Abstract
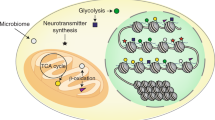
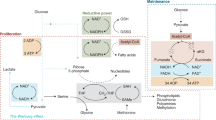
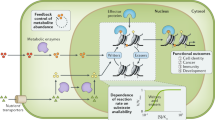
Metabolism and gene expression, which are two fundamental biological processes that are essential to all living organisms, reciprocally regulate each other to maintain homeostasis and regulate cell growth, survival and differentiation. Metabolism feeds into the regulation of gene expression via metabolic enzymes and metabolites, which can modulate chromatin directly or indirectly — through regulation of the activity of chromatin trans-acting proteins, including histone-modifying enzymes, chromatin-remodelling complexes and transcription regulators. Deregulation of these metabolic activities has been implicated in human diseases, prominently including cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Tan, M. et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146, 1016–1028 (2011). This study describes a large number of novel histone marks, including lysine crotonylation and tyrosine hydroxylation as novel histone modifications.
Huang, H., Sabari, B. R., Garcia, B. A., Allis, C. D. & Zhao, Y. SnapShot: histone modifications. Cell 159, 458–458.e1 (2014).
Tessarz, P. & Kouzarides, T. Histone core modifications regulating nucleosome structure and dynamics. Nat. Rev. Mol. Cell Biol. 15, 703–708 (2014).
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome — biological and translational implications. Nat. Rev. Cancer 11, 726–734 (2011).
Wu, H. & Zhang, Y. Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev. 25, 2436–2452 (2011).
Ruthenburg, A. J., Li, H., Patel, D. J. & Allis, C. D. Multivalent engagement of chromatin modifications by linked binding modules. Nat. Rev. Mol. Cell Biol. 8, 983–994 (2007).
Chi, P., Allis, C. D. & Wang, G. G. Covalent histone modifications — miswritten, misinterpreted and mis-erased in human cancers. Nat. Rev. Cancer 10, 457–469 (2010).
Dawson, M. A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Ronan, J. L., Wu, W. & Crabtree, G. R. From neural development to cognition: unexpected roles for chromatin. Nat. Rev. Genet. 14, 347–359 (2013).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Lu, C. et al. Histone H3K36 mutations promote sarcomagenesis through altered histone methylation landscape. Science 352, 844–849 (2016).
Cheng, X. Structural and functional coordination of DNA and histone methylation. Cold Spring Harb. Perspect. Biol. 6, a018747 (2014).
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Grillo, M. A. & Colombatto, S. S-adenosylmethionine and its products. Amino Acids 34, 187–193 (2008).
Mentch, S. J. et al. Histone methylation dynamics and gene regulation occur through the sensing of one-carbon metabolism. Cell. Metab. 22, 861–873 (2015).
Shiraki, N. et al. Methionine metabolism regulates maintenance and differentiation of human pluripotent stem cells. Cell. Metab. 19, 780–794 (2014).
Janke, R., Dodson, A. E. & Rine, J. Metabolism and epigenetics. Annu. Rev. Cell Dev. Biol. 31, 473–496 (2015).
Dolinoy, D. C., Huang, D. & Jirtle, R. L. Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development. Proc. Natl Acad. Sci. USA 104, 13056–13061 (2007).
Ye, C., Sutter, B. M., Wang, Y., Kuang, Z. & Tu, B. P. A metabolic function for phospholipid and histone methylation. Mol. Cell 66, 180–193.e8 (2017).
Metzger, E. et al. Phosphorylation of histone H3T6 by PKCbeta(I) controls demethylation at histone H3K4. Nature 464, 792–796 (2010).
Forneris, F., Binda, C., Vanoni, M. A., Mattevi, A. & Battaglioli, E. Histone demethylation catalysed by LSD1 is a flavin-dependent oxidative process. FEBS Lett. 579, 2203–2207 (2005).
Shi, Y. G. & Tsukada, Y. The discovery of histone demethylases. Cold Spring Harb. Perspect. Biol. 5, a017947 (2013).
Carey, B. W., Finley, L. W., Cross, J. R., Allis, C. D. & Thompson, C. B. Intracellular alpha-ketoglutarate maintains the pluripotency of embryonic stem cells. Nature 518, 413–416 (2015).
Hwang, I. Y. et al. Psat1-Dependent Fluctuations in alpha-Ketoglutarate Affect the Timing of ESC Differentiation. Cell. Metab. 24, 494–501 (2016).
Losman, J. A. & Kaelin, W. G. Jr. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 27, 836–852 (2013).
Salminen, A., Kauppinen, A., Hiltunen, M. & Kaarniranta, K. Krebs cycle intermediates regulate DNA and histone methylation: epigenetic impact on the aging process. Ageing Res. Rev. 16, 45–65 (2014).
Sciacovelli, M. et al. Fumarate is an epigenetic modifier that elicits epithelial-to-mesenchymal transition. Nature 537, 544–547 (2016).
Greer, E. L. & Shi, Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat. Rev. Genet. 13, 343–357 (2012).
Unoki, M. et al. Lysyl 5-hydroxylation, a novel histone modification, by Jumonji domain containing 6 (JMJD6). J. Biol. Chem. 288, 6053–6062 (2013).
Shimazu, T. et al. Suppression of oxidative stress by beta-hydroxybutyrate, an endogenous histone deacetylase inhibitor. Science 339, 211–214 (2013).
Canto, C., Menzies, K. J. & Auwerx, J. NAD(+) metabolism and the control of energy homeostasis: a balancing act between mitochondria and the nucleus. Cell Metab. 22, 31–53 (2015).
Sivanand, S., Viney, I. & Wellen, K. E. Spatiotemporal control of acetyl-CoA metabolism in chromatin regulation. Trends Biochem. Sci. 43, 61–74 (2018).
Cai, L., Sutter, B. M., Li, B. & Tu, B. P. Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol. Cell 42, 426–437 (2011). This study describes a link between lipid metabolism and histone methylation.
Chen, Y. et al. Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol. Cell. Proteom. 6, 812–819 (2007).
Dai, L. et al. Lysine 2-hydroxyisobutyrylation is a widely distributed active histone mark. Nature Chem. Biol. 10, 365–370 (2014).
Xie, Z. et al. Lysine succinylation and lysine malonylation in histones. Mol. Cell. Proteom. 11, 100–107 (2012). References 36–38 identify new types of histone modification.
Sabari, B. R., Zhang, D., Allis, C. D. & Zhao, Y. Metabolic regulation of gene expression through histone acylations. Nat. Rev. Mol. Cell Biol. 18, 90–101 (2017).
Sabari, B. R. et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol. Cell 58, 203–215 (2015).
Xie, Z. et al. Metabolic regulation of gene expression by histone lysine beta-hydroxybutyrylation. Mol. Cell 62, 194–206 (2016).
Liu, B. et al. Identification and characterization of propionylation at histone H3 lysine 23 in mammalian cells. J. Biol. Chem. 284, 32288–32295 (2009).
Goudarzi, A. et al. Dynamic competing histone H4 K5K8 acetylation and butyrylation are hallmarks of highly active gene promoters. Mol. Cell 62, 169–180 (2016).
Kebede, A. F. et al. Histone propionylation is a mark of active chromatin. Nat. Struct. Mol. Biol. 24, 1048–1056 (2017).
Cahill, G. F. Jr. Fuel metabolism in starvation. Annu. Rev. Nutr. 26, 1–22 (2006).
Laffel, L. Ketone bodies: a review of physiology, pathophysiology and application of monitoring to diabetes. Diabetes Metab. Res. Rev. 15, 412–426 (1999).
Robinson, A. M. & Williamson, D. H. Physiological roles of ketone bodies as substrates and signals in mammalian tissues. Physiol. Rev. 60, 143–187 (1980).
Yang, X. & Qian, K. Protein O-GlcNAcylation: emerging mechanisms and functions. Nat. Rev. Mol. Cell Biol. 18, 452–465 (2017).
Hardiville, S. & Hart, G. W. Nutrient regulation of gene expression by O-GlcNAcylation of chromatin. Curr. Opin. Chem. Biol. 33, 88–94 (2016).
Gambetta, M. C. & Muller, J. A critical perspective of the diverse roles of O-GlcNAc transferase in chromatin. Chromosoma 124, 429–442 (2015).
Kraus, W. L. & Lis, J. T. PARP goes transcription. Cell 113, 677–683 (2003).
Gupte, R., Liu, Z. & Kraus, W. L. PARPs and ADP-ribosylation: recent advances linking molecular functions to biological outcomes. Genes Dev. 31, 101–126 (2017).
Gibbs-Seymour, I., Fontana, P., Rack, J. G. M. & Ahel, I. HPF1/C4orf27 is a PARP-1-interacting protein that regulates PARP-1 ADP-ribosylation activity. Mol. Cell 62, 432–442 (2016).
Poirier, G. G., de Murcia, G., Jongstra-Bilen, J., Niedergang, C. & Mandel, P. Poly(ADP-ribosyl)ation of polynucleosomes causes relaxation of chromatin structure. Proc. Natl Acad. Sci. USA 79, 3423–3427 (1982).
Wright, R. H. et al. CDK2-dependent activation of PARP-1 is required for hormonal gene regulation in breast cancer cells. Genes Dev. 26, 1972–1983 (2012).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nature reviews. Mol. Cell Biol. 13, 411–424 (2012).
Bai, P. & Canto, C. The role of PARP-1 and PARP-2 enzymes in metabolic regulation and disease. Cell. Metab. 16, 290–295 (2012).
Gibson, B. A. et al. Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation. Science 353, 45–50 (2016).
Krishnakumar, R. & Kraus, W. L. PARP-1 regulates chromatin structure and transcription through a KDM5B-dependent pathway. Mol. Cell 39, 736–749 (2010).
Yang, W. & Lu, Z. Pyruvate kinase M2 at a glance. J. Cell Sci. 128, 1655–1660 (2015).
David, C. J., Chen, M., Assanah, M., Canoll, P. & Manley, J. L. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463, 364–368 (2010).
Desai, S. et al. Tissue-specific isoform switch and DNA hypomethylation of the pyruvate kinase PKM gene in human cancers. Oncotarget 5, 8202–8210 (2014).
Bluemlein, K. et al. No evidence for a shift in pyruvate kinase PKM1 to PKM2 expression during tumorigenesis. Oncotarget 2, 393–400 (2011).
Yang, W. et al. EGFR-induced and PKCε monoubiquitylation-dependent NF-κB activation upregulates PKM2 expression and promotes tumorigenesis. Mol. Cell 48, 771–784 (2012).
Kuan, C. T., Wikstrand, C. J. & Bigner, D. D. EGF mutant receptor vIII as a molecular target in cancer therapy. Endocr. Relat. Cancer 8, 83–96 (2001).
Yang, W. et al. ERK1/2-dependent phosphorylation and nuclear translocation of PKM2 promotes the Warburg effect. Nat. Cell Biol. 14, 1295–1304 (2012). This study describes an instrumental mechanism of the Warburg effect regulated by nuclear PKM2.
Lu, Z. & Hunter, T. Prolyl isomerase Pin1 in cancer. Cell Res. 24, 1033–1049 (2014).
Spoden, G. A. et al. The SUMO-E3 ligase PIAS3 targets pyruvate kinase M2. J. Cell. Biochem. 107, 293–302 (2009).
Lv, L. et al. Mitogenic and oncogenic stimulation of K433 acetylation promotes PKM2 protein kinase activity and nuclear localization. Mol. Cell 52, 340–352 (2013).
Wang, H. J. et al. JMJD5 regulates PKM2 nuclear translocation and reprograms HIF-1alpha-mediated glucose metabolism. Proc. Natl Acad. Sci. USA 111, 279–284 (2014).
Yang, W. et al. PKM2 phosphorylates histone H3 and promotes gene transcription and tumorigenesis. Cell 150, 685–696 (2012). This study discovers that a metabolic enzyme can directly phosphorylate histone proteins.
Yang, W. et al. Nuclear PKM2 regulates beta-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011).
Hosios, A. M., Fiske, B. P., Gui, D. Y. & Vander Heiden, M. G. Lack of evidence for PKM2 protein kinase activity. Mol. Cell 59, 850–857 (2015).
Li, S. et al. Serine and SAM responsive complex SESAME regulates histone modification crosstalk by sensing cellular metabolism. Mol. Cell 60, 408–421 (2015).
Yu, Q. et al. Regulation of SESAME-mediated H3T11 phosphorylation by glycolytic enzymes and metabolites. PLoS ONE 12, e0175576 (2017).
Gao, X., Wang, H., Yang, J. J., Liu, X. & Liu, Z. R. Pyruvate kinase M2 regulates gene transcription by acting as a protein kinase. Mol. Cell 45, 598–609 (2012). This study discovers that PKM2 phosphorylates the STAT3 transcription factor.
Luo, W. et al. Pyruvate kinase M2 is a PHD3-stimulated coactivator for hypoxia-inducible factor 1. Cell 145, 732–744 (2011).
Kato, H., Fukuda, T., Parkison, C., McPhie, P. & Cheng, S. Y. Cytosolic thyroid hormone-binding protein is a monomer of pyruvate kinase. Proc. Natl Acad. Sci. USA 86, 7861–7865 (1989).
Morfouace, M. et al. Control of glioma cell death and differentiation by PKM2-Oct4 interaction. Cell Death Dis. 5, e1036 (2014).
Matsuda, S. et al. Nuclear pyruvate kinase M2 complex serves as a transcriptional coactivator of arylhydrocarbon receptor. Nucleic Acids Res. 44, 636–647 (2016).
Dasgupta, S. et al. Metabolic enzyme PFKFB4 activates transcriptional coactivator SRC-3 to drive breast cancer. Nature 556, 249–254 (2018).
Weinert, B. T. et al. Lysine succinylation is a frequently occurring modification in prokaryotes and eukaryotes and extensively overlaps with acetylation. Cell Rep. 4, 842–851 (2013).
Zhang, Z. et al. Identification of lysine succinylation as a new post-translational modification. Nat. Chem. Biol. 7, 58–63 (2011).
Wang, Y. et al. KAT2A coupled with the alpha-KGDH complex acts as a histone H3 succinyltransferase. Nature 552, 273–277 (2017). This study discovers that KAT2A, which was already known as a HAT, is also a histone H3 succinyltransferase.
Jiang, Y. et al. Local generation of fumarate promotes DNA repair through inhibition of histone H3 demethylation. Nat. Cell Biol. 17, 1158–1168 (2015).
Yogev, O. et al. Fumarase: a mitochondrial metabolic enzyme and a cytosolic/nuclear component of the DNA damage response. PLoS Biol. 8, e1000328 (2010).
Ooi, A. et al. An antioxidant response phenotype shared between hereditary and sporadic type 2 papillary renal cell carcinoma. Cancer Cell 20, 511–523 (2011).
Adam, J. et al. Renal cyst formation in Fh1-deficient mice is independent of the Hif/Phd pathway: roles for fumarate in KEAP1 succination and Nrf2 signaling. Cancer Cell 20, 524–537 (2011).
Wang, T. et al. O-GlcNAcylation of fumarase maintains tumour growth under glucose deficiency. Nat. Cell Biol. 19, 833–843 (2017).
Li, B. et al. Fructose-1,6-bisphosphatase opposes renal carcinoma progression. Nature 513, 251–255 (2014).
Zheng, L., Roeder, R. G. & Luo, Y. S phase activation of the histone H2B promoter by OCA-S, a coactivator complex that contains GAPDH as a key component. Cell 114, 255–266 (2003).
Hedstrom, L. IMP dehydrogenase: structure, mechanism, and inhibition. Chem. Rev. 109, 2903–2928 (2009).
Kozhevnikova, E. N. et al. Metabolic enzyme IMPDH is also a transcription factor regulated by cellular state. Mol. Cell 47, 133–139 (2012).
Frappier, L. & Verrijzer, C. P. Gene expression control by protein deubiquitinases. Curr. Opin. Genet. Dev. 21, 207–213 (2011).
van der Knaap, J. A. et al. GMP synthetase stimulates histone H2B deubiquitylation by the epigenetic silencer USP7. Mol. Cell 17, 695–707 (2005).
van der Knaap, J. A., Kozhevnikova, E., Langenberg, K., Moshkin, Y. M. & Verrijzer, C. P. Biosynthetic enzyme GMP synthetase cooperates with ubiquitin-specific protease 7 in transcriptional regulation of ecdysteroid target genes. Mol. Cell. Biol. 30, 736–744 (2010).
Katoh, Y. et al. Methionine adenosyltransferase II serves as a transcriptional corepressor of Maf oncoprotein. Mol. Cell 41, 554–566 (2011).
Li, X. et al. Nucleus-translocated ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Mol. Cell 66, 684–697.e9 (2017).
Mews, P. et al. Acetyl-CoA synthetase regulates histone acetylation and hippocampal memory. Nature 546, 381–386 (2017). References 98 and 99 report that nuclear ACSS2 regulates histone acetylation in specific sets of gene promoter regions under physiological and pathological conditions.
Sivanand, S. et al. Nuclear acetyl-CoA production by ACLY promotes homologous recombination. Mol. Cell 67, 252–265.e6 (2017).
Wellen, K. E. et al. ATP-citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
Sutendra, G. et al. A nuclear pyruvate dehydrogenase complex is important for the generation of acetyl-CoA and histone acetylation. Cell 158, 84–97 (2014).
Takahashi, H., McCaffery, J. M., Irizarry, R. A. & Boeke, J. D. Nucleocytosolic acetyl-coenzyme a synthetase is required for histone acetylation and global transcription. Mol. Cell 23, 207–217 (2006). References 100–103 reveal important roles of nuclear acetyl-CoA synthesizing enzymes in histone acetylation.
Bulusu, V. et al. Acetate recapturing by nuclear acetyl-CoA synthetase 2 prevents loss of histone acetylation during oxygen and serum limitation. Cell Rep. 18, 647–658 (2017).
Li, X., Qian, X. & Lu, Z. Local histone acetylation by ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Autophagy 13, 1790–1791 (2017).
Peleg, S. et al. Altered histone acetylation is associated with age-dependent memory impairment in mice. Science 328, 753–756 (2010).
Bennett, D. A. et al. Epigenomics of Alzheimer’s disease. Transl Res. 165, 200–220 (2015).
Fischer, A. Targeting histone-modifications in Alzheimer’s disease. What is the evidence that this is a promising therapeutic avenue? Neuropharmacology 80, 95–102 (2014).
Lu, X. et al. Histone acetyltransferase p300 mediates histone acetylation of PS1 and BACE1 in a cellular model of Alzheimer’s disease. PLoS ONE 9, e103067 (2014).
Egervari, G., Ciccocioppo, R., Jentsch, J. D. & Hurd, Y. L. Shaping vulnerability to addiction - the contribution of behavior, neural circuits and molecular mechanisms. Neurosci. Biobehav. Rev. 85, 117–125 (2018).
Egervari, G. et al. A functional 3’UTR polymorphism (rs2235749) of prodynorphin alters microRNA-365 binding in ventral striatonigral neurons to influence novelty seeking and positive reward traits. Neuropsychopharmacology 41, 2512–2520 (2016).
Egervari, G. et al. Striatal H3K27 acetylation linked to glutamatergic gene dysregulation in human heroin abusers holds promise as therapeutic target. Biol Psychiatry 81, 585–594 (2017).
Koo, J. W. et al. Epigenetic basis of opiate suppression of Bdnf gene expression in the ventral tegmental area. Nat. Neurosci. 18, 415–422 (2015).
Robison, A. J. & Nestler, E. J. Transcriptional and epigenetic mechanisms of addiction. Nat. Rev. Neurosci. 12, 623–637 (2011).
Xu, M. et al. An acetate switch regulates stress erythropoiesis. Nat. Med. 20, 1018–1026 (2014).
Chen, R. et al. The acetate/ACSS2 switch regulates HIF-2 stress signaling in the tumor cell microenvironment. PLoS ONE 10, e0116515 (2015).
Nagaraj, R. et al. Nuclear localization of mitochondrial TCA cycle enzymes as a critical step in mammalian zygotic genome activation. Cell 168, 210–223.e1 (2017).
Shi, W. Y., Yang, X., Huang, B., Shen, W. H. & Liu, L. NOK mediates glycolysis and nuclear PDC associated histone acetylation. Front. Biosci. 22, 1792–1804 (2017).
Lu, Z. & Hunter, T. Metabolic kinases moonlighting as protein kinases. Trends Biochem. Sci. 43, 301–310 (2018).
Acknowledgements
This work was supported by National Institute of Neurological Disorders and Stroke grant R01 NS089754 (to Z.L.), NCI grants 2R01 CA109035 and R01CA204996 (to Z.L.) and the US National Institutes of Health (NIH) National Cancer Institute (NCI) under award number P30CA016672, 2P50 CA127001 (Brain Cancer SPORE). Z.L. is a Ruby E. Rutherford Distinguished Professor.
Reviewer information
Nature Reviews Molecular Cell Biology thanks Y. Zhao and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
X.L., G.E., S.L.B. and Z.L. wrote the article. All authors researched data for the article, contributed to discussion of the content, wrote the article and reviewed and/or edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Cofactors
-
Non-protein chemical compounds or metallic ions that are required for the activity of an enzyme.
- β-Oxidation
-
The catabolic process by which fatty acid molecules are broken down in the cytosol in prokaryotes and in the mitochondria in eukaryotes to generate acetyl-CoA.
- Ketone bodies
-
Any of three related compounds (acetone, acetoacetic acid and β-hydroxybutyric acid) produced during the metabolism of fats.
- Streptozotocin
-
A naturally occurring alkylating anti-neoplastic agent that is particularly toxic to the insulin-producing β-cells of the pancreas in mammals.
- Homeotic genes
-
A group of genes that regulates the development of anatomical structures in various organisms.
- Linker histone
-
Member of a family of histones that bind to the nucleosomal core particle around the DNA entry and exit sites and serve as key components of chromatin. Also known as H1 histone.
- β-Catenin
-
A protein that regulates cell–cell adhesion and gene transcription by translocating to the nucleus and associating with T cell factor (TCF) and lymphoid enhancer factor (LEF) transcription factors; it is encoded by the CTNNB1 gene.
- Warburg effect
-
The elevated glucose uptake and lactate production observed in many cancer cell lines regardless of oxygen availability.
- Aryl hydrocarbon receptor
-
(AhR). A ligand-activated transcription factor involved in the regulation of biological responses to planar aromatic (aryl) hydrocarbons.
- Pentose phosphate pathway
-
A glycolysis-parallel metabolic pathway that generates NADPH, pentoses and ribose 5-phosphate for nucleotide synthesis.
- DNA-dependent protein kinase
-
(DNA-PK). A nuclear serine/threonine-protein kinase that is activated upon DNA damage.
- Histone H2A.Z
-
An evolutionarily conserved histone variant involved in transcription regulation and genome stability.
- Nonhomologous end joining
-
(NHEJ). A pathway that repairs double-strand DNA breaks by direct ligation without the need for a homologous template.
- Hereditary leiomyomatosis
-
An autosomal dominant condition in which susceptible individuals are at risk of the development of cutaneous leiomyomas, early onset multiple uterine leiomyomas and an aggressive form of type 2 papillary renal cell cancer.
- Gluconeogenesis
-
A metabolic pathway that results in the generation of glucose from non-carbohydrate carbon substrates such as lactate, glycerol and glucogenic amino acids.
- von Hippel–Lindau
-
An inherited disorder characterized by the formation of tumours and fluid-filled sacs (cysts) in many different parts of the body.
- Retinitis pigmentosa
-
A genetic disorder of the eyes that causes loss of vision.
- Ecdysone
-
A steroid hormone secreted by the prothoracic gland that, in its active form, stimulates metamorphosis and regulates moulting in insects.
- NuRD complex
-
The nucleosome remodelling and deacetylase complex.
- SWI/SNF complex
-
An evolutionarily conserved multisubunit chromatin-remodelling complex that uses the energy of ATP hydrolysis to mobilize nucleosomes and remodel chromatin.
- Haem
-
The component of haemoglobin (and other haemoproteins) responsible for binding oxygen.
- Haematocrit
-
The ratio of the volume of red blood cells to the total volume of blood.
- Homologous recombination
-
(HR). A type of genetic recombination in which nucleotide sequences are exchanged between two similar or identical molecules of DNA.
- Ataxia-telangiectasia mutated (ATM) kinase
-
A serine/threonine-protein kinase that is recruited and activated by DNA double-strand breaks.
- TP53-binding protein
-
(53BP1). A protein involved in DNA repair, which is encoded by the TP53BP1 gene.
- Breast cancer type 1 susceptibility protein
-
(BRCA1). A tumour suppressor protein involved in DNA repair.
- Retinoblastoma protein
-
A tumour suppressor protein that inhibits cell cycle progression and is dysfunctional in several major cancers.
Rights and permissions
About this article
Cite this article
Li, X., Egervari, G., Wang, Y. et al. Regulation of chromatin and gene expression by metabolic enzymes and metabolites. Nat Rev Mol Cell Biol 19, 563–578 (2018). https://doi.org/10.1038/s41580-018-0029-7
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41580-018-0029-7
This article is cited by
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
Fructose-1,6-bisphosphatase 1 dephosphorylates and inhibits TERT for tumor suppression
Nature Chemical Biology (2024)
-
Epigenetic modification of m6A regulator proteins in cancer
Molecular Cancer (2023)
-
Epigenetic meets metabolism: novel vulnerabilities to fight cancer
Cell Communication and Signaling (2023)
-
A distinct Acyl-CoA binding protein (ACBP6) shapes tissue plasticity during nutrient adaptation in Drosophila
Nature Communications (2023)