Abstract
Phosphoinositides are signalling lipids derived from phosphatidylinositol, a ubiquitous phospholipid in the cytoplasmic leaflet of eukaryotic membranes. Initially discovered for their roles in cell signalling, phosphoinositides are now widely recognized as key integrators of membrane dynamics that broadly impact on all aspects of cell physiology and on disease. The past decade has witnessed a vast expansion of our knowledge of phosphoinositide biology. On the endocytic and exocytic routes, phosphoinositides direct the inward and outward flow of membrane as vesicular traffic is coupled to the conversion of phosphoinositides. Moreover, recent findings on the roles of phosphoinositides in autophagy and the endolysosomal system challenge our view of lysosome biology. The non-vesicular exchange of lipids, ions and metabolites at membrane contact sites in between organelles has also been found to depend on phosphoinositides. Here we review our current understanding of how phosphoinositides shape and direct membrane dynamics to impact on cell physiology, and provide an overview of emerging concepts in phosphoinositide regulation.
Similar content being viewed by others
Introduction
A hallmark of eukaryotic cells is the elaborate membrane system that serves to create physically and functionally distinct organelles, which enable cells to compartmentalize metabolic networks and signalling cascades1. Different membrane-bounded organelles, such as the endoplasmic reticulum (ER), the Golgi complex, endosomes and lysosomes, or the plasma membrane, exchange macromolecules between them via vesicular and tubular carriers in processes of internalization (endocytosis) and secretion (exocytosis). Many organelles can also form membrane contact sites (MCSs) between each other to enable non-vesicular exchange of lipids, ions and metabolites2,3. Finally, organelles undergo fusion and fission as well as maturation processes that underlie the cell physiological response to changing environmental conditions, such as starvation or stress. These processes, collectively referred to as ‘membrane dynamics’, require a finely tuned regulation of the interaction of proteins with membranes. Over the past few decades, it has become clear that phosphoinositides — that is, phosphorylated derivatives of the myo-inositol-containing membrane phospholipid phosphatidylinositol (PtdIns; Fig. 1a,b) — are key determinants of compartmental membrane identity and act as spatiotemporal cues to direct membrane dynamics4,5,6. For example, PtdIns 3-phosphate (PtdIns(3)P) acts as marker lipid that recruits key cytosolic proteins to early endosomes to define their identity4,6,7.
a | Chemical structure of phosphatidylinositol (PtdIns), a glycerophospholipid with myo-inositol as a head group. Note that only the 2′-hydroxy group of the inositol ring is in the axial position (that is, perpendicular to the plane of the inositol ring), while all others are equatorial. PtdIns is depicted in the 1-stearoyl (R1), 2-arachidonyl (R2) configuration, the most frequent acyl chain composition found in phosphoinositides. b | Phosphoinositide metabolism. Addition and removal of phosphate groups at the 3′-hydroxy, 4′-hydroxy and 5′-hydroxy groups of PtdIns by phosphoinositide kinases (blue) and phosphatases (red) creates seven distinct phosphoinositide species. Enzymatic interconversion of phosphoinositides creates a network in which formation of one species depends on the availability of another, meaning that the cellular levels of certain phosphoinositides are interdependent. Phosphoinositides enriched in the endolysosomal system (shaded yellow) versus the secretory pathway and the plasma membrane (shaded blue) are indicated. c | Subcellular enrichment of phosphoinositides. The spatiotemporally controlled activity of phosphoinositide kinases and phosphatases creates a distinctive enrichment of phosphoinositides across the cellular compartments. The plasma membrane is highly enriched in PtdIns 4,5-bisphosphate (PtdIns(4,5)P2). The Golgi apparatus and the trans-Golgi network (TGN) are characterized by PtdIns 4-phosphate (PtdIns4P), which is also present at the plasma membrane. By contrast, the endosomal compartments are dominated by 3′-phosphoinositides, with PtdIns 3-phosphate (PtdIns3P) being the signature lipid of early endosomes. A fraction of PtdIns3P is converted to PtdIns 3,5-bisphosphate (PtdIns(3,5)P2) on late endosomes. Lysosomes are unique in having a diverse complement of phosphoinositides in their membranes, likely depending on their functional and metabolic state. On multiple endocytic routes, PtdIns 3,4-bisphosphate (PtdIns(3,4)P2) is formed on nascent endocytic carriers before conversion to PtdIns3P in endocytic vesicles. ER, endoplasmic reticulum; MTM, myotubularin; MVB, multivesicular body; PI3K; phosphatidylinositol 3-kinase; PI4K, phosphatidylinositol 4-kinase; PIPKI, phosphatidylinositol 4-phosphate 5-kinase; PtdIns(3,4,5)P2, phosphatidylinositol 3,4,5-trisphosphate; PtdIns5P, phosphatidylinositol 5-phosphate.
PtdIns is synthesized in the ER and is delivered to other organelles via lipid exchange proteins at MCSs and by vesicular or tubular carriers that move along the secretory pathway and within the endolysosomal system4. Reversible phosphorylation and dephosphorylation of the inositol ring by phosphoinositide kinases and phosphatases generate seven distinct phosphoinositide species. These species can be interconverted depending on the enzymatic activity, resulting in a metabolic network with multiple interdependencies (Fig. 1b). Phosphoinositides constitute less than 1% of the total phospholipid pool of cells, yet exhibit a highly localized subcellular distribution as a result of the tight spatiotemporal control of their synthesis and turnover4 (Fig. 1c). This directly impacts on the localization and activity of cytosolic effector proteins and membrane-integral proteins such as receptors, ion channels or transporters that interact with these lipids and elicit a physiological response (for example, a signalling cascade or assembly of a protein complex that can remodel membranes). Phosphoinositide-binding domains of effector proteins generally bind to their target lipid with low affinity. As a result, multivalent coincident recognition of additional factors, including small GTP-binding proteins and other lipids, as well as avidity effects caused by the presence of multiple lipid-binding sites or domains on effector proteins are required for their function1,4,6. Dual, coincident recognition of lipids and proteins also underlies the observed compartment specificity of phosphoinositide-binding domain-based lipid biosensors (Box 1; Table 1). An evolutionary advantage of this organizational principle is that each component of the phosphoinositide effector module can be regulated independently, thereby enabling the rapid cellular rewiring of membrane flux and metabolism to adapt to changing environmental conditions and to internal or external cues.
The importance of phosphoinositides in cell physiology and dynamic remodelling of membranes is best illustrated by the ever-growing list of human diseases linked to dysfunction of phosphoinositide-metabolizing enzymes (Tables 2,3). In spite of the apparent redundancy of phosphoinositide kinases (Table 3) and phosphatases (Table 2) at the level of the biochemical reaction catalysed, mutation of specific isoforms of these enzymes gives rise to diseases with a surprisingly high level of tissue or cell-type selectivity8. These range from neuromuscular, skeletal and kidney disorders caused by loss of function of phosphoinositide phosphatases (for example, myotubularin family members, FIG4 and OCRL) to overgrowth syndrome, immune deficiency and cancer as a result of hyperactivation of the class I PtdIns 3-kinase (PI3K)/AKT signalling pathway5,9. Pathogens, including bacteria and viruses, have also been found to exploit the regulatory potential of phosphoinositides for infection (Supplementary Box 1).
In this Review we discuss how the spatiotemporally controlled conversion of distinct phosphoinositides enables membrane flux in the exocytic and endocytic pathways. Furthermore, we summarize the rapid progress in our understanding of the role of phosphoinositides in the autophagy/lysosome system and the regulation of MCSs between organelles in response to altering environmental (for example, nutrient) conditions. Finally, we outline perspectives for future research and biomedicine. For a detailed discussion of the roles of phosphoinositides in cancer cell signalling and in the regulation of ion channels and other membrane proteins10,11 and of non-canonical phosphoinositide functions in the nucleus, the reader is referred to excellent recent reviews9,12,13.
Exocytic and endocytic membrane dynamics
Phosphoinositides were originally discovered more than half a century ago as rare phospholipid species that undergo turnover upon stimulation of hormone secretion14. These observations related to what is now known as the phosphoinositide cycle: agonists activate phospholipase C (PLC) to hydrolyse PtdIns 4,5-bisphosphate (PtdIns(4,5)P2) into the second messengers diacylglycerol (DAG) and inositol 1,4,5-trisphosphate (Ins(1,4,5)P3), followed by recycling of the components and resynthesis of PtdIns(4,5)P2 (ref.4). Studies initiated in yeast15 and continued over the past few decades in multiple models have established that in addition to cell signalling, phosphoinositides control virtually every step of membrane traffic along the secretory and endocytic pathways. This regulatory role of phosphoinositides is closely linked to their differential distribution between cellular subcompartments6 (Fig. 1c). From a bird’s-eye view, PtdIns 4-phosphates signpost the secretory pathway with the Golgi complex containing high levels of PtdIns 4-phosphate (PtdIns4P) and the plasma membrane being enriched in PtdIns(4,5)P2 and, to a lesser extent, PtdIns4P. By contrast, PtdIns 3-phosphates such as PtdIns3P and PtdIns 3,5-bisphosphate (PtdIns(3,5)P2) dominate the endolysosomal system. To preserve organellar membrane identity, traffic between compartments needs to be coupled to the interconversion of phosphoinositide species by functional units comprising phosphoinositide phosphatases and kinases that act in concert. In the following sections, we discuss how this enables phosphoinositides to direct the flow of membranes along the secretory pathway, on the endocytic route and in the sorting of cargo from endosomes.
Secretory membrane traffic
Secretory proteins synthesized in the ER are trafficked to the Golgi complex via COPII vesicles or tubules before being packaged into carriers destined for the plasma membrane at the trans-Golgi network (TGN). Protein secretion and the function of the TGN as a sorting station crucially depend on PtdIns4P. Vertebrates express four isoforms of PtdIns 4-kinases (PI4Ks), the PI3K-related type III isoforms PI4KIIIα and PI4KIIIβ, and the evolutionarily unrelated type II isoforms PI4KIIα and PI4KIIβ4. The Golgi pool of PtdIns4P is synthesized by PI4KIIα, PI4KIIβ and by PI4KIIIβ, which is recruited to the Golgi complex by the small GTPase ARF1 (ref.16) and the giantin-interacting protein ACBD17.
PtdIns4P-mediated control of secretory carrier formation
Although none of the phosphoinositides appears to be enriched in ER membranes, COPII-coated carrier formation at ER exit sites has been linked to PtdIns4P generation by PI4KIIIα18. It is not clear how this facilitates ER-to-Golgi transport but illustrates how early secretory cargo is committed to PtdIns4P-enriched membranes. The formation of TGN-derived carriers directed towards the plasma membrane or to endolysosomal compartments depends on PtdIns4P, as shown by acute depletion of this lipid19 (Box 1). Membrane recruitment of the clathrin adaptor proteins AP1, GGA1 and GGA2, which mediate transport from the TGN to endosomes and lysosomes, is promoted by PtdIns4P19,20. The importance of PtdIns4P for assembly of clathrin adaptors on TGN and endosomal membranes is further evidenced by the interaction of AP1 with PI4KIIβ21, GGA2 with PI4KIIIβ22 and AP3 with PI4KIIα23.
A comprehensive understanding of the presumably clathrin-independent formation of vesicles for constitutive secretion is still elusive, although PtdIns4P plays a central role. Effectors of PtdIns4P include Golgi-localized phosphoprotein 3 (GOLPH3), which associates with PtdIns4P and myosin 18A and thereby exerts a tensile force on Golgi membranes24. Extrusion of membrane tubules in preparation for secretory carrier formation from the TGN has also been linked to microtubule motor proteins25. The recent description of the DOPEY1–MON2 complex as an adaptor linking PtdIns4P-, phosphatidic acid- and ARF1-positive membranes to kinesin 1 (ref.26) suggests that also kinesin-dependent carrier formation relies on PtdIns4P. A complex of PI4KIIIβ, 14-3-3 protein-γ and the fission-promoting factors CTBP1/BARS27 and protein kinase D (PKD)28 is believed to couple extrusion of vesicular carriers to membrane scission.
A more complex regulatory function of PtdIns4P in secretion from the TGN is its role in linking the formation of secretory carriers to maintenance of the distinct lipid composition of post-Golgi membranes, which are enriched in glycosphingolipids, sphingomyelin and cholesterol. PtdIns4P in TGN membranes is required for the transfer of cholesterol and ceramide lipids to the TGN from the ER (see the section entitled Membrane contact sites), which in turn promote generation of PtdIns4P, as explained later: ceramide is used together with phosphatidylcholine to generate sphingomyelin, resulting in the formation of DAG, which in turn activates PKD29. PKD promotes secretory carrier formation at the TGN28 and also stimulates PI4KIIIβ activity30. Increased cholesterol levels promote palmitoylation and TGN recruitment of PI4KIIα, which together with PI4KIIIβ facilitates PtdIns4P synthesis from PtdIns31. Hence, PtdIns4P couples secretion from the TGN to acquisition of the distinct membrane composition characteristic of post-Golgi membranes. The physiological relevance of this interplay was recently demonstrated by defective myelination in mice lacking PI4KIIIβ expression in Schwann cells32, a cell type needing to sustain a large secretory load to form the cholesterol- and sphingolipid-rich myelin sheath around axons that facilitates saltatory conduction in peripheral nerves.
PtdIns4P is also at the nexus of tuning secretion in response to growth factor and nutrient availability. Serum depletion causes the translocation of the PtdIns4P 4-phosphatase SAC1 from the ER to the Golgi complex, thereby reducing the levels of PtdIns4P at the TGN and effectively limiting secretion33. Intriguingly, PtdIns4P may further serve as a sensor of cytoplasmic pH, whereby cytoplasmic acidification and the ensuing protonation of phosphoesters in the lipid head group of PtdIns4P disrupt its interaction with PtdIns4P-binding effector proteins; this will lead to downregulation of secretion under non-permissive metabolic conditions (that is, glucose starvation), leading to inactivation of a proton pump34.
Vesicle exocytosis
The ultimate step of the secretory pathway, vesicle exocytosis, is accompanied by the acquisition of plasma membrane identity by the secretory vesicle as it fuses with the plasma membrane. Regulated exocytosis in neurons and neuroendocrine cells has been demonstrated to scale with the levels of plasma membrane PtdIns(4,5)P2 (refs35,36), which acts in trans to facilitate secretory vesicle docking via synaptotagmin 1 (ref.37) and other factors. Constitutive exocytosis — which is necessary for the secretion of, for example, general plasma membrane proteins, collagen or glycosaminoglycans — requires the exocyst, an eight-subunit complex that tethers exocytic vesicles to PtdIns(4,5)P2 (ref.38) at the plasma membrane39. Engagement of the complex with exocytic vesicles is regulated by the small GTPase RAB11 and PtdIns4P40. Of note, the exocytosis of recycling vesicles that emanate from PtdIns3P-rich endosomal compartments requires conversion of PtdIns3P to PtdIns4P by the 3-phosphatase myotubularin 1 (MTM1) and PI4KIIα41 (see the section entitled Phosphoinositides in endocytic recycling). Interestingly, PI4KIIα has also been found on neurosecretory synaptic vesicles, suggesting that conserved mechanisms regulate the final steps of vesicle exocytosis42.
From these studies a model emerges whereby the formation and fission of secretory carriers is controlled by PtdIns4P, while the final steps of vesicle exocytosis depend on PtdIns(4,5)P2, the key identity determinant of the plasma membrane.
Endocytosis
PtdIns(4,5)P2 together with PtdIns4P defines the strongly anionic nature of the plasma membrane43. Plasma membrane PtdIns4P is synthesized by PI4KIIIα and its associated subunits EFR3 and TTC7 (ref.44). This complex is further stabilized by FAM126A, a protein found mutated in patients with hypomyelination of the brain45 (Table 3). PtdIns4P then serves as a substrate for the PtdIns4P 5-kinases (PIPKI)4 to form PtdIns(4,5)P2, a lipid required for endocytic vesicle formation.
In contrast to the plasma membrane, the endosomal system is dominated by 3-phosphoinositides46. A common theme across different endocytic routes therefore is that endocytic carrier formation is linked to conversion of membrane identity from plasma membrane PtdIns(4,5)P2 to endosomal PtdIns3P (Fig. 1c).
Clathrin-mediated endocytosis
Clathrin-mediated endocytosis (CME) is the main constitutive internalization route common to all cells47. During CME the sequential assembly of the clathrin coat concentrates cargo proteins into a clathrin-coated pit (CCP) via simultaneous detection of both cargo and PtdIns(4,5)P2 by clathrin adaptors48 (Box 1). The nascent CCP acquires increasing curvature and eventually buds off to form an endocytic vesicle (Fig. 2a) that undergoes fusion with early endosomes. During this process PtdIns(4,5)P2 is gradually turned over, concomitant with the acquisition of 3′-phosphoinositide identity that is characteristic of the endosomal system46.
a | In clathrin-mediated endocytosis, nucleation and growth of endocytic clathrin-coated pits (CCPs) are promoted by phosphatidylinositol 4,5-bisphosphate (PtdIns(4,5)P2). Many adaptors, which connect the clathrin lattice with the membrane and concentrate cargo proteins in the nascent pit, bind to PtdIns(4,5)P2. Maturing CCPs acquire a number of phosphoinositide-metabolizing enzymes, including PI3KC2α and the 5-phosphatases SHIP2 and synaptojanin 1 (p170). The resulting formation of phosphatidylinositol 3,4-bisphosphate (PtdIns(3,4)P2) through its effectors sorting nexin 9 (SNX9), SNX18 and FCHSD2 promotes constriction of the neck of invaginated CCPs. During uncoating of newly formed vesicles, a burst of recruitment of the 5-phosphatases synaptojanin 1 (p145) and OCRL depletes PtdIns(4,5)P2. The 4-phosphatases INPP4A, INPP4B and SAC2 complete conversion of membrane identity towards the endosomal signature lipid phosphatidylinositol 3-phosphate (PtdIns3P). b | Macropinocytosis, the internalization of large amounts of extracellular fluid, is largely driven by membrane remodelling through branched F-actin formation. Initial membrane ruffling requires the formation of PtdIns(4,5)P2 by PIPKIα/γ to promote actin polymerization. Receptor (that is, insulin or growth factor receptors) activation triggers the formation of phosphatidylinositol 3,4,5-trisphosphate (PtdIns(3,4,5)P3) by class I phosphatidylinositol 3-kinases (PI3Ks) to regulate Rho family guanine-nucleotide exchange factors (GEFs) and GTPase-activating proteins (GAPs) during expansion of the macropinocytic cup. Remodelling and resolution of the subcortical actin cytoskeleton requires depletion of PtdIns(4,5)P2 by phospholipase C (PLC) and 5-phosphatases starting from the base of the cup. These phosphoinositide conversions enable recruitment of the dual PtdIns3P- and phosphatidylinositol 4-phosphate (PtdIns4P)-binding protein Phafin 2, a protein also required for actin remodelling. c | The endosomal sorting of cargo depends on PtdIns3P. Endosomal PtdIns3P is largely synthesized by VPS34 complex II, which is activated by RAB5–GTP. Coincidence of cargo and PtdIns3P underlies the sorting of cargo into degradative and retrieval subdomains. This is mediated by proteins that can simultaneously bind to PtdIns3P and to specific sorting signals in cargo proteins. Endosomal sorting complex required for transport (ESCRT) 0 binds to ubiquitylated proteins and through self-association forms degradative subdomains. Retrieval signals are recognized by various members of the SNX family that operate in conjunction with different retrieval complexes. Exit from endosomes and recycling to the plasma membrane require conversion of PtdIns3P to PtdIns4P (and possibly PtdIns(4,5)P2). Proteins that interact with phosphoinositides are highlighted with coloured boxes, corresponding to the phosphoinositide species they bind to. The coloured bars highlight the phosphoinositide kinases and phosphatases, with colours corresponding to the phosphoinositide species they generate. BAR, Bin–Amphiphysin–Rvs; ER, endoplasmic reticulum; ILV, intraluminal vesicle; MTM1, myotubularin 1; MTMR, myotubularin-related protein; PIPKI, phosphatidylinositol 4-phosphate 5-kinase; PtdIns, phosphatidylinositol; TGN, trans-Golgi network; WASH, Wiskott-Aldrich syndrome protein and SCAR homologue.
Many endocytic proteins associate with PtdIns(4,5)P2, including the early-acting clathrin adaptor AP2, the Bin–Amphiphysin–Rvs (BAR) domain-containing proteins Fer/Cip4 homology domain-only protein 1 (FCHO1) and FCHO2, and numerous cargo adaptors49. PtdIns(4,5)P2, cargo and FCHO proteins trigger conformational opening of AP2 to enable its binding to cargo and to clathrin50,51,52. Nascent CCP formation and PtdIns(4,5)P2 synthesis are further linked via activation of PIPKI by AP2–cargo complexes53. This suggests a feedforward loop of adaptors binding to PtdIns(4,5)P2 and cargo to trigger increased local formation of PtdIns(4,5)P2 during CCP nucleation, until clathrin binding to AP2 displaces PIPKI54. The subsequent growth and maturation of CCPs are accompanied by the gradual depletion of PtdIns(4,5)P2 from the coat and the concomitant synthesis of PtdIns 3,4-bisphosphate (PtdIns(3,4)P2)55. This phosphoinositide conversion mechanism involves the recruitment of the PtdIns(4,5)P2 5-phosphatase synaptojanin 1 p170 (ref.56), an enzyme linked to Parkinson disease (Table 2), and the PtdIns 3,4,5-trisphosphate (PtdIns(3,4,5)P3) and PtdIns(4,5)P2 5-phosphatase SHIP2 (ref.57) (Table 2). Coincidentally, clathrin recruits the PtdIns(4,5)P2-activated class II PI3K PI3KC2α58 to synthesize PtdIns(3,4)P2 from PtdIns4P55,59. Depletion of either PI3KC2α or PtdIns(3,4)P2 stalls CCP dynamics and impairs the constriction of invaginated CCPs55,59. While PI3KC2α is essential in mice, human patients with inherited homozygous null mutations in the PIK3C2A gene display congenital syndromic features, including kidney failure and cataract, due to early senescence of cells in the associated tissues60 (Table 3). PtdIns(3,4)P2 promotes constriction of the endocytic CCP neck by recruitment and activation of the membrane curvature-inducing Phox homology (PX) domain–BAR domain-containing proteins sorting nexin 9 (SNX9) and SNX18 (refs55,61,62) and by stimulating actin polymerization63. Membrane constriction is a prerequisite for endocytic vesicle fission via assembly of the large GTPase dynamin64 mediated by the specific association of its pleckstrin homology (PH) domain with the remaining pool of PtdIns(4,5)P2 (refs65,66). Dynamin-mediated fission is paralleled by a burst of recruitment of synaptojanin 1 (ref.67) and OCRL68,69, a 5-phosphatase mutated in patients with oculocerebrorenal syndrome of Lowe, a rare multisystem disorder characterized by congenital cataracts, glaucoma, intellectual disability, seizures, postnatal growth retardation and renal tubular dysfunction (Table 2). OCRL/synaptojanin 1-mediated hydrolysis generates PtdIns4P70 (possibly in conjunction with PtdIns(3,4)P2 (ref.71)), which finally recruits the co-chaperone protein auxilin or GAK to trigger removal of the clathrin coat. The subsequent conversion of PtdIns(3,4)P2 to PtdIns3P by the endosomal 4-phosphatase INPP4A59,72 may facilitate fusion of the nascent endocytic vesicle with early endosomes marked by PtdIns3P.
Visualizing this phosphoinositide conversion pathway in living cells has remained a challenge. The application of hybrid clathrin-binding sensors that allegedly detect PtdIns3P and PtdIns(3,4)P2 has led to the alternative proposal that PI3KC2α can directly synthesize PtdIns3P at late-stage endocytic vesicles (after fission)73. Such a model, however, is at odds with the observation that loss of PI3KC2α stalls endocytosis before vesicle fission and that a mutant form of PI3KC2α capable of synthesizing PtdIns3P but not PtdIns(3,4)P2 fails to rescue defective CME in cells lacking the endogenous enzyme55,59. A recent study using a new live probe for PtdIns(3,4)P2 (ref.74) failed to detect its enrichment at CCPs, likely as a consequence of competition with endogenous endocytic PtdIns(3,4)P2-binding proteins. This study further demonstrated that the alleged double clathrin–PtdIns(3,4)P2 sensor73 does not depend on PtdIns(3,4)P2 for enrichment at CCPs74. Hence, novel approaches are needed to monitor local phosphoinositide pools at endocytic sites in live cells.
Clathrin-independent endocytosis
Several clathrin-independent endocytic mechanisms have been proposed (see75 for a detailed review). These pathways respond to context-dependent or tissue-specific demands75 and are often linked to receptor-mediated class I PI3K activation.
During fast endophilin-mediated endocytosis (FEME)76, activation of signalling receptors (for example, the β1-adrenergic or EGF receptors) triggers PtdIns(3,4,5)P3 synthesis by the receptor-associated class I PI3K isoforms PI3Kα and PI3Kβ5 (see also Table 3). Subsequent formation of PtdIns(3,4)P2 by the 5-phosphatases SHIP1 and SHIP2 facilitates plasma membrane binding of the PtdIns(3,4)P2-associated actin modulator lamellipodin77. Lamellipodin recruits the membrane-deforming N-BAR domain-containing protein endophilin to the leading edge of cells, from where receptors are internalized into tubular endosomes. In addition to lamellipodin and endophilin, this process depends on the F-BAR domain-containing proteins FBP17 and CIP4 (ref.78), which prime endophilin assembly, actin and dynamin76. Akin to CME, FEME may be terminated by conversion of PtdIns(3,4)P2 to PtdIns3P en route to endosomes via the 4-phosphatases INPP4A and INPP4B76 (Table 2).
Large volumes of extracellular fluid are internalized by macropinocytosis (see79 for a review; see Supplementary Box 1 for the closely related process of phagocytosis) (Fig. 2b). The initial large-scale membrane remodelling needed to engulf extracellular fluid is mediated by actin polymerization. PtdIns(4,5)P2 produced by PIPKIα and PIPKIγ is essential for stabilizing the actin network and for activation of actin nucleation-promoting factors that stimulate the ARP2/3 complex80. The extension and closure of the macropinocytic cup is then driven by class I PI3K-mediated synthesis of PtdIns(3,4,5)P3, which mediates the coordinated activation of Rho family GTPases through modulation of their guanine-nucleotide exchange factors (GEFs) and GTPase-activating proteins81,82. This is accompanied by depletion of PtdIns(4,5)P2 and resolution of the actin meshwork starting from the base of the cup, to which PI3Ks, the activation of PLCγ, recruitment of 5-phosphatases and focal exocytosis contribute79. The sequential activities of SHIP1/2 and INPP4A/B following closure of the macropinosome convert PtdIns(3,4,5)P3 to PtdIns3P, which may finally be cleared by the phosphoinositide 3-phosphatase myotubularin-related protein 6 (MTMR6)79,83. The dual PtdIns3P- and PtdIns4P-binding protein Phafin 2 couples these phosphoinositide conversions to rearrangement of the actin cytoskeleton to enable macropinosome internalization84. It is likely that analogous lipid conversions govern other clathrin-independent endocytic routes (for example, the CLIC–GEEC pathway75).
A common principle of endocytic vesicle formation is thus the mechanistic coupling of membrane remodelling to a series of phosphoinositide conversion reactions that result in the loss of plasma membrane PtdIns(4,5)P2 and the concomitant acquisition of endosomal PtdIns3P identity.
Dynamics of endosomes
Following internalization, endocytic vesicles coalesce in the early endosomal compartment to enable further sorting. Endocytic cargo can be recycled to the plasma membrane, trafficked retrogradely to the TGN or be sorted into late endosomes (also called ‘multivesicular bodies’) for lysosomal degradation (Fig. 2c). The signature phosphoinositide of the endosomal compartment is PtdIns3P, which is predominantly synthesized from PtdIns by the sole class III PI3K VPS34, with minor contributions from class II PI3Ks, likely depending on the cell type5,46. VPS34 operates in two distinct heterotetrameric complexes. The endosomal complex II comprises VPS34, VPS15–beclin 1 and UVRAG (Table 3). In VPS34 complex I UVRAG is replaced by ATG14L to regulate autophagy (see later)5,46.
Establishing early endosomal membrane identity
The small GTPase RAB5 is instrumental for the maintenance and function of the early endosomal compartment85. Initial RAB5 activation is linked to the formation of endocytic carriers by RME6 (also known as GAPVD1), a GEF that couples clathrin–AP2 uncoating to PtdIns(4,5)P2 depletion86. On early endosomes RAB5 activity is controlled by the RAB5 GEF Rabex 5, the RAB5–GTP-associated VPS34 complex II (ref.87) and its lipid product PtdIns3P, and the RAB5 effector Rabaptin 5, which forms a tight complex with Rabex 5. The resulting feedforward loop generates and maintains the canonical membrane environment enriched in RAB5–GTP and PtdIns3P that characterizes early endosomes88. Dual key recognition of RAB5–GTP and PtdIns3P underlies homotypic early endosome fusion, a reaction that is required to sustain endocytic traffic of internalized cargo and to maintain early endosomal membrane homeostasis88. Further RAB5 effectors include the 5-phosphatases OCRL and INPP5B as well as the PtdIns(3,4)P2 4-phosphatases INPP4A and INPP4B72,89,90, which ensure conversion of incoming phosphoinositides (on the incoming endocytic vesicles) to PtdIns3P. Besides membrane fusion, endosomal PtdIns3P forms the basis for cargo fate decisions. As discussed later, the arrival of internalized cargo in the PtdIns3P-enriched endosome triggers the formation of spatially separated subdomains that enable cargo sorting for retrieval or degradation91.
Phosphoinositides in endocytic recycling
During endocytic recycling, membrane is returned from PtdIns3P-containing endosomal compartments to the PtdIns4P/PtdIns(4,5)P2-enriched plasma membrane. The bulk of the membrane is retrieved from endosomes in the form of tubulovesicular carriers via protein complexes including retromer, retriever and ESCPE-1 (Fig. 2c). These retrieval complexes cluster cargo containing specific sorting motifs into retrieval subdomains. Once a critical density is reached, the underlying membrane is tubulated by the concerted action of BAR domain-containing SNX proteins and Wiskott–Aldrich syndrome protein and SCAR homologue (WASH)-dependent branched F-actin formation92,93. These retrieval complexes are targeted to endosomes by PtdIns3P. Retromer operates in the recycling of distinct sets of cargo proteins with either SNX3 or SNX27, both of which contain a PtdIns3P-binding PX domain92. In retriever and ESCPE-1, a similar role is fulfilled by SNX17 (ref.94) and SNX1/2, respectively93,95. Interestingly, loss of the WASH-associated endosomal COMMD/CCDC22/CCDC93 (CCC) complex leads to elevated endosomal PtdIns3P levels as a consequence of defective recruitment of the 3-phosphatase MTMR2 (ref.96), an enzyme mutated in Charcot–Marie–Tooth disease type 4B (Table 2), suggesting control of PtdIns3P levels through components of the recycling machinery. This increase in PtdIns3P levels depends on VPS34 and is accompanied by accumulation of the WASH complex and F-actin on endosomal structures96. Distinct pools of PtdIns3P, synthesized by PI3KC2α on RAB11-positive endosomes97 and at a specialized endosomal compartment at the base of primary cilia98, have also been implicated in endocytic recycling, but the mechanistic underpinnings of these observations remain unclear.
The function of the WASH complex is further subject to regulation by PtdIns4P on endosomes through PI4KIIα/β and ER–endosome contact sites (see the section entitled Membrane contact sites)99. This raises the intriguing possibility that endosomal retrieval itself may be linked to phosphoinositide conversion of PtdIns3P to PtdIns4P. Upon retrieval from endosomes, cargo can be trafficked to either of two PtdIns4P-rich destinations, the TGN or the plasma membrane92. Indeed, the subsequent exocytosis of recycling vesicles requires a phosphoinositide conversion module comprising the PtdIns3P phosphatase MTM1, PI4KIIα and the exocyst complex41. In the absence of MTM1, recycling vesicles fail to fuse with the plasma membrane, a phenotype that is recapitulated upon perturbation of PI4KIIα, RAB11 or exocyst and which likely contributes to the pathology of X-linked centronuclear myopathy in patients with loss of function of MTM1 (ref.41) (Table 2). Hence, recycling from endosomes to the plasma membrane is intrinsically coupled to conversion of membrane identity from PtdIns3P to PtdIns4P (Fig. 2c).
Phosphoinositides in degradative sorting and endosomal maturation
Ubiquitylated cargo destined for lysosomal delivery is sorted into vesicles that bud into the lumen of maturing endosomes. This formation of intralumenal vesicles is catalysed by four subcomplexes called ‘endosomal sorting complexes required for transport (ESCRT) 0–III’ (ref.100). ESCRT-0 is a heterotetramer of two copies each of HRS and STAM and is constitutively targeted to the early endosomal compartment through the PtdIns3P-binding FYVE domain of HRS101 (Table 1). As both HRS and STAM contain multiple ubiquitin-binding sites and can self-associate into larger complexes, the coincidence of ubiquitylated cargo and PtdIns3P leads to the formation of degradative subdomains on endosomes92. The sequential activity of ESCRT-I to ESCRT-III mediates the inward budding of the endosomal membrane and eventually the inclusion of degradative subdomains in intralumenal vesicles100 (Fig. 2c). Increasing formation of intralumenal vesicles results in endosomal maturation to late endosomes/multivesicular bodies and is accompanied by endosomal conversion of RAB5 to RAB7. This depends on the RAB7 GEF MON1–CCZ1 binding to RAB5–GTP to displace Rabex 5 and thereby disrupt the self-perpetuating loop of RAB5 activation102,103. Effectors of RAB7–GTP characterize the late endosomal compartment and include VPS34 complex II and the retromer complex, ensuring continued PtdIns3P formation and retrieval of cargo103. Late endosomal PtdIns3P serves as a substrate for the PIKfyve complex, a PtdIns3P 5-kinase that is essential for late endosomal and lysosomal function and homeostasis104,105 (see also Table 3), as discussed in the following section.
In conclusion, endosomal recycling to the cell surface versus degradative sorting to late endosomes and lysosomes is governed by loss versus consolidation of endosomal PtdIns3P identity, likely involving distinct endosomal lipid nanodomains.
Autolysosomal system
Lysosomes are dynamic organelles that serve as major degradative compartments for intracellular and exogenous substrates that are broken down into their constituent building blocks by lumenal acid hydrolases106. Lysosomes and lysosome-related organelles (for example, lytic granules in immune cells and melanosomes) also fulfil multiple other functions, including roles in secretion (for example, of hydrolases and cytotoxins) and as a metabolic signalling hub that integrates nutrient sensing and metabolic adaptation with lipid and metabolite exchange between organelles, for example via MCSs (see later). It is thus not surprising that lysosomes harbour a diverse complement of phosphoinositides46,107 that may be adapted to the functional state of the cell (Fig. 3).
Lysosomes harbour a diverse complement of phosphoinositides that regulate their functions. In fed cells (left), lysosomal phosphatidylinositol 3-phosphate (PtdIns3P) and phosphatidylinositol 3,5-bisphosphate (PtdIns(3,5)P2) synthesized by VPS34 and PIKfyve, respectively, facilitate nutrient signalling via mechanistic target of rapamycin complex 1 (mTORC1), possibly by associating with its regulatory associated protein of mTOR (Raptor) subunit (number 1). mTORC1 activity is facilitated by oxysterol-binding protein (OSBP)-mediated cholesterol transfer from the endoplasmic reticulum (ER) at membrane contact sites (MCSs). PtdIns3P also promotes the anterograde transport of lysosomes via kinesin recruitment by FYVE and coiled-coil domain-containing protein 1 (FYCO1) and its binding partner RAB7 (number 2). Under starvation (right), mTORC1 activity is repressed by local phosphatidylinositol 3,4-bisphosphate (PtdIns(3,4)P2) synthesis mediated by class II phosphatidylinositol 3-kinase-β (PI3KC2β). PtdIns(3,4)P2 facilitates reverse cholesterol transfer from lysosomes to the ER via OSBP-related protein 1 long isoform (ORP1L) at MCSs (number 3). Concomitantly, starvation induces autophagy. Delivery of vesicles containing PI4KIIIβ and ATG9A may aid phagophore membrane extrusion (number 4). Formation of PtdIns3P by VPS34 complex I is required to drive expansion of the phagophore membrane from the ER by recruiting multiple effector proteins, leading to LC3 lipidation by the ATG16L complex (ATG12–ATG5–ATG16L) and autophagosome formation (number 5). Membrane fusion between PtdIns3P-containing autophagosomes and lysosomes involves complex pathways of PtdIns3P–PtdIns(3,5)P2 and phosphatidylinositol 4-phosphate (PtdIns4P)–phosphatidylinositol 4,5-bisphosphate (PtdIns(4,5)P2) synthesis and turnover that involve phosphatidylinositol 4-kinase IIα (PI4KIIα) and the RAB7 effector protein PLEKHM1 (number 6). Starvation-triggered Ca2+ efflux via the PtdIns(3,5)P2-activated Ca2+-channel mucolipin 1 activates calcineurin and, thereby, induces transcription of autophagy/lysosomal genes via nuclear translocation of transcription factor EB (TFEB) (number 7). Prolonged starvation triggers autophagic lysosome reformation (ALR), a process that involves multiple lysosomal phosphoinositides. Synthesis of PtdIns(4,5)P2 on lysosomes by PtdIns4P 5-kinase (PIPKI) isoforms induces the formation of lysosomal tubules and the assembly of clathrin–AP2 coats that support the budding of protolysosomes from tubular intermediates via dynamin 2. Tubulation is further facilitated by VPS34 complex II-mediated synthesis of PtdIns3P, which serves to recruit the FYVE domain-containing protein spastizin, and by PtdIns(3,5)P2 synthesis via PIKfyve (number 8). The colour of proteins indicates the phosphoinositide species they bind to. ALFY, autophagy-linked FYVE protein; DFCP1, double FYVE domain-containing protein 1; ULK1, UNC-51-like autophagy-activating kinase 1; VAP, VAMP-associated protein; WIPI2, WD repeat domain phosphoinsitide-interacting protein 2.
Extracellular material is delivered to the lysosome lumen by the endocytic pathway via fusion with late endosomes as discussed earlier herein. Aggregated intracellular proteins, defective organelles or pathogens can be targeted for lysosomal degradation via macroautophagy (hereafter referred to as ‘autophagy’), a stress-inducible catabolic process that involves the formation of double membrane-bounded autophagosomes that eventually fuse with lysosomes108. Autophagy is unique in enabling the topological inversion of the cytoplasm and the creation of de novo membrane identity by synthesis of PtdIns3P from PtdIns on specialized sites of the ER.
Phosphoinositides in autophagy
Phosphoinositide regulation of autophagosome formation
The biogenesis of autophagosomes is orchestrated by multiple complexes containing autophagy-related proteins (ATG proteins) that first initiate the formation of a preautophagosomal structure, termed the ‘phagophore’, that emanates from specialized PtdIns3P-enriched sites109,110 on the ER referred to as the ‘omegasome’109. The phagophore elongates and closes to form a mature autophagosome that will ultimately fuse with the lysosome.
Autophagy is initiated by activation of the UNC-51-like kinase 1 (ULK1) complex — for example downstream of mechanistic target of rapamycin complex 1 (mTORC1) inactivation (Fig. 3, number 3). Translocation of the ULK1 complex to phagophore initiation sites at the ER may be aided by delivery of PI4KIIIβ via vesicles marked by the transmembrane protein ATG9A to produce a local pool of PtdIns4P (ref.111). Active ULK1 promotes the recruitment of VPS34 complex I (ref.112), which is responsible for the local production of PtdIns3P (Fig. 3, number 4). Autophagic pools of PtdIns3P may also be synthesized by class II PI3Ks, including PI3KC2α, under specific cell physiological conditions (for example, shear stress113). PtdIns3P acts as a signalling molecule for the recruitment of various PtdIns3P-binding proteins, such as double FYVE domain-containing protein 1 (DFCP1), autophagy-linked FYVE protein (ALFY) and the WD repeat domain phosphoinositide-interacting protein (WIPI) family of scaffold proteins110. WIPI2 via its PROPPIN domain binds to PtdIns3P at the omegasome and recruits the ATG12–ATG5–ATG16L1 complex114, which acts as an E3-like ligase for the conjugation of ubiquitin-like LC3 family proteins to phosphatidylethanolamine. In this mechanism, WIPI2 and VPS34 complex I mutually promote each other’s recruitment, resulting in a positive feedback loop between PtdIns3P and LC3 lipidation115, which is essential for expansion and closure of the phagophore (Fig. 3, number 5). Furthermore, at PtdIns3P-abundant sites, WIPI4 via association with the lipid transfer protein ATG2 (ref.116) tethers the ER to the expanding phagophore, allowing phospholipid transfer required for phagophore expansion (Fig. 4a, number 5).
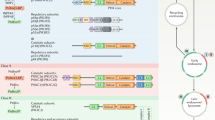
Phosphoinositides regulate the formation of membrane contact sites (MCSs) (panel a), while MCSs mediate lipid transfer and control phosphoinositide metabolism (panel b). a | During MCS formation, phosphoinositides in the membrane of one organelle recruit a specific phosphoinositide-binding protein anchored in the membrane of another organelle. Examples of this organization include endoplasmic reticulum (ER)–plasma membrane (PM) contacts mediated by extended synaptotagmin 1 (E-Syt1) (number 1); ER–trans-Golgi network (TGN) MCSs formed by ceramide transfer protein (CERT) and VAMP-associated protein (VAP) (number 2); ER–late endosome MCSs formed by protrudin with PDZD8, VAP and RAB7 (number 3); ER–lysosome MCSs formed by oxysterol-binding protein (OSBP)-related protein 1 long isoform (ORP1L) and VAP; and ER–omegasome contacts containing WD repeat domain phosphoinositide-interacting protein 4 (WIPI4) and autophagy-related protein ATG2. b | Vectorial transfer of lipids at MCSs. Lipid transfer can follow the concentration gradient or can be powered by the countertransport of another lipid. Examples of this lipid transfer include countertransport of cholesterol and phosphatidylinositol 4-phosphate (PtdIns4P) by OSBP at MCSs between the ER and the TGN (number 1); phosphatidylserine (PtdSer) and PtdIns4P by ORP10 at ER–endosome contacts (number 2); and PtdSer and PtdIns4P/phosphatidylinositol 4,5-bisphosphate (PtdIns(4,5)P2) by OSBP-related protein 5 (ORP5)/ORP8 at ER–plasma membrane MCSs (number 3). In addition, during infection with hepatitis C virus (HCV), Nir2 catalyses transport of phosphatidylinositol (PtdIns) from the ER to the plasma membrane, the Golgi complex, or HCV replicative organelles (HCV-RO) in exchange for phosphatidic acid (PtdOH) (number 4). In pancreatic β-cells, TMEM24 replenishes plasma membrane PtdIns lost during glucose-stimulated phospholipase C (PLC) signalling under conditions of low Ca2+ levels. PtdIns shuttled via Nir2 or TMEM24 is phosphorylated by phosphatidylinositol 4-kinase (PI4K) and PtdIns4P 5-kinase (PI4P5K) at the plasma membrane. FYVE, phosphatidylinositol 3-phosphate-binding domain named after FAB1, YOTB, VAC1 and EEA1; ORD, OSBP-related domain; PITP, phosphatidylinositol transfer protein domain; PH, pleckstrin homology domain; PtdIns3P, phosphatidylinositol 3-phosphate; PtdIns(3,4)P2, phosphatidylinositol 3,4-bisphosphate; PtdIns(3,4,5)P3, phosphatidylinositol 3,4,5-trisphosphate; SMP, synaptotagmin-like mitochondrial lipid-binding domain; StART, StAR-related lipid transfer domain.
While generation of PtdIns3P is necessary for the initiation of autophagy, its clearance is a prerequisite for the completion of the pathway by lysosomal degradation108,110. PtdIns3P hydrolysis during later stages of autophagy is mediated by members of the myotubularin family of 3-phosphatases (MTMR proteins). Depletion of these phosphatases causes cell type-specific and/or tissue type-specific defects in autophagy, and their dysfunction has been linked to human diseases, including X-linked centronuclear myopathy and Charcot–Marie–Tooth disease117 (Table 2).
Phosphoinositide regulation of autophagosome–lysosome fusion
Multiple phosphoinositides control the tethering and fusion of lysosomes with autophagosomes via incompletely understood mechanisms118 (Fig. 3, number 6). Autophagosomes are enriched in VPS34 complex I-derived PtdIns3P119, while lysosomes contain PtdIns3P synthesized by VPS34 complex II and PtdIns(3,5)P2 synthesized by the PIKfyve complex (comprising PIKfyve, VAC14 and FIG4). Lysosomal PtdIns(3,5)P2 is eventually hydrolysed by MTMR family members and by INPP5E104,105, a 5-phosphatase that can act on PtdIns(3,4,5)P3, PtdIns(4,5)P2 and PtdIns(3,5)P2 (ref.120). PIKfyve-mediated synthesis and turnover of lysosomal PtdIns(3,5)P2 are thus both key events in autophagosome–lysosome fusion. Potential effector proteins of lysosomal PtdIns(3,5)P2 include the actin-associated protein cortactin121, the lysosomal calcium release channel mucolipin122, twin pore channel 2 (TPC2)123 and the PX domain-containing protein SNX14 (ref.124). Given that INPP5E is mutated in Joubert syndrome in humans125 (Table 2) and that truncating mutations of SNX14 cause familial cerebellar atrophy, these phosphoinositide dynamics have important pathophysiological implications.
In addition to autophagosomal PtdIns3P and PtdIns(3,5)P2 on lysosomes, autophagosome–lysosome fusion requires synthesis of PtdIns4P by a lysosomal pool of PI4KIIα126. On lysosomes, PtdIns4P can be further phosphorylated to PtdIns(4,5)P2, which triggers the dissociation of the RAB7 effector PLEKHM1 by an unknown mechanism126. As lysosomal PtdIns(4,5)P2 inhibits the lysosomal calcium release channel mucolipin 1 — a protein required for sustained autophagy by regulating transcriptional activity of transcription factor EB (TFEB), which regulates the transcription of many lysosomal and autophagy genes127 (Fig. 3, number 7) — it likely needs to be restricted to nanodomains that may be generated by OCRL 5-phosphatase-mediated PtdIns(4,5)P2 hydrolysis128. Finally, recent data from mice and worms suggest that non-conventional synthesis of PtdIns(4,5)P2 via phosphorylation of PtdIns 5-phosphate (PtdIns5P) by PtdIns5P 4-kinases (Table 3) facilitates lipid catabolism by clearance of autophagosomes during fasting129. The function of this PtdIns(4,5)P2 pool remains to be elucidated.
From these data, a model emerges whereby parallel pathways of PtdIns3P–PtdIns(3,5)P2 and PtdIns4P–PtdIns(4,5)P2 synthesis and turnover control distinct, yet poorly characterized, steps in the formation of autolysosomes.
Lysosome dynamics and activity
Lysosomes harbour a diverse complement of phosphoinositides (Fig. 3) that may reflect their distinct roles depending on the functional or metabolic state of the cell. In addition to PtdIns3P, which controls autophagosome formation and dominates the endosomal system46, lysosomes have been shown to contain or be regulated by PtdIns(3,5)P2 and possibly PtdIns5P. These phosphoinositide species are thought to be specific to lysosomes, late endosomes and other lysosome-related organelles104,105. As described earlier herein, lysosomes also harbour minor pools of PtdIns4P and PtdIns(4,5)P2, phosphoinositides most highly enriched at the Golgi complex and at the plasma membrane, respectively,4,6,49 and PtdIns(3,4)P2, a rare phosphoinositide species that may control lysosome function in response to cessation of insulin growth factor signalling5,130. These different phosphoinositide pools likely correlate with the functional state and/or mark distinct subsets of lysosomes106 (for example, PtdIns(3,5)P2-containing degradative lysosomes harbouring a highly active vacuolar ATPase131).
Phosphoinositides couple lysosome dynamics and nutrient signalling
Lysosomes serve an essential role as metabolic signalling hubs106. Insulin and related growth factor signalling and the availability of nutrients (for example, amino acids) are integrated by mTORC1 to balance catabolic pathways with anabolic pathways (that is, autophagy with the synthesis of proteins, lipids and nucleotides). mTORC1 is a multiprotein assembly that is composed of the kinase mTOR, a distant relative of the PI3Ks, and regulatory-associated protein of mTOR (Raptor), as well as additional subunits. It is recruited to lysosomes by RAG small GTPases in response to cellular and lysosomal amino acid status106. Among the major direct substrates of active mTORC1 are translation-promoting ribosomal protein S6 kinase 1, the autophagy regulatory kinase ULK1 and TFEB. mTORC1-mediated phosphorylation of ULK1 and TFEB represses autophagy and autophagy/lysosomal gene expression127, while promoting protein synthesis under conditions of ample growth factor and nutrient supply106.
Phosphoinositides control lysosomal mTORC1 activity and, thereby, nutrient metabolism at several levels and at distinct subcellular sites. Insulin and related receptors at the plasma membrane, upon ligand binding, activate class I PI3Ks (for example, PI3Kα, an enzyme linked to cancer and somatic overgrowth; Table 3), resulting in the generation of PtdIns(3,4,5)P3 and the subsequent lipid-mediated activation of its effector protein kinase AKT. Sustained activation of AKT at the cell surface can be achieved by hydrolysis of PtdIns(3,4,5)P3 to PtdIns(3,4)P2 via the inositol 5-phosphatase SHIP2 (ref.5). Active AKT indirectly stimulates mTORC1 activity by phosphorylation-induced repression of lysosomal tuberous sclerosis complex 2 (TSC2), a negative regulator of the mTORC1-activating RHEB GTPase5,106.
Recent data indicate that mTORC1 activity and lysosome dynamics are further regulated locally by lysosomal phosphoinositides, for example PtdIns3P, PtdIns(3,4)P2 and PtdIns(3,5)P132. The PtdIns3P-generating kinase VPS34 appears to be required for full activation of mTORC1 (for example, during refeeding after starvation133,134). While the exact mechanism by which PtdIns3P activates mTORC1 is unclear, it has been postulated that PtdIns3P-mediated induction of mTORC1 activity requires the formation of MCSs between lysosomes and the ER (see the next section) and the concomitant translocation of lysosomal mTORC1 to the cell periphery, where insulin and growth factor signalling occur135. This mechanism involves the association of the ER membrane protein protrudin with lysosomal PtdIns3P and the recruitment of the PtdIns3P-binding protein FYVE and coiled-coiled domain-containing protein 1 (FYCO1) to lysosomes to facilitate kinesin-driven anterograde transport of lysosomes136 (Fig. 3, number 2). Loss of either protrudin or FYCO1 represses mTORC1 activity, while increasing the levels of active TFEB in the nucleus135, highlighting the close ties of phosphoinositides, mTORC1, and lysosome homeostasis and dynamics.
A second mechanism by which PtdIns3P can regulate lysosome function and dynamics involves the generation of PtdIns(3,5)P2 from PtdIns3P via PIKfyve. Neuronal depletion of PIKfyve or sustained pharmacological inhibition of PIKfyve stalls lysosome movement in neurites, likely contributing to the neurodegeneration observed in PIKfyve gene-knockout models137 (Table 3). In some cell types, including Schwann cells in the peripheral nervous system, and in yeast, PtdIns(3,5)P2 akin to PtdIns3P promotes mTORC1 activation138,139, possibly via binding of the mTORC1 subunit Raptor (Fig. 3, number 1). Interestingly, PtdIns(3,5)P2 has also been suggested to promote the membrane recruitment of the mTORC1-inhibitory TSC140. How these apparently conflicting roles of PtdIns(3,5)P2 in mTORC1 regulation in different cell types can be reconciled with each other will have to await further studies.
While PtdIns3P production by the class III PI3K VPS34 upregulates mTORC1 and induces lysosome dispersion, local generation of a lysosomal pool of PtdIns(3,4)P2 by the class II PI3K PI3KC2β plays an opposing role130. In growth factor-deprived conditions, PI3KC2β, an enzyme mutated in focal epilepsy in humans141 (Table 3), is recruited to the mTORC1 subunit Raptor on lysosomes and phosphorylates PtdIns4P to generate PtdIns(3,4)P2 (Fig. 3, number 3). Local PtdIns(3,4)P2 synthesis represses mTORC1 signalling via recruitment of inhibitory 14-3-3 proteins130 and by facilitating the oxysterol-binding protein (OSBP)-related protein 1 long isoform (ORP1L)-mediated transport of cholesterol142, an important cofactor for mTORC1 activation143, to the ER (Fig. 3, number 3). Concomitantly, lysosomal PtdIns(3,4)P2 also promotes the net retrograde transport of lysosomes towards the microtubule-organizing centre (away from the plasma membrane). PI3KC2β-mediated repression of mTORC1 signalling is relieved upon refeeding of cells by protein kinase N-mediated phosphorylation and complex formation of PI3KC2β with inhibitory 14-3-3 proteins in the cytoplasm144. Apart from 14-3-3 and ORP1L, the specific effector proteins of lysosomal PtdIns(3,4)P2 remain largely unknown. It is possible that PtdIns(3,4)P2 affects the activation cycle of small GTPases that link lysosomes to dynein or kinesin motors and, thereby, regulate lysosome positioning and mTORC1 activity106,107. These include ADP-ribosylation factor-like 8 (ARL8), which promotes kinesin association with lysosomes145,146, and RAB7, which connects lysosomes to dynein or kinesin motors147,148,149 How exactly lysosome positioning and mTORC1 activity are related mechanistically remains largely enigmatic.
Taken together, these examples illustrate how local phosphoinositide signalling couples nutrient signalling and metabolism to lysosome dynamics. The example of PtdIns(3,4)P2 also illustrates how a single lipid can have distinct opposing effects on a signalling pathway depending on whether it is present at the plasma membrane (that is, mTORC1 activation) or on lysosomes (that is, mTORC1 repression).
Phosphoinositides in lysosome reformation
During prolonged starvation lysosomes and autolysosomes tubulate and bud off nascent protolysosomes (that is, small and immature lysosomes that serve to replenish the pool of functional lysosomes150). This pathway, termed ‘autophagic lysosome reformation’ (Fig. 3, number 8), is controlled by multiple lysosomal phosphoinositide species, including PtdIns(4,5)P2 and PtdIns3P, which act at distinct stages of autophagic lysosome reformation. Prolonged starvation triggers synthesis of PtdIns(4,5)P2 on autolysosomes by PIPKI isoforms151. PtdIns(4,5)P2 induces lysosomal tubule formation via recruitment of kinesin 1 and the assembly of clathrin–AP2 coats, which support the budding of protolysosomes from tubular intermediates152. Starvation-induced lysosome tubulation is further facilitated by PtdIns(3,5)P2 synthesis153. Protolysosomal vesicles then pinch off from clathrin–AP2-coated tubules via the PtdIns(4,5)P2-regulated membrane fissioning enzyme dynamin 2 (ref.154) and VPS34 complex II-mediated synthesis of PtdIns3P155. Among the effectors of PtdIns3P in late steps of autophagic lysosome reformation is the FYVE domain-containing protein spastizin (also known as ZFYVE26 or FYVE-CENT), a protein mutated in hereditary spastic paraplegia in humans that facilitates the formation of lysosomal tubules156. In starved cells, spastizin together with its binding partners, the AP5 adaptor complex and SPG11, is recruited to membranes via coincident detection of PtdIns3P and inactive RAG GTPases157, suggesting crosstalk between autophagic lysosome reformation and mTORC1 signalling.
Membrane contact sites
In addition to the budding and fusion of vesicular and tubular carriers, organelles communicate with each other via MCSs — for example by exchanging lipids and ions, by channelling of small metabolites or by facilitating membrane fusion and fission processes2,3. At MCSs, the membranes of two distinct organelles lie closely apposed to each other (for example, 10–30 nm) owing to a physical connection through tethering factors, frequently in conjunction with a specific membrane lipid. Given the key role of phosphoinositides as determinants of organellar membrane identity, it is not surprising that these molecules also play important roles in the function of MCSs and in regulating their dynamics. Conversely, MCSs control the differential distribution and metabolism of phosphoinositides and of other lipids (for example, cholesterol, phosphatidylserine (PtdSer) or phosphatidic acid (PtdOH)), often in response to cell signalling or metabolic cues2,3 (Fig. 4). A particularly prominent role is played by the ER, which forms MCSs with essentially all other organelles, often involving dual key adaptors that recognize organelle-specific phosphoinositides and bind to ER-localized membrane proteins.
Phosphoinositides control MCS formation
A key organizing principle of intracellular membranes is that phosphoinositides in the membrane of one organelle recruit a specific phosphoinositide-binding protein anchored in the membrane of another organelle, leading to the formation of MCSs (Fig. 4a). A prime example is MCSs between the ER and the plasma membrane via extended synaptotagmins (E-Syts) and related tethers. E-Syts are ER membrane proteins that interact with the plasma membrane via their Ca2+- and lipid-binding C2 domains by specific recognition of PtdIns(4,5)P2 in trans (Fig. 4a, number 1). This binding occurs at resting Ca2+ concentrations and is facilitated by elevated Ca2+ levels, which trigger store-operated Ca2+ entry via association of the ER protein STIM1 with the plasma membrane Ca2+ channel ORAI158. STIM1–ORAI and E-Syts are part of a Ca2+-regulated signalling shunt159. In this shunt PtdIns(4,5)P2 facilitates E-Syt-mediated MCS formation, while simultaneously being hydrolysed by PLC to DAG and Ins(1,4,5)P3. Ins(1,4,5)P3 then activates Ca2+-conducting Ins(1,4,5)P3 receptors in the ER. E-Syts can transfer DAG from the plasma membrane to the ER160, where it may serve as a precursor for subsequent resynthesis of PtdIns. As E-Syts are dispensable for the formation and maintenance of ER–plasma membrane contacts2, it is likely that their function is compensated by other tethering factors, including the ER membrane-localized, PH-like GRAM domain protein GRAMD2A, TMEM24 (also known as C2CD2L), an ER protein that is concentrated at ER–plasma membrane MCSs via association with acidic plasma membrane lipids and that transfers PtdIns to the plasma membrane in stimulated pancreatic β-cells161, and/or ORP5 and ORP8, lipid transfer proteins that associate with plasma membrane PtdIns4P162 and PtdIns(4,5)P2 (ref.163).
Similar phosphoinositide-dependent mechanisms govern the formation of MCSs between the ER and the TGN and late endosomes or lysosomes, respectively2,3. At MCSs between the ER and the TGN, OSBP and related lipid transfer proteins (for example, ceramide transfer protein (CERT)164), coincidentally associate via their PH domains with PtdIns4P on the TGN and ER-localized VAMP-associated protein (VAP)2 (Fig. 4a, number 2). MCSs between the ER and late endosomes or lysosomes likely involve multiple phosphoinositide-binding proteins. One example is the recognition of lysosomal PtdIns3P by the FYVE domain-containing ER membrane protein protrudin and its associated factor PDZ domain-containing protein 8 (ref.165). Protrudin enables the transfer of kinesin 1 from the ER to lysosomes to facilitate lysosome dispersion136 and nutrient signalling. Moreover, OSBP166 and ORP1L associate with VAPs in the ER and with lysosomal phosphoinositides142 to regulate cholesterol homeostasis (see later) (Fig. 4a, numbers 3,4). As mentioned earlier, during autophagy PtdIns3P serves to tether early autophagic membranes to the ER via association with the PROPPIN domain of WIPI4, which in turn binds to the lipid transfer protein ATG2 to facilitate lipid transfer required for phagophore expansion116 (Fig. 4a, number 5).
Lipid transfer at MCSs
A major function of MCSs is to enable lipid flux between compartments mediated by lipid transfer proteins2. Vectorial transfer of lipids at MCSs can either follow their natural concentration gradient established by compartmentalized synthesis or be powered by the countertransport or consumption of another lipid (Fig. 4b).
PtdIns4P is synthesized mainly at the Golgi complex and the plasma membrane via specific PI4Ks, while its hydrolysis is largely mediated by the ER-localized 4-phosphatase SAC1 (ref.4). The resulting PtdIns4P gradient between the Golgi complex and the ER is utilized to power the export of lipids such as cholesterol from the ER, the main site of lipid synthesis, to the TGN and further to the cell surface. In this mechanism, OSBP connects the ER to PtdIns4P in the TGN through its FFAT motif and PH domain to enable the transfer of cholesterol between the apposed membranes via its OSBP-related domain. Concurrently, PtdIns4P is transported from the TGN to the ER, where it is hydrolysed to PtdIns by SAC1 (ref.167) (Fig. 4b, number 1). OSBP thus resembles a ferry bridge for the countertransport of lipids. The related CERT164 and PtdIns4P adaptor protein 2 (FAPP2)168 facilitate the vectorial transfer of ceramide (from the ER) and glucosylceramide (from the cis-Golgi network) to the TGN. CERT gain of function and the expected increase in ceramide transport is thought to underlie a specific form of severe intellectual disability169.
Similar PtdIns4P-based mechanisms also operate at other ER-based MCSs3. These include VAP–OSBP-based MCSs between the ER and lysosomes that fuel cholesterol transport to the limiting membrane of the lysosome to facilitate nutrient signalling166 and ER–endosome MCSs to control retromer–WASH-mediated retrograde membrane traffic from endosomes to the TGN99. Endosomal PtdIns4P also drives the countertransport of PtdSer from the ER to endosomes via the OSBP-related ORP10 (Fig. 4b, number 2), thereby promoting endosome fission by the dynamin-related ATPase EHD1 (ref.170). Moreover, the OSBP-related proteins ORP5 and ORP8 bind to PtdIns4P162 and PtdIns(4,5)P2 (ref.163) in the plasma membrane to countertransport PtdSer to the cell surface (Fig. 4b, number 3). Finally, PtdIns4P171 and its downstream products PtdIns(3,4)P2 and PtdIns(4,5)P2 (ref.142) have been postulated to enable the reverse flux of internalized cholesterol from lysosomes or phagolysosomes172 to the ER via ORP1L (Fig. 4a, number 4).
Recent data indicate that the vectorial transfer of lipids at MCSs is subject to regulation by cell signalling and metabolism. In yeast, starvation-induced cytosol acidification alters the protonation state of the head group of PtdIns4P, thereby impairing Osh1/OSBP-mediated sterol transfer and protein sorting at the TGN34. In mammalian cells, Ins(1,4,5)P3-triggered calcium efflux from ER stores downstream of receptor signalling causes OSBP to dissociate from the TGN, resulting in the depletion of cholesterol and associated glycolipids from the cell surface173, while promoting the formation of E-Syt-based MCSs between the ER and the plasma membrane158. Hence, distinct MCSs may be subject to antagonistically controlled mechanisms of regulation.
Such regulatory control of lipid flux across MCSs is critical for the integration of phosphoinositide metabolism with cell signalling. In response to the activation of PLC-coupled receptors, PtdIns(4,5)P2 is hydrolysed to DAG and Ins(1,4,5)P3, resulting in PtdIns(4,5)P2 depletion from the plasma membrane. To maintain phosphoinositide signalling competence during sustained agonist-induced PLC activation, the lipid transfer protein Nir2 is recruited to MCSs between the ER and the plasma membrane. At these VAP-based MCSs, Nir2 catalyses the transport of PtdIns from the ER, its site of synthesis, to the plasma membrane, while transferring PtdOH to the ER. PtdIns–PtdOH exchange between the ER and the cell surface then fuels the resynthesis of PtdIns4P from PtdIns to enable replenishment of PtdIns(4,5)P2, thereby ensuring sustained responsiveness to agonist stimulation174. Nir2 also facilitates the replenishment of the Golgi PtdIns pool to enable continued synthesis of PtdIns4P required to maintain the PtdIns4P–cholesterol cycle between these organelles175 (Fig. 4b, number 4). This pathway is highjacked by replicating hepatitis C virus to sustain viral replication during chronic infection by inducing the formation of OSBP–VAP-based as well as Nir2–VAP-based MCSs between hepatitis C virus-derived replication organelles and the host ER membrane176 (Supplementary Box 1). In pancreatic β-cells, glucose stimulation causes loss of PtdIns(4,5)P2 as a result of glucose-stimulated PLC activation. To replenish PtdIns(4,5)P2, its precursor PtdIns is transported from the ER to the plasma membrane by the lipid ER-anchored transport protein TMEM24 at MCSs (Fig. 4b, number 5). The activity of TMEM24 is controlled by Ca2+-regulated kinases and phosphatases that couple TMEM24-mediated PtdIns transfer to pulsatile insulin secretion161.
Finally, so far elusive MCSs enable ORP2-mediated cholesterol transfer from late endosomes to integrin- and focal adhesion kinase (FAK)-containing recycling endosomes. Cholesterol transfer promotes recycling endosomal synthesis of PtdIns(4,5)P2, FAK activation and, thereby, cell adhesion177.
Conclusions and perspectives
Phosphoinositides are involved in nearly all aspects of membrane function and dynamics. The development of new technologies to monitor and manipulate phosphoinositides (Box 1) has yielded insights into their nanoscale localization, turnover and dynamics as well as their reciprocal regulation by cell signalling. Hitherto unknown roles of phosphoinositides in the reorganization of organellar contacts, including the discovery of contacts between three organelles2, have emerged. For example, the fission of mitochondria178 or endosomes179 has been shown to be controlled by contacts of these organelles with PtdIns4P-containing vesicles and the ER.
We continue to discover new functions of phosphoinositide-metabolizing enzymes in human diseases ranging from cancer and metabolic disorders to rare inherited diseases caused by loss or gain of function of phosphoinositide kinases (Table 3) and phosphatases (Table 2) as well as their effector proteins and regulators (for example, lipid transfer proteins). Emerging concepts of the mechanisms that spatiotemporally control phosphoinositide conversion on the nanoscale not only provide us with a glimpse on the inner workings of the cell but also enable the development of novel treatment options for diseases linked to defective phosphoinositide metabolism. For example, pharmacological inhibition of PI3Ks such as PI3KC2β180 may serve as a therapy for patients with X-linked centronuclear myopathy (Table 2) caused by dysfunction of the PtdIns 3-phosphatase MTM1 (ref.41). The prominent role of PtdIns kinases and phosphatases in bacterial infection181 and in the replication of viruses, including severe acute respiratory syndrome coronavirus 2 (ref.182), suggests that specific inhibitors of these enzymes (for example, the PIKfyve inhibitor apilimod183) could serve as novel antibiotics or antiviral drugs to combat the rise of infectious diseases (Supplementary Box 1).
In spite of this progress, major questions related to the function of phosphoinositides in membrane dynamics remain unsolved. Our quantitative understanding of phosphoinositide biology especially with respect to transient and precarious lipid species (for example, PtdIns(3,4)P2 and PtdIns(3,5)P2) remains limited, largely owing to the lack of tools (Box 1) to determine the absolute concentrations of lipids in living cells and with subcellular resolution. How phosphoinositides are regulated by metabolism and, conversely, impinge on metabolic regulation and the intracellular flux of metabolites at the levels of cells, tissues or entire organisms remains largely enigmatic. Such information seems crucial as membrane traffic and contacts between organelles need to be adapted to cellular nutrient status (for example, to regulate metabolic flux of lipids or fatty acids). Conversely, many phosphoinositide kinases and phosphatases and their effectors are known to change their subcellular localization and/or activity in response to external and internal metabolic cues, identifying them as prime candidates for the metabolic rewiring of cell membranes and organelles130,134,155,184,185,186,187. New analytical tools to visualize phosphoinositide lipids in vivo and the development of CRISPR technology to tag endogenous lipid-modifying enzymes and phosphoinositide-binding effector proteins paired with correlative live microscopy approaches will help to resolve these issues.
References
Behnia, R. & Munro, S. Organelle identity and the signposts for membrane traffic. Nature 438, 597–604 (2005).
Prinz, W. A., Toulmay, A. & Balla, T. The functional universe of membrane contact sites. Nat. Rev. Mol. Cell Biol. 21, 7–24 (2020).
Wu, H., Carvalho, P. & Voeltz, G. K. Here, there, and everywhere: the importance of ER membrane contact sites. Science https://doi.org/10.1126/science.aan5835 (2018).
Balla, T. Phosphoinositides: tiny lipids with giant impact on cell regulation. Physiol. Rev. 93, 1019–1137 (2013).
Bilanges, B., Posor, Y. & Vanhaesebroeck, B. PI3K isoforms in cell signalling and vesicle trafficking. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-019-0129-z (2019).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Gillooly, D. J. et al. Localization of phosphatidylinositol 3-phosphate in yeast and mammalian cells. EMBO J. 19, 4577–4588 (2000).
Staiano, L., De Leo, M. G., Persico, M. & De Matteis, M. A. Mendelian disorders of PI metabolizing enzymes. Biochim. Biophys. Acta 1851, 867–881 (2015).
Goncalves, M. D., Hopkins, B. D. & Cantley, L. C. Phosphatidylinositol 3-kinase, growth disorders, and cancer. N. Engl. J. Med. 379, 2052–2062 (2018).
Falkenburger, B. H., Jensen, J. B., Dickson, E. J., Suh, B. C. & Hille, B. Phosphoinositides: lipid regulators of membrane proteins. J. Physiol. 588, 3179–3185 (2010).
Hille, B., Dickson, E. J., Kruse, M., Vivas, O. & Suh, B. C. Phosphoinositides regulate ion channels. Biochim. Biophys. Acta 1851, 844–856 (2015).
Vanhaesebroeck, B., Perry, M. W. D., Brown, J. R., Andre, F. & Okkenhaug, K. PI3K inhibitors are finally coming of age. Nat. Rev. Drug Discov. 20, 741–769 (2021).
Larijani, B., Pytowski, L. & Vaux, D. J. The enigma of phosphoinositides and their derivatives: their role in regulation of subcellular compartment morphology. Biochim. Biophys. Acta Biomembr. 1864, 183780 (2022).
Hokin, M. R. & Hokin, L. E. Enzyme secretion and the incorporation of P32 into phospholipides of pancreas slices. J. Biol. Chem. 203, 967–977 (1953).
Schu, P. V. et al. Phosphatidylinositol 3-kinase encoded by yeast VPS34 gene essential for protein sorting. Science 260, 88–91 (1993).
Godi, A. et al. ARF mediates recruitment of PtdIns-4-OH kinase-beta and stimulates synthesis of PtdIns(4,5)P2 on the Golgi complex. Nat. Cell Biol. 1, 280–287 (1999).
Sasaki, J., Ishikawa, K., Arita, M. & Taniguchi, K. ACBD3-mediated recruitment of PI4KB to picornavirus RNA replication sites. EMBO J. 31, 754–766 (2012).
Blumental-Perry, A. et al. Phosphatidylinositol 4-phosphate formation at ER exit sites regulates ER export. Dev. Cell 11, 671–682 (2006).
Szentpetery, Z., Varnai, P. & Balla, T. Acute manipulation of Golgi phosphoinositides to assess their importance in cellular trafficking and signaling. Proc. Natl Acad. Sci. USA 107, 8225–8230 (2010).
Wang, Y. J. et al. Phosphatidylinositol 4 phosphate regulates targeting of clathrin adaptor AP-1 complexes to the Golgi. Cell 114, 299–310 (2003).
Wieffer, M. et al. PI4K2beta/AP-1-based TGN-endosomal sorting regulates Wnt signaling. Curr. Biol. 23, 2185–2190 (2013).
Daboussi, L., Costaguta, G., Ghukasyan, R. & Payne, G. S. Conserved role for Gga proteins in phosphatidylinositol 4-kinase localization to the trans-Golgi network. Proc. Natl Acad. Sci. USA 114, 3433–3438 (2017).
Craige, B., Salazar, G. & Faundez, V. Phosphatidylinositol-4-kinase type II alpha contains an AP-3-sorting motif and a kinase domain that are both required for endosome traffic. Mol. Biol. Cell 19, 1415–1426 (2008).
Rahajeng, J. et al. Efficient Golgi forward trafficking requires GOLPH3-driven, PI4P-dependent membrane curvature. Dev. Cell 50, 573–585 e575 (2019).
De Matteis, M. A. & Luini, A. Exiting the Golgi complex. Nat. Rev. Mol. Cell Biol. 9, 273–284 (2008).
Mahajan, D., Tie, H. C., Chen, B. & Lu, L. Dopey1-Mon2 complex binds to dual-lipids and recruits kinesin-1 for membrane trafficking. Nat. Commun. 10, 3218 (2019).
Valente, C. et al. A 14-3-3gamma dimer-based scaffold bridges CtBP1-S/BARS to PI(4)KIIIbeta to regulate post-Golgi carrier formation. Nat. Cell Biol. 14, 343–354 (2012).
Liljedahl, M. et al. Protein kinase D regulates the fission of cell surface destined transport carriers from the trans-Golgi network. Cell 104, 409–420 (2001).
Baron, C. L. & Malhotra, V. Role of diacylglycerol in PKD recruitment to the TGN and protein transport to the plasma membrane. Science 295, 325–328 (2002).
Hausser, A. et al. Protein kinase D regulates vesicular transport by phosphorylating and activating phosphatidylinositol-4 kinase IIIbeta at the Golgi complex. Nat. Cell Biol. 7, 880–886 (2005).
Lu, D. et al. Phosphatidylinositol 4-kinase IIalpha is palmitoylated by Golgi-localized palmitoyltransferases in cholesterol-dependent manner. J. Biol. Chem. 287, 21856–21865 (2012).
Baba, T. et al. Myelination of peripheral nerves is controlled by PI4KB through regulation of Schwann cell Golgi function. Proc. Natl Acad. Sci. USA 117, 28102–28113 (2020).
Blagoveshchenskaya, A. et al. Integration of Golgi trafficking and growth factor signaling by the lipid phosphatase SAC1. J. Cell Biol. 180, 803–812 (2008).
Shin, J. J. H. et al. pH biosensing by PI4P regulates cargo sorting at the TGN. Dev. Cell 52, 461–476 e464 (2020).
Milosevic, I. et al. Plasmalemmal phosphatidylinositol-4,5-bisphosphate level regulates the releasable vesicle pool size in chromaffin cells. J. Neurosci. 25, 2557–2565 (2005).
Walter, A. M. et al. Phosphatidylinositol 4,5-bisphosphate optical uncaging potentiates exocytosis. eLife https://doi.org/10.7554/eLife.30203 (2017).
Chen, Y. et al. Synaptotagmin-1 interacts with PI(4,5)P2 to initiate synaptic vesicle docking in hippocampal neurons. Cell Rep. 34, 108842 (2021).
He, B., Xi, F., Zhang, X., Zhang, J. & Guo, W. Exo70 interacts with phospholipids and mediates the targeting of the exocyst to the plasma membrane. EMBO J. 26, 4053–4065 (2007).
Nishida-Fukuda, H. The exocyst: dynamic machine or static tethering complex? Bioessays 41, e1900056 (2019).
Mizuno-Yamasaki, E., Medkova, M., Coleman, J. & Novick, P. Phosphatidylinositol 4-phosphate controls both membrane recruitment and a regulatory switch of the Rab GEF Sec2p. Dev. Cell 18, 828–840 (2010).
Ketel, K. et al. A phosphoinositide conversion mechanism for exit from endosomes. Nature 529, 408–412 (2016). This work identifies an endosomal mechanism of PtdIns3P-to-PtdIns4P conversion to enable exocytosis that involves a lipid phosphatase mutated in a human muscle disease.
Guo, J. et al. Phosphatidylinositol 4-kinase type IIalpha is responsible for the phosphatidylinositol 4-kinase activity associated with synaptic vesicles. Proc. Natl Acad. Sci. USA 100, 3995–4000 (2003).
Hammond, G. R. et al. PI4P and PI(4,5)P2 are essential but independent lipid determinants of membrane identity. Science 337, 727–730 (2012). This study reports on an important function of PtdIns4P at the plasma membrane that is independent of its role as a precursor for PtdIns(4,5)P2.
Nakatsu, F. et al. PtdIns4P synthesis by PI4KIIIalpha at the plasma membrane and its impact on plasma membrane identity. J. Cell Biol. 199, 1003–1016 (2012).
Baskin, J. M. et al. The leukodystrophy protein FAM126A (hyccin) regulates PtdIns(4)P synthesis at the plasma membrane. Nat. Cell Biol. 18, 132–138 (2016).
Marat, A. L. & Haucke, V. Phosphatidylinositol 3-phosphates-at the interface between cell signalling and membrane traffic. EMBO J. 35, 561–579 (2016).
Kaksonen, M. & Roux, A. Mechanisms of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 19, 313–326 (2018).
Zoncu, R. et al. Loss of endocytic clathrin-coated pits upon acute depletion of phosphatidylinositol 4,5-bisphosphate. Proc. Natl Acad. Sci. USA 104, 3793–3798 (2007).
Posor, Y., Eichhorn-Grunig, M. & Haucke, V. Phosphoinositides in endocytosis. Biochim. Biophys. Acta 1851, 794–804 (2015).
Kelly, B. T. et al. Clathrin adaptors. AP2 controls clathrin polymerization with a membrane-activated switch. Science 345, 459–463 (2014).
Hollopeter, G. et al. The membrane-associated proteins FCHo and SGIP are allosteric activators of the AP2 clathrin adaptor complex. eLife https://doi.org/10.7554/eLife.03648 (2014).
Lehmann, M. et al. Nanoscale coupling of endocytic pit growth and stability. Sci. Adv. 5, eaax5775 (2019).
Krauss, M., Kukhtina, V., Pechstein, A. & Haucke, V. Stimulation of phosphatidylinositol kinase type I-mediated phosphatidylinositol (4,5)-bisphosphate synthesis by AP-2mu-cargo complexes. Proc. Natl Acad. Sci. USA 103, 11934–11939 (2006).
Thieman, J. R. et al. Clathrin regulates the association of PIPKIgamma661 with the AP-2 adaptor beta2 appendage. J. Biol. Chem. 284, 13924–13939 (2009).
Posor, Y. et al. Spatiotemporal control of endocytosis by phosphatidylinositol-3,4-bisphosphate. Nature 499, 233–237 (2013).
Perera, R. M., Zoncu, R., Lucast, L., De Camilli, P. & Toomre, D. Two synaptojanin 1 isoforms are recruited to clathrin-coated pits at different stages. Proc. Natl Acad. Sci. USA 103, 19332–19337 (2006).
Nakatsu, F. et al. The inositol 5-phosphatase SHIP2 regulates endocytic clathrin-coated pit dynamics. J. Cell Biol. 190, 307–315 (2010).
Lo, W. T. et al. Structural basis of phosphatidylinositol 3-kinase C2alpha function. Nat. Struct. Mol. Biol. 29, 218–228 (2022).
Wang, H. et al. Phosphatidylinositol 3,4-bisphosphate synthesis and turnover are spatially segregated in the endocytic pathway. J. Biol. Chem. 295, 1091–1104 (2020).
Gulluni, F. et al. PI(3,4)P2-mediated cytokinetic abscission prevents early senescence and cataract formation. Science 374, eabk0410 (2021).
Lo, W. T. et al. A coincidence detection mechanism controls PX-BAR domain-mediated endocytic membrane remodeling via an allosteric structural switch. Dev. Cell 43, 522–529 e524 (2017).
Schoneberg, J. et al. Lipid-mediated PX-BAR domain recruitment couples local membrane constriction to endocytic vesicle fission. Nat. Commun. 8, 15873 (2017).
Almeida-Souza, L. et al. A flat BAR protein promotes actin polymerization at the base of clathrin-coated pits. Cell 174, 325–337 e314 (2018).
Reubold, T. F. et al. Crystal structure of the dynamin tetramer. Nature 525, 404–408 (2015).
Bethoney, K. A., King, M. C., Hinshaw, J. E., Ostap, E. M. & Lemmon, M. A. A possible effector role for the pleckstrin homology (PH) domain of dynamin. Proc. Natl Acad. Sci. USA 106, 13359–13364 (2009).
Jost, M., Simpson, F., Kavran, J. M., Lemmon, M. A. & Schmid, S. L. Phosphatidylinositol-4,5-bisphosphate is required for endocytic coated vesicle formation. Curr. Biol. 8, 1399–1402 (1998).
Milosevic, I. et al. Recruitment of endophilin to clathrin-coated pit necks is required for efficient vesicle uncoating after fission. Neuron 72, 587–601 (2011).
Cauvin, C. et al. Rab35 GTPase triggers switch-like recruitment of the Lowe syndrome lipid phosphatase OCRL on newborn endosomes. Curr. Biol. 26, 120–128 (2016).
Nandez, R. et al. A role of OCRL in clathrin-coated pit dynamics and uncoating revealed by studies of Lowe syndrome cells. eLife 3, e02975 (2014).
He, K. et al. Dynamics of auxilin 1 and GAK in clathrin-mediated traffic. J. Cell Biol. https://doi.org/10.1083/jcb.201908142 (2020).
Massol, R. H., Boll, W., Griffin, A. M. & Kirchhausen, T. A burst of auxilin recruitment determines the onset of clathrin-coated vesicle uncoating. Proc. Natl Acad. Sci. USA 103, 10265–10270 (2006).
Shin, H. W. et al. An enzymatic cascade of Rab5 effectors regulates phosphoinositide turnover in the endocytic pathway. J. Cell Biol. 170, 607–618 (2005).
He, K. et al. Dynamics of phosphoinositide conversion in clathrin-mediated endocytic traffic. Nature 552, 410–414 (2017).
Goulden, B. D. et al. A high-avidity biosensor reveals plasma membrane PI(3,4)P2 is predominantly a class I PI3K signaling product. J. Cell Biol. 218, 1066–1079 (2019).
Sigismund, S., Lanzetti, L., Scita, G. & Di Fiore, P. P. Endocytosis in the context-dependent regulation of individual and collective cell properties. Nat. Rev. Mol. Cell Biol. 22, 625–643 (2021).
Boucrot, E. et al. Endophilin marks and controls a clathrin-independent endocytic pathway. Nature 517, 460–465 (2015).
Krause, M. et al. Lamellipodin, an Ena/VASP ligand, is implicated in the regulation of lamellipodial dynamics. Dev. Cell 7, 571–583 (2004).
Chan Wah Hak, L. et al. FBP17 and CIP4 recruit SHIP2 and lamellipodin to prime the plasma membrane for fast endophilin-mediated endocytosis. Nat. Cell Biol. 20, 1023–1031 (2018).
Walpole, G. F. W. & Grinstein, S. Endocytosis and the internalization of pathogenic organisms: focus on phosphoinositides. F1000Res https://doi.org/10.12688/f1000research.22393.1 (2020).
Botelho, R. J. et al. Localized biphasic changes in phosphatidylinositol-4,5-bisphosphate at sites of phagocytosis. J. Cell Biol. 151, 1353–1368 (2000).
Schlam, D. et al. Phosphoinositide 3-kinase enables phagocytosis of large particles by terminating actin assembly through Rac/Cdc42 GTPase-activating proteins. Nat. Commun. 6, 8623 (2015).
Ostrowski, P. P., Freeman, S. A., Fairn, G. & Grinstein, S. Dynamic podosome-like structures in nascent phagosomes are coordinated by phosphoinositides. Dev. Cell 50, 397–410 e393 (2019).
Maekawa, M. et al. Sequential breakdown of 3-phosphorylated phosphoinositides is essential for the completion of macropinocytosis. Proc. Natl Acad. Sci. USA 111, E978–E987 (2014).
Schink, K. O. et al. The phosphoinositide coincidence detector Phafin2 promotes macropinocytosis by coordinating actin organisation at forming macropinosomes. Nat. Commun. 12, 6577 (2021).
Zeigerer, A. et al. Rab5 is necessary for the biogenesis of the endolysosomal system in vivo. Nature 485, 465–470 (2012).
Semerdjieva, S. et al. Coordinated regulation of AP2 uncoating from clathrin-coated vesicles by rab5 and hRME-6. J. Cell Biol. 183, 499–511 (2008).
Tremel, S. et al. Structural basis for VPS34 kinase activation by Rab1 and Rab5 on membranes. Nat. Commun. 12, 1564 (2021).
Wandinger-Ness, A. & Zerial, M. Rab proteins and the compartmentalization of the endosomal system. Cold Spring Harb. Perspect. Biol. 6, a022616 (2014).
Liu, H. et al. The INPP4B tumor suppressor modulates EGFR trafficking and promotes triple-negative breast cancer. Cancer Discov. 10, 1226–1239 (2020).
Aki, S., Yoshioka, K., Takuwa, N. & Takuwa, Y. TGFbeta receptor endocytosis and Smad signaling require synaptojanin1, PI3K-C2alpha-, and INPP4B-mediated phosphoinositide conversions. Mol. Biol. Cell 31, 360–372 (2020).
Norris, A. et al. SNX-1 and RME-8 oppose the assembly of HGRS-1/ESCRT-0 degradative microdomains on endosomes. Proc. Natl Acad. Sci. USA 114, E307–E316 (2017).
Cullen, P. J. & Steinberg, F. To degrade or not to degrade: mechanisms and significance of endocytic recycling. Nat. Rev. Mol. Cell Biol. 19, 679–696 (2018).
Weeratunga, S., Paul, B. & Collins, B. M. Recognising the signals for endosomal trafficking. Curr. Opin. Cell Biol. 65, 17–27 (2020).
Ghai, R. et al. Phox homology band 4.1/ezrin/radixin/moesin-like proteins function as molecular scaffolds that interact with cargo receptors and Ras GTPases. Proc. Natl Acad. Sci. USA 108, 7763–7768 (2011).
Simonetti, B. et al. Molecular identification of a BAR domain-containing coat complex for endosomal recycling of transmembrane proteins. Nat. Cell Biol. 21, 1219–1233 (2019).
Singla, A. et al. Endosomal PI(3)P regulation by the COMMD/CCDC22/CCDC93 (CCC) complex controls membrane protein recycling. Nat. Commun. 10, 4271 (2019).
Campa, C. C. et al. Rab11 activity and PtdIns(3)P turnover removes recycling cargo from endosomes. Nat. Chem. Biol. 14, 801–810 (2018).
Franco, I. et al. PI3K class II alpha controls spatially restricted endosomal PtdIns3P and Rab11 activation to promote primary cilium function. Dev. Cell 28, 647–658 (2014).
Dong, R. et al. Endosome-ER contacts control actin nucleation and retromer function through VAP-dependent regulation of PI4P. Cell 166, 408–423 (2016).
Vietri, M., Radulovic, M. & Stenmark, H. The many functions of ESCRTs. Nat. Rev. Mol. Cell Biol. 21, 25–42 (2020).
Raiborg, C. et al. FYVE and coiled-coil domains determine the specific localisation of Hrs to early endosomes. J. Cell Sci. 114, 2255–2263 (2001).
Poteryaev, D., Datta, S., Ackema, K., Zerial, M. & Spang, A. Identification of the switch in early-to-late endosome transition. Cell 141, 497–508 (2010).
Borchers, A. C., Langemeyer, L. & Ungermann, C. Who’s in control? Principles of Rab GTPase activation in endolysosomal membrane trafficking and beyond. J. Cell Biol. https://doi.org/10.1083/jcb.202105120 (2021).
de Lartigue, J. et al. PIKfyve regulation of endosome-linked pathways. Traffic 10, 883–893 (2009).
Hasegawa, J., Strunk, B. S. & Weisman, L. S. PI5P and PI(3,5)P2: minor, but essential phosphoinositides. Cell Struct. Funct. 42, 49–60 (2017).
Lawrence, R. E. & Zoncu, R. The lysosome as a cellular centre for signalling, metabolism and quality control. Nat. Cell Biol. 21, 133–142 (2019).
Ebner, M., Koch, P. A. & Haucke, V. Phosphoinositides in the control of lysosome function and homeostasis. Biochem. Soc. Trans. 47, 1173–1185 (2019).
Saha, S., Panigrahi, D. P., Patil, S. & Bhutia, S. K. Autophagy in health and disease: a comprehensive review. Biomed. Pharmacother. 104, 485–495 (2018).
Axe, E. L. et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182, 685–701 (2008).
Palamiuc, L., Ravi, A. & Emerling, B. M. Phosphoinositides in autophagy: current roles and future insights. FEBS J. 287, 222–238 (2020).
Judith, D. et al. ATG9A shapes the forming autophagosome through Arfaptin 2 and phosphatidylinositol 4-kinase IIIbeta. J. Cell Biol. 218, 1634–1652 (2019).
Mercer, T. J. et al. Phosphoproteomic identification of ULK substrates reveals VPS15-dependent ULK/VPS34 interplay in the regulation of autophagy. EMBO J. 40, e105985 (2021).
Boukhalfa, A. et al. PI3KC2alpha-dependent and VPS34-independent generation of PI3P controls primary cilium-mediated autophagy in response to shear stress. Nat. Commun. 11, 294 (2020).
Dooley, H. C. et al. WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1. Mol. Cell 55, 238–252 (2014).
Fracchiolla, D., Chang, C., Hurley, J. H. & Martens, S. A PI3K-WIPI2 positive feedback loop allosterically activates LC3 lipidation in autophagy. J. Cell Biol. https://doi.org/10.1083/jcb.201912098 (2020).
Bakula, D. et al. WIPI3 and WIPI4 beta-propellers are scaffolds for LKB1-AMPK-TSC signalling circuits in the control of autophagy. Nat. Commun. 8, 15637 (2017).
Raess, M. A., Friant, S., Cowling, B. S. & Laporte, J. WANTED - dead or alive: myotubularins, a large disease-associated protein family. Adv. Biol. Regul. 63, 49–58 (2017).
Xu, H. & Ren, D. Lysosomal physiology. Annu. Rev. Physiol. 77, 57–80 (2015).
Noda, T., Matsunaga, K., Taguchi-Atarashi, N. & Yoshimori, T. Regulation of membrane biogenesis in autophagy via PI3P dynamics. Semin. Cell Dev. Biol. 21, 671–676 (2010).
Hasegawa, J. et al. Autophagosome-lysosome fusion in neurons requires INPP5E, a protein associated with Joubert syndrome. EMBO J. 35, 1853–1867 (2016).
Hong, N. H., Qi, A. & Weaver, A. M. PI(3,5)P2 controls endosomal branched actin dynamics by regulating cortactin-actin interactions. J. Cell Biol. 210, 753–769 (2015).
Dong, X. P. et al. PI(3,5)P2 controls membrane trafficking by direct activation of mucolipin Ca2+ release channels in the endolysosome. Nat. Commun. 1, 38 (2010).
Wang, X. et al. TPC proteins are phosphoinositide- activated sodium-selective ion channels in endosomes and lysosomes. Cell 151, 372–383 (2012).
Akizu, N. et al. Biallelic mutations in SNX14 cause a syndromic form of cerebellar atrophy and lysosome-autophagosome dysfunction. Nat. Genet. 47, 528–534 (2015).
Bielas, S. L. et al. Mutations in INPP5E, encoding inositol polyphosphate-5-phosphatase E, link phosphatidyl inositol signaling to the ciliopathies. Nat. Genet. 41, 1032–1036 (2009).
Baba, T., Toth, D. J., Sengupta, N., Kim, Y. J. & Balla, T. Phosphatidylinositol 4,5-bisphosphate controls Rab7 and PLEKMH1 membrane cycling during autophagosome-lysosome fusion. EMBO J. (2019).
Perera, R. M., Di Malta, C. & Ballabio, A. MiT/TFE family of transcription factors, lysosomes, and cancer. Annu. Rev. Cancer Biol. 3, 203–222 (2019).
De Leo, M. G. et al. Autophagosome-lysosome fusion triggers a lysosomal response mediated by TLR9 and controlled by OCRL. Nat. Cell Biol. 18, 839–850 (2016). This important study uncovers a lysosomal cargo response required to sustain autophagic flux via local confinement of PtdIns(4,5)P2 by the lipid phosphatase OCRL.
Lundquist, M. R. et al. Phosphatidylinositol-5-phosphate 4-kinases regulate cellular lipid metabolism by facilitating autophagy. Mol. Cell 70, 531–544 e539 (2018).
Marat, A. L. et al. mTORC1 activity repression by late endosomal phosphatidylinositol 3,4-bisphosphate. Science 356, 968–972 (2017). This work identifies class II PI3K-mediated synthesis of PtdIns(3,4)P2 as a local repressor of mTORC1 signalling on lysosomes in response to cessation of growth factor signalling.
Li, S. C. et al. The signaling lipid PI(3,5)P(2) stabilizes V(1)-V(o) sector interactions and activates the V-ATPase. Mol. Biol. Cell 25, 1251–1262 (2014).
Wallroth, A. & Haucke, V. Phosphoinositide conversion in endocytosis and the endolysosomal system. J. Biol. Chem. https://doi.org/10.1074/jbc.R117.000629 (2017).
Nobukuni, T. et al. Amino acids mediate mTOR/raptor signaling through activation of class 3 phosphatidylinositol 3OH-kinase. Proc. Natl Acad. Sci. USA 102, 14238–14243 (2005).
Yoon, M. S., Du, G., Backer, J. M., Frohman, M. A. & Chen, J. Class III PI-3-kinase activates phospholipase D in an amino acid-sensing mTORC1 pathway. J. Cell Biol. 195, 435–447 (2011).
Hong, Z. et al. PtdIns3P controls mTORC1 signaling through lysosomal positioning. J. Cell Biol. 216, 4217–4233 (2017).
Raiborg, C. et al. Repeated ER–endosome contacts promote endosome translocation and neurite outgrowth. Nature 520, 234–238 (2015). This study identifies a mechanism for late endosome/lysosome dispersion via membrane contacts with the ER-localized kinesin adaptor protrudin, FYCO1 and PtdIns3P.
Tsuruta, F. & Dolmetsch, R. E. PIKfyve mediates the motility of late endosomes and lysosomes in neuronal dendrites. Neurosci. Lett. 605, 18–23 (2015).
Sawade, L. et al. Rab35-regulated lipid turnover by myotubularins represses mTORC1 activity and controls myelin growth. Nat. Commun. 11, 2835 (2020).
Bridges, D. et al. Phosphatidylinositol 3,5-bisphosphate plays a role in the activation and subcellular localization of mechanistic target of rapamycin 1. Mol. Biol. Cell 23, 2955–2962 (2012).
Fitzian, K. et al. TSC1 binding to lysosomal PIPs is required for TSC complex translocation and mTORC1 regulation. Mol. Cell 81, 2705–2721 e2708 (2021).
Gozzelino, L. et al. Defective lipid signaling caused by mutations in PIK3C2B underlies focal epilepsy. Brain (in press, 2022).
Dong, J. et al. Allosteric enhancement of ORP1-mediated cholesterol transport by PI(4,5)P2/PI(3,4)P2. Nat. Commun. 10, 829 (2019).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9-Niemann-Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Wallroth, A., Koch, P. A., Marat, A. L., Krause, E. & Haucke, V. Protein kinase N controls a lysosomal lipid switch to facilitate nutrient signalling via mTORC1. Nat. Cell Biol. 21, 1093–1101 (2019).
Guardia, C. M., Farias, G. G., Jia, R., Pu, J. & Bonifacino, J. S. BORC functions upstream of kinesins 1 and 3 to coordinate regional movement of lysosomes along different microtubule tracks. Cell Rep. 17, 1950–1961 (2016).
Rosa-Ferreira, C. & Munro, S. Arl8 and SKIP act together to link lysosomes to kinesin-1. Dev. Cell 21, 1171–1178 (2011).
Cantalupo, G., Alifano, P., Roberti, V., Bruni, C. B. & Bucci, C. Rab-interacting lysosomal protein (RILP): the Rab7 effector required for transport to lysosomes. EMBO J. 20, 683–693 (2001).
Jordens, I. et al. The Rab7 effector protein RILP controls lysosomal transport by inducing the recruitment of dynein-dynactin motors. Curr. Biol. 11, 1680–1685 (2001).
Marwaha, R. et al. The Rab7 effector PLEKHM1 binds Arl8b to promote cargo traffic to lysosomes. J. Cell Biol. 216, 1051–1070 (2017).
Yu, L. et al. Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature 465, 942–946 (2010).
Du, W. et al. Kinesin 1 drives autolysosome tubulation. Dev. Cell 37, 326–336 (2016).
Rong, Y. et al. Clathrin and phosphatidylinositol-4,5-bisphosphate regulate autophagic lysosome reformation. Nat. Cell Biol. 14, 924–934 (2012).
Bissig, C., Hurbain, I., Raposo, G. & van Niel, G. PIKfyve activity regulates reformation of terminal storage lysosomes from endolysosomes. Traffic 18, 747–757 (2017).
Schulze, R. J. et al. Lipid droplet breakdown requires dynamin 2 for vesiculation of autolysosomal tubules in hepatocytes. J. Cell Biol. 203, 315–326 (2013).
Munson, M. J. et al. mTOR activates the VPS34-UVRAG complex to regulate autolysosomal tubulation and cell survival. EMBO J. 34, 2272–2290 (2015).
Chang, J., Lee, S. & Blackstone, C. Spastic paraplegia proteins spastizin and spatacsin mediate autophagic lysosome reformation. J. Clin. Invest. 124, 5249–5262 (2014).
Hirst, J., Hesketh, G. G., Gingras, A. C. & Robinson, M. S. Rag GTPases and phosphatidylinositol 3-phosphate mediate recruitment of the AP-5/SPG11/SPG15 complex. J. Cell Biol. https://doi.org/10.1083/jcb.202002075 (2021).
Giordano, F. et al. PI(4,5)P2-dependent and Ca2+-regulated ER-PM interactions mediated by the extended synaptotagmins. Cell 153, 1494–1509 (2013).
Chang, C.-L. et al. Feedback regulation of receptor-induced Ca2+ signaling mediated by E-Syt1 and Nir2 at endoplasmic reticulum-plasma membrane junctions. Cell Rep. 5, 813–825 (2013).
Saheki, Y. et al. Control of plasma membrane lipid homeostasis by the extended synaptotagmins. Nat. Cell Biol. 18, 504–515 (2016). Extended synaptotagmins are shown to act as PtdIns(4,5)P2- and Ca2+-regulated tethers that transfer glycerolipids between the ER and the plasma membrane.
Lees, J. A. et al. Lipid transport by TMEM24 at ER–plasma membrane contacts regulates pulsatile insulin secretion. Science 355, eaah6171 (2017).
Chung, J. et al. PI4P/phosphatidylserine countertransport at ORP5- and ORP8-mediated ER-plasma membrane contacts. Science 349, 428–432 (2015).
Sohn, M. et al. PI(4,5)P2 controls plasma membrane PI4P and PS levels via ORP5/8 recruitment to ER-PM contact sites. J. Cell Biol. 217, 1797–1813 (2018).
Hanada, K. et al. Molecular machinery for non-vesicular trafficking of ceramide. Nature 426, 803–809 (2003).
Shirane, M. et al. Protrudin and PDZD8 contribute to neuronal integrity by promoting lipid extraction required for endosome maturation. Nat. Commun. 11, 1–19 (2020).
Lim, C.-Y. et al. ER–lysosome contacts enable cholesterol sensing by mTORC1 and drive aberrant growth signalling in Niemann–Pick type C. Nat. Cell Biol. 21, 1206–1218 (2019).
Mesmin, B. et al. A four-step cycle driven by PI(4)P hydrolysis directs sterol/PI(4)P exchange by the ER-Golgi tether OSBP. Cell 155, 830–843 (2013). This is a landmark study that identifies OSBP as a lipid shuttle utilizing the gradient of PtdIns4P between the ER and the TGN to promote cholesterol export at MCSs.
D’Angelo, G. et al. Glycosphingolipid synthesis requires FAPP2 transfer of glucosylceramide. Nature 449, 62–67 (2007).
Murakami, H. et al. Intellectual disability-associated gain-of-function mutations in CERT1 that encodes the ceramide transport protein CERT. PLoS ONE 15, e0243980 (2020).
Kawasaki, A. et al. PI4P/PS countertransport by ORP10 at ER–endosome membrane contact sites regulates endosome fission. J. Cell Biol. 221, e202103141 (2021).
Rocha, N. et al. Cholesterol sensor ORP1L contacts the ER protein VAP to control Rab7–RILP–p150Glued and late endosome positioning. J. Cell Biol. 185, 1209–1225 (2009).
Levin-Konigsberg, R. et al. Phagolysosome resolution requires contacts with the endoplasmic reticulum and phosphatidylinositol-4-phosphate signalling. Nat. Cell Biol. 21, 1234–1247 (2019).
Malek, M. et al. Inositol triphosphate-triggered calcium release blocks lipid exchange at endoplasmic reticulum-Golgi contact sites. Nat. Commun. 12, 2673 (2021).
Kim, Y. J., Guzman-Hernandez, M.-L., Wisniewski, E. & Balla, T. Phosphatidylinositol-phosphatidic acid exchange by Nir2 at ER-PM contact sites maintains phosphoinositide signaling competence. Dev. Cell 33, 549–561 (2015).
Cockcroft, S. & Lev, S. Mammalian PITPs at the Golgi and ER-Golgi membrane contact sites. Contact 3, 2515256420964170 (2020).
Wang, H. & Tai, A. W. Nir2 is an effector of VAPs necessary for efficient hepatitis C virus replication and phosphatidylinositol 4-phosphate enrichment at the viral replication organelle. J. Virol. 93, e00742-19 (2019).
Takahashi, K. et al. ORP2 couples LDL-cholesterol transport to FAK activation by endosomal cholesterol/PI(4,5)P2 exchange. EMBO J. 40, e106871 (2021).
Nagashima, S. et al. Golgi-derived PI(4)P-containing vesicles drive late steps of mitochondrial division. Science 367, 1366–1371 (2020).
Gong, B. et al. A Golgi-derived vesicle potentiates PtdIns4P to PtdIns3P conversion for endosome fission. Nat. Cell Biol. 23, 782–795 (2021). Nagashima et al. (2020) and Gong et al. (2021) show that TGN-derived PtdIns4P-containing vesicles are recruited to ER-based ternary MCSs to trigger fission of mitochondria and endosomes, respectively.
Sabha, N. et al. PIK3C2B inhibition improves function and prolongs survival in myotubular myopathy animal models. J. Clin. Invest. 126, 3613–3625 (2016).
Walpole, G. F. W., Grinstein, S. & Westman, J. The role of lipids in host-pathogen interactions. IUBMB Life 70, 384–392 (2018).
Wang, R. et al. Genetic screens identify host factors for SARS-CoV-2 and common cold coronaviruses. Cell 184, 106–119 e114 (2021).
Kang, Y. L. et al. Inhibition of PIKfyve kinase prevents infection by Zaire ebolavirus and SARS-CoV-2. Proc. Natl Acad. Sci. USA 117, 20803–20813 (2020).
Alliouachene, S. et al. Inactivation of the class II PI3K-C2beta potentiates insulin signaling and sensitivity. Cell Rep. 13, 1881–1894 (2015).
Braccini, L. et al. PI3K-C2gamma is a Rab5 effector selectively controlling endosomal Akt2 activation downstream of insulin signalling. Nat. Commun. 6, 7400 (2015).
Mazloumi Gavgani, F. et al. Class I phosphoinositide 3-kinase PIK3CA/p110alpha and PIK3CB/p110beta isoforms in endometrial cancer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms19123931 (2018).
Zhang, M. et al. INPP4B protects from metabolic syndrome and associated disorders. Commun. Biol. 4, 416 (2021).
Pemberton, J. G. et al. Defining the subcellular distribution and metabolic channeling of phosphatidylinositol. J. Cell Biol. https://doi.org/10.1083/jcb.201906130 (2020).
Hammond, G. R., Machner, M. P. & Balla, T. A novel probe for phosphatidylinositol 4-phosphate reveals multiple pools beyond the Golgi. J. Cell Biol. 205, 113–126 (2014).
Balla, A., Tuymetova, G., Tsiomenko, A., Varnai, P. & Balla, T. A plasma membrane pool of phosphatidylinositol 4-phosphate is generated by phosphatidylinositol 4-kinase type-III alpha: studies with the PH domains of the oxysterol binding protein and FAPP1. Mol. Biol. Cell 16, 1282–1295 (2005).
Lemmon, M. A., Ferguson, K. M., O’Brien, R., Sigler, P. B. & Schlessinger, J. Specific and high-affinity binding of inositol phosphates to an isolated pleckstrin homology domain. Proc. Natl Acad. Sci. USA 92, 10472–10476 (1995).
Stauffer, T. P., Ahn, S. & Meyer, T. Receptor-induced transient reduction in plasma membrane PtdIns(4,5)P2 concentration monitored in living cells. Curr. Biol. 8, 343–346 (1998).
Manna, D., Albanese, A., Park, W. S. & Cho, W. Mechanistic basis of differential cellular responses of phosphatidylinositol 3,4-bisphosphate- and phosphatidylinositol 3,4,5-trisphosphate-binding pleckstrin homology domains. J. Biol. Chem. 282, 32093–32105 (2007).
Watton, S. J. & Downward, J. Akt/PKB localisation and 3’ phosphoinositide generation at sites of epithelial cell-matrix and cell-cell interaction. Curr. Biol. 9, 433–436 (1999).
Varnai, P., Rother, K. I. & Balla, T. Phosphatidylinositol 3-kinase-dependent membrane association of the Bruton’s tyrosine kinase pleckstrin homology domain visualized in single living cells. J. Biol. Chem. 274, 10983–10989 (1999).
Hammond, G. R. & Balla, T. Polyphosphoinositide binding domains: key to inositol lipid biology. Biochim. Biophys. Acta 1851, 746–758 (2015).
Nicholson, G. et al. Distinctive genetic and clinical features of CMT4J: a severe neuropathy caused by mutations in the PI(3,5)P(2) phosphatase FIG4. Brain 134, 1959–1971 (2011).
Elong Edimo, W., Schurmans, S., Roger, P. P. & Erneux, C. SHIP2 signaling in normal and pathological situations: its impact on cell proliferation. Adv. Biol. Regul. 54, 142–151 (2014).
Quadri, M. et al. Mutation in the SYNJ1 gene associated with autosomal recessive, early-onset Parkinsonism. Hum. Mutat. 34, 1208–1215 (2013).
Previtali, S. C., Quattrini, A. & Bolino, A. Charcot-Marie-Tooth type 4B demyelinating neuropathy: deciphering the role of MTMR phosphatases. Expert. Rev. Mol. Med. 9, 1–16 (2007).
De Matteis, M. A., Staiano, L., Emma, F. & Devuyst, O. The 5-phosphatase OCRL in Lowe syndrome and Dent disease 2. Nat. Rev. Nephrol. 13, 455–470 (2017).
Ngeow, J. & Eng, C. PTEN in hereditary and sporadic cancer. Cold Spring Harb. Perspect. Med. https://doi.org/10.1101/cshperspect.a036087 (2020).
Madsen, R. R., Vanhaesebroeck, B. & Semple, R. K. Cancer-associated PIK3CA mutations in overgrowth disorders. Trends Mol. Med. 24, 856–870 (2018).
Graupera, M. et al. Angiogenesis selectively requires the p110 alpha isoform of PI3K to control endothelial cell migration. Nature 453, 662–666 (2008).
Foukas, L. C. et al. Critical role for the p110alpha phosphoinositide-3-OH kinase in growth and metabolic regulation. Nature 441, 366–370 (2006).
Lucas, C. L., Chandra, A., Nejentsev, S., Condliffe, A. M. & Okkenhaug, K. PI3Kdelta and primary immunodeficiencies. Nat. Rev. Immunol. 16, 702–714 (2016).
Takeda, A. J. et al. Human PI3Kgamma deficiency and its microbiota-dependent mouse model reveal immunodeficiency and tissue immunopathology. Nat. Commun. 10, 4364 (2019).
Tiosano, D. et al. Mutations in PIK3C2A cause syndromic short stature, skeletal abnormalities, and cataracts associated with ciliary dysfunction. PLoS Genet. 15, e1008088 (2019).
Ikonomov, O. C., Sbrissa, D. & Shisheva, A. Small molecule PIKfyve inhibitors as cancer therapeutics: translational promises and limitations. Toxicol. Appl. Pharm. https://doi.org/10.1016/j.taap.2019.114771 (2019).
Lima, K. et al. PIP4K2A and PIP4K2C transcript levels are associated with cytogenetic risk and survival outcomes in acute myeloid leukemia. Cancer Genet. 233-234, 56–66 (2019).
Narkis, G. et al. Lethal contractural syndrome type 3 (LCCS3) is caused by a mutation in PIP5K1C, which encodes PIPKI gamma of the phophatidylinsitol pathway. Am. J. Hum. Genet. 81, 530–539 (2007).
Li, S. L. et al. Mutations in PIP5K3 are associated with Francois-Neetens Mouchetee fleck corneal dystrophy. Am. J. Hum. Genet. 77, 54–63 (2005).
Idevall-Hagren, O. & De Camilli, P. Detection and manipulation of phosphoinositides. Biochim. Biophys. Acta 1851, 736–745 (2015).
Clark, J. et al. Quantification of PtdInsP3 molecular species in cells and tissues by mass spectrometry. Nat. Methods 8, 267–272 (2011).
Malek, M. et al. PTEN regulates PI(3,4)P2 signaling downstream of class I PI3K. Mol. Cell 68, 566–580 e510 (2017).
Cheung, H. Y. F. et al. Targeted phosphoinositides analysis using high-performance ion chromatography-coupled selected reaction monitoring mass spectrometry. J. Proteome Res. 20, 3114–3123 (2021).
Li, P. & Lammerhofer, M. Isomer selective comprehensive lipidomics analysis of phosphoinositides in biological samples by liquid chromatography with data independent acquisition tandem mass spectrometry. Anal. Chem. 93, 9583–9592 (2021).
Suh, B. C., Inoue, T., Meyer, T. & Hille, B. Rapid chemically induced changes of PtdIns(4,5)P2 gate KCNQ ion channels. Science 314, 1454–1457 (2006).
Varnai, P., Thyagarajan, B., Rohacs, T. & Balla, T. Rapidly inducible changes in phosphatidylinositol 4,5-bisphosphate levels influence multiple regulatory functions of the lipid in intact living cells. J. Cell Biol. 175, 377–382 (2006).
Idevall-Hagren, O., Dickson, E. J., Hille, B., Toomre, D. K. & De Camilli, P. Optogenetic control of phosphoinositide metabolism. Proc. Natl Acad. Sci. USA 109, E2316–E2323 (2012). Together with Suh et al. (2006) and Varnai et al. (2006), this work provides elegant toolsets for the acute perturbation of phosphoinositide levels by induced recruitment of a lipid phosphatase via a ligand or by light.
Laketa, V. et al. Membrane-permeant phosphoinositide derivatives as modulators of growth factor signaling and neurite outgrowth. Chem. Biol. 16, 1190–1196 (2009).
Subramanian, D. et al. Activation of membrane-permeant caged PtdIns(3)P induces endosomal fusion in cells. Nat. Chem. Biol. 6, 324–326 (2010).
Laketa, V. et al. PIP(3) induces the recycling of receptor tyrosine kinases. Sci. Signal. 7, ra5 (2014).
Muller, R., Kojic, A., Citir, M. & Schultz, C. Synthesis and cellular labeling of multifunctional phosphatidylinositol bis- and trisphosphate derivatives. Angew. Chem. Int. Ed. 60, 19759–19765 (2021).
Acknowledgements
The authors gratefully acknowledge support of their own work by grants from the Deutsche Forschungsgemeinschaft (TRR186/A08, HA2686/15-1 and HA2686/22-1 to V.H.), the European Union (H2020-MSCA-ITN-2015:675392 to V.H.), the Leibniz Association (SAW K216/2016 to V.H.) and the European Research Council (ERC-AdG to V.H.). W.J. was supported by a Leibniz-German Academic Exchange Service (DAAD) Research Fellowship (57423756) and the Postdoctoral Fellowship Program (Nurturing Next-generation Researchers) of the National Research Foundation of Korea (2018R1A6A3A03010583).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Molecular Cell Biology thanks Harald Stenmark and Maria Antonietta De Matteis for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Endosomes
-
Membrane-bound compartments on the endocytic route that can be distinguished as early, late and recycling endosomes. Early endosomes act as sorting stations and are marked by RAB5 and phosphatidylinositol 3-phosphate. They can mature into RAB7-positive late endosomes (also called ‘multivesicular bodies’) by inward budding. Recycling endosomes are tubular in nature and recycle cargo to the cell surface.
- PtdIns 3-kinase
-
(PI3K). A family of lipid kinases that phosphorylate the 3′ position of the inositol head group of phosphatidylinositol (PtdIns) lipids. Class I PI3Ks act in receptor signalling, whereas class II and class III PI3Ks primarily control intracellular membrane dynamics.
- Click chemistry
-
A class of biocompatible organic reactions that rapidly and selectively react (‘click’) with each other under mild, aqueous conditions.
- Autophagy
-
A stress-inducible catabolic process that involves the formation of double membrane-bounded autophagosomes that fuse with lysosomes to mediate catabolic turnover of proteins, organelles or pathogens.
- Phospholipase C
-
(PLC). A group of hydrolytic enzymes that cleave phosphatidylinositol 4,5-bisphosphate into inositol 3,4,5-trisphosphate and diacylglycerol.
- COPII
-
Coatomer complex II, a type of coat protein that promotes the formation of secretory vesicles or tubules from exit sites of the endoplasmic reticulum to effect cargo transport to the endoplasmic reticulum–Golgi intermediate compartment.
- trans-Golgi network
-
(TGN). A highly dynamic series of interconnected tubules and vesicles at the trans face of the Golgi stack. The TGN functions in the sorting and processing of glycoproteins and glycolipids at the interface of the biosynthetic and endosomal pathways (for example, protein secretion and the sorting of lysosomal enzymes).
- Clathrin adaptor proteins
-
A collective term for proteins/protein complexes that recruit clathrin — a triskelial scaffold protein comprising three heavy chains and three associated light chains — to membranes and aid polymerization, often via binding to phosphoinositide lipids (for example, adaptor complexes AP1 and AP2, and monomeric GGA1, GGA2 and GGA3).
- DOPEY1–MON2 complex
-
A protein complex comprising the peripheral Golgi membrane proteins DOPEY1 and MON2, a relative of the SEC7 family of guanine-nucleotide exchange factors for ARF GTPases, proposed to act as a phosphatidylinositol 4-phosphate-dependent kinesin adaptor for membrane traffic at the trans-Golgi network.
- 14-3-3 protein
-
A family of conserved regulatory proteins expressed in all eukaryotic cells that can bind to functionally diverse, usually phosphorylated signalling proteins to regulate their function.
- Glycosphingolipids
-
A subclass of glycolipids that contain the amino alcohol sphingosine. They are found in the cell membranes of organisms from bacteria to humans and are the major glycolipids of animals.
- Sphingomyelin
-
An abundant phosphosphingolipid in animal cell membranes; it is especially enriched in the membranous myelin sheath that surrounds some nerve cell axons. Its hydrolysis releases ceramide and phosphocholine.
- Ceramide
-
Synthesized in the endoplasmic reticulum, the precursor of sphingomyelin. Glucosylceramide is the precursor of glycosphingolipids and is synthesized in the cis-Golgi network.
- Schwann cells
-
The principal glia of the peripheral nervous system that function to support neurons by forming a myelin sheath around axons for insulation. Schwann cells are also important for nerve regeneration.
- Synaptotagmin 1
-
A major type I transmembrane protein enriched in synaptic vesicles that acts as a calcium sensor for regulated exocytosis in central nervous system neurons.
- Clathrin-mediated endocytosis
-
(CME). The internalization of plasma membrane and receptors present therein into small vesicles that is mediated by a protein coat containing clathrin, adaptors and accessory proteins. During endocytosis clathrin triskelia polymerize into hexagons and pentagons to promote endocytic vesicle formation.
- Phox homology (PX) domain
-
A structurally conserved phosphoinositide-binding domain consisting of approximately 120 amino acids found in a wide range of proteins.
- Dynamin
-
A large mechanochemical GTPase that oligomerizes at the neck of endocytic vesicles or tubules to promote membrane fission.
- Pleckstrin homology (PH) domain
-
Sequence of approximately 100 amino acids that can mediate specific binding to phosphoinositide lipids and that is present in many signalling molecules. Only a minority of PH domains actually bind lipids, with the PH domain representing a conserved structural fold in proteins without necessarily a specific biological function.
- Macropinocytosis
-
An evolutionarily conserved endocytic pathway that allows internalization of extracellular fluid via large endocytic vesicles called ‘macropinosomes’.
- Nucleation-promoting factors
-
Factors such as WASP, N-WASP and Wiskott-Aldrich syndrome protein and SCAR homologue (WASH) that stimulate the intrinsically low activity of the ARP2/3 complex to nucleate actin filaments.
- ARP2/3 complex
-
Actin-related protein 2/3 complex, a seven-subunit protein complex that acts to promote the nucleation of branched actin filaments in eukaryotic cells.
- CLIC–GEEC pathway
-
A major clathrin-independent pinocytic pathway mediated by uncoated tubulovesicular carriers called ‘clathrin-independent carriers’ (CLICs) that mature into tubular early endocytic compartments called ‘glycosylphosphatidylinositol-anchored protein enriched compartments’ (GEECs).
- Multivesicular bodies
-
An alternative term for RAB7-positive late endosomes that form by inward budding of vesicles into the endosome lumen.
- Retromer
-
An evolutionarily conserved heterotetrameric complex involved in recycling of cargo from endosomes. It is composed of a membrane-associated sorting nexin (SNX3 or SNX27), and a vacuolar protein sorting trimer containing VPS26, VPS29 and VPS35.
- Retriever
-
Structurally and functionally related to retromer, the retriever complex comprises sorting nexin 17 (SNX17) and the CCC complex.
- ESCPE-1
-
Endosomal sorting nexin (SNX)–Bin–Amphiphysin–Rvs (BAR) sorting complex for promoting exit 1, a heterodimer of either SNX5 or SNX6 with either SNX1 or SNX2 that mediates retrieval of a subset of cargoes independently of retromer.
- Charcot–Marie–Tooth disease
-
A group of hereditary motor and sensory neuropathies that damage the peripheral nerves. Charcot–Marie–Tooth disease type 4B is a rare subtype of the disease caused by mutations in the phosphoinositide 3-phosphatase myotubularin-related protein 2 (MTMR2).
- Primary cilia
-
Non-motile type of cilia comprising an axoneme of nine doublet microtubules that are found on nearly all eukaryotic cells and function as microscopic sensory antennae.
- X-linked centronuclear myopathy
-
A severe human disease characterized by muscle fibre defects that result from mutations in the gene encoding the phosphoinositide 3-phosphatase myotubularin 1 (MTM1).
- FYVE domain
-
A phosphatidylinositol 3-phosphate-binding domain of approximately 60–65 amino acids that is named after the four cysteine-rich proteins FAB1, YOTB, VAC1 and EEA1, in which it has been found.
- PIKfyve
-
An evolutionarily conserved complex comprising the lipid kinase PIKfyve, the scaffold protein VAC14, and the putative 5-phosphatase FIG4. It mediates synthesis of lysosomal phosphatidylinositol 3,5-bisphosphate and phosphatidylinositol 5-phosphate in eukaryotic cells.
- Mechanistic target of rapamycin complex 1
-
(mTORC1). A multiprotein assembly composed of the kinase mTOR, a distant relative of the phosphatidylinositol 3-kinases, regulatory associated protein of mTOR (Raptor), mammalian lethal with SEC13 protein 8 (MLST8) and DEP domain-containing mTOR-interacting protein (DEPTOR) that promotes anabolism.
- PROPPIN domain
-
β-Propeller that binds polyphosphoinositides, a domain containing WD40 motifs and that has been identified in autophagy proteins such as yeast Atg18 and mammalian WD repeat domain phosphoinositide-interacting proteins (WIPI proteins). Via its β-propeller fold, it binds bind to phosphatidylinositol 3-phosphate and phosphatidylinositol 3,5-bisphosphate.
- LC3 family proteins
-
Proteins comprising microtubule-associated protein 1A/1B light chain 3 (LC3) and the closely related GABARAP proteins; they share structural homology with ubiquitin. They play key roles in autophagy.
- Cortactin
-
An actin nucleation-promoting factor that binds to F-actin filaments and to the ARP2/3 complex to regulate cell shape and movement.
- Joubert syndrome
-
A rare autosomal recessive disorder that is characterized by a distinctive cerebellar and brainstem malformation resulting in ataxia, mental retardation and retina degeneration. Among the genetic causes of the disease are loss-of-function mutations in the phosphoinositide 5-phosphatase INPP5E.
- Familial cerebellar atrophy
-
Cerebellar degeneration caused by inherited gene changes.
- Vacuolar ATPase
-
An ATP-driven proton pump that is closely related to the mitochondrial FoF1-ATPases and that is responsible for the lumenal acidification of endosomes, lysosomes and related organelles.
- RAG small GTPases
-
A unique family of evolutionarily conserved, heterodimeric, lysosome-localized small GTPases that promote anabolic processes through activation of mechanistic target of rapamycin complex 1 signalling in the presence of abundant amino acids.
- Hereditary spastic paraplegia
-
A group of rare inherited disorders that cause weakness and stiffness in the leg muscles.
- C2 domains
-
A membrane-binding domain homologous to the C2 domain of protein kinase C with mostly only moderate lipid specificity. Some C2 domains associate with membranes in a Ca2+-dependent manner.
- Store-operated Ca2+ entry
-
The regulated entry of Ca2+ into cells in response to the depletion of Ca2+ in the endoplasmic reticulum.
- FFAT motif
-
A peptide sequence (with, for example, two phenylalanines in an acidic tract) that binds to VAMP-associated proteins (VAPs) to facilitate the formation of endoplasmic reticulum-based membrane contact sites.
- Focal adhesion kinase
-
(FAK). A cytoplasmic non-receptor tyrosine kinase that localizes to focal adhesions and contributes to integrin-mediated cell signalling.
Rights and permissions
About this article
Cite this article
Posor, Y., Jang, W. & Haucke, V. Phosphoinositides as membrane organizers. Nat Rev Mol Cell Biol 23, 797–816 (2022). https://doi.org/10.1038/s41580-022-00490-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-022-00490-x
This article is cited by
-
Membranes are functionalized by a proteolipid code
BMC Biology (2024)
-
Rab GTPases and phosphoinositides fine-tune SNAREs dependent targeting specificity of intracellular vesicle traffic
Nature Communications (2024)
-
Lipids as mediators of cancer progression and metastasis
Nature Cancer (2024)
-
Glucose controls lipolysis through Golgi PtdIns4P-mediated regulation of ATGL
Nature Cell Biology (2024)
-
Nuclear phosphoinositide signaling promotes YAP/TAZ-TEAD transcriptional activity in breast cancer
The EMBO Journal (2024)