Abstract
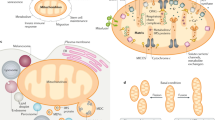
Mitochondria are cellular organelles responsible for generation of chemical energy in the process called oxidative phosphorylation. They originate from a bacterial ancestor and maintain their own genome, which is expressed by designated, mitochondrial transcription and translation machineries that differ from those operating for nuclear gene expression. In particular, the mitochondrial protein synthesis machinery is structurally and functionally very different from that governing eukaryotic, cytosolic translation. Despite harbouring their own genetic information, mitochondria are far from being independent of the rest of the cell and, conversely, cellular fitness is closely linked to mitochondrial function. Mitochondria depend heavily on the import of nuclear-encoded proteins for gene expression and function, and hence engage in extensive inter-compartmental crosstalk to regulate their proteome. This connectivity allows mitochondria to adapt to changes in cellular conditions and also mediates responses to stress and mitochondrial dysfunction. With a focus on mammals and yeast, we review fundamental insights that have been made into the biogenesis, architecture and mechanisms of the mitochondrial translation apparatus in the past years owing to the emergence of numerous near-atomic structures and a considerable amount of biochemical work. Moreover, we discuss how cellular mitochondrial protein expression is regulated, including aspects of mRNA and tRNA maturation and stability, roles of auxiliary factors, such as translation regulators, that adapt mitochondrial translation rates, and the importance of inter-compartmental crosstalk with nuclear gene expression and cytosolic translation and how it enables integration of mitochondrial translation into the cellular context.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Martijn, J., Vosseberg, J., Guy, L., Offre, P. & Ettema, T. J. G. Deep mitochondrial origin outside the sampled alphaproteobacteria. Nature 557, 101–105 (2018).
Spinelli, J. B. & Haigis, M. C. The multifaceted contributions of mitochondria to cellular metabolism. Nat. Cell Biol. 20, 745–754 (2018).
Pearce, S., Nezich, C. L. & Spinazzola, A. Mitochondrial diseases: translation matters. Mol. Cell Neurosci. 55, 1–12 (2013).
Boczonadi, V. & Horvath, R. Mitochondria: impaired mitochondrial translation in human disease. Int. J. Biochem. Cell Biol. 48, 77–84 (2014).
Sun, N., Youle, R. J. & Finkel, T. The mitochondrial basis of aging. Mol. Cell 61, 654–666 (2016).
Greber, B. J. & Ban, N. Structure and function of the mitochondrial ribosome. Annu. Rev. Biochem. 85, 103–132 (2016).
Zhang, L. et al. Antibiotic susceptibility of mammalian mitochondrial translation. FEBS Lett. 579, 6423–6427 (2005).
Waltz, F. & Giege, P. Striking diversity of mitochondria-specific translation processes across eukaryotes. Trends Biochem. Sci. 45, 149–162 (2020).
Ojala, D., Montoya, J. & Attardi, G. tRNA punctuation model of RNA processing in human mitochondria. Nature 290, 470–474 (1981). This paper is the first description of the tRNA punctuation model of mitochondrial RNA processing.
Rackham, O. et al. Hierarchical RNA processing is required for mitochondrial ribosome assembly. Cell Rep. 16, 1874–1890 (2016).
Siira, S. J. et al. Concerted regulation of mitochondrial and nuclear non-coding RNAs by a dual-targeted RNase Z. EMBO Rep. https://doi.org/10.15252/embr.201846198 (2018).
Merten, S., Synenki, R. M., Locker, J., Christianson, T. & Rabinowitz, M. Processing of precursors of 21S ribosomal RNA from yeast mitochondria. Proc. Natl Acad. Sci. USA 77, 1417–1421 (1980).
Bogenhagen, D. F., Martin, D. W. & Koller, A. Initial steps in RNA processing and ribosome assembly occur at mitochondrial DNA nucleoids. Cell Metab. 19, 618–629 (2014).
Dalla Rosa, I. et al. MPV17L2 is required for ribosome assembly in mitochondria. Nucleic Acids Res. 42, 8500–8515 (2014).
Jourdain, A. A. et al. GRSF1 regulates RNA processing in mitochondrial RNA granules. Cell Metab. 17, 399–410 (2013).
Antonicka, H. & Shoubridge, E. A. Mitochondrial RNA granules are centers for posttranscriptional RNA processing and ribosome biogenesis. Cell Rep. 10, 920–932 (2015).
Tu, Y. T. & Barrientos, A. The human mitochondrial DEAD-box protein DDX28 resides in RNA granules and functions in mitoribosome assembly. Cell Rep. 10, 854–864 (2015).
Maiti, P., Kim, H. J., Tu, Y. T. & Barrientos, A. Human GTPBP10 is required for mitoribosome maturation. Nucleic Acids Res. 46, 11423–11437 (2018).
Sloan, K. E. et al. Tuning the ribosome: the influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol. 14, 1138–1152 (2017).
Van Haute, L. et al. METTL15 introduces N4-methylcytidine into human mitochondrial 12S rRNA and is required for mitoribosome biogenesis. Nucleic Acids Res. 47, 10267–10281 (2019).
De Silva, D., Tu, Y. T., Amunts, A., Fontanesi, F. & Barrientos, A. Mitochondrial ribosome assembly in health and disease. Cell Cycle 14, 2226–2250 (2015).
Brown, A. et al. Structures of the human mitochondrial ribosome in native states of assembly. Nat. Struct. Mol. Biol. 24, 866–869 (2017).
Itoh, Y., Naschberger, A., Mortezaei, N., Herrmann, J. M. & Amunts, A. Analysis of translating mitoribosome reveals functional characteristics of translation in mitochondria of fungi. Preprint at bioRxiv https://doi.org/10.1101/2020.01.31.929331 (2020).
Davis, J. H. & Williamson, J. R. Structure and dynamics of bacterial ribosome biogenesis. Philos. Trans. R Soc. Lond. B Biol. Sci. https://doi.org/10.1098/rstb.2016.0181 (2017).
Kargas, V. et al. Mechanism of completion of peptidyltransferase centre assembly in eukaryotes. eLife https://doi.org/10.7554/eLife.44904 (2019).
Ma, C. et al. Structural snapshot of cytoplasmic pre-60S ribosomal particles bound by Nmd3, Lsg1, Tif6 and Reh1. Nat. Struct. Mol. Biol. 24, 214–220 (2017).
Malyutin, A. G., Musalgaonkar, S., Patchett, S., Frank, J. & Johnson, A. W. Nmd3 is a structural mimic of eIF5A, and activates the cpGTPase Lsg1 during 60S ribosome biogenesis. EMBO J. 36, 854–868 (2017).
Jomaa, A. et al. Functional domains of the 50S subunit mature late in the assembly process. Nucleic Acids Res. 42, 3419–3435 (2014).
Davis, J. H. et al. Modular assembly of the bacterial large ribosomal subunit. Cell 167, 1610–1622.e15 (2016).
Klinge, S. & Woolford, J. L. Jr. Ribosome assembly coming into focus. Nat. Rev. Mol. Cell Biol. 20, 116–131 (2019).
Ramrath, D. J. F. et al. Evolutionary shift toward protein-based architecture in trypanosomal mitochondrial ribosomes. Science https://doi.org/10.1126/science.aau7735 (2018). This study is the first visualization of an extreme case of mitoribosomal diversification with a minimal rRNA core and a giant ribosomal protein shell.
Saurer, M. et al. Mitoribosomal small subunit biogenesis in trypanosomes involves an extensive assembly machinery. Science 365, 1144–1149 (2019).
Jaskolowski, M. et al. Structural insights into the mechanism of mitoribosomal large subunit biogenesis. Mol. Cell 79, 629–644 (2020).
Soufari, H. et al. Structure of the mature kinetoplastids mitoribosome and insights into its large subunit biogenesis. Proc. Natl Acad. Sci. USA 117, 29851–29861 (2020).
Tobiasson, V. et al. Interconnected assembly factors regulate the biogenesis of mitoribosomal large subunit. Preprint at bioRxiv https://doi.org/10.1101/2020.06.28.176446 (2020).
Bogenhagen, D. F., Ostermeyer-Fay, A. G., Haley, J. D. & Garcia-Diaz, M. Kinetics and mechanism of mammalian mitochondrial ribosome assembly. Cell Rep. 22, 1935–1944 (2018).
Zeng, R., Smith, E. & Barrientos, A. Yeast mitoribosome large subunit assembly proceeds by hierarchical incorporation of protein clusters and modules on the inner membrane. Cell Metab. 27, 645–656.e7 (2018).
Kaur, J. & Stuart, R. A. Truncation of the Mrp20 protein reveals new ribosome-assembly subcomplex in mitochondria. EMBO Rep. 12, 950–955 (2011).
Chen, S. S., Sperling, E., Silverman, J. M., Davis, J. H. & Williamson, J. R. Measuring the dynamics of E. coli ribosome biogenesis using pulse-labeling and quantitative mass spectrometry. Mol. Biosyst. 8, 3325–3334 (2012).
De Silva, D., Fontanesi, F. & Barrientos, A. The DEAD box protein Mrh4 functions in the assembly of the mitochondrial large ribosomal subunit. Cell Metab. 18, 712–725 (2013).
Rodnina, M. V. Translation in prokaryotes. Cold Spring Harb. Perspect Biol. https://doi.org/10.1101/cshperspect.a032664 (2018).
Weisser, M. & Ban, N. Extensions, extra factors, and extreme complexity: ribosomal structures provide insights into eukaryotic translation. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a032367 (2019).
Spencer, A. C. & Spremulli, L. L. Interaction of mitochondrial initiation factor 2 with mitochondrial fMet-tRNA. Nucleic Acids Res. 32, 5464–5470 (2004).
Roll-Mecak, A., Shin, B. S., Dever, T. E. & Burley, S. K. Engaging the ribosome: universal IFs of translation. Trends Biochem. Sci. 26, 705–709 (2001).
Carter, A. P. et al. Crystal structure of an initiation factor bound to the 30S ribosomal subunit. Science 291, 498–501 (2001).
Lomakin, I. B. & Steitz, T. A. The initiation of mammalian protein synthesis and mRNA scanning mechanism. Nature 500, 307–311 (2013).
Hussain, T. et al. Structural changes enable start codon recognition by the eukaryotic translation initiation complex. Cell 159, 597–607 (2014).
Weisser, M., Voigts-Hoffmann, F., Rabl, J., Leibundgut, M. & Ban, N. The crystal structure of the eukaryotic 40S ribosomal subunit in complex with eIF1 and eIF1A. Nat. Struct. Mol. Biol. 20, 1015–1017 (2013).
Atkinson, G. C. et al. Evolutionary and genetic analyses of mitochondrial translation initiation factors identify the missing mitochondrial IF3 in S. cerevisiae. Nucleic Acids Res. 40, 6122–6134 (2012).
Gaur, R. et al. A single mammalian mitochondrial translation initiation factor functionally replaces two bacterial factors. Mol. Cell 29, 180–190 (2008). This paper is the first description that a mitochondria-specific insertion in an initiation factor compensates for the loss of bacterial IF1.
Yassin, A. S. et al. Insertion domain within mammalian mitochondrial translation initiation factor 2 serves the role of eubacterial initiation factor 1. Proc. Natl Acad. Sci. USA 108, 3918–3923 (2011).
Kummer, E. et al. Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM. Nature 560, 263–267 (2018). This paper is the first structural report on a complete, reconstituted mitochondrial translation complex.
Kozak, M. Point mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Cell 44, 283–292 (1986).
Hinnebusch, A. G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 83, 779–812 (2014).
Herrmann, J. M., Woellhaf, M. W. & Bonnefoy, N. Control of protein synthesis in yeast mitochondria: the concept of translational activators. Biochim. Biophys. Acta 1833, 286–294 (2013).
Temperley, R. J., Wydro, M., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Human mitochondrial mRNAs — like members of all families, similar but different. Biochim. Biophys. Acta 1797, 1081–1085 (2010).
Montoya, J., Ojala, D. & Attardi, G. Distinctive features of the 5′-terminal sequences of the human mitochondrial mRNAs. Nature 290, 465–470 (1981).
Greber, B. J. et al. Ribosome. The complete structure of the 55S mammalian mitochondrial ribosome. Science 348, 303–308 (2015).
Christian, B. E. & Spremulli, L. L. Preferential selection of the 5′-terminal start codon on leaderless mRNAs by mammalian mitochondrial ribosomes. J. Biol. Chem. 285, 28379–28386 (2010).
Ayyub, S. A. et al. Fidelity of translation in the presence of mammalian mitochondrial initiation factor 3. Mitochondrion 39, 1–8 (2018).
Bhargava, K. & Spremulli, L. L. Role of the N- and C-terminal extensions on the activity of mammalian mitochondrial translational initiation factor 3. Nucleic Acids Res. 33, 7011–7018 (2005).
Haque, M. E. & Spremulli, L. L. Roles of the N- and C-terminal domains of mammalian mitochondrial initiation factor 3 in protein biosynthesis. J. Mol. Biol. 384, 929–940 (2008).
Rudler, D. L. et al. Fidelity of translation initiation is required for coordinated respiratory complex assembly. Sci. Adv. 5, eaay2118 (2019).
Koripella, R. K. et al. Structure of human mitochondrial translation initiation factor 3 bound to the small ribosomal subunit. iScience 12, 76–86 (2019).
Khawaja, A. et al. Distinct pre-initiation steps in human mitochondrial translation. Nat. Commun. 11, 2932 (2020).
Bilbille, Y. et al. The human mitochondrial tRNAMet: structure/function relationship of a unique modification in the decoding of unconventional codons. J. Mol. Biol. 406, 257–274 (2011).
Nakano, S. et al. NSUN3 methylase initiates 5-formylcytidine biogenesis in human mitochondrial tRNA(Met). Nat. Chem. Biol. 12, 546–551 (2016).
Haag, S. et al. NSUN3 and ABH1 modify the wobble position of mt-tRNAMet to expand codon recognition in mitochondrial translation. EMBO J. 35, 2104–2119 (2016).
Van Haute, L. et al. Deficient methylation and formylation of mt-tRNAMet wobble cytosine in a patient carrying mutations in NSUN3. Nat. Commun. 7, 12039 (2016).
Schwartzbach, C. J. & Spremulli, L. L. Bovine mitochondrial protein synthesis elongation factors. Identification and initial characterization of an elongation factor Tu-elongation factor Ts complex. J. Biol. Chem. 264, 19125–19131 (1989).
Woriax, V. L., Bullard, J. M., Ma, L., Yokogawa, T. & Spremulli, L. L. Mechanistic studies of the translational elongation cycle in mammalian mitochondria. Biochim. Biophys. Acta 1352, 91–101 (1997).
Cai, Y. C., Bullard, J. M., Thompson, N. L. & Spremulli, L. L. Interaction of mitochondrial elongation factor Tu with aminoacyl-tRNA and elongation factor Ts. J. Biol. Chem. 275, 20308–20314 (2000).
Jeppesen, M. G., Navratil, T., Spremulli, L. L. & Nyborg, J. Crystal structure of the bovine mitochondrial elongation factor Tu.Ts complex. J. Biol. Chem. 280, 5071–5081 (2005).
Hammarsund, M. et al. Identification and characterization of two novel human mitochondrial elongation factor genes, hEFG2 and hEFG1, phylogenetically conserved through evolution. Hum. Genet. 109, 542–550 (2001).
Tsuboi, M. et al. EF-G2mt is an exclusive recycling factor in mammalian mitochondrial protein synthesis. Mol. Cell 35, 502–510 (2009). This paper is the first description of mtEFG1 and mtEFG2 having divergent functions in the translation cycle.
Bhargava, K., Templeton, P. & Spremulli, L. L. Expression and characterization of isoform 1 of human mitochondrial elongation factor G. Protein Expr. Purif. 37, 368–376 (2004).
Chung, H. K. & Spremulli, L. L. Purification and characterization of elongation factor G from bovine liver mitochondria. J. Biol. Chem. 265, 21000–21004 (1990).
Eberly, S. L., Locklear, V. & Spremulli, L. L. Bovine mitochondrial ribosomes. Elongation factor specificity. J. Biol. Chem. 260, 8721–8725 (1985).
Koripella, R. K. et al. Structures of the human mitochondrial ribosome bound to EF-G1 reveal distinct features of mitochondrial translation elongation. Nat. Commun. 11, 3830 (2020).
Kummer, E. & Ban, N. Structural insights into mammalian mitochondrial translation elongation catalyzed by mtEFG1. EMBO J. https://doi.org/10.15252/embj.2020104820 (2020).
Zhou, J., Lancaster, L., Donohue, J. P. & Noller, H. F. Spontaneous ribosomal translocation of mRNA and tRNAs into a chimeric hybrid state. Proc. Natl Acad. Sci. USA 116, 7813–7818 (2019).
Peng, B. Z. et al. Active role of elongation factor G in maintaining the mRNA reading frame during translation. Sci. Adv. 5, eaax8030 (2019).
Bauerschmitt, H., Funes, S. & Herrmann, J. M. The membrane-bound GTPase Guf1 promotes mitochondrial protein synthesis under suboptimal conditions. J. Biol. Chem. 283, 17139–17146 (2008).
Yang, F. et al. Mitochondrial EF4 links respiratory dysfunction and cytoplasmic translation in Caenorhabditis elegans. Biochim. Biophys. Acta 1837, 1674–1683 (2014).
Jia, L. et al. Yeast Oxa1 interacts with mitochondrial ribosomes: the importance of the C-terminal region of Oxa1. EMBO J. 22, 6438–6447 (2003).
Szyrach, G., Ott, M., Bonnefoy, N., Neupert, W. & Herrmann, J. M. Ribosome binding to the Oxa1 complex facilitates co-translational protein insertion in mitochondria. EMBO J. 22, 6448–6457 (2003).
Ott, M. et al. Mba1, a membrane-associated ribosome receptor in mitochondria. EMBO J. 25, 1603–1610 (2006).
Haque, M. E. et al. Properties of the C-terminal tail of human mitochondrial inner membrane protein Oxa1L and its interactions with mammalian mitochondrial ribosomes. J. Biol. Chem. 285, 28353–28362 (2010).
Escobar-Alvarez, S. et al. Inhibition of human peptide deformylase disrupts mitochondrial function. Mol. Cell Biol. 30, 5099–5109 (2010).
Lee, M. D. et al. A new human peptide deformylase inhibitable by actinonin. Biochem. Biophys. Res. Commun. 312, 309–315 (2003).
Nguyen, K. T. et al. Characterization of a human peptide deformylase: implications for antibacterial drug design. Biochemistry 42, 9952–9958 (2003).
Serero, A., Giglione, C., Sardini, A., Martinez-Sanz, J. & Meinnel, T. An unusual peptide deformylase features in the human mitochondrial N-terminal methionine excision pathway. J. Biol. Chem. 278, 52953–52963 (2003).
Leszczyniecka, M. et al. MAP1D, a novel methionine aminopeptidase family member is overexpressed in colon cancer. Oncogene 25, 3471–3478 (2006).
Greber, B. J. et al. Architecture of the large subunit of the mammalian mitochondrial ribosome. Nature 505, 515–519 (2014).
Pfeffer, S., Woellhaf, M. W., Herrmann, J. M. & Forster, F. Organization of the mitochondrial translation machinery studied in situ by cryoelectron tomography. Nat. Commun. 6, 6019 (2015).
Englmeier, R., Pfeffer, S. & Forster, F. Structure of the human mitochondrial ribosome studied in situ by cryoelectron tomography. Structure 25, 1574–1581 e1572 (2017). Together with Pfeffer et al. (2015), this paper is the first visualization of mitochondrial ribosomes inside the mitochondrion.
Lee, R. G. et al. Cardiolipin is required for membrane docking of mitochondrial ribosomes and protein synthesis. J. Cell Sci. https://doi.org/10.1242/jcs.240374 (2020).
Barrell, B. G., Bankier, A. T. & Drouin, J. A different genetic code in human mitochondria. Nature 282, 189–194 (1979).
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981).
Akabane, S., Ueda, T., Nierhaus, K. H. & Takeuchi, N. Ribosome rescue and translation termination at non-standard stop codons by ICT1 in mammalian mitochondria. PLoS Genet. 10, e1004616 (2014).
Soleimanpour-Lichaei, H. R. et al. mtRF1a is a human mitochondrial translation release factor decoding the major termination codons UAA and UAG. Mol. Cell 27, 745–757 (2007). This study discovers that mtRF1a is the translation termination factor recognizing canonical stop codons.
Nozaki, Y., Matsunaga, N., Ishizawa, T., Ueda, T. & Takeuchi, N. HMRF1L is a human mitochondrial translation release factor involved in the decoding of the termination codons UAA and UAG. Genes Cell 13, 429–438 (2008).
Richter, R. et al. A functional peptidyl-tRNA hydrolase, ICT1, has been recruited into the human mitochondrial ribosome. EMBO J. 29, 1116–1125 (2010).
Kogure, H. et al. Identification of residues required for stalled-ribosome rescue in the codon-independent release factor YaeJ. Nucleic Acids Res. 42, 3152–3163 (2014).
Wesolowska, M. T., Richter-Dennerlein, R., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Overcoming stalled translation in human mitochondria. Front. Microbiol. 5, 374 (2014).
Feaga, H. A., Quickel, M. D., Hankey-Giblin, P. A. & Keiler, K. C. Human cells require non-stop ribosome rescue activity in mitochondria. PLoS Genet. 12, e1005964 (2016).
Lind, C., Sund, J. & Aqvist, J. Codon-reading specificities of mitochondrial release factors and translation termination at non-standard stop codons. Nat. Commun. 4, 2940 (2013).
Temperley, R., Richter, R., Dennerlein, S., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Hungry codons promote frameshifting in human mitochondrial ribosomes. Science 327, 301 (2010).
Young, D. J. et al. Bioinformatic, structural, and functional analyses support release factor-like MTRF1 as a protein able to decode nonstandard stop codons beginning with adenine in vertebrate mitochondria. RNA 16, 1146–1155 (2010).
Desai, N. et al. Elongational stalling activates mitoribosome-associated quality control. Science 370, 1105–1110 (2020). This paper is the first description of a mitoribosomal rescue pathway that alleviates ribosome stalling.
Pearce, S. F. et al. Maturation of selected human mitochondrial tRNAs requires deadenylation. eLife https://doi.org/10.7554/eLife.27596 (2017).
Gopalakrishna, S. et al. C6orf203 is an RNA-binding protein involved in mitochondrial protein synthesis. Nucleic Acids Res. 47, 9386–9399 (2019).
Christian, B. E. & Spremulli, L. L. Evidence for an active role of IF3mt in the initiation of translation in mammalian mitochondria. Biochemistry 48, 3269–3278 (2009).
Lavdovskaia, E. et al. Dual function of GTPBP6 in biogenesis and recycling of human mitochondrial ribosomes. Nucleic Acids Res. 48, 12929–12942 (2020).
Zhang, Y. et al. HflX is a ribosome-splitting factor rescuing stalled ribosomes under stress conditions. Nat. Struct. Mol. Biol. 22, 906–913 (2015).
Chujo, T. et al. LRPPRC/SLIRP suppresses PNPase-mediated mRNA decay and promotes polyadenylation in human mitochondria. Nucleic Acids Res. 40, 8033–8047 (2012).
Nagao, A., Hino-Shigi, N. & Suzuki, T. Measuring mRNA decay in human mitochondria. Methods Enzymol. 447, 489–499 (2008).
Piechota, J. et al. Differential stability of mitochondrial mRNA in HeLa cells. Acta Biochim. Pol. 53, 157–168 (2006).
Jourdain, A. A. et al. The FASTK family of proteins: emerging regulators of mitochondrial RNA biology. Nucleic Acids Res. 45, 10941–10947 (2017).
Lagouge, M. et al. SLIRP regulates the rate of mitochondrial protein synthesis and protects LRPPRC from degradation. PLoS Genet. 11, e1005423 (2015).
Ruzzenente, B. et al. LRPPRC is necessary for polyadenylation and coordination of translation of mitochondrial mRNAs. EMBO J. 31, 443–456 (2012).
Kehrein, K. et al. Organization of mitochondrial gene expression in two distinct ribosome-containing assemblies. Cell Rep. 10, 843–853 (2015).
Aibara, S., Singh, V., Modelska, A. & Amunts, A. Structural basis of mitochondrial translation. eLife https://doi.org/10.7554/eLife.58362 (2020).
Popow, J. et al. FASTKD2 is an RNA-binding protein required for mitochondrial RNA processing and translation. RNA 21, 1873–1884 (2015).
Hillen, H. S., Temiakov, D. & Cramer, P. Structural basis of mitochondrial transcription. Nat. Struct. Mol. Biol. 25, 754–765 (2018).
Barchiesi, A. & Vascotto, C. Transcription, processing, and decay of mitochondrial RNA in health and disease. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20092221 (2019).
Gustafsson, C. M., Falkenberg, M. & Larsson, N. G. Maintenance and expression of mammalian mitochondrial DNA. Annu. Rev. Biochem. 85, 133–160 (2016).
Holzmann, J. et al. RNase P without RNA: identification and functional reconstitution of the human mitochondrial tRNA processing enzyme. Cell 135, 462–474 (2008). This paper identifies the unusual protein-only RNase P in mitochondria.
Brzezniak, L. K., Bijata, M., Szczesny, R. J. & Stepien, P. P. Involvement of human ELAC2 gene product in 3′ end processing of mitochondrial tRNAs. RNA Biol. 8, 616–626 (2011).
Rossmanith, W. Localization of human RNase Z isoforms: dual nuclear/mitochondrial targeting of the ELAC2 gene product by alternative translation initiation. PLoS ONE 6, e19152 (2011).
Sanchez, M. I. et al. RNA processing in human mitochondria. Cell Cycle 10, 2904–2916 (2011).
Nagaike, T. et al. Identification and characterization of mammalian mitochondrial tRNA nucleotidyltransferases. J. Biol. Chem. 276, 40041–40049 (2001).
Mercer, T. R. et al. The human mitochondrial transcriptome. Cell 146, 645–658 (2011).
Ali, A. T. et al. Nuclear genetic regulation of the human mitochondrial transcriptome. eLife https://doi.org/10.7554/eLife.41927 (2019).
Borowski, L. S., Dziembowski, A., Hejnowicz, M. S., Stepien, P. P. & Szczesny, R. J. Human mitochondrial RNA decay mediated by PNPase–hSuv3 complex takes place in distinct foci. Nucleic Acids Res. 41, 1223–1240 (2013).
Szczesny, R. J. et al. Human mitochondrial RNA turnover caught in flagranti: involvement of hSuv3p helicase in RNA surveillance. Nucleic Acids Res. 38, 279–298 (2010).
Nagaike, T., Suzuki, T., Katoh, T. & Ueda, T. Human mitochondrial mRNAs are stabilized with polyadenylation regulated by mitochondria-specific poly(A) polymerase and polynucleotide phosphorylase. J. Biol. Chem. 280, 19721–19727 (2005).
Wang, D. D. et al. Helicase SUV3, polynucleotide phosphorylase, and mitochondrial polyadenylation polymerase form a transient complex to modulate mitochondrial mRNA polyadenylated tail lengths in response to energetic changes. J. Biol. Chem. 289, 16727–16735 (2014).
Jourdain, A. A. et al. A mitochondria-specific isoform of FASTK is present in mitochondrial RNA granules and regulates gene expression and function. Cell Rep. 10, 1110–1121 (2015).
Pietras, Z. et al. Dedicated surveillance mechanism controls G-quadruplex forming non-coding RNAs in human mitochondria. Nat. Commun. 9, 2558 (2018). This paper is the first description of the specific role of GRSF1 in removal of G quadruplex-containing light-strand transcripts to preserve the mitochondrial transcriptome.
Wang, G. et al. PNPASE regulates RNA import into mitochondria. Cell 142, 456–467 (2010).
Silva, S., Camino, L. P. & Aguilera, A. Human mitochondrial degradosome prevents harmful mitochondrial R loops and mitochondrial genome instability. Proc. Natl Acad. Sci. USA 115, 11024–11029 (2018).
Dhir, A. et al. Mitochondrial double-stranded RNA triggers antiviral signalling in humans. Nature 560, 238–242 (2018).
Siira, S. J. et al. LRPPRC-mediated folding of the mitochondrial transcriptome. Nat. Commun. 8, 1532 (2017).
Sasarman, F. et al. LRPPRC and SLIRP interact in a ribonucleoprotein complex that regulates posttranscriptional gene expression in mitochondria. Mol. Biol. Cell 21, 1315–1323 (2010). Together with Siira et al. (2017), this paper discovers LRPPRC–SLIRP as a general mitochondrial RNA chaperone.
Wilson, W. C. et al. A human mitochondrial poly(A) polymerase mutation reveals the complexities of post-transcriptional mitochondrial gene expression. Hum. Mol. Genet. 23, 6345–6355 (2014).
Wolf, A. R. & Mootha, V. K. Functional genomic analysis of human mitochondrial RNA processing. Cell Rep. 7, 918–931 (2014).
Boehm, E. et al. FASTKD1 and FASTKD4 have opposite effects on expression of specific mitochondrial RNAs, depending upon their endonuclease-like RAP domain. Nucleic Acids Res. 45, 6135–6146 (2017).
Boehm, E. et al. Role of FAST kinase domains 3 (FASTKD3) in post-transcriptional regulation of mitochondrial gene expression. J. Biol. Chem. 291, 25877–25887 (2016).
Antonicka, H., Sasarman, F., Nishimura, T., Paupe, V. & Shoubridge, E. A. The mitochondrial RNA-binding protein GRSF1 localizes to RNA granules and is required for posttranscriptional mitochondrial gene expression. Cell Metab. 17, 386–398 (2013). Together with Jourdain et al. (2013) and Antonicka and Shoubridge (2015), this paper is the first description of mitochondrial RNA granules.
Rackham, O. et al. Long noncoding RNAs are generated from the mitochondrial genome and regulated by nuclear-encoded proteins. RNA 17, 2085–2093 (2011).
Levy, S. & Schuster, G. Polyadenylation and degradation of RNA in the mitochondria. Biochem. Soc. Trans. 44, 1475–1482 (2016).
Slomovic, S., Portnoy, V., Liveanu, V. & Schuster, G. RNA polyadenylation in prokaryotes and organelles; different tails tell different tales. Crit. Rev. Plant. Sci. 25, 65–77 (2007).
Hammani, K. & Giege, P. RNA metabolism in plant mitochondria. Trends Plant. Sci. 19, 380–389 (2014).
Tomecki, R., Dmochowska, A., Gewartowski, K., Dziembowski, A. & Stepien, P. P. Identification of a novel human nuclear-encoded mitochondrial poly(A) polymerase. Nucleic Acids Res. 32, 6001–6014 (2004).
Hirsch, M. & Penman, S. Post-transcriptional addition of polyadenylic acid to mitochondrial RNA by a cordycepin-insensitive process. J. Mol. Biol. 83, 131–142 (1974).
Fiedler, M., Rossmanith, W., Wahle, E. & Rammelt, C. Mitochondrial poly(A) polymerase is involved in tRNA repair. Nucleic Acids Res. 43, 9937–9949 (2015).
Bratic, A. et al. Mitochondrial polyadenylation is a one-step process required for mRNA integrity and tRNA maturation. PLoS Genet. 12, e1006028 (2016).
Slomovic, S., Laufer, D., Geiger, D. & Schuster, G. Polyadenylation and degradation of human mitochondrial RNA: the prokaryotic past leaves its mark. Mol. Cell Biol. 25, 6427–6435 (2005).
Toompuu, M. et al. Polyadenylation and degradation of structurally abnormal mitochondrial tRNAs in human cells. Nucleic Acids Res. 46, 5209–5226 (2018).
Kuznetsova, I. et al. Simultaneous processing and degradation of mitochondrial RNAs revealed by circularized RNA sequencing. Nucleic Acids Res. 45, 5487–5500 (2017).
Temperley, R. J. et al. Investigation of a pathogenic mtDNA microdeletion reveals a translation-dependent deadenylation decay pathway in human mitochondria. Hum. Mol. Genet. 12, 2341–2348 (2003).
Wydro, M., Bobrowicz, A., Temperley, R. J., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Targeting of the cytosolic poly(A) binding protein PABPC1 to mitochondria causes mitochondrial translation inhibition. Nucleic Acids Res. 38, 3732–3742 (2010).
Slomovic, S. & Schuster, G. Stable PNPase RNAi silencing: its effect on the processing and adenylation of human mitochondrial RNA. RNA 14, 310–323 (2008).
Rorbach, J., Nicholls, T. J. & Minczuk, M. PDE12 removes mitochondrial RNA poly(A) tails and controls translation in human mitochondria. Nucleic Acids Res. 39, 7750–7763 (2011).
Poulsen, J. B. et al. Human 2′-phosphodiesterase localizes to the mitochondrial matrix with a putative function in mitochondrial RNA turnover. Nucleic Acids Res. 39, 3754–3770 (2011).
Couvillion, M. T., Soto, I. C., Shipkovenska, G. & Churchman, L. S. Synchronized mitochondrial and cytosolic translation programs. Nature 533, 499–503 (2016). This paper describes unidirectional coupling of cytosolic and mitochondrial translation in yeast.
Isaac, R. S., McShane, E. & Churchman, L. S. The multiple levels of mitonuclear coregulation. Annu. Rev. Genet. 52, 511–533 (2018).
Weraarpachai, W. et al. Mutation in TACO1, encoding a translational activator of COX I, results in cytochrome c oxidase deficiency and late-onset Leigh syndrome. Nat. Genet. 41, 833–837 (2009).
Richman, T. R. et al. Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice. Nat. Commun. 7, 11884 (2016).
Richter-Dennerlein, R. et al. Mitochondrial protein synthesis adapts to influx of nuclear-encoded protein. Cell 167, 471–483.e10 (2016).
Wang, C. et al. MITRAC15/COA1 promotes mitochondrial translation in a ND2 ribosome–nascent chain complex. EMBO Rep. 21, e48833 (2020).
Mick, D. U. et al. MITRAC links mitochondrial protein translocation to respiratory-chain assembly and translational regulation. Cell 151, 1528–1541 (2012). This paper is the first description of the role of OXPHOS assembly factors in feedback regulation of mitochondrial translation to the import of nuclear-encoded OXPHOS subunits in humans.
Weraarpachai, W. et al. Mutations in C12orf62, a factor that couples COX I synthesis with cytochrome c oxidase assembly, cause fatal neonatal lactic acidosis. Am. J. Hum. Genet. 90, 142–151 (2012).
Bogenhagen, D. F. & Haley, J. D. Pulse-chase SILAC-based analyses reveal selective oversynthesis and rapid turnover of mitochondrial protein components of respiratory complexes. J. Biol. Chem. 295, 2544–2554 (2020).
Quiros, P. M., Mottis, A. & Auwerx, J. Mitonuclear communication in homeostasis and stress. Nat. Rev. Mol. Cell Biol. 17, 213–226 (2016).
Michel, S., Canonne, M., Arnould, T. & Renard, P. Inhibition of mitochondrial genome expression triggers the activation of CHOP-10 by a cell signaling dependent on the integrated stress response but not the mitochondrial unfolded protein response. Mitochondrion 21, 58–68 (2015).
Houtkooper, R. H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Quiros, P. M. et al. Multi-omics analysis identifies ATF4 as a key regulator of the mitochondrial stress response in mammals. J. Cell Biol. 216, 2027–2045 (2017).
Ferreira, N. et al. Stress signaling and cellular proliferation reverse the effects of mitochondrial mistranslation. EMBO J. 38, e102155 (2019).
Munch, C. & Harper, J. W. Mitochondrial unfolded protein response controls matrix pre-RNA processing and translation. Nature 534, 710–713 (2016).
Fiorese, C. J. et al. The transcription factor ATF5 mediates a mammalian mitochondrial UPR. Curr. Biol. 26, 2037–2043 (2016).
Weidberg, H. & Amon, A. MitoCPR — a surveillance pathway that protects mitochondria in response to protein import stress. Science https://doi.org/10.1126/science.aan4146 (2018).
Wang, X. & Chen, X. J. A cytosolic network suppressing mitochondria-mediated proteostatic stress and cell death. Nature 524, 481–484 (2015).
Wrobel, L. et al. Mistargeted mitochondrial proteins activate a proteostatic response in the cytosol. Nature 524, 485–488 (2015).
Boos, F. et al. Mitochondrial protein-induced stress triggers a global adaptive transcriptional programme. Nat. Cell Biol. 21, 442–451 (2019). This paper identifies the integration of mitochondrial precursor accumulation stress into the general heat shock response.
Suhm, T. et al. Mitochondrial translation efficiency controls cytoplasmic protein homeostasis. Cell Metab. 27, 1309–1322.e6 (2018).
Zhao, Q. et al. A mitochondrial specific stress response in mammalian cells. EMBO J. 21, 4411–4419 (2002).
Aldridge, J. E., Horibe, T. & Hoogenraad, N. J. Discovery of genes activated by the mitochondrial unfolded protein response (mtUPR) and cognate promoter elements. PLoS ONE 2, e874 (2007).
Fakruddin, M. et al. Defective mitochondrial tRNA taurine modification activates global proteostress and leads to mitochondrial disease. Cell Rep. 22, 482–496 (2018).
Song, J., Herrmann, J. M. & Becker, T. Quality control of the mitochondrial proteome. Nat. Rev. Mol. Cell Biol. 22, 54–70 (2021).
Fessler, E. et al. A pathway coordinated by DELE1 relays mitochondrial stress to the cytosol. Nature 579, 433–437 (2020).
Guo, X. et al. Mitochondrial stress is relayed to the cytosol by an OMA1–DELE1–HRI pathway. Nature 579, 427–432 (2020).
Khan, N. A. et al. mTORC1 regulates mitochondrial integrated stress response and mitochondrial myopathy progression. Cell Metab. 26, 419–428.e5 (2017).
Morita, M. et al. mTORC1 controls mitochondrial activity and biogenesis through 4E-BP-dependent translational regulation. Cell Metab. 18, 698–711 (2013).
Donnelly, N., Gorman, A. M., Gupta, S. & Samali, A. The eIF2α kinases: their structures and functions. Cell Mol. Life Sci. 70, 3493–3511 (2013).
Martinus, R. D. et al. Selective induction of mitochondrial chaperones in response to loss of the mitochondrial genome. Eur. J. Biochem. 240, 98–103 (1996).
Rainbolt, T. K., Atanassova, N., Genereux, J. C. & Wiseman, R. L. Stress-regulated translational attenuation adapts mitochondrial protein import through Tim17A degradation. Cell Metab. 18, 908–919 (2013).
Richter, U., Lahtinen, T., Marttinen, P., Suomi, F. & Battersby, B. J. Quality control of mitochondrial protein synthesis is required for membrane integrity and cell fitness. J. Cell Biol. 211, 373–389 (2015).
Cunningham, J. T. et al. mTOR controls mitochondrial oxidative function through a YY1–PGC-1α transcriptional complex. Nature 450, 736–740 (2007).
Torrence, M. E. et al. The mTORC1-mediated activation of ATF4 promotes protein and glutathione synthesis downstream of growth signals. Preprint bioRxiv https://doi.org/10.1101/2020.10.03.324186 (2020).
Ben-Sahra, I., Hoxhaj, G., Ricoult, S. J. H., Asara, J. M. & Manning, B. D. mTORC1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science 351, 728–733 (2016).
Park, Y., Reyna-Neyra, A., Philippe, L. & Thoreen, C. C. mTORC1 balances cellular amino acid supply with demand for protein synthesis through post-transcriptional control of ATF4. Cell Rep. 19, 1083–1090 (2017).
Molenaars, M. et al. A conserved mito-cytosolic translational balance links two longevity pathways. Cell Metab. 31, 549–563.e7 (2020).
Fernandez-Marcos, P. J. & Auwerx, J. Regulation of PGC-1α, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884S–890S (2011).
Gleyzer, N., Vercauteren, K. & Scarpulla, R. C. Control of mitochondrial transcription specificity factors (TFB1M and TFB2M) by nuclear respiratory factors (NRF-1 and NRF-2) and PGC-1 family coactivators. Mol. Cell Biol. 25, 1354–1366 (2005).
Virbasius, J. V. & Scarpulla, R. C. Activation of the human mitochondrial transcription factor A gene by nuclear respiratory factors: a potential regulatory link between nuclear and mitochondrial gene expression in organelle biogenesis. Proc. Natl Acad. Sci. USA 91, 1309–1313 (1994).
Cam, H. et al. A common set of gene regulatory networks links metabolism and growth inhibition. Mol. Cell 16, 399–411 (2004).
Scarpulla, R. C. Transcriptional paradigms in mammalian mitochondrial biogenesis and function. Physiol. Rev. 88, 611–638 (2008).
Lehtonen, J. M. et al. FGF21 is a biomarker for mitochondrial translation and mtDNA maintenance disorders. Neurology 87, 2290–2299 (2016).
Suomalainen, A. & Battersby, B. J. Mitochondrial diseases: the contribution of organelle stress responses to pathology. Nat. Rev. Mol. Cell Biol. 19, 77–92 (2018).
Hill, S. & Van Remmen, H. Mitochondrial stress signaling in longevity: a new role for mitochondrial function in aging. Redox Biol. 2, 936–944 (2014).
Ferrari, A., Del’Olio, S. & Barrientos, A. The diseased mitoribosome. FEBS Lett. https://doi.org/10.1002/1873-3468.14024 (2020).
Levinger, L., Morl, M. & Florentz, C. Mitochondrial tRNA 3′ end metabolism and human disease. Nucleic Acids Res. 32, 5430–5441 (2004).
Di Nottia, M. et al. A homozygous MRPL24 mutation causes a complex movement disorder and affects the mitoribosome assembly. Neurobiol. Dis. 141, 104880 (2020).
Serre, V. et al. Mutations in mitochondrial ribosomal protein MRPL12 leads to growth retardation, neurological deterioration and mitochondrial translation deficiency. Biochim. Biophys. Acta 1832, 1304–1312 (2013).
Emdadul Haque, M., Grasso, D., Miller, C., Spremulli, L. L. & Saada, A. The effect of mutated mitochondrial ribosomal proteins S16 and S22 on the assembly of the small and large ribosomal subunits in human mitochondria. Mitochondrion 8, 254–261 (2008).
Lake, N. J. et al. Biallelic mutations in MRPS34 lead to instability of the small mitoribosomal subunit and Leigh syndrome. Am. J. Hum. Genet. 101, 239–254 (2017).
Richman, T. R. et al. Mutation in MRPS34 compromises protein synthesis and causes mitochondrial dysfunction. PLoS Genet. 11, e1005089 (2015).
Menezes, M. J. et al. Mutation in mitochondrial ribosomal protein S7 (MRPS7) causes congenital sensorineural deafness, progressive hepatic and renal failure and lactic acidemia. Hum. Mol. Genet. 24, 2297–2307 (2015).
Jackson, C. B. et al. A variant in MRPS14 (uS14m) causes perinatal hypertrophic cardiomyopathy with neonatal lactic acidosis, growth retardation, dysmorphic features and neurological involvement. Hum. Mol. Genet. 28, 639–649 (2019).
Huang, G., Li, H. & Zhang, H. Abnormal expression of mitochondrial ribosomal proteins and their encoding genes with cell apoptosis and diseases. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21228879 (2020).
Greber, B. J. et al. The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature 515, 283–286 (2014).
Brown, A. et al. Structure of the large ribosomal subunit from human mitochondria. Science 346, 718–722 (2014).
Rorbach, J. et al. Human mitochondrial ribosomes can switch their structural RNA composition. Proc. Natl Acad. Sci. USA 113, 12198–12201 (2016).
Suzuki, T. et al. Proteomic analysis of the mammalian mitochondrial ribosome. Identification of protein components in the 28S small subunit. J. Biol. Chem. 276, 33181–33195 (2001).
Sharma, M. R. et al. Structure of the mammalian mitochondrial ribosome reveals an expanded functional role for its component proteins. Cell 115, 97–108 (2003).
Amunts, A., Brown, A., Toots, J., Scheres, S. H. W. & Ramakrishnan, V. Ribosome. The structure of the human mitochondrial ribosome. Science 348, 95–98 (2015). Together with Greber et al. (2015), this paper is the first near-atomic description of the architecture of the mammalian mitoribosome.
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Desai, N., Brown, A., Amunts, A. & Ramakrishnan, V. The structure of the yeast mitochondrial ribosome. Science 355, 528–531 (2017).
Waltz, F., Soufari, H., Bochler, A., Giege, P. & Hashem, Y. Cryo-EM structure of the RNA-rich plant mitochondrial ribosome. Nat. Plants 6, 377–383 (2020).
Tobiasson, V. & Amunts, A. Ciliate mitoribosome illuminates evolutionary steps of mitochondrial translation. eLife https://doi.org/10.7554/eLife.59264 (2020).
Sloan, D. B. et al. Cytonuclear integration and co-evolution. Nat. Rev. Genet. 19, 635–648 (2018).
Petrov, A. S. et al. Structural patching fosters divergence of mitochondrial ribosomes. Mol. Biol. Evol. 36, 207–219 (2019).
Rodel, G. Two yeast nuclear genes, CBS1 and CBS2, are required for translation of mitochondrial transcripts bearing the 5′-untranslated COB leader. Curr. Genet. 11, 41–45 (1986).
Islas-Osuna, M. A., Ellis, T. P., Marnell, L. L., Mittelmeier, T. M. & Dieckmann, C. L. Cbp1 is required for translation of the mitochondrial cytochrome b mRNA of Saccharomyces cerevisiae. J. Biol. Chem. 277, 37987–37990 (2002).
Mittelmeier, T. M. & Dieckmann, C. L. In vivo analysis of sequences required for translation of cytochrome b transcripts in yeast mitochondria. Mol. Cell Biol. 15, 780–789 (1995).
Gruschke, S. et al. Cbp3–Cbp6 interacts with the yeast mitochondrial ribosomal tunnel exit and promotes cytochrome b synthesis and assembly. J. Cell Biol. 193, 1101–1114 (2011).
Salvatori, R. et al. Molecular wiring of a mitochondrial translational feedback loop. Mol. Cell 77, 887–900.e5 (2020).
Tucker, E. J. et al. Mutations in the UQCC1-interacting protein, UQCC2, cause human complex III deficiency associated with perturbed cytochrome b protein expression. PLoS Genet. 9, e1004034 (2013).
Manthey, G. M. & McEwen, J. E. The product of the nuclear gene PET309 is required for translation of mature mRNA and stability or production of intron-containing RNAs derived from the mitochondrial COX1 locus of Saccharomyces cerevisiae. EMBO J. 14, 4031–4043 (1995).
Tavares-Carreon, F. et al. The pentatricopeptide repeats present in Pet309 are necessary for translation but not for stability of the mitochondrial COX1 mRNA in yeast. J. Biol. Chem. 283, 1472–1479 (2008).
Roloff, G. A. & Henry, M. F. Mam33 promotes cytochrome c oxidase subunit I translation in Saccharomyces cerevisiae mitochondria. Mol. Biol. Cell 26, 2885–2894 (2015).
Decoster, E., Simon, M., Hatat, D. & Faye, G. The MSS51 gene product is required for the translation of the COX1 mRNA in yeast mitochondria. Mol. Gen. Genet. 224, 111–118 (1990).
Poutre, C. G. & Fox, T. D. PET111, a Saccharomyces cerevisiae nuclear gene required for translation of the mitochondrial mRNA encoding cytochrome c oxidase subunit II. Genetics 115, 637–647 (1987).
Costanzo, M. C. & Fox, T. D. Specific translational activation by nuclear gene products occurs in the 5′ untranslated leader of a yeast mitochondrial mRNA. Proc. Natl Acad. Sci. USA 85, 2677–2681 (1988).
De Silva, D. et al. The DEAD-box helicase Mss116 plays distinct roles in mitochondrial ribogenesis and mRNA-specific translation. Nucleic Acids Res. 45, 6628–6643 (2017).
Brown, N. G., Costanzo, M. C. & Fox, T. D. Interactions among three proteins that specifically activate translation of the mitochondrial COX3 mRNA in Saccharomyces cerevisiae. Mol. Cell Biol. 14, 1045–1053 (1994).
Green-Willms, N. S., Butler, C. A., Dunstan, H. M. & Fox, T. D. Pet111p, an inner membrane-bound translational activator that limits expression of the Saccharomyces cerevisiae mitochondrial gene COX2. J. Biol. Chem. 276, 6392–6397 (2001).
Manthey, G. M., Przybyla-Zawislak, B. D. & McEwen, J. E. The Saccharomyces cerevisiae Pet309 protein is embedded in the mitochondrial inner membrane. Eur. J. Biochem. 255, 156–161 (1998).
Naithani, S., Saracco, S. A., Butler, C. A. & Fox, T. D. Interactions among COX1, COX2, and COX3 mRNA-specific translational activator proteins on the inner surface of the mitochondrial inner membrane of Saccharomyces cerevisiae. Mol. Biol. Cell 14, 324–333 (2003).
Zamudio-Ochoa, A., Camacho-Villasana, Y., Garcia-Guerrero, A. E. & Perez-Martinez, X. The Pet309 pentatricopeptide repeat motifs mediate efficient binding to the mitochondrial COX1 transcript in yeast. RNA Biol. 11, 953–967 (2014).
Siep, M., van Oosterum, K., Neufeglise, H., van der Spek, H. & Grivell, L. A. Mss51p, a putative translational activator of cytochrome c oxidase subunit-1 (COX1) mRNA, is required for synthesis of Cox1p in Saccharomyces cerevisiae. Curr. Genet. 37, 213–220 (2000).
Barrientos, A., Korr, D. & Tzagoloff, A. Shy1p is necessary for full expression of mitochondrial COX1 in the yeast model of Leigh’s syndrome. EMBO J. 21, 43–52 (2002).
Perez-Martinez, X., Broadley, S. A. & Fox, T. D. Mss51p promotes mitochondrial Cox1p synthesis and interacts with newly synthesized Cox1p. EMBO J. 22, 5951–5961 (2003).
Barrientos, A., Zambrano, A. & Tzagoloff, A. Mss51p and Cox14p jointly regulate mitochondrial Cox1p expression in Saccharomyces cerevisiae. EMBO J. 23, 3472–3482 (2004).
Mick, D. U. et al. Shy1 couples Cox1 translational regulation to cytochrome c oxidase assembly. EMBO J. 26, 4347–4358 (2007).
Mick, D. U. et al. Coa3 and Cox14 are essential for negative feedback regulation of COX1 translation in mitochondria. J. Cell Biol. 191, 141–154 (2010).
Zambrano, A. et al. Aberrant translation of cytochrome c oxidase subunit 1 mRNA species in the absence of Mss51p in the yeast Saccharomyces cerevisiae. Mol. Biol. Cell 18, 523–535 (2007).
Pierrel, F. et al. Coa1 links the Mss51 post-translational function to Cox1 cofactor insertion in cytochrome c oxidase assembly. EMBO J. 26, 4335–4346 (2007).
Fontanesi, F., Clemente, P. & Barrientos, A. Cox25 teams up with Mss51, Ssc1, and Cox14 to regulate mitochondrial cytochrome c oxidase subunit 1 expression and assembly in Saccharomyces cerevisiae. J. Biol. Chem. 286, 555–566 (2011).
Perez-Martinez, X., Butler, C. A., Shingu-Vazquez, M. & Fox, T. D. Dual functions of Mss51 couple synthesis of Cox1 to assembly of cytochrome c oxidase in Saccharomyces cerevisiae mitochondria. Mol. Biol. Cell 20, 4371–4380 (2009).
Helfenbein, K. G., Ellis, T. P., Dieckmann, C. L. & Tzagoloff, A. ATP22, a nuclear gene required for expression of the F0 sector of mitochondrial ATPase in Saccharomyces cerevisiae. J. Biol. Chem. 278, 19751–19756 (2003).
Rak, M. et al. Regulation of mitochondrial translation of the ATP8/ATP6 mRNA by Smt1p. Mol. Biol. Cell 27, 919–929 (2016).
Rak, M. & Tzagoloff, A. F1-dependent translation of mitochondrially encoded Atp6p and Atp8p subunits of yeast ATP synthase. Proc. Natl Acad. Sci. USA 106, 18509–18514 (2009).
Acknowledgements
The authors thank A. Filipovska, T. Lenarcic, M.Saurer and M. Jaskolowski for critical reading of the manuscript and helpful comments. They also thank all referees for helpful comments. The authors apologize to everyone whose work unfortunately could not be included in this review due to space restrictions. E.K. was supported by a European Molecular Biology Organization (EMBO) long-term fellowship (1196-2014). This work was supported by a Swiss National Science Foundation grant (310030B_163478) and via the National Centre of Excellence in RNA and Disease and project funding 138262 to N.B.
Author information
Authors and Affiliations
Contributions
E.K. conceptualized and wrote the manuscript and prepared the figures. N.B. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Molecular Cell Biology thanks J. Rorbach, A. Amunts and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Pseudouridylation
-
RNA modification whereby the nucleoside uridine is converted into an isomer, in which the nucleobase uracil is attached to the ribose via a carbon–carbon instead of a nitrogen–carbon bond.
- Small nucleolar RNAs
-
Short RNA molecules that are usually found in complex with proteins and that target specific sites of ribosomal RNA (rRNA) to trigger modification of these rRNA sites by the associated protein component.
- Peptidyl transferase centre
-
An active site of the large ribosomal subunit that catalyses peptide bond formation during protein synthesis.
- Ribosomal P site
-
The peptidyl (P) site is one of three ribosomal tRNA binding sites and contains the tRNA carrying the nascent polypeptide chain.
- Formylated methionine
-
(fMet-tRNAMet). A derivative of the amino acid methionine (Met-tRNAMet) that carries an additional formyl modification at its amino group.
- Ribosomal A site
-
The aminoacyl (A) site is one of three ribosomal tRNA binding sites and contains the newly delivered, aminoacylated tRNA.
- Ribosomal decoding centre
-
A site on the small ribosomal subunit at which the correct base pairing of the mRNA codon and the aminoacylated tRNA is probed.
- Shine–Dalgarno sequence
-
A ribosomal binding site in bacterial mRNAs with the consensus sequence AGGAGG that is located upstream of the start codon and helps to align the ribosome with the start of the open reading frame.
- Kozak sequence
-
A consensus sequence of the most frequent nucleobases surrounding the mRNA start codon in eukaryotes, which aids start codon recognition by the ribosome during translation initiation.
- Pentatricopeptide repeat
-
(PPR). A conserved protein fold that usually occurs in tandem to mediate binding to RNAs or proteins.
- Ribosomal GTPase-associated centre
-
A site on the large ribosomal subunit that contains multiple protein and RNA elements involved in binding and activation of translational GTPases.
- Back-translocation
-
Backward movement of the tRNA–mRNA complex on the ribosome that may help to overcome ribosome stalling or that may reduce translation speed to facilitate co-translational protein folding.
- –1 frameshifting
-
Movement of the ribosome on the mRNA in the 5′ direction by one nucleotide.
- G-quadruplex structures
-
Guanosine-rich RNA or DNA sequences that fold into a stable secondary structure, in which four guanine nucleobases self-assemble via Hoogsteen hydrogen base pairing into a planar array.
- Poly(A)-assisted RNA degradation
-
A 3′–5′ RNA decay pathway, in which a short poly(A) tail serves as a degron to initiate exonucleolytic RNA cleavage, for example in order to remove aberrant RNA fragments generated by endoribonucleases, and to control the abundance of regulatory non-coding RNAs.
- Leigh syndrome
-
A severe neurological disorder driven by the degeneration of the central nervous system, which results, among others, in cognitive and movement disabilities.
- OXPHOS assembly factors
-
Protein factors that aid the assembly but are not part of the oxidative phosphorylation (OXPHOS) complex. Assembly factors may, for example, stabilize assembly intermediates, modify OXPHOS subunits or mediate delivery of cofactors.
- Mitochondrial membrane potential
-
The potential across the inner mitochondrial membrane due to an electrochemical proton gradient that is generated by oxidative phosphorylation.
Rights and permissions
About this article
Cite this article
Kummer, E., Ban, N. Mechanisms and regulation of protein synthesis in mitochondria. Nat Rev Mol Cell Biol 22, 307–325 (2021). https://doi.org/10.1038/s41580-021-00332-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-021-00332-2
This article is cited by
-
Focusing on mitochondria in the brain: from biology to therapeutics
Translational Neurodegeneration (2024)
-
SUCLG1 restricts POLRMT succinylation to enhance mitochondrial biogenesis and leukemia progression
The EMBO Journal (2024)
-
Peroxisome proliferator-activated receptor gamma coactivator-1 (PGC-1) family in physiological and pathophysiological process and diseases
Signal Transduction and Targeted Therapy (2024)
-
Nuclear and mitochondrial genetic variants associated with mitochondrial DNA copy number
Scientific Reports (2024)
-
Mitochondrial stress activates YAP/TAZ through RhoA oxidation to promote liver injury
Cell Death & Disease (2024)