Abstract
The suprachiasmatic nucleus (SCN) of the hypothalamus is remarkable. Despite numbering only about 10,000 neurons on each side of the third ventricle, the SCN is our principal circadian clock, directing the daily cycles of behaviour and physiology that set the tempo of our lives. When this nucleus is isolated in organotypic culture, its autonomous timing mechanism can persist indefinitely, with precision and robustness. The discovery of the cell-autonomous transcriptional and post-translational feedback loops that drive circadian activity in the SCN provided a powerful exemplar of the genetic specification of complex mammalian behaviours. However, the analysis of circadian time-keeping is moving beyond single cells. Technical and conceptual advances, including intersectional genetics, multidimensional imaging and network theory, are beginning to uncover the circuit-level mechanisms and emergent properties that make the SCN a uniquely precise and robust clock. However, much remains unknown about the SCN, not least the intrinsic properties of SCN neurons, its circuit topology and the neuronal computations that these circuits support. Moreover, the convention that the SCN is a neuronal clock has been overturned by the discovery that astrocytes are an integral part of the timepiece. As a test bed for examining the relationships between genes, cells and circuits in sculpting complex behaviours, the SCN continues to offer powerful lessons and opportunities for contemporary neuroscience.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Czeisler, C. A. SLEEP. Measuring the passage of brain time. Science 353, 648–649 (2016).
Potter, G. D. et al. Circadian rhythm and sleep disruption: causes, metabolic consequences, and countermeasures. Endocr. Rev. 37, 584–608 (2016).
Dallmann, R., Brown, S. A. & Gachon, F. Chronopharmacology: new insights and therapeutic implications. Annu. Rev. Pharmacol. Toxicol. 54, 339–361 (2014).
Reppert, S. M. & Weaver, D. R. Coordination of circadian timing in mammals. Nature 418, 935–941 (2002).
Kyriacou, C. P. & Hastings, M. H. Circadian clocks: genes, sleep, and cognition. Trends Cogn. Sci. 14, 259–267 (2010).
Santhi, N. et al. Sex differences in the circadian regulation of sleep and waking cognition in humans. Proc. Natl Acad. Sci. USA 113, E2730–E2739 (2016). This study is an elegant use of forced-desynchrony protocols to reveal circadian contributions to the control of sleep timing and quality, and waking cognition in humans.
de Vivo, L. et al. Ultrastructural evidence for synaptic scaling across the wake/sleep cycle. Science 355, 507–510 (2017).
Zhang, R., Lahens, N. F., Ballance, H. I., Hughes, M. E. & Hogenesch, J. B. A circadian gene expression atlas in mammals: implications for biology and medicine. Proc. Natl Acad. Sci. USA 111, 16219–16224 (2014). This paper provides an extensive systematic and comprehensive analysis of circadian gene expression profiles across multiple tissues of mice and the relationship of circadian gene expression to current and future clinical practice.
Mure, L. S. et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 359, E2730 (2018). This study is an extensive systematic and comprehensive analysis of circadian gene expression profiles across multiple tissues of baboons that will be a vital comparative resource for studies of human circadian biology.
Roh, J. H. et al. Disruption of the sleep-wake cycle and diurnal fluctuation of beta-amyloid in mice with Alzheimer’s disease pathology. Sci. Transl. Med. 4, 150ra122 (2012).
Xie, L. et al. Sleep drives metabolite clearance from the adult brain. Science 342, 373–377 (2013). This article provides a demonstration of the central role of sleep in brain homeostasis.
Frank, E. et al. Influencing circadian and sleep-wake regulation for prevention and intervention in mood and anxiety disorders: what makes a good homeostat? Ann. NY Acad. Sci. 1334, 1–25 (2014).
Hastings, M. H. & Goedert, M. Circadian clocks and neurodegenerative diseases: time to aggregate? Curr. Opin. Neurobiol. 23, 880–887 (2013).
Panda, S. Circadian physiology of metabolism. Science 354, 1008–1015 (2016).
Weaver, D. R. The suprachiasmatic nucleus: a 25-year retrospective. J. Biol. Rhythms 13, 100–112 (1998). This paper is a catalogue of the early studies that established the SCN as the central circadian pacemaker of mammals.
Ralph, M. R., Foster, R. G., Davis, F. C. & Menaker, M. Transplanted suprachiasmatic nucleus determines circadian period. Science 247, 975–978 (1990).
King, V. M. et al. A hVIPR transgene as a novel tool for the analysis of circadian function in the mouse suprachiasmatic nucleus. Eur. J. Neurosci. 17, 822–832 (2003).
Colwell, C. S. Linking neural activity and molecular oscillations in the SCN. Nat. Rev. Neurosci. 12, 553–569 (2011).
LeGates, T. A., Fernandez, D. C. & Hattar, S. Light as a central modulator of circadian rhythms, sleep and affect. Nat. Rev. Neurosci. 15, 443–454 (2014).
Mohawk, J. A., Green, C. B. & Takahashi, J. S. Central and peripheral circadian clocks in mammals. Annu. Rev. Neurosci. 35, 445–462 (2012).
Hastings, M. H., Reddy, A. B. & Maywood, E. S. A clockwork web: circadian timing in brain and periphery, in health and disease. Nat. Rev. Neurosci. 4, 649–661 (2003).
Gerber, A. et al. Blood-borne circadian signal stimulates daily oscillations in actin dynamics and SRF activity. Cell 152, 492–503 (2013).
Morf, J. & Schibler, U. Body temperature cycles: gatekeepers of circadian clocks. Cell Cycle 12, 539–540 (2013).
Gizowski, C., Zaelzer, C. & Bourque, C. W. Clock-driven vasopressin neurotransmission mediates anticipatory thirst prior to sleep. Nature 537, 685–688 (2016).
Kalsbeek, A. et al. SCN outputs and the hypothalamic balance of life. J. Biol. Rhythms 21, 458–469 (2006).
Gjorgjieva, J., Drion, G. & Marder, E. Computational implications of biophysical diversity and multiple timescales in neurons and synapses for circuit performance. Curr. Opin. Neurobiol. 37, 44–52 (2016).
Yan, G. et al. Network control principles predict neuron function in the Caenorhabditis elegans connectome. Nature 550, 519–523 (2017).
Callaway, E. & Ledford, H. Medicine Nobel awarded for work on circadian clocks. Nature 550, 18 (2017).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Cho, H. et al. Regulation of circadian behaviour and metabolism by REV-ERB-alpha and REV-ERB-beta. Nature 485, 123–127 (2012).
Solt, L. A. et al. Regulation of circadian behaviour and metabolism by synthetic REV-ERB agonists. Nature 485, 62–68 (2012).
Akhtar, R. A. et al. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr. Biol. 12, 540–550 (2002).
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002).
Pizarro, A., Hayer, K., Lahens, N. F. & Hogenesch, J. B. CircaDB: a database of mammalian circadian gene expression profiles. Nucleic Acids Res. 41, D1009–D1013 (2013).
Abrahamson, E. E. & Moore, R. Y. Suprachiasmatic nucleus in the mouse: retinal innervation, intrinsic organization and efferent projections. Brain Res. 916, 172–191 (2001).
Noguchi, T. et al. Calcium circadian rhythmicity in the suprachiasmatic nucleus: cell autonomy and network modulation. eNeuro 4, 0160–17 (2017).
Liu, A. C. et al. Intercellular coupling confers robustness against mutations in the SCN circadian clock network. Cell 129, 605–616 (2007).
Patton, A. P., Chesham, J. E. & Hastings, M. H. Combined pharmacological and genetic manipulations unlock unprecedented temporal elasticity and reveal phase-specific modulation of the molecular circadian clock of the mouse suprachiasmatic nucleus. J. Neurosci. 36, 9326–9341 (2016).
Meijer, J. H. & Michel, S. Neurophysiological analysis of the suprachiasmatic nucleus: a challenge at multiple levels. Methods Enzymol. 552, 75–102 (2015).
Houben, T., Coomans, C. P. & Meijer, J. H. Regulation of circadian and acute activity levels by the murine suprachiasmatic nuclei. PLoS ONE 9, e110172 (2014).
Sato, T. & Kawamura, H. Circadian rhythms in multiple unit activity inside and outside the suprachiasmatic nucleus in the diurnal chipmunk (Eutamias sibiricus). Neurosci. Res. 1, 45–52 (1984).
Saper, C. B., Scammell, T. E. & Lu, J. Hypothalamic regulation of sleep and circadian rhythms. Nature 437, 1257–1263 (2005).
Fernandez, D. C., Chang, Y. T., Hattar, S. & Chen, S. K. Architecture of retinal projections to the central circadian pacemaker. Proc. Natl Acad. Sci. USA 113, 6047–6052 (2016).
Ebling, F. J. The role of glutamate in the photic regulation of the suprachiasmatic nucleus. Prog. Neurobiol. 50, 109–132 (1996).
Jones, J. R., Tackenberg, M. C. & McMahon, D. G. Manipulating circadian clock neuron firing rate resets molecular circadian rhythms and behavior. Nat. Neurosci. 18, 373–375 (2015).
Sakamoto, K. et al. Clock and light regulation of the CREB coactivator CRTC1 in the suprachiasmatic circadian clock. J. Neurosci. 33, 9021–9027 (2013).
Jagannath, A. et al. The CRTC1-SIK1 pathway regulates entrainment of the circadian clock. Cell 154, 1100–1111 (2013).
Wegner, S., Belle, M. D. C., Hughes, A. T. L., Diekman, C. O. & Piggins, H. D. Delayed cryptochrome degradation asymmetrically alters the daily rhythm in suprachiasmatic clock neuron excitability. J. Neurosci. 37, 7824–7836 (2017).
Chiang, C. K. et al. The proteomic landscape of the suprachiasmatic nucleus clock reveals large-scale coordination of key biological processes. PLoS Genet. 10, e1004695 (2014).
Deery, M. J. et al. Proteomic analysis reveals the role of synaptic vesicle cycling in sustaining the suprachiasmatic circadian clock. Curr. Biol. 19, 2031–2036 (2009).
Herzog, E. D., Hermanstyne, T., Smyllie, N. J. & Hastings, M. H. Regulating the suprachiasmatic nucleus (SCN) circadian clockwork: interplay between cell-autonomous and circuit-level mechanisms. Cold Spring Harb. Perspect. Biol. 9, a027706 (2017).
Flourakis, M. et al. A conserved bicycle model for circadian clock control of membrane excitability. Cell 162, 836–848 (2015).
Whitt, J. P., Montgomery, J. R. & Meredith, A. L. BK channel inactivation gates daytime excitability in the circadian clock. Nat. Commun. 7, 10837 (2016).
Hermanstyne, T. O., Granados-Fuentes, D., Mellor, R. L., Herzog, E. D. & Nerbonne, J. M. Acute knockdown of Kv4.1 regulates repetitive firing rates and clock gene expression in the suprachiasmatic nucleus and daily rhythms in locomotor behavior. eNeuro https://doi.org/10.1523/ENEURO.0377-16.2017 (2017).
Granados-Fuentes, D., Hermanstyne, T. O., Carrasquillo, Y., Nerbonne, J. M. & Herzog, E. D. IA channels encoded by Kv1.4 and Kv4.2 regulate circadian period of PER2 expression in the suprachiasmatic nucleus. J. Biol. Rhythms 30, 396–407 (2015).
Brancaccio, M., Maywood, E. S., Chesham, J. E., Loudon, A. S. & Hastings, M. H. A Gq-Ca(2+) axis controls circuit-level encoding of circadian time in the suprachiasmatic nucleus. Neuron 78, 714–728 (2013).
Yamaguchi, S. et al. Synchronization of cellular clocks in the suprachiasmatic nucleus. Science 302, 1408–1412 (2003).
Maywood, E. S. et al. Analysis of core circadian feedback loop in suprachiasmatic nucleus of mCry1-luc transgenic reporter mouse. Proc. Natl Acad. Sci. USA 110, 9547–9552 (2013).
Yoo, S. H. et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl Acad. Sci. USA 101, 5339–5346 (2004).
Parsons, M. J. et al. The regulatory factor ZFHX3 modifies circadian function in SCN via an AT motif-driven axis. Cell 162, 607–621 (2015).
Smyllie, N. J. et al. Visualizing and quantifying intracellular behavior and abundance of the core circadian clock protein PERIOD2. Curr. Biol. 26, 1880–1886 (2016).
Brancaccio, M., Patton, A. P., Chesham, J. E., Maywood, E. S. & Hastings, M. H. Astrocytes control circadian timekeeping in the suprachiasmatic nucleus via glutamatergic signaling. Neuron 93, 1420–1435 e5 (2017). This article presents a demonstration of the central role of glutamatergic signalling by astrocytes in controlling circadian pacemaking in the SCN.
Cohen, S. & Greenberg, M. E. Communication between the synapse and the nucleus in neuronal development, plasticity, and disease. Annu. Rev. Cell Dev. Biol. 24, 183–209 (2008).
Schmal, C., Herzog, E. D. & Herzel, H. Measuring relative coupling strength in circadian systems. J. Biol. Rhythms 33, 84–98 (2018).
Koinuma, S. et al. Regional circadian period difference in the suprachiasmatic nucleus of the mammalian circadian center. Eur. J. Neurosci. 38, 2832–2841 (2013).
Enoki, R. et al. Topological specificity and hierarchical network of the circadian calcium rhythm in the suprachiasmatic nucleus. Proc. Natl Acad. Sci. USA 109, 21498–21503 (2012).
Enoki, R., Ono, D., Kuroda, S., Honma, S. & Honma, K. I. Dual origins of the intracellular circadian calcium rhythm in the suprachiasmatic nucleus. Sci. Rep. 7, 41733 (2017).
Pauls, S. D., Honma, K., Honma, S. & Silver, R. Deconstructing circadian rhythmicity with models and manipulations. Trends Neurosci. 39, 405–419 (2016).
Paszek, P. et al. Population robustness arising from cellular heterogeneity. Proc. Natl Acad. Sci. USA 107, 11644–11649 (2010).
Evans, J. A., Leise, T. L., Castanon-Cervantes, O. & Davidson, A. J. Dynamic interactions mediated by nonredundant signaling mechanisms couple circadian clock neurons. Neuron 80, 973–983 (2013). This study is an elegant demonstration of the complementary roles of GABA and neuropeptidergic signalling in sculpting circuit-level circadian time-keeping in the SCN.
Smyllie, N. J., Chesham, J. E., Hamnett, R., Maywood, E. S. & Hastings, M. H. Temporally chimeric mice reveal flexibility of circadian period-setting in the suprachiasmatic nucleus. Proc. Natl Acad. Sci. USA. 113, 3657–62 (2016).
Doi, M. et al. Circadian regulation of intracellular G-protein signalling mediates intercellular synchrony and rhythmicity in the suprachiasmatic nucleus. Nat. Commun. 2, 327 (2011).
Hastings, M. H., Brancaccio, M. & Maywood, E. S. Circadian pacemaking in cells and circuits of the suprachiasmatic nucleus. J. Neuroendocrinol. 26, 2–10 (2014).
Maywood, E. S., Chesham, J. E., O’Brien, J. A. & Hastings, M. H. A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc. Natl Acad. Sci. USA 108, 14306–14311 (2011).
Albus, H., Vansteensel, M. J., Michel, S., Block, G. D. & Meijer, J. H. A. GABAergic mechanism is necessary for coupling dissociable ventral and dorsal regional oscillators within the circadian clock. Curr. Biol. 15, 886–893 (2005).
Freeman, G. M. et al. GABA networks destabilize genetic oscillations in the circadian pacemaker. Neuron 78, 799–806 (2013).
Yoshikawa, T. et al. Localization of photoperiod responsive circadian oscillators in the mouse suprachiasmatic nucleus. Sci. Rep. 7, 8210 (2017).
Azzi, A. et al. Network dynamics mediate circadian clock plasticity. Neuron 93, 441–450 (2017).
Farajnia, S., van Westering, T. L., Meijer, J. H. & Michel, S. Seasonal induction of GABAergic excitation in the central mammalian clock. Proc. Natl Acad. Sci. USA 111, 9627–9632 (2014).
Brown, L. A. et al. Meta-analysis of transcriptomic datasets identifies genes enriched in the mammalian circadian pacemaker. Nucleic Acids Res. 45, 9860–9873 (2017).
Park, J. et al. Single-cell transcriptional analysis reveals novel neuronal phenotypes and interaction networks involved in the central circadian clock. Front. Neurosci. 10, 481 (2016). This study is an innovative use of single-cell transcriptomics to categorize distinct cell populations (modules) in the SCN defined by their respective signalling pathways.
Yamaguchi, Y. et al. Mice genetically deficient in vasopressin V1a and V1b receptors are resistant to jet lag. Science 342, 85–90 (2013).
Yamaguchi, Y. Arginine vasopressin signaling in the suprachiasmatic nucleus on the resilience of circadian clock to jet lag. Neurosci. Res. 129, 57–61 (2017).
Zhou, Q. Y. & Cheng, M. Y. Prokineticin 2 and circadian clock output. FEBS J. 272, 5703–5709 (2005).
Prosser, H. M. et al. Prokineticin receptor 2 (Prokr2) is essential for the regulation of circadian behavior by the suprachiasmatic nuclei. Proc. Natl Acad. Sci. USA 104, 648–653 (2007).
Qu, Z. et al. Loss of ZBTB20 impairs circadian output and leads to unimodal behavioral rhythms. eLife 5, e17171 (2016).
Doi, M. et al. Gpr176 is a Gz-linked orphan G-protein-coupled receptor that sets the pace of circadian behaviour. Nat. Commun. 7, 10583 (2016).
Kon, N. et al. CaMKII is essential for the cellular clock and coupling between morning and evening behavioral rhythms. Genes Dev. 28, 1101–1110 (2014).
Wilcox, A. G., Vizor, L., Parsons, M. J., Banks, G. & Nolan, P. M. Inducible knockout of mouse Zfhx3 emphasizes its key role in setting the pace and amplitude of the adult circadian clock. J. Biol. Rhythms. 32, 433–443 (2017).
Bedont, J. L. et al. Lhx1 controls terminal differentiation and circadian function of the suprachiasmatic nucleus. Cell Rep. 7, 609–622 (2014).
Hatori, M. et al. Lhx1 maintains synchrony among circadian oscillator neurons of the SCN. eLife 3, e03357 (2014).
Bedont, J. L. et al. An LHX1-regulated transcriptional network controls sleep/wake coupling and thermal resistance of the central circadian clockworks. Curr. Biol. 27, 128–136 (2017).
Carmona-Alcocer, V. et al. Ontogeny of circadian rhythms and synchrony in the suprachiasmatic nucleus. J. Neurosci. 38, 1326–1334 (2017).
Ono, D., Honma, S. & Honma, K. Cryptochromes are critical for the development of coherent circadian rhythms in the mouse suprachiasmatic nucleus. Nat. Commun. 4, 1666 (2013).
Gu, C. & Yang, H. Differences in intrinsic amplitudes of neuronal oscillators improve synchronization in the suprachiasmatic nucleus. Chaos 27, 093108 (2017).
Enoki, R. et al. Synchronous circadian voltage rhythms with asynchronous calcium rhythms in the suprachiasmatic nucleus. Proc. Natl Acad. Sci. USA 114, E2476–E2485 (2017).
Pauls, S. et al. Differential contributions of intra-cellular and inter-cellular mechanisms to the spatial and temporal architecture of the suprachiasmatic nucleus circadian circuitry in wild-type, cryptochrome-null and vasoactive intestinal peptide receptor 2-null mutant mice. Eur. J. Neurosci. 40, 2528–2540 (2014).
Edwards, M. D., Brancaccio, M., Chesham, J. E., Maywood, E. S. & Hastings, M. H. Rhythmic expression of cryptochrome induces the circadian clock of arrhythmic suprachiasmatic nuclei through arginine vasopressin signaling. Proc. Natl Acad. Sci. USA 113, 2732–2737 (2016).
Taylor, S. R., Wang, T. J., Granados-Fuentes, D. & Herzog, E. D. Resynchronization dynamics reveal that the ventral entrains the dorsal suprachiasmatic nucleus. J. Biol. Rhythms 32, 35–47 (2017).
Abel, J. H. et al. Functional network inference of the suprachiasmatic nucleus. Proc. Natl Acad. Sci. USA 113, 4512–4517 (2016). This article provides a thorough demonstration of the small-world properties of the SCN network.
Fan, J. et al. Vasoactive intestinal polypeptide (VIP)-expressing neurons in the suprachiasmatic nucleus provide sparse GABAergic outputs to local neurons with circadian regulation occurring distal to the opening of postsynaptic GABAA ionotropic receptors. J. Neurosci. 35, 1905–1920 (2015).
Bentley, B. et al. The multilayer connectome of Caenorhabditis elegans. PLoS Comput. Biol. 12, e1005283 (2016). This study is an insightful, topologically based analysis of the neural connectome of Caenorhabditis elegans across multiple levels of organization, including synaptic, aminergic and neuropeptidergic levels.
Barabasi, A. L. Network science. Philos. Trans. A Math. Phys. Eng. Sci. 371, 20120375 (2013).
Low-Zeddies, S. S. & Takahashi, J. S. Chimera analysis of the clock mutation in mice shows that complex cellular integration determines circadian behavior. Cell 105, 25–42 (2001).
Lee, I. T. et al. Neuromedin s-producing neurons act as essential pacemakers in the suprachiasmatic nucleus to couple clock neurons and dictate circadian rhythms. Neuron 85, 1086–1102 (2015). This study is a tour de force of intersectional genetic approaches to interrogate the pacemaking properties of the NMS-positive cells of the SCN.
Mieda, M. et al. Cellular clocks in AVP NEURONS of the SCN are critical for interneuronal coupling regulating circadian behavior rhythm. Neuron 85, 1103–1116 (2015).
Mieda, M., Okamoto, H. & Sakurai, T. Manipulating the cellular circadian period of arginine vasopressin neurons alters the behavioral circadian period. Curr. Biol. 26, 2535–2542 (2016).
Meng, Q. J. et al. Entrainment of disrupted circadian behavior through inhibition of casein kinase 1 (CK1) enzymes. Proc. Natl Acad. Sci. USA 107, 15240–15245 (2010).
Lee, H. M. et al. The period of the circadian oscillator is primarily determined by the balance between casein kinase 1 and protein phosphatase 1. Proc. Natl Acad. Sci. USA 108, 16451–16456 (2011).
van Oosterhout, F. et al. Amplitude of the SCN clock enhanced by the behavioral activity rhythm. PLoS ONE 7, e39693 (2012).
Grippo, R. M., Purohit, A. M., Zhang, Q., Zweifel, L. S. & Guler, A. D. Direct midbrain dopamine input to the suprachiasmatic nucleus accelerates circadian entrainment. Curr. Biol. 27, 2465–2475 e3 (2017).
Meng, Q. J. et al. Setting clock speed in mammals: the CK1 epsilon tau mutation in mice accelerates circadian pacemakers by selectively destabilizing PERIOD proteins. Neuron 58, 78–88 (2008).
Prosser, R. A., Edgar, D. M., Heller, H. C. & Miller, J. D. A possible glial role in the mammalian circadian clock. Brain Res. 643, 296–301 (1994).
Lavialle, M. & Serviere, J. Developmental study in the circadian clock of the golden hamster: a putative role of astrocytes. Brain Res. Dev. Brain Res. 86, 275–282 (1995).
Santos, J. W. Q. et al. Circadian variation in GFAP immunoreactivity in the mouse suprachiasmatic nucleus. Biol. Rhythm Res. 36, 141–150 (2005).
Prolo, L. M., Takahashi, J. S. & Herzog, E. D. Circadian rhythm generation and entrainment in astrocytes. J. Neurosci. 25, 404–408 (2005).
Barca-Mayo, O. et al. Astrocyte deletion of Bmal1 alters daily locomotor activity and cognitive functions via GABA signalling. Nat. Commun. 8, 14336 (2017).
Tso, C. F. et al. Astrocytes regulate daily rhythms in the suprachiasmatic nucleus and behavior. Curr. Biol. 27, 1055–1061 (2017).
Moldavan, M. et al. Localization and expression of GABA transporters in the suprachiasmatic nucleus. Eur. J. Neurosci. 42, 3018–3032 (2015).
Moldavan, M., Cravetchi, O. & Allen, C. N. GABA transporters regulate tonic and synaptic GABAA receptor-mediated currents in the suprachiasmatic nucleus neurons. J. Neurophysiol. 118, 3092–3106 (2017).
Chai, H. et al. Neural circuit-specialized astrocytes: transcriptomic, proteomic, morphological, and functional evidence. Neuron 95, 531–549 e9 (2017).
Clasadonte, J. & Prevot, V. The special relationship: glia-neuron interactions in the neuroendocrine hypothalamus. Nat. Rev. Endocrinol. 14, 25–44 (2018).
Duhart, J. M. et al. Suprachiasmatic astrocytes modulate the circadian clock in response to TNF-alpha. J. Immunol. 191, 4656–4664 (2013).
Marpegan, L. et al. Circadian regulation of ATP release in astrocytes. J. Neurosci. 31, 8342–8350 (2011).
Burkeen, J. F., Womac, A. D., Earnest, D. J. & Zoran, M. J. Mitochondrial calcium signaling mediates rhythmic extracellular ATP accumulation in suprachiasmatic nucleus astrocytes. J. Neurosci. 31, 8432–8440 (2011).
Schwarz, Y., Zhao, N., Kirchhoff, F. & Bruns, D. Astrocytes control synaptic strength by two distinct v-SNARE-dependent release pathways. Nat. Neurosci. 20, 1529–1539 (2017).
Shinohara, K., Honma, S., Katsuno, Y., Abe, H. & Honma, K. Circadian release of amino acids in the suprachiasmatic nucleus in vitro. Neuroreport 9, 137–140 (1998).
Hastings, M. H., Roberts, A. C. & Herbert, J. Neurotoxic lesions of the anterior hypothalamus disrupt the photoperiodic but not the circadian system of the Syrian hamster. Neuroendocrinology 40, 316–324 (1985).
Marpegan, L., Krall, T. J. & Herzog, E. D. Vasoactive intestinal polypeptide entrains circadian rhythms in astrocytes. J. Biol. Rhythms 24, 135–143 (2009).
Lavialle, M. et al. Structural plasticity of perisynaptic astrocyte processes involves ezrin and metabotropic glutamate receptors. Proc. Natl Acad. Sci. USA 108, 12915–12919 (2011).
Becquet, D., Girardet, C., Guillaumond, F., Francois-Bellan, A. M. & Bosler, O. Ultrastructural plasticity in the rat suprachiasmatic nucleus. Possible involvement in clock entrainment. Glia 56, 294–305 (2008).
Kutsuwada, T. et al. Molecular diversity of the NMDA receptor channel. Nature 358, 36–41 (1992).
Hu, Y. et al. GWAS of 89,283 individuals identifies genetic variants associated with self-reporting of being a morning person. Nat. Commun. 7, 10448 (2016).
Lane, J. M. et al. Genome-wide association analysis identifies novel loci for chronotype in 100,420 individuals from the UK Biobank. Nat. Commun. 7, 10889 (2016).
van den Pol, A. N., Finkbeiner, S. M. & Cornell-Bell, A. H. Calcium excitability and oscillations in suprachiasmatic nucleus neurons and glia in vitro. J. Neurosci. 12, 2648–2664 (1992).
Poskanzer, K. E. & Yuste, R. Astrocytes regulate cortical state switching in vivo. Proc. Natl Acad. Sci. USA 113, E2675–E2684 (2016).
Papouin, T., Dunphy, J. M., Tolman, M., Dineley, K. T. & Haydon, P. G. Septal cholinergic neuromodulation tunes the astrocyte-dependent gating of hippocampal NMDA receptors to wakefulness. Neuron 94, 840–854 e7 (2017).
Clasadonte, J., Scemes, E., Wang, Z., Boison, D. & Haydon, P. G. Connexin 43-mediated astroglial metabolic networks contribute to the regulation of the sleep-wake cycle. Neuron 95, 1365–1380 e5 (2017).
Absinta, M. et al. Human and nonhuman primate meninges harbor lymphatic vessels that can be visualized noninvasively by MRI. eLife 6, e29738 (2017).
Iliff, J. J. et al. A paravascular pathway facilitates CSF flow through the brain parenchyma and the clearance of interstitial solutes, including amyloid beta. Sci. Transl. Med. 4, 147ra111 (2012).
Dijk, D. J. et al. Amplitude reduction and phase shifts of melatonin, cortisol and other circadian rhythms after a gradual advance of sleep and light exposure in humans. PLoS ONE 7, e30037 (2012).
Morris, C. J. et al. Endogenous circadian system and circadian misalignment impact glucose tolerance via separate mechanisms in humans. Proc. Natl Acad. Sci. USA 112, E2225–E2234 (2015).
Voogel, A. J., Koopman, M. G., Hart, A. A., van Montfrans, G. A. & Arisz, L. Circadian rhythms in systemic hemodynamics and renal function in healthy subjects and patients with nephrotic syndrome. Kidney Int. 59, 1873–1880 (2001).
Acknowledgements
The authors are grateful to A. Patton, N. Smyllie and W. Schafer (MRC Laboratory of Molceular Biology (LMB)) and C. Partch (University of California, Santa Cruz) for very helpful discussions on the text and to P. Margiotta (MRC LMB) for graphic support. The work of the authors is supported by MRC funding to M.H.H. (MC_U105170643).
Author information
Authors and Affiliations
Contributions
The authors all researched data for the article, provided a substantial contribution to discussion of its content, wrote the article and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41583_2018_26_MOESM1_ESM.mov
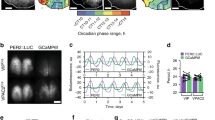
Supplementary information (video): Representative combined time-lapse recording of PER2:; Luciferase-reported TTFL bioluminescence rhythm (cyan) and GCaMP-reported [Ca2+]i rhythms (green) in SCN organotypic slice. Note the phase difference between the two reporters and the phase dispersion between cells leading to regular progressive waves across the SCN circuit. Reproduced with permission from REF. 56, Elsevier.
Glossary
- Emergent properties
-
Properties expressed at a circuit level that do not occur in isolated cells (for example, synchrony of cellular circadian gene expression, ensemble period and phase divergence).
- Amplitude
-
The range from peak level to trough level of a circadian rhythm (for example, the range for gene expression or intensity of behavioural activity).
- Ensemble period
-
The circadian period of oscillation shared by all SCN cells in the intact, synchronized circuit.
- Synchrony
-
The temporal order of individual oscillators expressing a common period and stable phase relationships.
- Phase dispersion
-
The range of phases present in a population of oscillators (for example, between rhythmic cells in an intact SCN circuit).
- Circadian time
-
(CT). Internal temporal scale of a tissue or an individual expressing a free-running circadian cycle, with predicted dawn denoted as CT0 and the period of the entire cycle divided into 24 circadian hours.
- Entrainment
-
A process whereby a rhythmic physical or biological cue sets the period and the phase of a circadian oscillation.
- Small-world network
-
A naturally occurring network topology in which most nodes are directly connected to only a few neighbours (which facilitates local information processing as modules) with a few richly connected nodes (hubs) that facilitate integration between nodes and modules.
- Hubs
-
Nodes in a network with higher-than-average connections.
- Nodes
-
Individual elements in a network.
- Cell-autonomous clock
-
The mechanism within most mammalian cells that enables them to express, independent of external cues, circadian cycles of gene expression and cellular metabolism.
- Cre
-
Cre-recombinase enzyme that catalyses the site-specific recombination of DNA between loxP sites in a target gene.
- Rich club
-
A restricted group of interconnected hubs in a network.
Rights and permissions
About this article
Cite this article
Hastings, M.H., Maywood, E.S. & Brancaccio, M. Generation of circadian rhythms in the suprachiasmatic nucleus. Nat Rev Neurosci 19, 453–469 (2018). https://doi.org/10.1038/s41583-018-0026-z
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41583-018-0026-z
This article is cited by
-
Single-cell transcriptomics reveals that glial cells integrate homeostatic and circadian processes to drive sleep–wake cycles
Nature Neuroscience (2024)
-
Sleep and circadian rhythmicity as entangled processes serving homeostasis
Nature Reviews Neuroscience (2024)
-
Circadian regulation of cancer stem cells and the tumor microenvironment during metastasis
Nature Cancer (2024)
-
System-level time computation and representation in the suprachiasmatic nucleus revealed by large-scale calcium imaging and machine learning
Cell Research (2024)
-
Chronic phase advances reduces recognition memory and increases vascular cognitive dementia-like impairments in aged mice
Scientific Reports (2024)