heuvel, M. P. et al. Multimodal analysis of cortical chemoarchitecture and macroscale fMRI resting-state functional connectivity. Hum. Brain Mapp. 37, 3103–3113 (2016).
Arnatkeviciute, A. et al. Hub connectivity, neuronal diversity, and gene expression in the Caenorhabditis elegans connectome. PLOS Comput. Biol. 14, e1005989 (2018).
Scholtens, L. H. et al. Linking macroscale graph analytical organization to microscale neuroarchitectonics in the macaque connectome. J. Neurosci. 34, 12192–12205 (2014).
van den Heuvel, M. P. et al. Bridging cytoarchitectonics and connectomics in human cerebral cortex. J. Neurosci. 35, 13943–13948 (2015).
Elston, G. N. Cortex, cognition and the cell: new insights into the pyramidal neuron and prefrontal function. Cereb. Cortex 13, 1124–1138 (2003).
Vaishnavi, S. N. et al. Regional aerobic glycolysis in the human brain. Proc. Natl Acad. Sci. USA 107, 17757–17762 (2011).
Fulcher, B. D. & Fornito, A. A transcriptional signature of hub connectivity in the mouse connectome. Proc. Natl Acad. Sci. USA 113, 1435–1440 (2016).
Iturria-Medina, Y. & Evans, A. C. On the central role of brain connectivity in neurodegenerative disease progression. Front. Aging Neurosci. 7, 90 (2015).
Jones, D. T. et al. Tau, amyloid, and cascading network failure across the Alzheimer’s disease spectrum. Cortex 97, 143–159 (2017).
Jones, D. T. et al. Cascading network failure across the Alzheimer’s disease spectrum. Brain 139, 547–562 (2016).
Pahwa, S., Scoglio, C. & Scala, A. Abruptness of cascade failures in power grids. Sci. Rep. 4, 3694 (2014).
de Lange, S. C. et al. Shared vulnerability for connectome alterations across psychiatric and neurological brain disorders. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/360586v1 (2018).
Senden, M. et al. Task-related effective connectivity reveals that the cortical rich club gates cortex-wide communication. Hum. Brain Mapp. 39, 1246–1262 (2018).
Vertes, P. E. et al. Simple models of human brain functional networks. Proc. Natl Acad. Sci. USA 109, 5868–5873 (2012).
Fair, D. A. et al. Functional brain networks develop from a “local to distributed” organization. PLOS Comput. Biol. 5, e1000381 (2009).
Wolf, D. A. & Mash, E. J. (eds) Behavioral and Emotional Disorders in Adolescents: Nature, Assessment, and Treatment (Guilford Press, 2008).
Luby, J. L. et al. Early childhood depression and alterations in the trajectory of gray matter maturation in middle childhood and early adolescence. JAMA Psychiatry 73, 31–38 (2016).
Avena-Koenigsberger, A. et al. Network morphospace. J. R. Soc. Interface 12, 20140881 (2015).
Menke, R. A. L. et al. Increased functional connectivity common to symptomatic amyotrophic lateral sclerosis and those at genetic risk. J. Neurol. Neurosurg. Psychiatry 87, 580–588 (2016).
Grefkes, C. & Fink, G. R. Reorganization of cerebral networks after stroke: new insights from neuroimaging with connectivity approaches. Brain 134, 1264–1276 (2011).
Li, Y. X. et al. Alterations in spontaneous brain activity and functional network reorganization following surgery in children with medically refractory epilepsy: a resting-state functional magnetic resonance imaging study. Front. Neurol. 8, 374 (2017).
Mohammadi, B. et al. Functional neuroimaging at different disease stages reveals distinct phases of neuroplastic changes in amyotrophic lateral sclerosis. Hum. Brain Mapp. 32, 750–758 (2011).
Filippi, M. et al. Brain network connectivity differs in early-onset neurodegenerative dementia. Neurology 89, 1764–1772 (2017).
Douaud, G. et al. Integration of structural and functional magnetic resonance imaging in amyotrophic lateral sclerosis. Brain 134, 3470–3479 (2011).
Hillary, F. G. & Grafman, J. H. Injured brains and adaptive networks: the benefits and costs of hyperconnectivity. Trends Cogn. Sci. 21, 385–401 (2017).
de Haan, W. et al. Altering neuronal excitability to preserve network connectivity in a computational model of Alzheimer’s disease. PLOS Comput. Biol. 13, e1005707 (2017).
Liu, J. et al. Enhanced interhemispheric functional connectivity compensates for anatomical connection damages in subcortical stroke. Stroke 46, 1045–1051 (2015).
Zhang, H. Y. et al. Resting brain connectivity: changes during the progress of Alzheimer disease. Radiology 256, 598–606 (2010).
Dima, D., Roberts, R. E. & Frangou, S. Connectomic markers of disease expression, genetic risk and resilience in bipolar disorder. Transl Psychiatry 6, e706 (2016).
Braun, U. et al. From maps to multi-dimensional network mechanisms of mental disorders. Neuron 97, 14–31 (2018).
Cuthbert, B. N. The RDoC framework: facilitating transition from ICD/DSM to dimensional approaches that integrate neuroscience and psychopathology. World Psychiatry 13, 28–35 (2014).
van der Burgh, H. K. et al. Deep learning predictions of survival based on MRI in amyotrophic lateral sclerosis. Neuroimage Clin. 13, 361–369 (2017).
Scholtens, L. H. & van den Heuvel, M. P. Multimodal connectomics in psychiatry: bridging scales from micro to macro. Biol. Psychiatry 3, 767–776 (2018).
Kotov, R. et al. New dimensions in the quantitative classification of mental illness. Arch. Gen. Psychiatry 68, 1003–1011 (2011).
Miranda-Dominguez, O. et al. Connectotyping: model based fingerprinting of the functional connectome. PLOS ONE 9, e111048 (2014).
O’Donoghue, S. et al. Anatomical dysconnectivity in bipolar disorder compared with schizophrenia: A selective review of structural network analyses using diffusion MRI. J. Affect. Disord. 209, 217–228 (2017).
Finn, E. S. et al. Functional connectome fingerprinting: identifying individuals using patterns of brain connectivity. Nat. Neurosci. 18, 1664–1671 (2015).
Thompson, P. M. et al. ENIGMA and the individual: Predicting factors that affect the brain in 35 countries worldwide. Neuroimage 145, 389–408 (2017).
Jack, C. R. Jr et al. The Alzheimer’s Disease Neuroimaging Initiative (ADNI): MRI methods. J. Magn. Reson. Imaging 27, 685–691 (2008).
Mirzaei, G., Adeli, A. & Adeli, H. Imaging and machine learning techniques for diagnosis of Alzheimer’s disease. Rev. Neurosci. 27, 857–870 (2016).
Simpraga, S. et al. EEG machine learning for accurate detection of cholinergic intervention and Alzheimer’s disease. Sci. Rep. 7, 5775 (2017).
Brown, M. R. et al. ADHD-200 Global Competition: diagnosing ADHD using personal characteristic data can outperform resting state fMRI measurements. Front. Syst. Neurosci. 6, 69 (2012).
Schnack, H. G. et al. Can structural MRI aid in clinical classification? A machine learning study in two independent samples of patients with schizophrenia, bipolar disorder and healthy subjects. Neuroimage 84, 299–306 (2014).
Salvador, R. et al. Evaluation of machine learning algorithms and structural features for optimal MRI-based diagnostic prediction in psychosis. PLOS ONE 12, e0175683 (2017).
Weng, S. F. et al. Can machine-learning improve cardiovascular risk prediction using routine clinical data? PLOS ONE 12, e0174944 (2017).
Ramasubbu, R. et al. Accuracy of automated classification of major depressive disorder as a function of symptom severity. Neuroimage Clin. 12, 320–331 (2016).
Doucet, G. E. et al. The role of intrinsic brain functional connectivity in vulnerability and resilience to bipolar disorder. Am. J. Psychiatry 174, 1214–1222 (2017).
Schmidt, A. et al. Structural network disorganization in subjects at clinical high risk for psychosis. Schizophr. Bull. 43, 583–591 (2017).
Collin, G. et al. Affected anatomical rich club and structural-functional coupling in young offspring of schizophrenia and bipolar disorder patients. Biol. Psychiatry 82, 746–755 (2017).
Deco, G. et al. Rethinking segregation and integration: contributions of whole-brain modelling. Nat. Rev. Neurosci. 16, 430–439 (2015).
Sanz Leon, P. et al. The Virtual Brain: a simulator of primate brain network dynamics. Front. Neuroinform. 7, 10 (2013).
Grayson, D. S. et al. The rhesus monkey connectome predicts disrupted functional networks resulting from pharmacogenetic inactivation of the amygdala. Neuron 91, 453–466 (2016).
Silasi, G. & Murphy, T. H. Stroke and the connectome: how connectivity guides therapeutic intervention. Neuron 83, 1354–1368 (2014).
Alstott, J. et al. Modeling the impact of lesions in the human brain. PLOS Comput. Biol. 5, e1000408 (2009).
Aerts, H. et al. Brain networks under attack: robustness properties and the impact of lesions. Brain 139, 3063–3083 (2016).
Ellegood, J. et al. Analysis of neuroanatomical differences in mice with genetically modified serotonin transporters assessed by structural magnetic resonance imaging. Mol. Autism 9, 24 (2018).
Mechling, A. E. et al. Deletion of the mu opioid receptor gene in mice reshapes the reward-aversion connectome. Proc. Natl Acad. Sci. USA 113, 11603–11608 (2016).
Grandjean, J. et al. Chronic psychosocial stress in mice leads to changes in brain functional connectivity and metabolite levels comparable to human depression. Neuroimage 142, 544–552 (2016).
Whitfield-Gabrieli, S. et al. Brain connectomics predict response to treatment in social anxiety disorder. Mol. Psychiatry 21, 680–685 (2016).
Bullmore, E. & Sporns, O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009).
Sanz-Arigita, E. J. et al. Loss of ‘small-world’ networks in Alzheimer’s disease: graph analysis of FMRI resting-state functional connectivity. PLOS ONE 5, e13788 (2010).
Stam, C. J. & Reijneveld, J. C. Graph theoretical analysis of complex networks in the brain. Nonlinear Biomed. Phys. 1, 3 (2007).
Chen, G. et al. Modular reorganization of brain resting state networks and its independent validation in Alzheimer’s disease patients. Front. Hum. Neurosci. 7, 456 (2013).
Prescott, J. W. et al. The Alzheimer structural connectome: changes in cortical network topology with increased amyloid plaque burden. Radiology 273, 175–184 (2014).
Castellazzi, G. et al. A comprehensive assessment of resting state networks: bidirectional modification of functional integrity in cerebro-cerebellar networks in dementia. Front. Neurosci. 8, 223 (2014).
Pineda-Pardo, J. A. et al. Guiding functional connectivity estimation by structural connectivity in MEG: an application to discrimination of conditions of mild cognitive impairment. Neuroimage 101, 765–777 (2014).
Zhu, D. J. et al. Connectome-scale assessments of structural and functional connectivity in MCI. Hum. Brain Mapp. 35, 2911–2923 (2014).
Wang, J. H. et al. Disrupted functional brain connectome in individuals at risk for Alzheimer’s disease. Biol. Psychiatry 73, 472–481 (2013).
Odish, O. F. F. et al. Dynamics of the connectome in Huntington’s disease: a longitudinal diffusion MRI study. Neuroimage Clin. 9, 32–43 (2015).
McColgan, P. et al. Selective vulnerability of Rich Club brain regions is an organizational principle of structural connectivity loss in Huntington’s disease. Brain 138, 3327–3344 (2015).
Tinaz, S. et al. Changes in functional organization and white matter integrity in the connectome in Parkinson’s disease. Neuroimage Clin. 13, 395–404 (2017).
Lee, S. E. et al. Network degeneration and dysfunction in presymptomatic C9ORF72 expansion carriers. Neuroimage Clin. 14, 286–297 (2017).
de Albuquerque, M. et al. Longitudinal evaluation of cerebral and spinal cord damage in amyotrophic lateral sclerosis. Neuroimage Clin. 14, 269–276 (2017).
Shu, N. et al. Disrupted topological organization of structural and functional brain connectomes in clinically isolated syndrome and multiple sclerosis. Sci. Rep. 6, 29383 (2016).
Liu, Y. et al. Functional brain network alterations in clinically isolated syndrome and multiple sclerosis: a graph-based connectome study. Radiology 282, 534–541 (2017).
Kocevar, G. et al. Graph theory-based brain connectivity for automatic classification of multiple sclerosis clinical courses. Front. Neurosci. 10, 478 (2016).
Burns, S. P. et al. A network analysis of the dynamics of seizure. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2012, 4684–4687 (2012).
Bernhardt, B. C., Bonilha, L. & Gross, D. W. Network analysis for a network disorder: the emerging role of graph theory in the study of epilepsy. Epilepsy Behav. 50, 162–170 (2015).
Bonilha, L. et al. The brain connectome as a personalized biomarker of seizure outcomes after temporal lobectomy. Neurology 84, 1846–1853 (2015).
Just, M. A. et al. Cortical activation and synchronization during sentence comprehension in high-functioning autism: evidence of underconnectivity. Brain 127, 1811–1821 (2004).
Mevel, K. & Fransson, P. The functional brain connectome of the child and autism spectrum disorders. Acta Paediatr. 105, 1024–1035 (2016).
Zalesky, A. et al. Disrupted axonal fiber connectivity in schizophrenia. Biol. Psychiatry 69, 80–89 (2011).
Kelly, S. et al. Widespread white matter microstructural differences in schizophrenia across 4322 individuals: results from the ENIGMA Schizophrenia DTI Working Group. Mol. Psychiatry 23, 1261–1269 (2017).
Klauser, P. et al. White matter disruptions in schizophrenia are spatially widespread and topologically converge on brain network hubs. Schizophr. Bull. 43, 425–435 (2017).
Collin, G. et al. Impaired rich club connectivity in unaffected siblings of schizophrenia patients. Schizophr. Bull. 40, 438–448 (2014).
Calhoun, V. D. et al. Exploring the psychosis functional connectome: aberrant intrinsic networks in schizophrenia and bipolar disorder. Front. Psychiatry 2, 75 (2011).
Colibazzi, T. et al. Aberrant temporal connectivity in persons at clinical high risk for psychosis. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 2, 696–705 (2017).
Yu, Q. et al. Disrupted correlation between low frequency power and connectivity strength of resting state brain networks in schizophrenia. Schizophr. Res. 143, 165–171 (2013).
Lo, C. Y. Z. et al. Randomization and resilience of brain functional networks as systems-level endophenotypes of schizophrenia. Proc. Natl Acad. Sci. USA 112, 9123–9128 (2015).
Sun, Y. et al. Modular-level alterations of structure-function coupling in schizophrenia connectome. Hum. Brain Mapp. 38, 2008–2025 (2017).
Sarrazin, S. et al. Corpus callosum area in patients with bipolar disorder with and without psychotic features: an international multicentre study. J. Psychiatry Neurosci. 40, 352–359 (2015).
Collin, G. et al. Brain network analysis reveals affected connectome structure in bipolar I disorder. Hum. Brain Mapp. 37, 122–134 (2016).
Ajilore, O. et al. Connectome signatures of neurocognitive abnormalities in euthymic bipolar I disorder. Neuropsychopharmacology 39, S227–S228 (2014).
Lois, G. et al. Large-scale network functional interactions during distraction and reappraisal in remitted bipolar and unipolar patients. Bipolar Disord. 19, 487–495 (2017).
Wang, Y. et al. Topologically convergent and divergent functional connectivity patterns in unmedicated unipolar depression and bipolar disorder. Transl Psychiatry 7, e1165 (2017).
Korgaonkar, M. S. et al. Abnormal structural networks characterize major depressive disorder: a connectome analysis. Biol. Psychiatry 76, 567–574 (2014).
Satterthwaite, T. D. et al. Dimensional depression severity in women with major depression and post-traumatic stress disorder correlates with fronto-amygdalar hypoconnectivty. Mol. Psychiatry 21, 894–902 (2016).
Drysdale, A. T. et al. Resting-state connectivity biomarkers define neurophysiological subtypes of depression. Nat. Med. 23, 28–38 (2017).
Tagliazucchi, E. & van Someren, E. J. W. The large-scale functional connectivity correlates of consciousness and arousal during the healthy and pathological human sleep cycle. Neuroimage 160, 55–72 (2017).
Shine, J. M. et al. The dynamics of functional brain networks: integrated network states during cognitive task performance. Neuron 92, 544–554 (2016).
Xia, C. H. et al. Linked dimensions of psychopathology and connectivity in functional brain networks. Nat. Commun. 9, 3003 (2018).
Elliot, M. L. et al. A connectome-wide functional signature of transdiagnostic risk for mental illness. Biol. Psychiatry 84, 452–459 (2017).
Yoon, Y. B. et al. Altered fronto-temporal functional connectivity in individuals at ultra-high-risk of developing psychosis. PLOS ONE 10, e0135347 (2015).
Mitchell, P. et al. Structural dysconnectivity of key cognitive and emotional hubs in young people at high genetic risk for bipolar disorder. Biol. Psychiatry 81, S318–S319 (2017).
Kaufmann, T. et al. Delayed stabilization and individualization in connectome development are related to psychiatric disorders. Nat. Neurosci. 20, 513–515 (2017).
Mueller, S. et al. Individual variability in functional connectivity architecture of the human brain. Neuron 77, 586–595 (2013).
Wook Yoo, S. et al. A network flow-based analysis of cognitive reserve in normal ageing and Alzheimer’s disease. Sci. Rep. 5, 10057 (2015).
Bozzali, M. et al. The impact of cognitive reserve on brain functional connectivity in Alzheimer’s disease. J. Alzheimers Dis. 44, 243–250 (2015).
Brickman, A. M. et al. White matter hyperintensities and cognition: testing the reserve hypothesis. Neurobiol. Aging 32, 1588–1598 (2011).
Ganella, E. P. et al. Risk and resilience brain networks in treatment-resistant schizophrenia. Schizophr. Res. 193, 284–292 (2017).
Acknowledgements
The authors thank A. Griffa and A. Zalesky for helpful comments and discussions on earlier versions of the manuscript. M.P.v.d.H. was funded by VIDI (NWO-VIDI 452-16-015) and ALWopen (ALWOP.179) grants from the Netherlands Organization for Scientific Research and by a fellowship from MQ. O.S. was supported by the US National Institutes of Health (grant R01 AT009036-01).
Reviewer information
Nature Reviews Neuroscience thanks D. Fair and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
The authors both researched data for article, provided substantial contributions to discussion of its content, wrote the article and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
van den Heuvel, M.P., Sporns, O. A cross-disorder connectome landscape of brain dysconnectivity. Nat Rev Neurosci 20, 435–446 (2019). https://doi.org/10.1038/s41583-019-0177-6
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41583-019-0177-6
This article is cited by
-
Genetic overlap between multivariate measures of human functional brain connectivity and psychiatric disorders
Nature Mental Health (2024)
-
Abundant pleiotropy across neuroimaging modalities identified through a multivariate genome-wide association study
Nature Communications (2024)
-
Modeling brain network flexibility in networks of coupled oscillators: a feasibility study
Scientific Reports (2024)
-
Altered whole-brain functional network in patients with frontal low-grade gliomas: a resting-state functional MRI study
Neuroradiology (2024)
-
Atypical local brain connectivity in pediatric autism spectrum disorder? A coordinate-based meta-analysis of regional homogeneity studies
European Archives of Psychiatry and Clinical Neuroscience (2024)