Abstract
Transcriptional regulation of catabolic pathways is a central mechanism by which cells respond to physiological cues to generate the energy required for anabolic pathways, transport of molecules and mechanical work. Nuclear receptors are members of a superfamily of transcription factors that transduce hormonal, nutrient, metabolite and redox signals into specific metabolic gene programmes, and thus hold a major status as regulators of cellular energy generation. Nuclear receptors also regulate the expression of genes involved in cellular processes that are implicated in energy production, including mitochondrial biogenesis and autophagy. Recent advances in genome-wide approaches have considerably expanded the repertoire of both nuclear receptors and metabolic genes under their direct transcriptional control. To fine-tune the expression of their target genes, nuclear receptors must act cooperatively with other transcription factors and coregulator proteins, integrate signals from key metabolic sensory systems such as the AMP-activated protein kinase (AMPK) and mechanistic target of rapamycin (mTOR) complexes and synchronize their activities with the biological clock. Therefore, nuclear receptors must function as more than molecular switches for small lipophilic ligands — as initially ascribed — but rather must be capable of orchestrating a large ensemble of input signals. Therefore, a primary role for several nuclear receptors is to serve as the focal point of transcriptional hubs in energy metabolism: their molecular task is to receive and transduce multiple systemic and intracellular metabolic signals to maintain energy homeostasis from individual cells to the whole organism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Judge, A. & Dodd, M. S. Metabolism. Essays Biochem. 64, 607–647 (2020). This work is a comprehensive review of the biochemical pathways involved in energy metabolism.
Pavlova, N. N., Zhu, J. & Thompson, C. B. The hallmarks of cancer metabolism: still emerging. Cell Metab. 34, 355–377 (2022).
Giguère, V., Yang, N., Segui, P. & Evans, R. M. Identification of a new class of steroid hormone receptors. Nature 331, 91–94 (1988). This article describes the identification of the first orphan nuclear receptor, predicting the existence of previously unknown hormone and small lipophilic compound response systems.
Wang, L. H. et al. Coup transcription factor is a member of the steroid-receptor superfamily. Nature 340, 163–166 (1989).
Giguère, V. Orphan nuclear receptors: from gene to function. Endocr. Rev. 20, 689–725 (1999).
Mullican, S. E., Dispirito, J. R. & Lazar, M. A. The orphan nuclear receptors at their 25-year reunion. J. Mol. Endocrinol. 51, T115–T140 (2013).
Markov, G. V. & Laudet, V. Origin and evolution of the ligand-binding ability of nuclear receptors. Mol. Cell. Endocrinol. 334, 21–30 (2011).
Sural, S. & Hobert, O. Nematode nuclear receptors as integrators of sensory information. Curr. Biol. 31, 4361–4366 e4362 (2021).
Robinson-Rechavi, M., Escriva Garcia, H. & Laudet, V. The nuclear receptor superfamily. J. Cell Sci. 116, 585–586 (2003).
Robinson-Rechavi, M., Maina, C. V., Gissendanner, C. R., Laudet, V. & Sluder, A. Explosive lineage-specific expansion of the orphan nuclear receptor HNF4 in nematodes. J. Mol. Evol. 60, 577–586 (2005).
Bodofsky, S., Koitz, F. & Wightman, B. Conserved and exapted functions of nuclear receptors in animal development. Nucl. Receptor Res. 4, 101305 (2017).
Wisely, G. B. et al. Hepatocyte nuclear factor 4 is a transcription factor that constitutively binds fatty acids. Structure 10, 1225–1234 (2002).
Van Gilst, M. R., Hadjivassiliou, H., Jolly, A. & Yamamoto, K. R. Nuclear hormone receptor NHR-49 controls fat consumption and fatty acid composition in C. elegans. PLoS Biol. 3, e53 (2005).
Palanker, L., Tennessen, J. M., Lam, G. & Thummel, C. S. Drosophila HNF4 regulates lipid mobilization and β-oxidation. Cell Metab. 9, 228–239 (2009). This genetic and metabolic study in Drosophila defines the ancestral function of HNF4 as a key regulator of lipid mobilization and β-oxidation.
Tennessen, J. M., Baker, K. D., Lam, G., Evans, J. & Thummel, C. S. The Drosophila estrogen-related receptor directs a metabolic switch that supports developmental growth. Cell Metab. 13, 139–148 (2011).
Pathare, P. P., Lin, A., Bornfeldt, K. E., Taubert, S. & Van Gilst, M. R. Coordinate regulation of lipid metabolism by novel nuclear receptor partnerships. PLoS Genet. 8, e1002645 (2012).
Storelli, G., Nam, H. J., Simcox, J., Villanueva, C. J. & Thummel, C. S. Drosophila HNF4 directs a switch in lipid metabolism that supports the transition to adulthood. Dev. Cell 48, 200–214 e206 (2019).
Beebe, K. et al. Drosophila estrogen-related receptor directs a transcriptional switch that supports adult glycolysis and lipogenesis. Genes Dev. 34, 701–714 (2020).
Voss, T. C. & Hager, G. L. Dynamic regulation of transcriptional states by chromatin and transcription factors. Nat. Rev. Genet. 15, 69–81 (2014).
Boyle, A. P. et al. High-resolution mapping and characterization of open chromatin across the genome. Cell 132, 311–322 (2008).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21 29 21–21 29 29 (2015).
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Carroll, J. S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122, 33–43 (2005).
Laganière, J. et al. Location analysis of estrogen receptor α target promoters reveals that FOXA1 defines a domain of the estrogen response. Proc. Natl Acad. Sci. USA 102, 11651–11656 (2005). This study, together with Carroll et al. (2005), shows the requirement of the pioneer factor FOXA1 for a nuclear receptor binding to enhancers and promoters in the genome.
Kain, J. et al. Pioneer factor Foxa2 enables ligand-dependent activation of type II nuclear receptors FXR and LXRα. Mol. Metab. 53, 101291 (2021).
Sahu, B. et al. FoxA1 specifies unique androgen and glucocorticoid receptor binding events in prostate cancer cells. Cancer Res. 73, 1570–1580 (2013).
Jin, H. J., Zhao, J. C., Wu, L., Kim, J. & Yu, J. Cooperativity and equilibrium with FOXA1 define the androgen receptor transcriptional program. Nat. Commun. 5, 3972 (2014).
Reizel, Y. et al. Collapse of the hepatic gene regulatory network in the absence of FoxA factors. Genes Dev. 34, 1039–1050 (2020).
Horisawa, K. et al. The dynamics of transcriptional activation by hepatic reprogramming factors. Mol. Cell 79, 660–676 e668 (2020).
Voss, T. C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011).
Swinstead, E. E. et al. Steroid receptors reprogram FoxA1 occupancy through dynamic chromatin transitions. Cell 165, 593–605 (2016).
Adachi, K. et al. Esrrb unlocks silenced enhancers for reprogramming to naive pluripotency. Cell Stem Cell 23, 266–275 e266 (2018).
Madsen, M. S., Siersbaek, R., Boergesen, M., Nielsen, R. & Mandrup, S. Peroxisome proliferator-activated receptor γ and C/EBPα synergistically activate key metabolic adipocyte genes by assisted loading. Mol. Cell. Biol. 34, 939–954 (2014).
Ratman, D. et al. Chromatin recruitment of activated AMPK drives fasting response genes co-controlled by GR and PPARα. Nucleic Acids Res. 44, 10539–10553 (2016).
Biddie, S. C. et al. Transcription factor AP1 potentiates chromatin accessibility and glucocorticoid receptor binding. Mol. Cell 43, 145–155 (2011).
Perissi, V., Aggarwal, A., Glass, C. K., Rose, D. W. & Rosenfeld, M. G. A corepressor/coactivator exchange complex required for transcriptional activation by nuclear receptors and other regulated transcription factors. Cell 116, 511–526 (2004).
Shabtai, Y. et al. A coregulator shift, rather than the canonical switch, underlies thyroid hormone action in the liver. Genes Dev. 35, 367–378 (2021).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat. Methods 4, 651–657 (2007).
Cohen, D. M. & Steger, D. J. Nuclear receptor function through genomics: lessons from the glucocorticoid receptor. Trends Endocrinol. Metab. 28, 531–540 (2017).
Siersbaek, R. et al. Dynamic rewiring of promoter-anchored chromatin loops during adipocyte differentiation. Mol. Cell 66, 420–435 e425 (2017). This elegant work using a variety of genomics approaches demonstrates that rewiring of promoter-anchored loops is coupled to changes in the activity of the connected promoters and enhancers occupied by a cluster of metabolic transcription factors and nuclear receptors.
Lefterova, M. I. et al. Cell-specific determinants of peroxisome proliferator-activated receptor γ function in adipocytes and macrophage. Mol. Cell. Biol. 30, 2078–2089 (2010).
Hu, W. et al. Individual-specific functional epigenomics reveals genetic determinants of adverse metabolic effects of glucocorticoids. Cell Metab. 33, 1592–1609 e1597 (2021).
Deblois, G. et al. ERRα mediates metabolic adaptations driving lapatinib resistance in breast cancer. Nat. Commun. 7, 12156 (2016). This work reveals a molecular mechanism by which nuclear receptor-induced metabolic reprogramming promotes survival of drug-resistant cancer cells.
Qin, Y., Grimm, S. A., Roberts, J. D., Chrysovergis, K. & Wade, P. A. Alterations in promoter interaction landscape and transcriptional network underlying metabolic adaptation to diet. Nat. Commun. 11, 962 (2020).
Fang, B. & Lazar, M. A. Dissecting the Rev-erbα cistrome and the mechanisms controlling circadian transcription in liver. Cold Spring Harb. Symp. Quant. Biol. 80, 233–238 (2015).
Xia, H., Dufour, C. R. & Giguère, V. ERRα as a bridge between transcription and function: role in liver metabolism and disease. Front. Endocrinol. 10, 206 (2019).
Soccio, R. E. et al. Genetic variation determines PPARγ function and anti-diabetic drug response In vivo. Cell 162, 33–44 (2015). This tour-de-force work combining mouse genetics and genome-wide association studies in human shows that genetic variation in PPARγ genomic occupancy determines metabolic disease risk and drug response, providing a proof of concept for personalized medicine related to nuclear receptor genomic occupancy.
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Hah, N. et al. A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell 145, 622–634 (2011).
Step, S. E. et al. Anti-diabetic rosiglitazone remodels the adipocyte transcriptome by redistributing transcription to PPARγ-driven enhancer. Genes Dev. 28, 1018–1028 (2014).
Wang, D. et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature 474, 390–394 (2011).
Goldstein, I. & Hager, G. L. Transcriptional and chromatin regulation during fasting- the genomic era. Trends Endocrinol. Metab. 26, 699–710 (2015).
Grontved, L. et al. C/EBP maintains chromatin accessibility in liver and facilitates glucocorticoid receptor recruitment to steroid response elements. EMBO J. 32, 1568–1583 (2013).
Langlais, D., Couture, C., Balsalobre, A. & Drouin, J. The Stat3/GR interaction code: predictive value of direct/indirect DNA recruitment for transcription outcome. Mol. Cell 47, 38–49 (2012).
Vockley, C. M. et al. Direct GR binding sites potentiate clusters of TF binding across the human genome. Cell 166, 1269–1281 e1219 (2016).
Giudici, M., Goni, S., Fan, R. & Treuter, E. Nuclear receptor coregulators in metabolism and disease. Handb. Exp. Pharmacol. 233, 95–135 (2016).
Zhang, Y. et al. Discrete functions of nuclear receptor Rev-erbα couple metabolism to the clock. Science 348, 1488–1492 (2015). This work shows that REV-ERBα uses distinct mechanisms, direct binding and tethering, to recognize its target genes in the genome and thus differentially regulates biological clock and metabolic genes.
Murayama, Y. et al. Glucocorticoid receptor suppresses gene expression of Rev-erbα (Nr1d1) through interaction with the CLOCK complex. FEBS Lett. 593, 423–432 (2019).
Boergesen, M. et al. Genome-wide profiling of liver X receptor, retinoid X receptor, and peroxisome proliferator-activated receptor α in mouse liver reveals extensive sharing of binding sites. Mol. Cell. Biol. 32, 852–867 (2012).
Siersbaek, R. et al. Molecular architecture of transcription factor hotspots in early adipogenesis. Cell Rep. 7, 1434–1442 (2014).
Grontved, L. et al. Transcriptional activation by the thyroid hormone receptor through ligand-dependent receptor recruitment and chromatin remodelling. Nat. Commun. 6, 7048 (2015).
Dasgupta, S., Lonard, D. M. & O’Malley, B. W. Nuclear receptor coactivators: master regulators of human health and disease. Annu. Rev. Med. 65, 279–292 (2014).
Mouchiroud, L., Eichner, L. J., Shaw, R. J. & Auwerx, J. Transcriptional coregulators: fine-tuning metabolism. Cell Metab. 20, 26–40 (2014).
Puigserver, P. et al. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 92, 829–839 (1998).
Vainshtein, A., Tryon, L. D., Pauly, M. & Hood, D. A. Role of PGC-1α during acute exercise-induced autophagy and mitophagy in skeletal muscle. Am. J. Physiol. Cell Physiol. 308, C710–C719 (2015).
Gidlund, E. K. et al. Rapidly elevated levels of PGC-1α-β protein in human skeletal muscle after exercise: exploring regulatory factors in a randomized controlled trial. J. Appl. Physiol. 119, 374–384 (2015).
Rodgers, J. T. et al. Nutrient control of glucose homeostasis through a complex of PGC-1α and SIRT1. Nature 434, 113–118 (2005). This article shows that NAD+-dependent induction of SIRT1 deacetylates PGC1α at specific lysine residues and leads to the induction of gluconeogenic genes and hepatic glucose output, thus identifying a molecular mechanism whereby sirtuins function in glucose homeostasis.
Jager, S., Handschin, C., St-Pierre, J. & Spiegelman, B. M. AMP-activated protein kinase (AMPK) action in skeletal muscle via direct phosphorylation of PGC-1α. Proc. Natl Acad. Sci. USA 104, 12017–12022 (2007).
Coste, A. et al. The genetic ablation of SRC-3 protects against obesity and improves insulin sensitivity by reducing the acetylation of PGC-1α. Proc. Natl Acad. Sci. USA 105, 17187–17192 (2008).
Schreiber, S. N., Knutti, D., Brogli, K., Uhlmann, T. & Kralli, A. The transcriptional coactivator PGC-1 regulates the expression and activity of the orphan nuclear receptor estrogen-related receptor α (ERRα). J. Biol. Chem. 278, 9013–9018 (2003).
Laganiere, J. et al. A polymorphic autoregulatory hormone response element in the human estrogen-related receptor α (ERRα) promoter dictates peroxisome proliferator-activated receptor γ coactivator-1α control of ERRα expression. J. Biol. Chem. 279, 18504–18510 (2004).
Mootha, V. K. et al. ERRα and Gabpa/b specify PGC-1α-dependent oxidative phosphorylation gene expression that is altered in diabetic muscle. Proc. Natl Acad. Sci. USA 101, 6570–6575 (2004). The results in this article demonstrate that, together, ERRα and PGC1α control the expression of OXPHOS pathway gene expression, suggesting that modulation of ERRα transcriptional activity may ameliorate insulin resistance in individuals with type 2 diabetes mellitus.
Gaillard, S. et al. Receptor-selective coactivators as tools to define the biology of specific receptor-coactivator pairs. Mol. Cell 24, 797–803 (2006).
Chen, W., Yang, Q. & Roeder, R. G. Dynamic interactions and cooperative functions of PGC-1α and MED1 in TRα-mediated activation of the brown-fat-specific UCP-1 gene. Mol. Cell 35, 755–768 (2009).
Zhou, J. et al. MED1 mediator subunit is a key regulator of hepatic autophagy and lipid metabolism. Autophag 17, 4043–4061 (2021).
Ito, K. et al. Critical roles of transcriptional coactivator MED1 in the formation and function of mouse adipose tissues. Genes Dev. 35, 729–748 (2021).
Stashi, E. et al. SRC-2 is an essential coactivator for orchestrating metabolism and circadian rhythm. Cell Rep. 6, 633–645 (2014).
Tachmatzidi, E. C., Galanopoulou, O. & Talianidis, I. Transcription control of liver development. Cells 10, 2026 (2021).
Charest-Marcotte, A. et al. The homeobox protein Prox1 is a negative modulator of ERRα/PGC-1α bioenergetic functions. Genes Dev. 24, 537–542 (2010). This article identifies PROX1 as an important coregulator of the PGC1α–ERRα complex involved in the transcriptional regulation of genes encoding components of the OXPHOS, TCA cycle and ETC pathways. This vast cluster of functionally linked energy metabolism genes is referred to in this study as the ‘ERRα bioenergetic regulon’.
Dufour, C. R. et al. Genomic convergence among ERRα, Prox1 and Bmal1 in the control of metabolic clock outputs. PLoS Genet. 7, e1002143 (2011).
Song, K. H., Li, T. & Chiang, J. Y. A Prospero-related homeodomain protein is a novel co-regulator of hepatocyte nuclear factor 4α that regulates the cholesterol 7α-hydroxylase gene. J. Biol. Chem. 281, 10081–10088 (2006).
Qin, J. et al. Prospero-related homeobox (Prox1) is a corepressor of human liver receptor homolog-1 and suppresses the transcription of the cholesterol 7-α-hydroxylase gene. Mol. Endocrinol. 18, 2424–2439 (2004).
Stein, S. et al. SUMOylation-dependent LRH-1/PROX1 interaction promotes atherosclerosis by decreasing hepatic reverse cholesterol transport. Cell Metab. 20, 603–613 (2014).
Steffensen, K. R. et al. Functional conservation of interactions between a homeodomain cofactor and a mammalian FTZ-F1 homologue. EMBO Rep. 5, 613–619 (2004).
Armour, S. M. et al. An HDAC3-PROX1 corepressor module acts on HNF4α to control hepatic triglycerides. Nat. Commun. 8, 549 (2017). This article shows the importance of specific combinations of nuclear receptors and coregulators in the fine-tuning of metabolism.
Goto, T. et al. Liver-specific Prox1 inactivation causes hepatic injury and glucose intolerance in mice. FEBS Lett. 591, 624–635 (2017).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Lecompte, S. et al. Genetic and molecular insights into the role of PROX1 in glucose metabolism. Diabetes 62, 1738–1745 (2013).
Liang, N., Jakobsson, T., Fan, R. & Treuter, E. The nuclear receptor-co-repressor complex in control of liver metabolism and disease. Front. Endocrinol. 10, 411 (2019).
Perez-Schindler, J. et al. The corepressor NCoR1 antagonizes PGC-1α and estrogen-related receptor α in the regulation of skeletal muscle function and oxidative metabolism. Mol. Cell. Biol. 32, 4913–4924 (2012).
Jo, Y. S. et al. Phosphorylation of the nuclear receptor corepressor 1 by protein kinase B switches its corepressor targets in the liver in mice. Hepatology 62, 1606–1618 (2015).
Herzig, S. & Shaw, R. J. AMPK: guardian of metabolism and mitochondrial homeostasis. Nat. Rev. Mol. Cell Biol. 19, 121–135 (2018).
Lien, F. et al. Metformin interferes with bile acid homeostasis through AMPK-FXR crosstalk. J. Clin. Invest. 124, 1037–1051 (2014).
Chopra, A. R. et al. Cellular energy depletion resets whole-body energy by promoting coactivator-mediated dietary fuel absorption. Cell Metab. 13, 35–43 (2011).
Hong, Y. H., Varanasi, U. S., Yang, W. & Leff, T. AMP-activated protein kinase regulates HNF4α transcriptional activity by inhibiting dimer formation and decreasing protein stability. J. Biol. Chem. 278, 27495–27501 (2003).
Ding, J. et al. AMPK phosphorylates PPARδ to mediate its stabilization, inhibit glucose and glutamine uptake and colon tumor growth. J. Biol. Chem. 297, 100954 (2021).
Audet-Walsh, E. et al. The PGC-1α/ERRα axis represses one-carbon metabolism and promotes sensitivity to anti-folate therapy in breast cancer. Cell Rep. 14, 920–931 (2016).
Yan, M. et al. Chronic AMPK activation via loss of FLCN induces functional beige adipose tissue through PGC-1α/ERRα. Genes Dev. 30, 1034–1046 (2016).
Possik, E. et al. Folliculin regulates AMPK-dependent autophagy and metabolic stress survival. PLoS Genet. 10, e1004273 (2014).
Gonzalez, A. & Hall, M. N. Nutrient sensing and TOR signaling in yeast and mammals. EMBO J. 36, 397–408 (2017).
Szwed, A., Kim, E. & Jacinto, E. Regulation and metabolic functions of mTORC1 and mTORC2. Physiol. Rev. 101, 1371–1426 (2021).
Giguère, V. Canonical signaling and nuclear activity of mTOR: a teamwork effort to regulate metabolism and cell growth. FEBS J. 285, 1572–1588 (2018).
Shimizu, N. et al. Crosstalk between glucocorticoid receptor and nutritional sensor mTOR in skeletal muscle. Cell Metab. 13, 170–182 (2011).
Chaveroux, C. et al. Molecular and genetic crosstalks between mTOR and ERRα are key determinants of rapamycin-induced non-alcoholic fatty liver. Cell Metab. 17, 586–598 (2013).
Audet-Walsh, E. et al. Nuclear mTOR acts as a transcriptional integrator of the androgen signaling pathway in prostate cancer. Genes Dev. 31, 1228–1242 (2017). Chaveroux et al. (2013) and Audet-Walsh et al. (Genes Dev., 2017) are two publications revealing that nuclear mTOR occupies specific sites in the genome and works in concert with nuclear receptors to regulate the expression of metabolic genes in both normal and cancer cells.
Sengupta, S., Peterson, T. R., Laplante, M., Oh, S. & Sabatini, D. M. mTORC1 controls fasting-induced ketogenesis and its modulation by ageing. Nature 468, 1100–1104 (2010).
Kim, K., Pyo, S. & Um, S. H. S6 kinase 2 deficiency enhances ketone body production and increases peroxisome proliferator-activated receptor α activity in the liver. Hepatology 55, 1727–1737 (2012).
Reinke, H. & Asher, G. Crosstalk between metabolism and circadian clocks. Nat. Rev. Mol. Cell Biol. 20, 227–241 (2019).
Panda, S. Circadian physiology of metabolism. Science 354, 1008–1015 (2016).
Kim, Y. H. & Lazar, M. A. Transcriptional control of circadian rhythms and metabolism: a matter of time and space. Endocr. Rev. 41, 707–732 (2020).
Zhao, X. et al. Nuclear receptors rock around the clock. EMBO Rep. 15, 518–528 (2014).
Nader, N., Chrousos, G. P. & Kino, T. Circadian rhythm transcription factor CLOCK regulates the transcriptional activity of the glucocorticoid receptor by acetylating its hinge region lysine cluster: potential physiological implications. FASEB J. 23, 1572–1583 (2009).
Grimaldi, B. et al. PER2 controls lipid metabolism by direct regulation of PPARγ. Cell Metab. 12, 509–520 (2010).
Lamia, K. A. et al. Cryptochromes mediate rhythmic repression of the glucocorticoid receptor. Nature 480, 552–556 (2011). This article demonstrates that two core components of the biological clock, CRY1 and CRY2, interact with GR in a ligand-dependent fashion to alter the transcriptional response to glucocorticoid and regulate metabolic homeostasis.
Jordan, S. D. et al. CRY1/2 selectively repress PPARδ and limit exercise capacity. Cell Metab. 26, 243–255 e246 (2017).
Thorens, B. & Mueckler, M. Glucose transporters in the 21st century. Am. J. Physiol. Endocrinol. Metab. 298, E141–E145 (2010).
Leto, D. & Saltiel, A. R. Regulation of glucose transport by insulin: traffic control of GLUT4. Nat. Rev. Mol. Cell Biol. 13, 383–396 (2012).
Oosterveer, M. H. et al. LRH-1-dependent glucose sensing determines intermediary metabolism in liver. J. Clin. Invest. 122, 2817–2826 (2012).
Baba, T. et al. Glycolytic genes are targets of the nuclear receptor Ad4BP/SF-1. Nat. Commun. 5, 3634 (2014).
Wende, A. R., Huss, J. M., Schaeffer, P. J., Giguère, V. & Kelly, D. P. PGC-1α coactivates PDK4 gene expression via the orphan nuclear receptor ERRα: a mechanism for transcriptional control of muscle glucose metabolism. Mol. Cell. Biol. 25, 10684–10694 (2005).
Zhang, Y. et al. Estrogen-related receptors stimulate pyruvate dehydrogenase kinase isoform 4 gene expression. J. Biol. Chem. 281, 39897–39906 (2006).
Niu, B. et al. In vivo genome-wide binding interactions of mouse and human constitutive androstane receptors reveal novel gene targets. Nucleic Acids Res. 46, 8385–8403 (2018).
Connaughton, S. et al. Regulation of pyruvate dehydrogenase kinase isoform 4 (PDK4) gene expression by glucocorticoids and insulin. Mol. Cell. Endocrinol. 315, 159–167 (2010).
Nielsen, R. et al. Genome-wide profiling of PPARγ:RXR and RNA polymerase II occupancy reveals temporal activation of distinct metabolic pathways and changes in RXR dimer composition during adipogenesis. Genes Dev. 22, 2953–2967 (2008).
Hunter, A. L. et al. Adipocyte NR1D1 dictates adipose tissue expansion during obesity. eLife 10, e63324 (2021).
Gan, Z. et al. The nuclear receptor PPARβ/δ programs muscle glucose metabolism in cooperation with AMPK and MEF2. Genes Dev. 25, 2619–2630 (2011).
Park, S. et al. ERRα-regulated lactate metabolism contributes to resistance to targeted therapies in breast cancer. Cell Rep. 15, 323–335 (2016).
Fan, W. & Evans, R. PPARs and ERRs: molecular mediators of mitochondrial metabolism. Curr. Opin. Cell Biol. 33, 49–54 (2015).
Sladek, R., Bader, J.-A. & Giguère, V. The orphan nuclear receptor estrogen-related receptor α is a transcriptional regulator of the human medium-chain acyl coenzyme A dehydrogenase gene. Mol. Cell. Biol. 17, 5400–5409 (1997).
Vega, R. B. & Kelly, D. P. A role for estrogen-related receptor α in the control of mitochondrial fatty acid β-oxidation during brown adipocyte differentiation. J. Biol. Chem. 272, 31693–31699 (1997).
Maehara, K. et al. Functional interference between estrogen-related receptor α and peroxisome proliferator-activated receptor α/9-cis-retinoic acid receptor α heterodimer complex in the nuclear receptor response element-1 of the medium chain acyl-coenzyme A dehydrogenase gene. J. Mol. Endocrinol. 31, 47–60 (2003).
Gulick, T., Cresci, S., Caira, T., Moore, D. D. & Kelly, D. P. The peroxisome proliferator activated receptor regulates mitochondrial fatty acid oxidative enzyme gene expression. Proc. Natl Acad. Sci. USA 91, 11012–11016 (1994).
Alaynick, W. A. et al. ERRγ directs and maintains the transition to oxidative metabolism in the post-natal heart. Cell Metab. 6, 16–24 (2007).
Huss, J. M., Torra, I. P., Staels, B., Giguère, V. & Kelly, D. P. Estrogen-related receptor α directs peroxisome proliferator-activated receptor α signaling in the transcriptional control of energy metabolism in cardiac and skeletal muscle. Mol. Cell. Biol. 24, 9079–9091 (2004).
Seth, A. et al. The transcriptional corepressor RIP140 regulates oxidative metabolism in skeletal muscle. Cell Metab. 6, 236–245 (2007).
Dufour, C. R. et al. Genome-wide orchestration of cardiac functions by orphan nucler receptors ERRα and γ. Cell Metab. 5, 345–356 (2007).
Ribet, C. et al. Peroxisome proliferator-activated receptor-α control of lipid and glucose metabolism in human white adipocytes. Endocrinology 151, 123–133 (2010).
McMullen, P. D. et al. A map of the PPARα transcription regulatory network for primary human hepatocytes. Chem. Biol. Interact. 209, 14–24 (2014).
He, Y. et al. The role of retinoic acid in hepatic lipid homeostasis defined by genomic binding and transcriptome profiling. BMC Genomics 14, 575 (2013).
Tanaka, T. et al. Activation of peroxisome proliferator-activated receptor δ induces fatty acid β-oxidation in skeletal muscle and attenuates metabolic syndrome. Proc. Natl Acad. Sci. USA 100, 15924–15929 (2003).
Chen, L. et al. HNF4 regulates fatty acid oxidation and Is required for renewal of intestinal stem cells in mice. Gastroenterology 158, 985–999 e989 (2020).
Rodriguez, J. C., Ortiz, J. A., Hegardt, F. G. & Haro, D. Chicken ovalbumin upstream-promoter transcription factor (COUP-TF) could act as a transcriptional activator or repressor of the mitochondrial 3-hydroxy-3-methylglutaryl-CoA synthase gene. Biochem. J. 326, 587–592 (1997).
Nakamura, K., Moore, R., Negishi, M. & Sueyoshi, T. Nuclear pregnane X receptor cross-talk with FoxA2 to mediate drug-induced regulation of lipid metabolism in fasting mouse liver. J. Biol. Chem. 282, 9768–9776 (2007).
Chen, F. et al. Metabolomic approaches reveal the role of CAR in energy metabolism. J. Proteome Res. 18, 239–251 (2019).
Rodriguez, J. C., Ortiz, J. A., Hegardt, F. G. & Haro, D. The hepatocyte nuclear factor 4 (HNF-4) represses the mitochondrial HMG-CoA synthase gene. Biochem. Biophys. Res. Commun. 242, 692–696 (1998).
Rodriguez, J. C., Gil-Gomez, G., Hegardt, F. G. & Haro, D. Peroxisome proliferator-activated receptor mediates induction of the mitochondrial 3-hydroxy-3-methylglutaryl-CoA synthase gene by fatty acids. J. Biol. Chem. 269, 18767–18772 (1994).
Loft, A. et al. A macrophage-hepatocyte glucocorticoid receptor axis coordinates fasting ketogenesis. Cell Metab. 34, 473–486 e479 (2022).
Svensson, K., Albert, V., Cardel, B., Salatino, S. & Handschin, C. Skeletal muscle PGC-1α modulates systemic ketone body homeostasis and ameliorates diabetic hyperketonemia in mice. FASEB J. 30, 1976–1986 (2016).
Dhillon, P. et al. The nuclear receptor ESRRA protects from kidney disease by coupling metabolism and differentiation. Cell Metab. 33, 379–394 e378 (2021).
Tran, A. et al. Estrogen-related receptor α (ERRα) is a key regulator of intestinal homeostasis and protects against colitis. Sci. Rep. 11, 15073 (2021).
Sjogren, R. J. O. et al. Branched-chain amino acid metabolism is regulated by ERRα in primary human myotubes and is further impaired by glucose loading in type 2 diabetes. Diabetologia 64, 2077–2091 (2021).
Contreras, A. V. et al. PPARα via HNF4α regulates the expression of genes encoding hepatic amino acid catabolizing enzymes to maintain metabolic homeostasis. Genes Nutr. 10, 452 (2015).
Tobon-Cornejo, S. et al. PPARα/RXRα downregulates amino acid catabolism in the liver via interaction with HNF4α promoting its proteasomal degradation. Metabolism 116, 154705 (2021).
Wu, S. P. et al. Increased COUP-TFII expression in adult hearts induces mitochondrial dysfunction resulting in heart failure. Nat. Commun. 6, 8245 (2015).
Singh, B. K. et al. Thyroid hormone receptor and ERRα coordinately regulate mitochondrial fission, mitophagy, biogenesis, and function. Sci. Signal. 11, eaam5855 (2018).
Koenis, D. S. et al. Nuclear receptor Nur77 limits the macrophage inflammatory response through transcriptional reprogramming of mitochondrial metabolism. Cell Rep. 24, 2127–2140 e2127 (2018).
Giguère, V. Transcriptional control of energy homeostasis by the estrogen-related receptors. Endocr. Rev. 29, 677–696 (2008).
Eichner, L. J. & Giguère, V. Estrogen related receptors (ERRs): a new dawn in transcriptional control of mitochondrial gene networks. Mitochondrion 11, 544–552 (2011).
Sonoda, J. et al. Nuclear receptor ERRα and coactivator PGC-1β are effectors of IFN-γ induced host defense. Genes Dev. 21, 1909–1920 (2007).
Villena, J. A. et al. Orphan nuclear receptor estrogen-related receptor α is essential for adaptive thermogenesis. Proc. Natl Acad. Sci. USA 104, 1418–1423 (2007).
Eichner, L. J. et al. miR-378* mediates metabolic shift in breast cancer cells via the PGC-1β/ERRγ transcriptional pathway. Cell Metab. 12, 352–361 (2010).
Narkar, V. A. et al. Exercise and PGC-1α-independent synchronization of type I muscle metabolism and vasculature by ERRγ. Cell Metab. 13, 283–293 (2011).
Rangwala, S. M. et al. Estrogen-related receptor γ is a key regulator of muscle mitochondrial activity and oxidative capacity. J. Biol. Chem. 285, 22619–22629 (2010).
Cho, Y., Hazen, B. C., Russell, A. P. & Kralli, A. Peroxisome proliferator-activated receptor γ coactivator 1 (PGC-1)- and estrogen-related receptor (ERR)-induced regulator in muscle 1 (Perm1) is a tissue-specific regulator of oxidative capacity in skeletal muscle cells. J. Biol. Chem. 288, 25207–25218 (2013).
Hong, E.-J., Levasseur, M.-P., Dufour, C. R., Perry, M.-C. & Giguère, V. Loss of estrogen-related receptor α promotes hepatocellular carcinogenesis development via metabolic and inflammatory disturbances. Proc. Natl Acad. Sci. USA 110, 17975–17980 (2013).
Pei, L. et al. Dependence of hippocampal function on ERRγ-regulated mitochondrial metabolism. Cell Metab. 21, 628–636 (2015).
Audet-Walsh, E. et al. Androgen-dependent repression of ERRγ reprograms metabolism in prostate cancer. Cancer Res. 77, 378–389 (2017).
Gantner, M. L., Hazen, B. C., Eury, E., Brown, E. L. & Kralli, A. Complementary roles of estrogen-related receptors in brown adipocyte thermogenic function. Endocrinology 157, 4770–4781 (2016).
Fan, W. et al. PPARδ promotes running endurance by preserving glucose. Cell Metab. 25, 1186–1193 e1184 (2017).
Fan, W. et al. ERRγ promotes angiogenesis, mitochondrial biogenesis, and oxidative remodeling in PGC1α/β-deficient muscle. Cell Rep. 22, 2521–2529 (2018).
Vega, R. B. & Kelly, D. P. Cardiac nuclear receptors: architects of mitochondrial structure and function. J. Clin. Invest. 127, 1155–1164 (2017).
Wang, T. et al. Estrogen-related receptor α (ERRα) and ERRγ are essential coordinators of cardiac metabolism and function. Mol. Cell. Biol. 35, 1281–1298 (2015).
Dyar, K. A. et al. Transcriptional programming of lipid and amino acid metabolism by the skeletal muscle circadian clock. PLoS Biol. 16, e2005886 (2018).
DeNicola, G. M. & Cantley, L. C. Cancer’s fuel choice: new flavors for a picky eater. Mol. Cell 60, 514–523 (2015).
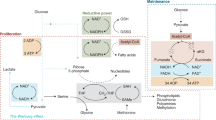
Liberti, M. V. & Locasale, J. W. The Warburg effect: how does it benefit cancer cells? Trends Biochem. Sci. 41, 211–218 (2016).
Vazquez, F. et al. PGC-1α expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell 23, 287–301 (2013).
LeBleu, V. S. et al. PGC-1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 16, 992–1003 (2014).
Srivastava, N. et al. Inhibition of cancer cell proliferation by PPARγ is mediated by a metabolic switch that increases reactive oxygen species levels. Cell Metab. 20, 650–661 (2014).
Park, S. et al. Inhibition of ERRα prevents mitochondrial pyruvate uptake exposing NADPH-generating pathways as targetable vulnerabilities in breast cancer. Cell Rep. 27, 3587–3601 e3584 (2019).
Sies, H. et al. Defining roles of specific reactive oxygen species (ROS) in cell biology and physiology. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-022-00456-z (2022).
Scholtes, C. & Giguère, V. Transcriptional regulation of ROS homeostasis by the ERR subfamily of nuclear receptors. Antioxid. (Basel) 10, 437 (2021).
Vernier, M. et al. Estrogen-related receptors are targetable ROS sensors. Genes Dev. 34, 544–559 (2020).
Brown, E. L. et al. Estrogen-related receptors mediate the adaptive response of brown adipose tissue to adrenergic stimulation. iScience 2, 221–237 (2018).
Nie, Y. & Wong, C. Suppressing the activity of ERRα in 3T3-L1 adipocytes reduces mitochondrial biogenesis but enhances glycolysis and basal glucose uptake. J. Cell. Mol. Med. 13, 3051–3060 (2009).
Schreiber, S. N. et al. The estrogen-related receptor α (ERRα) functions in PPARγ coactivator 1α (PGC-1α)-induced mitochondrial biogenesis. Proc. Natl Acad. Sci. USA 101, 6472–6477 (2004).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Koh, J. H. et al. PPARβ Is essential for maintaining normal levels of PGC-1α and mitochondria and for the increase in muscle mitochondria induced by exercise. Cell Metab. 25, 1176–1185 e1175 (2017).
Mattingly, K. A. et al. Estradiol stimulates transcription of nuclear respiratory factor-1 and increases mitochondrial biogenesis. Mol. Endocrinol. 22, 609–622 (2008).
Li, L. et al. The nuclear orphan receptor COUP-TFII plays an essential role in adipogenesis, glucose homeostasis, and energy metabolism. Cell Metab. 9, 77–87 (2009).
Giacomello, M., Pyakurel, A., Glytsou, C. & Scorrano, L. The cell biology of mitochondrial membrane dynamics. Nat. Rev. Mol. Cell Biol. 21, 204–224 (2020).
Choudhary, V. et al. Novel role of androgens in mitochondrial fission and apoptosis. Mol. Cancer Res. 9, 1067–1077 (2011).
Kim, H. J. et al. Liver-specific deletion of RORα aggravates diet-induced nonalcoholic steatohepatitis by inducing mitochondrial dysfunction. Sci. Rep. 7, 16041 (2017).
Li, Y. et al. Peroxisome proliferator-activated receptor δ regulates mitofusin 2 expression in the heart. J. Mol. Cell. Cardiol. 46, 876–882 (2009).
Cartoni, R. et al. Mitofusins 1/2 and ERRα expression are increased in human skeletal muscle after physical exercise. J. Physiol. 567, 349–358 (2005).
Soriano, F. X. et al. Evidence for a mitochondrial regulatory pathway defined by peroxisome proliferator-activated receptor-γ coactivator-1α, estrogen-related receptor-α, and mitofusin 2. Diabetes 55, 1783–1791 (2006).
Dikic, I. & Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 19, 349–364 (2018).
Kitada, M. & Koya, D. Autophagy in metabolic disease and ageing. Nat. Rev. Endocrinol. 17, 647–661 (2021).
Kim, K. H. et al. Hepatic FXR/SHP axis modulates systemic glucose and fatty acid homeostasis in aged mice. Hepatology 66, 498–509 (2017).
Blessing, A. M. et al. Transcriptional regulation of core autophagy and lysosomal genes by the androgen receptor promotes prostate cancer progression. Autophagy 13, 506–521 (2017).
Mylka, V. et al. The autophagy receptor SQSTM1/p62 mediates anti-inflammatory actions of the selective NR3C1/glucocorticoid receptor modulator compound A (CpdA) in macrophages. Autophagy 14, 2049–2064 (2018).
Silwal, P., Paik, S., Jeon, S. M. & Jo, E. K. Nuclear receptors as autophagy-based antimicrobial therapeutics. Cells 9, 1979 (2020).
Lee, D. H. et al. Mir214-3p and Hnf4α/Hnf4α reciprocally regulate Ulk1 expression and autophagy in nonalcoholic hepatic steatosis. Autophagy 17, 2415–2431 (2021).
Seok, S. et al. Transcriptional regulation of autophagy by an FXR-CREB axis. Nature 516, 108–111 (2014). This study identifies FXR as a key physiological switch regulating autophagy via its repression of autophagy genes, resulting in FXR-dependent sustained nutrient regulation of autophagy during feeding–fasting cycles in mice.
Byun, S. et al. A postprandial FGF19-SHP-LSD1 regulatory axis mediates epigenetic repression of hepatic autophagy. EMBO J. 36, 1755–1769 (2017).
Sinha, R. A. et al. Thyroid hormone stimulates hepatic lipid catabolism via activation of autophagy. J. Clin. Invest. 122, 2428–2438 (2012).
Zhou, J. et al. Thyroid hormone receptor α regulates autophagy, mitochondrial biogenesis, and fatty acid use in skeletal muscle. Endocrinology 162, bqab112 (2021).
Woldt, E. et al. Rev-erb-α modulates skeletal muscle oxidative capacity by regulating mitochondrial biogenesis and autophagy. Nat. Med. 19, 1039–1046 (2013). This work reports that REV-ERBα deficiency induces skeletal muscle autophagy and plays a key role in regulating the oxidative capacity of the muscle and exercise endurance.
Kim, S. Y. et al. ESRRA (estrogen-related receptor α) is a key coordinator of transcriptional and post-translational activation of autophagy to promote innate host defense. Autophagy 14, 152–168 (2018).
Kim, S. et al. ESRRA (estrogen related receptor α) is a critical regulator of intestinal homeostasis through activation of autophagic flux via gut microbiota. Autophagy 17, 2856–2875 (2021).
Tavera-Mendoza, L. E. et al. Vitamin D receptor regulates autophagy in the normal mammary gland and in luminal breast cancer cells. Proc. Natl Acad. Sci. USA 114, E2186–E2194 (2017).
Patch, R. J. et al. Indazole-based ligands for estrogen-related receptor α as potential anti-diabetic agents. Eur. J. Med. Chem. 138, 830–853 (2017).
Patch, R. J. et al. Identification of diaryl ether-based ligands for estrogen-related receptor α as potential antidiabetic agents. J. Med. Chem. 54, 788–808 (2011).
Baar, K. et al. Adaptations of skeletal muscle to exercise: rapid increase in the transcriptional coactivator PGC-1. Federation Am. Societies Exp. Biol. 16, 1879–1886 (2002).
Wilson, B. J., Tremblay, A. M., Deblois, G., Sylvain-Drolet, G. & Giguère, V. An acetylation switch modulates the transcriptional activity of estrogen-related recetpor α. Mol. Endocrinol. 24, 1349–1358 (2010).
Xia, H. et al. Insulin action and resistance is dependent on a GSK3β-FBXW7-ERRα transcriptional axis. Nat. Commun. 13, 2105 (2022).
Francque, S. M. et al. A randomized, controlled trial of the Pan-PPAR agonist lanifibranor in NASH. N. Engl. J. Med. 385, 1547–1558 (2021).
Cariou, B., Charbonnel, B. & Staels, B. Thiazolidinediones and PPARγ agonists: time for a reassessment. Trends Endocrinol. Metab. 23, 205–215 (2012).
Narkar, V. A. et al. AMPK and PPARδ agonists are exercise mimetic. Cell 134, 405–415 (2008).
Fan, W., Atkins, A. R., Yu, R. T., Downes, M. & Evans, R. M. Road to exercise mimetics: targeting nuclear receptors in skeletal muscle. J. Mol. Endocrinol. 51, T87–T100 (2013). Narkar et al. (2008) and Fan et al. (2013) demonstrate and discuss the possibility that orally active drugs targeting nuclear receptors and their coregulators can be used to enhance training adaptation or even to increase endurance without exercise.
Nuclear Receptors Nomenclature Committee. A unified nomenclature system for the nuclear receptors superfamily. Cell 97, 161–163 (1999).
Giguère, V. Editorial: what’s in a name, or the impact of misnomers in endocrine research. Mol. Endocrinol. 29, 789–790 (2015).
Giguère, V., Hollenberg, S. H., Rosenfeld, M. G. & Evans, R. M. Functional domains of the human glucocorticoid receptor. Cell 46, 645–652 (1986).
Chandra, V. et al. Structure of the intact PPAR-γ-RXR-α nuclear receptor complex on DNA. Nature 456, 350–356 (2008).
Giguère, V. et al. Isoform-specific amino-terminal domains dictate DNA-binding properties of RORα, a novel family of orphan nuclear receptors. Genes Dev. 8, 538–553 (1994).
Tremblay, A., Tremblay, G. B., Labrie, F. & Giguère, V. Ligand-independent recruitment of SRC-1 by estrogen receptor b through phosphorylation of activation function AF-1. Mol. Cell 3, 513–519 (1999).
Yu, X. et al. Structural insights of transcriptionally active, full-length androgen receptor coactivator complexes. Mol. Cell 79, 812–823 e814 (2020).
Beinsteiner, B. et al. A structural signature motif enlightens the origin and diversification of nuclear receptors. PLoS Genet. 17, e1009492 (2021).
Johnson, T. A., Paakinaho, V., Kim, S., Hager, G. L. & Presman, D. M. Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms. Nat. Commun. 12, 1987 (2021).
Presman, D. M. et al. DNA binding triggers tetramerization of the glucocorticoid receptor in live cells. Proc. Natl Acad. Sci. USA 113, 8236–8241 (2016).
Evans, R. M. & Mangelsdorf, D. J. Nuclear receptors, RXR, and the Big Bang. Cell 157, 255–266 (2014).
Rivers, C. A. et al. Glucocorticoid receptor-tethered mineralocorticoid receptors increase glucocorticoid-induced transcriptional responses. Endocrinology 160, 1044–1056 (2019).
De Bosscher, K., Desmet, S. J., Clarisse, D., Estebanez-Perpina, E. & Brunsveld, L. Nuclear receptor crosstalk - defining the mechanisms for therapeutic innovation. Nat. Rev. Endocrinol. 16, 363–377 (2020).
Chandra, V. et al. Multidomain integration in the structure of the HNF-4α nuclear receptor complex. Nature 495, 394–398 (2013).
Ellard, S. & Colclough, K. Mutations in the genes encoding the transcription factors hepatocyte nuclear factor 1α (HNF1A) and 4α (HNF4A) in maturity-onset diabetes of the young. Hum. Mutat. 27, 854–869 (2006).
Tin, A. et al. Target genes, variants, tissues and transcriptional pathways influencing human serum urate levels. Nat. Genet. 51, 1459–1474 (2019).
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Molecular Cell Biology thanks David Mangelsdorf and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Ketone bodies
-
Small, water-soluble circulating lipids produced by the adult liver that have an important role as an energy source during starvation, especially for the brain.
- Catabolism
-
A series of chemical reactions involved in the degradation of complex macromolecules resulting in the production of energy and small molecules. Catabolic reactions are opposed by anabolic reactions, which consume energy, leading to the synthesis of complex macromolecules from small molecules.
- Redox
-
Abbreviation for ‘reduction–oxidation’. Refers to a chemical reaction in which the oxidation states of atoms are changed. Oxidation is an increase in oxidation state, and reduction is a decrease in oxidation state. Redox homeostasis is the regulation of reactive oxygen species formation and removal inside the cell.
- Autophagy
-
A lysosome-dependent catabolic process whereby cytoplasmic components are degraded and recycled. Autophagy plays an essential role in maintaining cellular energy homeostasis in response to intracellular stress.
- DNase I hypersensitive site sequencing
-
A molecular biology technique used to identify the location of regulatory regions, based on the genome-wide sequencing of regions sensitive to cleavage by DNase I.
- Assay for transposase-accessible chromatin using sequencing
-
A molecular biology technique used to assess genome-wide chromatin accessibility.
- Pioneer factors
-
A class of transcription factors that can associate with compacted chromatin to facilitate the binding of additional transcription factors.
- Maturity-onset diabetes of the young, type 1
-
A rare hereditary form of diabetes caused by a loss-of-function mutation in the HNF4A gene that usually develops in adolescence or early adulthood. It causes high blood glucose levels from reduced production of insulin.
- Mediator complex
-
A multiprotein complex that mediates the interaction between transcription factors, coregulators and the general transcription machinery.
- Chromatin immunoprecipitation followed by deep sequencing
-
(ChIP–seq). Molecular biology technique used to globally identify the binding sites of DNA-associated proteins in a genome.
- High-throughput chromosome conformation capture
-
(Hi-C). A chromosome conformation capture technique used to analyse the spatial organization of chromatin in a cell. Hi-C quantifies the number of interactions between genomic loci that are adjacent in three-dimensional space but separated by lengthy DNA segments in the linear genome.
- Global run-on followed by sequencing
-
A molecular biology technique used to identify the genes that are being actively transcribed. It provides a detailed catalogue of genes engaged in transcription with quantitative levels of expression.
- Superenhancers
-
A group of enhancers located close to each other (less than 12 kb) in the genome and functionally linked together by the mediator complex.
- Sirtuin
-
Evolutionarily conserved family of epigenetic regulators that modulate the activity of their targets by removing covalently attached acetyl groups. They are metabolic sensors sensitive to NAD+ levels that maintain physiological homeostasis.
- General control non-depressible 5
-
(GCN5). Catalytic subunit of several histone acetyltransferase (HAT) complexes reported to play a range of different functions associated with transcription.
- One-carbon metabolism
-
A group of reactions with a set of enzymes that have in common the transfer of one-carbon groups and that play a role in multiple physiological processes, such as biosynthesis, amino acid homeostasis or epigenetic maintenance.
- Brown fat
-
A type of fat tissue very rich in mitochondria that plays a central role in regulating whole-body energy homeostasis and temperature by its capacity to dissipate macronutrient energy as heat.
- Regulon
-
Group of several genes that are activated or inactivated in response to the same signal by the same regulatory protein.
- Metabolic flexibility
-
The capacity to alter metabolism in response to exercise or available fuel. An organism displays metabolic flexibility if it is capable of metabolizing either carbohydrate or fat efficiently, depending on the availability of those fuels. Metabolic inflexibility is the inability to generate enough energy in response to physiological demands.
- Slow twitch muscle type
-
One of the two main types of muscle fibres. This type contains more mitochondria and depends on aerobic respiration. Their role is sustained, smaller movements and postural control.
- Reactive oxygen species
-
(ROS). Reactive molecules and free radicals or non-radicals derived from molecular oxygen.
- Free radical
-
A chemical species unstable with unpaired electrons.
Rights and permissions
About this article
Cite this article
Scholtes, C., Giguère, V. Transcriptional control of energy metabolism by nuclear receptors. Nat Rev Mol Cell Biol 23, 750–770 (2022). https://doi.org/10.1038/s41580-022-00486-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-022-00486-7
This article is cited by
-
NCOR1/2 and glucocorticoid receptor orchestrate hepatic function
Nature Metabolism (2024)
-
Mitotic bookmarking redundancy by nuclear receptors in pluripotent cells
Nature Structural & Molecular Biology (2024)
-
PPARγ Agonist Rosiglitazone and Antagonist GW9662: Antihypertensive Effects on Chronic Intermittent Hypoxia-Induced Hypertension in Rats
Journal of Cardiovascular Translational Research (2024)
-
Sub-nanosized vanadate hybrid clusters maintain glucose homeostasis and restore treatment response in inflammatory disease in obese mice
Nano Research (2024)
-
Dynamic interplay of nuclear receptors in tumor cell plasticity and drug resistance: Shifting gears in malignant transformations and applications in cancer therapeutics
Cancer and Metastasis Reviews (2024)