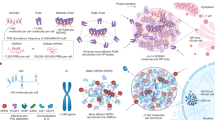
Abstract
Long non-coding RNAs (lncRNAs) outnumber protein-coding transcripts, but their functions remain largely unknown. In this Review, we discuss the emerging roles of lncRNAs in the control of gene transcription. Some of the best characterized lncRNAs have essential transcription cis-regulatory functions that cannot be easily accomplished by DNA-interacting transcription factors, such as XIST, which controls X-chromosome inactivation, or imprinted lncRNAs that direct allele-specific repression. A growing number of lncRNA transcription units, including CHASERR, PVT1 and HASTER (also known as HNF1A-AS1) act as transcription-stabilizing elements that fine-tune the activity of dosage-sensitive genes that encode transcription factors. Genetic experiments have shown that defects in such transcription stabilizers often cause severe phenotypes. Other lncRNAs, such as lincRNA-p21 (also known as Trp53cor1) and Maenli (Gm29348) contribute to local activation of gene transcription, whereas distinct lncRNAs influence gene transcription in trans. We discuss findings of lncRNAs that elicit a function through either activation of their transcription, transcript elongation and processing or the lncRNA molecule itself. We also discuss emerging evidence of lncRNA involvement in human diseases, and their potential as therapeutic targets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lipshitz, H. D., Peattie, D. A. & Hogness, D. S. Novel transcripts from the ultrabithorax domain of the bithorax complex. Genes. Dev. 1, 307–322 (1987).
Cumberledge, S., Zaratzian, A. & Sakonju, S. Characterization of two RNAs transcribed from the cis-regulatory region of the abd-A domain within the Drosophila bithorax complex. Proc. Natl Acad. Sci. USA 87, 3259–3263 (1990).
Mattick, J. S. et al. Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat. Rev. Mol. Cell Biol. 24, 430–447 (2023).
Anderson, D. M. et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 160, 595–606 (2015).
Morgado-Palacin, L. et al. The TINCR ubiquitin-like microprotein is a tumor suppressor in squamous cell carcinoma. Nat. Commun. 14, 1328 (2023).
Frankish, A. et al. GENCODE: reference annotation for the human and mouse genomes in 2023. Nucleic Acids Res. 51, D942–D949 (2023).
Gil, N. & Ulitsky, I. Regulation of gene expression by cis-acting long non-coding RNAs. Nat. Rev. Genet. 21, 102–117 (2020).
Rinn, J. L. & Chang, H. Y. Long noncoding RNAs: molecular modalities to organismal functions. Annu. Rev. Biochem. 89, 283–308 (2020).
Moran, I. et al. Human β-cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab. 16, 435–448 (2012).
Cabili, M. N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes. Dev. 25, 1915–1927 (2011).
Hon, C. C. et al. An atlas of human long non-coding RNAs with accurate 5′ ends. Nature 543, 199–204 (2017).
Akerman, I. et al. Human pancreatic β cell lncRNAs control cell-specific regulatory networks. Cell Metab. 25, 400–411 (2017).
Dimitrova, N. et al. LincRNA-p21 activates p21 in cis to promote Polycomb target gene expression and to enforce the G1/S checkpoint. Mol. Cell 54, 777–790 (2014).
Olivero, C. E. et al. p53 activates the long noncoding RNA Pvt1b to inhibit myc and suppress tumorigenesis. Mol. Cell 77, 761–774.e8 (2020).This study identifies a p53-induced isoform of the lncRNA Pvt1, which acts in cis to suppress Myc transcription in response to stress and thus limits cellular proliferation.
Winkler, L. et al. Functional elements of the cis-regulatory lincRNA-p21. Cell Rep. 39, 110687 (2022).Systematic genetic dissection of the lincRNA-p21 locus reveals that transcription initiation of lincRNA-p21 is sufficient for stimulation of p21 expression in cis.
Gil, N. et al. Complex regulation of Eomes levels mediated through distinct functional features of the Meteor long non-coding RNA locus. Cell Rep. 42, 112569 (2023).
Allou, L. et al. Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator. Nature 592, 93–98 (2021).Genetically engineered mouse models reveal that transcript elongation of the lncRNA Maenli promotes En1 expression and supports limb development. This study demonstrates that a human developmental limb disorder is likely caused by a monogenic lncRNA defect.
Elling, R. et al. Genetic models reveal cis and trans immune-regulatory activities for lincRNA-Cox2. Cell Rep. 25, 1511–1524.e6 (2018).
Isoda, T. et al. Non-coding transcription instructs chromatin folding and compartmentalization to dictate enhancer–promoter communication and T cell fate. Cell 171, 103–119.e18 (2017).
Perry, R. B., Hezroni, H., Goldrich, M. J. & Ulitsky, I. Regulation of neuroregeneration by long noncoding RNAs. Mol. Cell 72, 553–567.e5 (2018).
De Santa, F. et al. A large fraction of extragenic RNA Pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Paralkar, V. R. et al. Lineage and species-specific long noncoding RNAs during erythro-megakaryocytic development. Blood 123, 1927–1937 (2014).
Paralkar, V. R. et al. Unlinking an lncRNA from its associated cis element. Mol. Cell 62, 104–110 (2016).
Espinosa, J. M. Revisiting lncRNAs: how do you know yours is not an eRNA? Mol. Cell 62, 1–2 (2016).
Wang, K. C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Pradeepa, M. M. et al. Psip1/p52 regulates posterior Hoxa genes through activation of lncRNA Hottip. PLoS Genet. 13, e1006677 (2017).
Unfried, J. P. & Ulitsky, I. Substoichiometric action of long noncoding RNAs. Nat. Cell Biol. 24, 608–615 (2022).
Henninger, J. E. et al. RNA-mediated feedback control of transcriptional condensates. Cell 184, 207–225.e24 (2021).
Sharp, P. A., Chakraborty, A. K., Henninger, J. E. & Young, R. A. RNA in formation and regulation of transcriptional condensates. RNA 28, 52–57 (2022).
Oksuz, O. et al. Transcription factors interact with RNA to regulate genes. Mol. Cell 83, 2449–2463.e13 (2023).
Ntini, E. et al. Long ncRNA A-ROD activates its target gene DKK1 at its release from chromatin. Nat. Commun. 9, 1636 (2018).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Groff, A. F. et al. In vivo characterization of Linc-p21 reveals functional cis-regulatory DNA elements. Cell Rep. 16, 2178–2186 (2016).
Furuhata, R. et al. LincRNA-p21 exon 1 expression correlates with Cdkn1a expression in vivo. Genes. Cell 27, 14–24 (2022).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Gil, N. & Ulitsky, I. Production of spliced long noncoding RNAs specifies regions with increased enhancer activity. Cell Syst. 7, 537–547.e3 (2018).
Haerty, W. & Ponting, C. P. Unexpected selection to retain high GC content and splicing enhancers within exons of multiexonic lncRNA loci. RNA 21, 333–346 (2015).
Schuler, A., Ghanbarian, A. T. & Hurst, L. D. Purifying selection on splice-related motifs, not expression level nor RNA folding, explains nearly all constraint on human lincRNAs. Mol. Biol. Evol. 31, 3164–3183 (2014).
Tan, J. Y. & Marques, A. C. The activity of human enhancers is modulated by the splicing of their associated lncRNAs. PLoS Comput. Biol. 18, e1009722 (2022).
Canzio, D. et al. Antisense lncRNA transcription mediates DNA demethylation to drive stochastic protocadherin α promoter choice. Cell 177, 639–653.e15 (2019).
Heinz, S. et al. Transcription elongation can affect genome 3D structure. Cell 174, 1522–1536.e22 (2018).
Zhang, S., Ubelmesser, N., Barbieri, M. & Papantonis, A. Enhancer–promoter contact formation requires RNAPII and antagonizes loop extrusion. Nat. Genet. 55, 832–840 (2023).
Banigan, E. J. et al. Transcription shapes 3D chromatin organization by interacting with loop extrusion. Proc. Natl Acad. Sci. USA 120, e2210480120 (2023).
Lai, F. et al. Activating RNAs associate with mediator to enhance chromatin architecture and transcription. Nature 494, 497–501 (2013).
Li, W. et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature 498, 516–520 (2013).
Melo, C. A. et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell 49, 524–535 (2013).
Orom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Kalantari, R., Chiang, C. M. & Corey, D. R. Regulation of mammalian transcription and splicing by nuclear RNAi. Nucleic Acids Res. 44, 524–537 (2016).
Khvorova, A. Modulation of DNA transcription: the future of ASO therapeutics? Cell 185, 2011–2013 (2022).
Marasco, L. E. et al. Counteracting chromatin effects of a splicing-correcting antisense oligonucleotide improves its therapeutic efficacy in spinal muscular atrophy. Cell 185, 2057–2070.e15 (2022).
Lee, J. H. et al. Enhancer RNA m6A methylation facilitates transcriptional condensate formation and gene activation. Mol. Cell 81, 3368–3385.e9 (2021).
Rahnamoun, H. et al. RNAs interact with BRD4 to promote enhanced chromatin engagement and transcription activation. Nat. Struct. Mol. Biol. 25, 687–697 (2018).
Liang, L. et al. Complementary Alu sequences mediate enhancer–promoter selectivity. Nature 619, 868–875 (2023).
Barshad, G. et al. RNA polymerase II dynamics shape enhancer–promoter interactions. Nat. Genet. 55, 1370–1380 (2023).
Dao, L. T. M. et al. Genome-wide characterization of mammalian promoters with distal enhancer functions. Nat. Genet. 49, 1073–1081 (2017).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Ulitsky, I., Shkumatava, A., Jan, C. H., Sive, H. & Bartel, D. P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Liu, N. et al. Direct promoter repression by BCL11A controls the fetal to adult hemoglobin switch. Cell 173, 430–442.e17 (2018).
Pang, B., van Weerd, J. H., Hamoen, F. L. & Snyder, M. P. Identification of non-coding silencer elements and their regulation of gene expression. Nat. Rev. Mol. Cell Biol. 24, 383–395 (2023).
Maamar, H., Cabili, M. N., Rinn, J. & Raj, A. linc-HOXA1 is a noncoding RNA that represses Hoxa1 transcription in cis. Genes. Dev. 27, 1260–1271 (2013).
Su, G. et al. Enhancer architecture-dependent multilayered transcriptional regulation orchestrates RA signaling-induced early lineage differentiation of ESCs. Nucleic Acids Res. 49, 11575–11595 (2021).
Yin, Y. et al. Opposing roles for the lncRNA haunt and its genomic locus in regulating HOXA gene activation during embryonic stem cell differentiation. Cell Stem Cell 16, 504–516 (2015).
Anderson, K. M. et al. Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature 539, 433–436 (2016).
Han, X. et al. The lncRNA Hand2os1/Uph locus orchestrates heart development through regulation of precise expression of Hand2. Development 146, dev176198 (2019).
Ritter, N. et al. The lncRNA locus handsdown regulates cardiac gene programs and is essential for early mouse development. Dev. Cell 50, 644–657.e8 (2019).
Grote, P. et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 24, 206–214 (2013).
Zemmour, D., Pratama, A., Loughhead, S. M., Mathis, D. & Benoist, C. Flicr, a long noncoding RNA, modulates Foxp3 expression and autoimmunity. Proc. Natl Acad. Sci. USA 114, E3472–E3480 (2017).
Beucher, A. et al. The HASTER lncRNA promoter is a cis-acting transcriptional stabilizer of HNF1A. Nat. Cell Biol. 24, 1528–1540 (2022).This study shows that the promoter of the lncRNA HASTER ensures that levels of the transcripion factor HNF1A are maintained within a narrow homeostatic range. Haster deficiency causes abnormal HNF1A genomic occupancy and diabetes in mice.
Rom, A. et al. Regulation of CHD2 expression by the Chaserr long noncoding RNA gene is essential for viability. Nat. Commun. 10, 5092 (2019).
Bond, A. M. et al. Balanced gene regulation by an embryonic brain ncRNA is critical for adult hippocampal GABA circuitry. Nat. Neurosci. 12, 1020–1027 (2009).
Amandio, A. R., Necsulea, A., Joye, E., Mascrez, B. & Duboule, D. Hotair is dispensible for mouse development. PLoS Genet. 12, e1006232 (2016).
Cho, S. W. et al. Promoter of lncRNA gene PVT1 is a tumor-suppressor DNA boundary element. Cell 173, 1398–1412.e22 (2018).
Chen, F. L. et al. The long noncoding RNA Playrr regulates Pitx2 dosage and protects against cardiac arrhythmias. Preprint at bioRxiv https://doi.org/10.1101/2022.09.20.508562 (2022).
Szafranski, P., Gambin, T., Karolak, J. A., Popek, E. & Stankiewicz, P. Lung-specific distant enhancer cis regulates expression of FOXF1 and lncRNA FENDRR. Hum. Mutat. 42, 694–698 (2021).
Ali, T. & Grote, P. Beyond the RNA-dependent function of lncRNA genes. eLife 9, e60583 (2020).
Ghildiyal, R. et al. Loss of long noncoding RNA NXTAR in prostate cancer augments androgen receptor expression and enzalutamide resistance. Cancer Res. 82, 155–168 (2022).
Kribelbauer, J. F., Rastogi, C., Bussemaker, H. J. & Mann, R. S. Low-affinity binding sites and the transcription factor specificity paradox in eukaryotes. Annu. Rev. Cell Dev. Biol. 35, 357–379 (2019).
Golson, M. L. & Kaestner, K. H. Fox transcription factors: from development to disease. Development 143, 4558–4570 (2016).
Servitja, J. M. et al. Hnf1α (MODY3) controls tissue-specific transcriptional programs and exerts opposed effects on cell growth in pancreatic islets and liver. Mol. Cell Biol. 29, 2945–2959 (2009).
Fernandez Garcia, M. et al. Structural features of transcription factors associating with nucleosome binding. Mol. Cell 75, 921–932.e6 (2019).
Huang, P. et al. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 475, 386–389 (2011).
Ali, T. et al. Fendrr synergizes with Wnt signalling to regulate fibrosis related genes during lung development via its RNA:dsDNA triplex element. Nucleic Acids Res. 51, 6227–6237 (2023).
Colombo, T., Farina, L., Macino, G. & Paci, P. PVT1: a rising star among oncogenic long noncoding RNAs. Biomed. Res. Int. 2015, 304208 (2015).
Tesfaye, E. et al. The p53 transcriptional response across tumor types reveals core and senescence-specific signatures modulated by long noncoding RNAs. Proc. Natl Acad. Sci. USA 118, e2025539118 (2021).
Kotzin, J. J. et al. The long noncoding RNA morrbid regulates CD8 T cells in response to viral infection. Proc. Natl Acad. Sci. USA 116, 11916–11925 (2019).
Kotzin, J. J. et al. The long non-coding RNA morrbid regulates Bim and short-lived myeloid cell lifespan. Nature 537, 239–243 (2016).
Zhao, Y. et al. Natural temperature fluctuations promote COOLAIR regulation of FLC. Genes. Dev. 35, 888–898 (2021).
Jegu, T., Aeby, E. & Lee, J. T. The X chromosome in space. Nat. Rev. Genet. 18, 377–389 (2017).
Deng, X., Berletch, J. B., Nguyen, D. K. & Disteche, C. M. X chromosome regulation: diverse patterns in development, tissues and disease. Nat. Rev. Genet. 15, 367–378 (2014).
Brockdorff, N., Bowness, J. S. & Wei, G. Progress toward understanding chromosome silencing by Xist RNA. Genes. Dev. 34, 733–744 (2020).
Augui, S., Nora, E. P. & Heard, E. Regulation of X-chromosome inactivation by the X-inactivation centre. Nat. Rev. Genet. 12, 429–442 (2011).
Galupa, R. & Heard, E. X-chromosome inactivation: a crossroads between chromosome architecture and gene regulation. Annu. Rev. Genet. 52, 535–566 (2018).
Furlan, G. & Galupa, R. Mechanisms of choice in X-chromosome inactivation. Cells 11, 535 (2022).
Mutzel, V. & Schulz, E. G. Dosage sensing, threshold responses, and epigenetic memory: a systems biology perspective on random X-chromosome inactivation. Bioessays 42, e1900163 (2020).
Jacobson, E. C., Pandya-Jones, A. & Plath, K. A lifelong duty: how Xist maintains the inactive X chromosome. Curr. Opin. Genet. Dev. 75, 101927 (2022).
van Bemmel, J. G. et al. The bipartite TAD organization of the X-inactivation center ensures opposing developmental regulation of Tsix and Xist. Nat. Genet. 51, 1024–1034 (2019).
Gjaltema, R. A. F. et al. Distal and proximal cis-regulatory elements sense X chromosome dosage and developmental state at the Xist locus. Mol. Cell 82, 190–208.e17 (2022).
Rosspopoff, O. et al. Species-specific regulation of XIST by the JPX/FTX orthologs. Nucleic Acids Res. 51, 2177–2194 (2023).Functional similarities and differences between the human and mouse lncRNAs orthologues JPX and Jpx and FTX and Ftx highlight the complementary roles of lncRNA transcription and the mature lncRNAs in XCI.
Quesada-Espinosa, J. F. et al. First female with Allan–Herndon–Dudley syndrome and partial deletion of X-inactivation center. Neurogenetics 22, 343–346 (2021).
Sun, S. et al. Jpx RNA activates Xist by evicting CTCF. Cell 153, 1537–1551 (2013).
Loda, A., Collombet, S. & Heard, E. Gene regulation in time and space during X-chromosome inactivation. Nat. Rev. Mol. Cell Biol. 23, 231–249 (2022).
Ogawa, Y. & Lee, J. T. Xite, X-inactivation intergenic transcription elements that regulate the probability of choice. Mol. Cell 11, 731–743 (2003).
Galupa, R. et al. A conserved noncoding locus regulates random monoallelic xist expression across a topological boundary. Mol. Cell 77, 352–367.e8 (2020).
Lee, J. T. & Lu, N. Targeted mutagenesis of Tsix leads to nonrandom X inactivation. Cell 99, 47–57 (1999).
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Lee, J. T., Strauss, W. M., Dausman, J. A. & Jaenisch, R. A 450 kb transgene displays properties of the mammalian X-inactivation center. Cell 86, 83–94 (1996).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015).
Markaki, Y. et al. Xist nucleates local protein gradients to propagate silencing across the X chromosome. Cell 184, 6212 (2021).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Minajigi, A. et al. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015).
Raposo, A. C., Casanova, M., Gendrel, A. V. & da Rocha, S. T. The tandem repeat modules of Xist lncRNA: a Swiss army knife for the control of X-chromosome inactivation. Biochem. Soc. Trans. 49, 2549–2560 (2021).
Carter, A. C. et al. Spen links RNA-mediated endogenous retrovirus silencing and X chromosome inactivation. eLife 9, e54508 (2020).
Dossin, F. et al. SPEN integrates transcriptional and epigenetic control of X-inactivation. Nature 578, 455–460 (2020).
Jachowicz, J. W. et al. Xist spatially amplifies SHARP/SPEN recruitment to balance chromosome-wide silencing and specificity to the X chromosome. Nat. Struct. Mol. Biol. 29, 239–249 (2022).
Zylicz, J. J. et al. The implication of early chromatin changes in X chromosome inactivation. Cell 176, 182–197.e23 (2019).
Bousard, A. et al. The role of Xist-mediated Polycomb recruitment in the initiation of X-chromosome inactivation. EMBO Rep. 20, e48019 (2019).
Jansz, N. et al. Smchd1 targeting to the inactive X is dependent on the Xist–HnrnpK–PRC1 pathway. Cell Rep. 25, 1912–1923.e9 (2018).
Pintacuda, G. et al. hnRNPK recruits PCGF3/5–PRC1 to the Xist RNA B-repeat to establish polycomb-mediated chromosomal silencing. Mol. Cell 68, 955–969.e10 (2017).
Wang, C. Y., Colognori, D., Sunwoo, H., Wang, D. & Lee, J. T. PRC1 collaborates with SMCHD1 to fold the X-chromosome and spread Xist RNA between chromosome compartments. Nat. Commun. 10, 2950 (2019).
Wang, C. Y., Jegu, T., Chu, H. P., Oh, H. J. & Lee, J. T. SMCHD1 merges chromosome compartments and assists formation of super-structures on the inactive X. Cell 174, 406–421.e25 (2018).
Colognori, D., Sunwoo, H., Wang, D., Wang, C. Y. & Lee, J. T. Xist repeats A and B account for two distinct phases of X inactivation establishment. Dev. Cell 54, 21–32.e5 (2020).
Pandya-Jones, A. et al. A protein assembly mediates Xist localization and gene silencing. Nature 587, 145–151 (2020).Assembly of multiple RNA-binding proteins on Xist E-repeats promotes homotypic and heterotypic interactions that result in the formation of a condensate, which is essential for gene silencing. Once formed, this condensate can sustain XCI in absence of Xist.
Sunwoo, H., Colognori, D., Froberg, J. E., Jeon, Y. & Lee, J. T. Repeat E anchors Xist RNA to the inactive X chromosomal compartment through CDKN1A-interacting protein (CIZ1). Proc. Natl Acad. Sci. USA 114, 10654–10659 (2017).
Strehle, M. & Guttman, M. Xist drives spatial compartmentalization of DNA and protein to orchestrate initiation and maintenance of X inactivation. Curr. Opin. Cell Biol. 64, 139–147 (2020).
de Napoles, M. et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev. Cell 7, 663–676 (2004).
Sun, B. K., Deaton, A. M. & Lee, J. T. A transient heterochromatic state in Xist preempts X inactivation choice without RNA stabilization. Mol. Cell 21, 617–628 (2006).
Zhao, J., Sun, B. K., Erwin, J. A., Song, J. J. & Lee, J. T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Simon, M. D. et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 504, 465–469 (2013).
Tucci, V., Isles, A. R., Kelsey, G., Ferguson-Smith, A. C. & Erice Imprinting, G. Genomic imprinting and physiological processes in mammals. Cell 176, 952–965 (2019).
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harb. Perspect. Biol. 6, a018382 (2014).
Guenzl, P. M. & Barlow, D. P. Macro lncRNAs: a new layer of cis-regulatory information in the mammalian genome. RNA Biol. 9, 731–741 (2012).
Latos, P. A. & Barlow, D. P. Regulation of imprinted expression by macro non-coding RNAs. RNA Biol. 6, 100–106 (2009).
Kota, S. K. et al. ICR noncoding RNA expression controls imprinting and DNA replication at the Dlk1–Dio3 domain. Dev. Cell 31, 19–33 (2014).
Schertzer, M. D. et al. lncRNA-induced spread of Polycomb controlled by genome architecture, RNA abundance, and CpG island DNA. Mol. Cell 75, 523–537.e10 (2019).
Seidl, C. I., Stricker, S. H. & Barlow, D. P. The imprinted Air ncRNA is an atypical RNAPII transcript that evades splicing and escapes nuclear export. EMBO J. 25, 3565–3575 (2006).
Terranova, R. et al. Polycomb group proteins Ezh2 and Rnf2 direct genomic contraction and imprinted repression in early mouse embryos. Dev. Cell 15, 668–679 (2008).
Tibbit, C. J. et al. Antisense activity across the Nesp promoter is required for Nespas-mediated silencing in the imprinted Gnas cluster. Noncoding RNA 1, 246–265 (2015).
Quinodoz, S. A. et al. RNA promotes the formation of spatial compartments in the nucleus. Cell 184, 5775–5790.e30 (2021).
MacDonald, W. A. & Mann, M. R. W. Long noncoding RNA functionality in imprinted domain regulation. PLoS Genet. 16, e1008930 (2020).
Pauler, F. M., Koerner, M. V. & Barlow, D. P. Silencing by imprinted noncoding RNAs: is transcription the answer? Trends Genet. 23, 284–292 (2007).
Hao, N., Palmer, A. C., Dodd, I. B. & Shearwin, K. E. Directing traffic on DNA — how transcription factors relieve or induce transcriptional interference. Transcription 8, 120–125 (2017).
Andergassen, D. et al. The Airn lncRNA does not require any DNA elements within its locus to silence distant imprinted genes. PLoS Genet. 15, e1008268 (2019).
Latos, P. A. et al. Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science 338, 1469–1472 (2012).
Santoro, F. et al. Imprinted Igf2r silencing depends on continuous Airn lncRNA expression and is not restricted to a developmental window. Development 140, 1184–1195 (2013).
Sleutels, F., Zwart, R. & Barlow, D. P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002).
Golding, M. C. et al. Depletion of Kcnq1ot1 non-coding RNA does not affect imprinting maintenance in stem cells. Development 138, 3667–3678 (2011).
Mancini-Dinardo, D., Steele, S. J., Levorse, J. M., Ingram, R. S. & Tilghman, S. M. Elongation of the Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genes. Dev. 20, 1268–1282 (2006).
Meng, L., Person, R. E. & Beaudet, A. L. Ube3a-ATS is an atypical RNA polymerase II transcript that represses the paternal expression of Ube3a. Hum. Mol. Genet. 21, 3001–3012 (2012).
Meng, L. et al. Truncation of Ube3a-ATS unsilences paternal Ube3a and ameliorates behavioral defects in the Angelman syndrome mouse model. PLoS Genet. 9, e1004039 (2013).
Meng, L. et al. Towards a therapy for Angelman syndrome by targeting a long non-coding RNA. Nature 518, 409–412 (2015).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Lewis, A. et al. Epigenetic dynamics of the Kcnq1 imprinted domain in the early embryo. Development 133, 4203–4210 (2006).
Lewis, A. et al. Imprinting on distal chromosome 7 in the placenta involves repressive histone methylation independent of DNA methylation. Nat. Genet. 36, 1291–1295 (2004).
Mohammad, F., Mondal, T., Guseva, N., Pandey, G. K. & Kanduri, C. Kcnq1ot1 noncoding RNA mediates transcriptional gene silencing by interacting with Dnmt1. Development 137, 2493–2499 (2010).
Nagano, T. et al. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322, 1717–1720 (2008).
Pandey, R. R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008).
Redrup, L. et al. The long noncoding RNA Kcnq1ot1 organises a lineage-specific nuclear domain for epigenetic gene silencing. Development 136, 525–530 (2009).
Umlauf, D. et al. Imprinting along the Kcnq1 domain on mouse chromosome 7 involves repressive histone methylation and recruitment of Polycomb group complexes. Nat. Genet. 36, 1296–1300 (2004).
Wagschal, A. et al. G9a histone methyltransferase contributes to imprinting in the mouse placenta. Mol. Cell Biol. 28, 1104–1113 (2008).
Braceros, A. K. et al. Proximity-dependent recruitment of Polycomb repressive complexes by the lncRNA Airn. Cell Rep. 42, 112803 (2023).
Long, Y. et al. RNA is essential for PRC2 chromatin occupancy and function in human pluripotent stem cells. Nat. Genet. 52, 931–938 (2020).
Lleres, D. et al. CTCF modulates allele-specific sub-TAD organization and imprinted gene activity at the mouse Dlk1–Dio3 and Igf2–H19 domains. Genome Biol. 20, 272 (2019).
Hansen, A. S. et al. Distinct classes of chromatin loops revealed by deletion of an RNA-binding region in CTCF. Mol. Cell 76, 395–411.e13 (2019).
Saldana-Meyer, R. et al. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cell 76, 412–422.e5 (2019).
Kurukuti, S. et al. CTCF binding at the H19 imprinting control region mediates maternally inherited higher-order chromatin conformation to restrict enhancer access to Igf2. Proc. Natl Acad. Sci. USA 103, 10684–10689 (2006).
Murrell, A., Heeson, S. & Reik, W. Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nat. Genet. 36, 889–893 (2004).
Kopp, F. & Mendell, J. T. Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393–407 (2018).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Li, L. et al. Targeted disruption of Hotair leads to homeotic transformation and gene derepression. Cell Rep. 5, 3–12 (2013).
Davidovich, C. et al. Toward a consensus on the binding specificity and promiscuity of PRC2 for RNA. Mol. Cell 57, 552–558 (2015).
Schorderet, P. & Duboule, D. Structural and functional differences in the long non-coding RNA hotair in mouse and human. PLoS Genet. 7, e1002071 (2011).
Selleri, L. et al. A hox-embedded long noncoding RNA: is it all hot air? PLoS Genet. 12, e1006485 (2016).
Smith, K. P., Hall, L. L. & Lawrence, J. B. Nuclear hubs built on RNAs and clustered organization of the genome. Curr. Opin. Cell Biol. 64, 67–76 (2020).
Hutchinson, J. N. et al. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8, 39 (2007).
West, J. A. et al. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 55, 791–802 (2014).
Ballarino, M. et al. Deficiency in the nuclear long noncoding RNA charme causes myogenic defects and heart remodeling in mice. EMBO J. 37, e99697 (2018).
Desideri, F. et al. Intronic determinants coordinate charme lncRNA nuclear activity through the interaction with MATR3 and PTBP1. Cell Rep. 33, 108548 (2020).
Taliani, V. et al. The long noncoding RNA Charme supervises cardiomyocyte maturation by controlling cell differentiation programs in the developing heart. eLife 12, e81360 (2023).
Daneshvar, K. et al. lncRNA DIGIT and BRD3 protein form phase-separated condensates to regulate endoderm differentiation. Nat. Cell Biol. 22, 1211–1222 (2020).
Chang, K. C. et al. MaTAR25 lncRNA regulates the Tensin1 gene to impact breast cancer progression. Nat. Commun. 11, 6438 (2020).
Creamer, K. M., Kolpa, H. J. & Lawrence, J. B. Nascent RNA scaffolds contribute to chromosome territory architecture and counter chromatin compaction. Mol. Cell 81, 3509–3525.e5 (2021).
Mele, M. & Rinn, J. L. “Cat’s cradling” the 3D genome by the act of lncRNA transcription. Mol. Cell 62, 657–664 (2016).
Andergassen, D. et al. In vivo firre and Dxz4 deletion elucidates roles for autosomal gene regulation. eLife 8, e47214 (2019).
Hacisuleyman, E. et al. Topological organization of multichromosomal regions by the long intergenic noncoding RNA Firre. Nat. Struct. Mol. Biol. 21, 198–206 (2014).
Blank-Giwojna, A., Postepska-Igielska, A. & Grummt, I. lncRNA KHPS1 activates a poised enhancer by triplex-dependent recruitment of epigenomic regulators. Cell Rep. 26, 2904–2915.e4 (2019).
Postepska-Igielska, A. et al. lncRNA Khps1 regulates expression of the proto-oncogene SPHK1 via triplex-mediated changes in chromatin structure. Mol. Cell 60, 626–636 (2015).
O’Leary, V. B. et al. PARTICLE, a triplex-forming long ncRNA, regulates locus-specific methylation in response to low-dose irradiation. Cell Rep. 11, 474–485 (2015).
Grote, P. & Herrmann, B. G. The long non-coding RNA Fendrr links epigenetic control mechanisms to gene regulatory networks in mammalian embryogenesis. RNA Biol. 10, 1579–1585 (2013).
Kalwa, M. et al. The lncRNA HOTAIR impacts on mesenchymal stem cells via triple helix formation. Nucleic Acids Res. 44, 10631–10643 (2016).
Leisegang, M. S. et al. HIF1α-AS1 is a DNA:DNA:RNA triplex-forming lncRNA interacting with the HUSH complex. Nat. Commun. 13, 6563 (2022).
Trembinski, D. J. et al. Aging-regulated anti-apoptotic long non-coding RNA Sarrah augments recovery from acute myocardial infarction. Nat. Commun. 11, 2039 (2020).
Zhang, X. et al. KCNQ1OT1 promotes genome-wide transposon repression by guiding RNA-DNA triplexes and HP1 binding. Nat. Cell Biol. 24, 1617–1629 (2022).
Uroda, T. et al. Conserved pseudoknots in lncRNA MEG3 are essential for stimulation of the p53 pathway. Mol. Cell 75, 982–995.e9 (2019).
Mondal, T. et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA–DNA triplex structures. Nat. Commun. 6, 7743 (2015).
Ariel, F. et al. Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin-driven chromatin loop. Mol. Cell 55, 383–396 (2014).
Ariel, F. et al. R-loop mediated trans action of the APOLO long noncoding RNA. Mol. Cell 77, 1055–1065.e4 (2020).
Szafranski, P. et al. Small noncoding differentially methylated copy-number variants, including lncRNA genes, cause a lethal lung developmental disorder. Genome Res. 23, 23–33 (2013).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013).
van Dijk, M. et al. HELLP babies link a novel lincRNA to the trophoblast cell cycle. J. Clin. Invest. 122, 4003–4011 (2012).
Kvon, E. Z., Waymack, R., Gad, M. & Wunderlich, Z. Enhancer redundancy in development and disease. Nat. Rev. Genet. 22, 324–336 (2021).
Miguel-Escalada, I. et al. Pancreas agenesis mutations disrupt a lead enhancer controlling a developmental enhancer cluster. Dev. Cell 57, 1922–1936.e9 (2022).
Osterwalder, M. et al. Enhancer redundancy provides phenotypic robustness in mammalian development. Nature 554, 239–243 (2018).
Chenier, S. et al. CHD2 haploinsufficiency is associated with developmental delay, intellectual disability, epilepsy and neurobehavioural problems. J. Neurodev. Disord. 6, 9 (2014).
Cohen, A. S. A. et al. Haploinsufficiency of the basic helix–loop–helix transcription factor HAND2 causes congenital heart defects. Am. J. Med. Genet. A 182, 1263–1267 (2020).
Dirkx, E. et al. Nfat and miR-25 cooperate to reactivate the transcription factor Hand2 in heart failure. Nat. Cell Biol. 15, 1282–1293 (2013).
Tamura, M. et al. Overdosage of Hand2 causes limb and heart defects in the human chromosomal disorder partial trisomy distal 4q. Hum. Mol. Genet. 22, 2471–2481 (2013).
Yamagata, K. et al. Mutations in the hepatocyte nuclear factor-1α gene in maturity-onset diabetes of the young (MODY3). Nature 384, 455–458 (1996).
Luco, R. F. et al. A conditional model reveals that induction of hepatocyte nuclear factor-1α in Hnf1α-null mutant β-cells can activate silenced genes postnatally, whereas overexpression is deleterious. Diabetes 55, 2202–2211 (2006).
Gage, P. J., Suh, H. & Camper, S. A. Dosage requirement of Pitx2 for development of multiple organs. Development 126, 4643–4651 (1999).
Tumer, Z. & Bach-Holm, D. Axenfeld–Rieger syndrome and spectrum of PITX2 and FOXC1 mutations. Eur. J. Hum. Genet. 17, 1527–1539 (2009).
Cole, M. D. The myc oncogene: its role in transformation and differentiation. Annu. Rev. Genet. 20, 361–384 (1986).
George, M. R. et al. Minimal in vivo requirements for developmentally regulated cardiac long intergenic non-coding RNAs. Development 146, dev185314 (2019).
Atla, G. et al. Genetic regulation of RNA splicing in human pancreatic islets. Genome Biol. 23, 196 (2022).
Holdt, L. M. & Teupser, D. Long noncoding RNA ANRIL: lnc-ing genetic variation at the chromosome 9p21 locus to molecular mechanisms of atherosclerosis. Front. Cardiovasc. Med. 5, 145 (2018).
de Goede, O. M. et al. Population-scale tissue transcriptomics maps long non-coding RNAs to complex disease. Cell 184, 2633–2648.e19 (2021).Identification of numerous lncRNAs as candidate mediators of genetic association signals that underly susceptibility for prevalent human diseases.
Cory, S., Graham, M., Webb, E., Corcoran, L. & Adams, J. M. Variant (6;15) translocations in murine plasmacytomas involve a chromosome 15 locus at least 72 kb from the c-myc oncogene. EMBO J. 4, 675–681 (1985).
Graham, M., Adams, J. M. & Cory, S. Murine T lymphomas with retroviral inserts in the chromosomal 15 locus for plasmacytoma variant translocations. Nature 314, 740–743 (1985).
Hu, X. et al. A functional genomic approach identifies FAL1 as an oncogenic long noncoding RNA that associates with BMI1 and represses p21 expression in cancer. Cancer Cell 26, 344–357 (2014).
Leucci, E. et al. Melanoma addiction to the long non-coding RNA SAMMSON. Nature 531, 518–522 (2016).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304.e6 (2018).
Olivero, C. E. & Dimitrova, N. Identification and characterization of functional long noncoding RNAs in cancer. FASEB J. 34, 15360–15646 (2020).
Hilton, L. K. et al. The double-hit signature identifies double-hit diffuse large B-cell lymphoma with genetic events cryptic to FISH. Blood 134, 1528–1532 (2019).
Gutschner, T., Hammerle, M. & Diederichs, S. MALAT1 — a paradigm for long noncoding RNA function in cancer. J. Mol. Med. 91, 791–801 (2013).
Gupta, R. A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Martinez-Terroba, E. et al. Overexpressed malat1 drives metastasis through inflammatory reprogramming of lung adenocarcinoma microenvironment. Preprint at bioRxiv https://doi.org/10.1101/2023.03.20.533534 (2023).
Hibi, K. et al. Loss of H19 imprinting in esophageal cancer. Cancer Res. 56, 480–482 (1996).
Kondo, M. et al. Frequent loss of imprinting of the H19 gene is often associated with its overexpression in human lung cancers. Oncogene 10, 1193–1198 (1995).
Rainier, S. et al. Relaxation of imprinted genes in human cancer. Nature 362, 747–749 (1993).
Tseng, Y. Y. & Bagchi, A. The PVT1–MYC duet in cancer. Mol. Cell Oncol. 2, e974467 (2015).
Cai, Z. et al. Targeting bim via a lncRNA morrbid regulates the survival of preleukemic and leukemic cells. Cell Rep. 31, 107816 (2020).
Cai, Z. et al. Role of lncRNA Morrbid in PTPN11(Shp2)E76K-driven juvenile myelomonocytic leukemia. Blood Adv. 4, 3246–3251 (2020).
Huang, Y. et al. The role of lincRNA-p21 in regulating the biology of cancer cells. Hum. Cell 35, 1640–1649 (2022).
Borensztein, M. et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 24, 226–233 (2017).
Marahrens, Y., Panning, B., Dausman, J., Strauss, W. & Jaenisch, R. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes. Dev. 11, 156–166 (1997).
Takagi, N. & Abe, K. Detrimental effects of two active X chromosomes on early mouse development. Development 109, 189–201 (1990).
Yang, L., Yildirim, E., Kirby, J. E., Press, W. & Lee, J. T. Widespread organ tolerance to Xist loss and X reactivation except under chronic stress in the gut. Proc. Natl Acad. Sci. USA 117, 4262–4272 (2020).
Yildirim, E. et al. Xist RNA is a potent suppressor of hematologic cancer in mice. Cell 152, 727–742 (2013).
Syrett, C. M. et al. Loss of Xist RNA from the inactive X during B cell development is restored in a dynamic YY1-dependent two-step process in activated B cells. PLoS Genet. 13, e1007050 (2017).
Spaziano, A. & Cantone, I. X-chromosome reactivation: a concise review. Biochem. Soc. Trans. 49, 2797–2805 (2021).
Syrett, C. M. et al. Altered X-chromosome inactivation in T cells may promote sex-biased autoimmune diseases. JCI Insight 4, e12671 (2019).Findings in mouse and human suggesting that XIST dysregulation and abnormal X- chromosome inactivation in T cells underlie the increased prevalence of systemic lupus erythematosus in women.
Wang, J. et al. Unusual maintenance of X chromosome inactivation predisposes female lymphocytes for increased expression from the inactive X. Proc. Natl Acad. Sci. USA 113, E2029–E2038 (2016).
Yu, B. et al. B cell-specific XIST complex enforces X-inactivation and restrains atypical B cells. Cell 184, 1790–1803.e17 (2021).
Dou, D. R. et al. XIST ribonucleoproteins promote female sex-biased autoimmunity. Preprint at bioRxiv https://doi.org/10.1101/2022.11.05.515306 (2022).
Li, Y. et al. A noncoding RNA modulator potentiates phenylalanine metabolism in mice. Science 373, 662–673 (2021).
Dindot, S. V. et al. An ASO therapy for Angelman syndrome that targets an evolutionarily conserved region at the start of the UBE3A-AS transcript. Sci. Transl. Med. 15, eabf4077 (2023).
Wolter, J. M. et al. Cas9 gene therapy for Angelman syndrome traps Ube3a-ATS long non-coding RNA. Nature 587, 281–284 (2020).
Jiang, J. et al. Translating dosage compensation to trisomy 21. Nature 500, 296–300 (2013).
Abulwerdi, F. A. et al. Selective small-molecule targeting of a triple helix encoded by the long noncoding RNA, MALAT1. ACS Chem. Biol. 14, 223–235 (2019).
Donlic, A., Zafferani, M., Padroni, G., Puri, M. & Hargrove, A. E. Regulation of MALAT1 triple helix stability and in vitro degradation by diphenylfurans. Nucleic Acids Res. 48, 7653–7664 (2020).
Zafferani, M. et al. Multiassay profiling of a focused small molecule library reveals predictive bidirectional modulation of the lncRNA MALAT1 triplex stability in vitro. ACS Chem. Biol. 17, 2437–2447 (2022).
Aguilar, R. et al. Targeting Xist with compounds that disrupt RNA structure and X inactivation. Nature 604, 160–166 (2022).
Rosa, S., Duncan, S. & Dean, C. Mutually exclusive sense-antisense transcription at FLC facilitates environmentally induced gene repression. Nat. Commun. 7, 13031 (2016).
Li, P., Tao, Z. & Dean, C. Phenotypic evolution through variation in splicing of the noncoding RNA COOLAIR. Genes. Dev. 29, 696–701 (2015).
Wu, Z., Fang, X., Zhu, D. & Dean, C. Autonomous pathway: flowering locus C repression through an antisense-mediated chromatin-silencing mechanism. Plant. Physiol. 182, 27–37 (2020).
Yang, M. et al. In vivo single-molecule analysis reveals COOLAIR RNA structural diversity. Nature 609, 394–399 (2022).Temperature-dependent structural alterations in the lncRNA COOLAIR underly its role as a transcription repressive switch during seasonal transition in plants.
Sun, Q., Csorba, T., Skourti-Stathaki, K., Proudfoot, N. J. & Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340, 619–621 (2013).
Acknowledgements
The authors thank T. Graff, M. Cuenca-Ardura and B. Payer for critical reading of this manuscript. This work was supported by European Research Council (789055) and Spanish Ministry of Science and Innovation (PID2021-122522OB-I00) grants to J.F., and by National Institute of Health (R01CA262286 and R37CA230580) grants to N.D.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Molecular Cell Biology thanks Takayuki Nojima, Igor Ulitsky and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- CpG islands
-
Genomic regions of 500 nucleotides or longer with >50% CpG dinucleotide repeat content. CpG islands are associated with the transcription start sites of most housekeeping genes and as many as 40% of tissue-specific genes; they are bound by regulatory proteins.
- CTCF
-
A zinc-finger transcription factor (TF), also known as CCCTC-binding factor, that binds specific DNA sequences and participates in the formation of chromatin loops that influence gene transcription by defining the boundaries of topologically associated domains (TADs) and bringing enhancers into proximity with promoters.
- DNA–DNA–RNA triplex
-
A structure in which single-stranded RNA invades the major groove of double-stranded DNA and binds by forming Hoogsteen hydrogen bonds. DNA–DNA–lncRNA triplexes can be identified by pull-downs with a triplex-specific antibody.
- Enhancer RNAs
-
(eRNAs). Non-coding RNAs that are bidirectionally transcribed from enhancer regions, and are typically ≤500 nucleotides and unstable (half-life ≤ 2 min).
- Enhancers
-
Genomic regions that are recognized by transcription factors (TFs) and activate and increase the transcription of genes in cis, sometimes from considerable distances. Active enhancers are flanked by nucleosomes that carry post-translational histone modifications such as histone H3 acetylated at lysine 27 (H3K27ac) and H3 methylated at lysine 4 (H3K4me1).
- Expression quantitative trait loci
-
(eQTL). Genetic loci in which different alleles of a DNA variant influence expression levels of coding or non-coding transcripts.
- Focal deletions
-
Cancer-associated genomic deletions smaller than 5 Mb that affect both alleles.
- Genome-wide association studies
-
(GWAS). Studies that survey DNA variants genome-wide to identify those showing association with a disease or trait. GWAS have been used to discover susceptibility variants for prevalent polygenic diseases. A large fraction of significant associations are found in non-coding genomic regions, indicating that they are mediated by genetic variants that influence regulatory functions.
- lncRNAs that act in cis
-
Long non-coding RNAs (lncRNAs) that act on the same chromosome from which they are transcribed, including the regulation of a neighbouring gene, of multiple genes or of the entire chromosome.
- Silencers
-
Genomic regions that are bound by repressive transcription factors (TFs) and decrease the transcription of genes in cis.
- Splicing quantitative trait loci
-
Genetic loci in which different alleles influence RNA splicing patterns.
- Topologically associated domains
-
(TADs). Genomic regions defined by having a higher frequency of long-range chromatin contacts, such as between genes and their regulatory elements, than the frequency of contacts with elements outside the region.
- Transcriptional condensates
-
Chromatin-associated, dynamic nuclear assemblies comprising a heterogeneous mix of RNAs, transcription factors (TFs) and co-regulators that modulate transcriptional output.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ferrer, J., Dimitrova, N. Transcription regulation by long non-coding RNAs: mechanisms and disease relevance. Nat Rev Mol Cell Biol 25, 396–415 (2024). https://doi.org/10.1038/s41580-023-00694-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-023-00694-9