Abstract
Klebsiella pneumoniae is a common cause of antimicrobial-resistant opportunistic infections in hospitalized patients. The species is naturally resistant to penicillins, and members of the population often carry acquired resistance to multiple antimicrobials. However, knowledge of K. pneumoniae ecology, population structure or pathogenicity is relatively limited. Over the past decade, K. pneumoniae has emerged as a major clinical and public health threat owing to increasing prevalence of healthcare-associated infections caused by multidrug-resistant strains producing extended-spectrum β-lactamases and/or carbapenemases. A parallel phenomenon of severe community-acquired infections caused by ‘hypervirulent’ K. pneumoniae has also emerged, associated with strains expressing acquired virulence factors. These distinct clinical concerns have stimulated renewed interest in K. pneumoniae research and particularly the application of genomics. In this Review, we discuss how genomics approaches have advanced our understanding of K. pneumoniae taxonomy, ecology and evolution as well as the diversity and distribution of clinically relevant determinants of pathogenicity and antimicrobial resistance. A deeper understanding of K. pneumoniae population structure and diversity will be important for the proper design and interpretation of experimental studies, for interpreting clinical and public health surveillance data and for the design and implementation of novel control strategies against this important pathogen.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Adeolu, M., Alnajar, S., Naushad, S. & Gupta, R. S. Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. Int. J. Syst. Evol. Microbiol. 66, 5575–5599 (2016).
Pendleton, J. N., Gorman, S. P. & Gilmore, B. F. Clinical relevance of the ESKAPE pathogens. Expert. Rev. Anti Infect. Ther. 11, 297–308 (2013).
Okomo, U. et al. Aetiology of invasive bacterial infection and antimicrobial resistance in neonates in sub-Saharan Africa: a systematic review and meta-analysis in line with the STROBE-NI reporting guidelines. Lancet Infect. Dis. 19, 1219–1234 (2019).
Zaidi, A. K. M. et al. Hospital-acquired neonatal infections in developing countries. Lancet 365, 1175–1188 (2005).
World Health Organization. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics (WHO, 2017).
Cassini, A. et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: a population-level modelling analysis. Lancet Infect. Dis. 19, 56–66 (2019).
Musicha, P. et al. Trends in antimicrobial resistance in bloodstream infection isolates at a large urban hospital in Malawi (1998–2016): a surveillance study. Lancet. Infect. Dis. 17, 1042–1052 (2017).
Bagley, S. T. Habitat association of Klebsiella species. Infect. Contr. 6, 52–58 (1985).
Holt, K. E. et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc. Natl Acad. Sci. USA 112, E3574–E3581 (2015). This large-scale comparative genomics study of diverse K. pneumoniae and related members of the species complex from seven countries establishes the global genomic framework and the scale and heterogeneity of the pan-genome, and identifies genetic loci that are statistically associated with invasive disease versus asymptomatic colonization in humans.
Wyres, K. L. et al. Distinct evolutionary dynamics of horizontal gene transfer in drug resistant and virulent clones of Klebsiella pneumoniae. PLoS Genet. 15, e1008114 (2019).
Gu, D. et al. A fatal outbreak of ST11 carbapenem-resistant hypervirulent Klebsiella pneumoniae in a Chinese hospital: a molecular epidemiological study. Lancet Infect. Dis. 3099, 1–10 (2017). This study presents an initial report of MDR-ST11 harbouring a KpVP-1 virulence plasmid variant, which displays enhanced survival in a human neutrophil assay and enhanced virulence in the Galleria mellonella infection model in comparison with typical MDR-ST11.
Lam, M. M. C. et al. Convergence of virulence and MDR in a single plasmid vector in MDR Klebsiella pneumoniae ST15. J. Antimicrob. Chemother. 74, 1218–1222 (2019).
Long, S. W. et al. Whole-genome sequencing of human clinical Klebsiella pneumoniae isolates reveals misidentification and misunderstandings of Klebsiella pneumoniae, Klebsiella variicola, and Klebsiella quasipneumoniae. mSphere 2, e00290–e00317 (2017).
Gorrie, C. L. et al. Gastrointestinal carriage is a major reservoir of K. pneumoniae infection in intensive care patients. Clin. Infect. Dis. 65, 208–215 (2017).
Rodrigues, C., Passet, V., Rakotondrasoa, A. & Brisse, S. Identification of Klebsiella pneumoniae, Klebsiella quasipneumoniae, Klebsiella variicola and related phylogroups by MALDI-TOF mass spectrometry. Front. Microbiol. 9, 1–7 (2018).
Long, S. W. et al. Population genomic analysis of 1,777 extended-spectrum β-lactamase-producing Klebsiella pneumoniae isolates, Houston, Texas: unexpected abundance of clonal group 307. mBio 8, e00489–e00517 (2017).
Henson, S. P. et al. Molecular epidemiology of Klebsiella pneumoniae invasive infections over a decade at Kilifi County Hospital in Kenya. Int. J. Med. Microbiol. 307, 422–429 (2017).
Heinz, E. et al. Resistance mechanisms and population structure of highly drug resistant Klebsiella in Pakistan during the introduction of the carbapenemase NDM-1. Sci. Rep. 9, 2392 (2019).
Gorrie, C. L. et al. Antimicrobial resistant Klebsiella pneumoniae carriage and infection in specialized geriatric care wards linked to acquisition in the referring hospital. Clin. Infect. Dis. 67, 161–170 (2018).
Wyres, K. L. & Holt, K. E. Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr. Opin. Microbiol. 45, 131–139 (2018).
Marques, C. et al. Evidence of Klebsiella pneumoniae sharing between healthy companion animals and co-habiting humans. J. Clin. Microbiol. 57, e01537–e01618 (2019).
Zadoks, R. N. et al. Sources of Klebsiella and Raoultella species on dairy farms: be careful where you walk. J. Dairy Sci. 94, 1045–1051 (2011).
Conlan, S., Kong, H. H. & Segre, J. A. Species-level analysis of DNA sequence data from the NIH human microbiome project. PLoS One 7, e47075 (2012).
Martin, R. M. et al. Molecular epidemiology of colonizing and infecting isolates of Klebsiella pneumoniae. mSphere 1, e00261–e00316 (2016).
Ludden, C. et al. A One Health study of the genetic relatedness of Klebsiella pneumoniae and their mobile elements in the East of England. Clin. Infect. Dis. 70, 219–226 (2020).
Chung, D. R. et al. Fecal carriage of serotype K1 Klebsiella pneumoniae ST23 strains closely related to liver abscess isolates in Koreans living in Korea. Eur. J. Clin. Microbiol. Infect. Dis. 31, 481–486 (2012).
Lin, Y.-T. et al. Seroepidemiology of Klebsiella pneumoniae colonizing the intestinal tract of healthy Chinese and overseas Chinese adults in Asian countries. BMC Microbiol. 12, 13 (2012).
Löhr, I. H. et al. Long-term faecal carriage in infants and intra-household transmission of CTX-M-15-producing Klebsiella pneumoniae following a nosocomial outbreak. J. Antimicrob. Chemother. 68, 1043–1048 (2013).
Mo, Y. et al. Carriage duration of carbapenemase-producing Enterobacteriaceae in a hospital cohort—implications for infection control measures. medRxiv. https://doi.org/10.1101/19001479 (2019).
Podschun, R. & Ullmann, U. Klebsiella spp. as nosocomial pathogens: epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin. Microbiol. Rev. 11, 589–603 (1998).
Shimasaki, T. et al. Increased relative abundance of Klebsiella pneumoniae carbapenemase-producing Klebsiella pneumoniae within the gut microbiota is associated with risk of bloodstream infection in long-term acute care hospital patients. Clin. Infect. Dis. 68, 2053–2059 (2019).
Xu, L., Sun, X. & Ma, X. Systematic review and meta-analysis of mortality of patients infected with carbapenem-resistant Klebsiella pneumoniae. Ann. Clin. Microbiol. Antimicrob. 16, 18 (2017).
Bassetti, M., Peghin, M., Vena, A. & Giacobbe, D. R. Treatment of infections due to MDR Gram-negative bacteria. Front. Med. 6, 74 (2019).
Manohar, P., Tamhankar, A. J., Lundborg, C. S. & Nachimuthu, R. Therapeutic characterization and efficacy of bacteriophage cocktails infecting Escherichia coli, Klebsiella pneumoniae, and Enterobacter species. Front. Microbiol. 10, 574 (2019).
Meatherall, B. L., Gregson, D., Ross, T., Pitout, J. D. D. & Laupland, K. B. Incidence, risk factors, and outcomes of Klebsiella pneumoniae bacteremia. Am. J. Med. 122, 866–873 (2009).
Russo, T. A. & Marr, C. M. Hypervirulent Klebsiella pneumoniae. Clin. Microbiol. Rev. 32, e00001–e00019 (2019).
Ko, W. C. et al. Community-acquired Klebsiella pneumoniae bacteremia: global differences in clinical patterns. Emerg. Infect. Dis. 8, 160–166 (2002).
Kim, J. K., Chung, D. R., Wie, S. H., Yoo, J. H. & Park, S. W. Risk factor analysis of invasive liver abscess caused by the K1 serotype Klebsiella pneumoniae. Eur. J. Clin. Microbiol. Infect. Dis. 28, 109–111 (2009).
Brockhurst, M. A. et al. The ecology and evolution of pangenomes. Curr. Biol. 29, R1094–R1103 (2019).
Bialek-Davenet, S. et al. Genomic definition of hypervirulent and multidrug-resistant Klebsiella pneumoniae clonal groups. Emerg. Infect. Dis. 20, 1812–1820 (2014). This work establishes the cgMLST scheme for K. pneumoniae and the associated species complex, which is based on 694 core genes and is available through the BIGSdb-Kp online database.
Diancourt, L., Passet, V., Verhoef, J., Grimont, P. A. & Brisse, S. Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J. Clin. Microbiol. 43, 4178–4182 (2005).
Brisse, S. et al. Virulent clones of Klebsiella pneumoniae: identification and evolutionary scenario based on genomic and phenotypic characterization. PLoS One 4, e4982 (2009).
Breurec, S. et al. Klebsiella pneumoniae resistant to third-generation cephalosporins in five African and two Vietnamese major towns: multiclonal population structure with two major international clonal groups, CG15 and CG258. Clin. Microbiol. Infect. 19, 349–355 (2013).
McInerney, J. O., McNally, A. & O’Connell, M. J. Why prokaryotes have pangenomes. Nat. Microbiol. 2, 17040 (2017).
Wyres, K. L. et al. Extensive capsule locus variation and large-scale genomic recombination within the Klebsiella pneumoniae clonal group 258. Genome Biol. Evol. 7, 1267–1279 (2015).
Lam, M. M. C. et al. Population genomics of hypervirulent Klebsiella pneumoniae clonal group 23 reveals early emergence and rapid global dissemination. Nat. Commun. 9, 2703 (2018).
Bowers, J. R. et al. Genomic analysis of the emergence and rapid global dissemination of the clonal group 258 Klebsiella pneumoniae pandemic. PLoS One 10, e0133727 (2015).
Navon-Venezia, S., Kondratyeva, K. & Carattoli, A. Klebsiella pneumoniae: a major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol. Rev. 41, 252–275 (2017).
Conlan, S. et al. Plasmid dynamics in KPC-positive Klebsiella pneumoniae during long-term patient colonization. mBio 7, e00742–e00816 (2016).
Shen, J., Lv, L., Wang, X., Xiu, Z. & Chen, G. Comparative analysis of CRISPR–Cas systems in Klebsiella genomes. J. Basic. Microbiol. 57, 325–336 (2017).
Ellington, M. J. et al. Contrasting patterns of longitudinal population dynamics and antimicrobial resistance mechanisms in two priority bacterial pathogens over 7 years in a single center. Genome Biol. 20, 184 (2019).
Wyres, K. L. & Holt, K. E. Klebsiella pneumoniae population genomics and antimicrobial-resistant clones. Trends Microbiol. 24, 944–956 (2016).
Wyres, K. L. et al. Emergence and rapid global dissemination of CTX-M-15-associated Klebsiella pneumoniae strain ST307. J. Antimicrob. Chemother. 74, 577–581 (2019).
David, S. et al. Epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread. Nat. Microbiol. 4, 1919–1929 (2019). This study is a genomic analysis of >1,700 CRKp and carbapenem-susceptible K. pneumoniae infections isolated from patients in 244 hospitals in 32 countries across Europe, providing the first large-scale systematic sample for comparison of clone and resistance gene distributions.
Turton, J. F. et al. Virulence genes in isolates of Klebsiella pneumoniae from the UK during 2016, including among carbapenemase gene-positive hypervirulent K1-ST23 and ‘non-hypervirulent’ types ST147, ST15 and ST383. J. Med. Microbiol. 67, 118–128 (2017).
Siu, L. K. et al. Molecular typing and virulence analysis of serotype K1 Klebsiella pneumoniae strains isolated from liver abscess patients and stool samples from noninfectious subjects in Hong Kong, Singapore, and Taiwan. J. Clin. Microbiol. 49, 3761–3765 (2011).
Lin, J. C. et al. Genotypes and virulence in serotype K2 Klebsiella pneumoniae from liver abscess and non-infectious carriers in Hong Kong, Singapore and Taiwan. Gut Pathog. 12, 21 (2014).
Shi, Q. et al. Diversity of virulence level phenotype of hypervirulent Klebsiella pneumoniae from different sequence type lineage. BMC Microbiol. 18, 1–6 (2018).
Lee, I. R. et al. Differential host susceptibility and bacterial virulence factors driving Klebsiella liver abscess in an ethnically diverse population. Sci. Rep. 13, 29316 (2016).
Zhang, Y. et al. High prevalence of hypervirulent Klebsiella pneumoniae infection in China: geographic distribution, clinical characteristics, and antimicrobial resistance. Antimicrob. Agents Chemother. 60, 6115–6120 (2016).
Wyres, K. L. et al. Genomic surveillance for hypervirulence and multi-drug resistance in invasive Klebsiella pneumoniae from South and Southeast Asia. Genomic med. 12, 11 (2019).
Heinz, E., Brindle, R., Morgan-McCalla, A., Peters, K. & Thomson, N. R. Caribbean multi-centre study of Klebsiella pneumoniae: whole genome sequencing, antimicrobial resistance and virulence factors. Microb. Genomics. 5, e000266 (2019).
Musicha, P. et al. Genomic analysis of Klebsiella pneumoniae isolates from Malawi reveals acquisition of multiple ESBL determinants across diverse lineages. J. Antimicrob. Chemother. 74, 1223–1232 (2019).
Zhang, R. et al. Presence of NDM in non-E. coli Enterobacteriaceae in the poultry production environment. J. Antimicrob. Chemother. 74, 2209–2213 (2019).
Marques, C. et al. Klebsiella pneumoniae causing urinary tract infections in companion animals and humans: population structure, antimicrobial resistance and virulence genes. J. Antimicrob. Chemother. 74, 594–602 (2019).
Anzai, E. K. et al. First case report of non-human primates (Alouatta clamitans) with the hypervirulent Klebsiella pneumoniae serotype K1 strain ST23: a possible emerging wildlife pathogen. J. Med. Primatol. 46, 337–342 (2017).
Bowring, B. G., Fahy, V. A., Morris, A. & Collins, A. M. An unusual culprit: Klebsiella pneumoniae causing septicaemia outbreaks in neonatal pigs? Vet. Microbiol. 203, 267–270 (2017).
Runcharoen, C. et al. Whole genome sequencing reveals high-resolution epidemiological links between clinical and environmental Klebsiella pneumoniae. Genome Med. 9, 6 (2017).
Davis, G. S. et al. Intermingled Klebsiella pneumoniae populations between retail meats and human urinary tract infections. Clin. Infect. Dis. 61, 892–899 (2015).
Singh, A., Lekshmi, M., Prakasan, S., Nayak, B. & Kumar, S. Multiple antibiotic-resistant, extended spectrum-β-lactamase (ESBL)-producing Enterobacteria in fresh seafood. Microorganisms 5, E53 (2017).
Zekar, F. M. et al. From farms to markets: Gram-negative bacteria resistant to third-generation cephalosporins in fruits and vegetables in a region of North Africa. Front. Microbiol. 8, 1569 (2017).
Yaici, L. et al. Spread of ESBL/AmpC-producing Escherichia coli and Klebsiella pneumoniae in the community through ready-to-eat sandwiches in Algeria. Int. J. Food Microbiol. 245, 66–72 (2017).
Projahn, M. et al. Contamination of chicken meat with extended-spectrum β-lactamase-producing Klebsiella pneumoniae and Escherichia coli during scalding and defeathering of broiler carcasses. Food Microbiol. 77, 185–191 (2019).
Rodrigues, C. et al. Description of Klebsiella africanensis sp. nov., Klebsiella variicola subsp. tropicalensis subsp. nov. and Klebsiella variicola subsp. variicola subsp. nov. Res. Microbiol. 170, 165–170 (2019).
Ford, P. & Avison, M. Evolutionary mapping of the SHV β-lactamase and evidence for two separate IS26-dependent bla SHV mobilization events from the Klebsiella pneumoniae chromosome. J. Antimicrob. Chemother. 54, 69–75 (2004).
Liakopoulos, A., Mevius, D. & Ceccarelli, D. A review of SHV extended-spectrum β-lactamases: neglected yet ubiquitous. Front. Microbiol. 5, 1374 (2016).
Turner, M. et al. Plasmid-borne bla SHV genes in Klebsiella pneumoniae are associated with strong promoters. J. Antimicrob. Chemother. 64, 960–964 (2009).
Ito, R. et al. Widespread fosfomycin resistance in Gram-negative bacteria attributable to the chromosomal fosA gene. mBio 8, e00749–e00817 (2017).
Li, J. et al. The nature and epidemiology of OqxAB, a multidrug efflux pump. Antimicrob. Resist. Infect. Control. 8, 44 (2019).
Bernardini, A. et al. The intrinsic resistome of Klebsiella pneumoniae. Int. J. Antimicrob. Agents. 53, 29–33 (2019).
Jana, B. et al. The secondary resistome of multidrug-resistant Klebsiella pneumoniae. Sci. Rep. 7, 42483 (2017).
Nicolas-Chanoine, M.-H., Mayer, N., Guyot, K., Dumont, E. & Pagès, J.-M. Interplay between membrane permeability and enzymatic barrier leads to antibiotic-dependent resistance in Klebsiella pneumoniae. Front. Microbiol. 9, 1422 (2018).
Xu, Q. et al. Efflux pumps AcrAB and OqxAB contribute to nitrofurantoin resistance in an uropathogenic Klebsiella pneumoniae isolate. Int. J. Antimicrob. Agents 54, 223–227 (2019).
He, F. et al. Tigecycline susceptibility and the role of efflux pumps in tigecycline resistance in KPC-producing Klebsiella pneumoniae. PLoS one 10, e0119064 (2015).
Fajardo-Lubián, A., Ben Zakour, N. L., Agyekum, A., Qi, Q. & Iredell, J. R. Host adaptation and convergent evolution increases antibiotic resistance without loss of virulence in a major human pathogen. PLoS Pathog. 15, e1007218 (2019). This study shows that deletions or mutations within OmpK35/36 porins are widespread and have emerged in multiple independent lineages of K. pneumoniae, facilitating enhanced resistance to β-lactams (including carbapenems) without significantly reducing the ability to colonize the gut or to cause pneumonia (in murine models).
Wong, J. L. C. et al. OmpK36-mediated carbapenem resistance attenuates ST258 Klebsiella pneumoniae in vivo. Nat. Commun. 10, 3957 (2019). This study experimentally confirms predictions that specific di-amino acid insertions in OmpK36 constrict its central pore, restricting diffusion of nutrients and carbapenems, and demonstrates using competition assays that these mutations do reduce fitness in the context of the murine pneumonia model.
Lunha, K. et al. High-level carbapenem-resistant OXA-48-producing Klebsiella pneumoniae with a novel OmpK36 variant and low-level, carbapenem-resistant, non-porin-deficient, OXA-181-producing Escherichia coli from Thailand. Diagn. Microbiol. Infect. Dis. 85, 221–226 (2016).
Cain, A. K. et al. Morphological, genomic and transcriptomic responses of Klebsiella pneumoniae to the last-line antibiotic colistin. Sci. Rep. 8, 9868 (2018).
Chang, H.-H. et al. Origin and proliferation of multiple-drug resistance in bacterial pathogens. Microbiol. Mol. Biol. Rev. 79, 101–116 (2015).
Lehtinen, S. et al. Evolution of antibiotic resistance is linked to any genetic mechanism affecting bacterial duration of carriage. Proc. Natl Acad. Sci. USA 114, 1075–1080 (2017).
Conlan, S. et al. Single-molecule sequencing to track plasmid diversity of hospital-associated carbapenemase-producing Enterobacteriaceae. Sci. Transl Med. 6, 254ra126 (2014).
Martin, J. et al. Covert dissemination of carbapenemase-producing Klebsiella pneumoniae (KPC) in a successfully controlled outbreak: long- and short-read whole-genome sequencing demonstrate multiple genetic modes of transmission. J. Antimicrob. Chemother. 72, 3025–3034 (2017).
Sheppard, A. E. et al. Nested Russian doll-like genetic mobility drives rapid dissemination of the carbapenem resistance gene bla KPC. Antimicrob. Agents Chemother. 60, 3767–3778 (2016). This study, using a genomic comparison of carbapenem-resistant Enterobacteriaceae isolated from a single health-care institution, highlights the importance of strain, plasmid and transposon transmission for the dissemination of bla KPC carbapenemase genes.
Buckner, M. M. C. et al. Clinically relevant plasmid–host interactions indicate that transcriptional and not genomic modifications ameliorate fitness costs of Klebsiella pneumoniae carbapenemase-carrying plasmids. mBio 9, e02303–e02317 (2018). This study demonstrates that the pKpQIL plasmid of ST258 alters K. pneumoniae gene expression and shows differences in transfer efficiencies of pKpQIL and related plasmids depending on the genetic background of donor and recipient strains.
Hardiman, C. et al. Horizontal transfer of carbapenemase-encoding plasmids and comparison with hospital epidemiological data. Antimicrob. Agents Chemother. 60, 4910–4919 (2016).
Lepuschitz, S. et al. Whole genome sequencing reveals resemblance between ESBL-producing and carbapenem resistant Klebsiella pneumoniae isolates from Austrian rivers and clinical isolates from hospitals. Sci. Total. Environ. 662, 227–235 (2019).
Paczosa, M. K. & Mecsas, J. Klebsiella pneumoniae: going on the offense with a strong defense. Microbiol. Mol. Biol. Rev. 80, 629–661 (2016).
Bengoechea, J. A. & Sa Pessoa, J. Klebsiella pneumoniae infection biology: living to counteract host defences. FEMS Microbiol. Rev. 43, 123–144 (2019).
Follador, R. et al. The diversity of Klebsiella pneumoniae surface polysaccharides. Microb. Genom. 2, e000073 (2016).
Wyres, K. L. et al. Identification of Klebsiella capsule synthesis loci from whole genome data. Microb. Genom. 2, e000102 (2016).
Bachman, M. A., Lenio, S., Schmidt, L., Oyler, J. E. & Weiser, J. N. Interaction of lipocalin 2, transferrin, and siderophores determines the replicative niche of Klebsiella pneumoniae during pneumonia. mBio 3, e00224-11 (2012).
Pan, Y.-J. et al. Genetic analysis of capsular polysaccharide synthesis gene clusters in 79 capsular types of Klebsiella spp. Sci. Rep. 5, 15573 (2015).
Whitfield, C. Biosynthesis and assembly of capsular polysaccharides in Escherichia coli. Annu. Rev. Biochem. 75, 39–68 (2006).
Ørskov, I. D. A. & Fife-Asbury, M. A. New Klebsiella capsular antigen, K82, and the deletion of five of those previously assigned. Int. J. Syst. Bacteriol. 27, 386–387 (1977).
Wick, R. R., Heinz, E., Holt, K. E. & Wyres, K. L. Kaptive Web: user-friendly capsule and lipopolysaccharide serotype prediction for Klebsiella genomes. J. Clin. Microbiol. 56, e00197–e00218 (2018).
Clarke, B. R. et al. Molecular basis for the structural diversity in serogroup O2-antigen polysaccharides in Klebsiella pneumoniae. J. Biol. Chem. 293, 4666–4679 (2018).
Guachalla, L. M. et al. Discovery of monoclonal antibodies cross-reactive to novel subserotypes of K. pneumoniae O3. Sci. Rep. 7, 6635 (2017).
Mostowy, R. J. & Holt, K. E. Diversity-generating machines: genetics of bacterial sugar-coating. Trends Microbiol. 26, 1008–1021 (2018).
Khater, F. et al. In silico analysis of usher encoding genes in Klebsiella pneumoniae and characterization of their role in adhesion and colonization. PLoS One 10, e0116215 (2015).
Murphy, C. N., Mortensen, M. S., Krogfelt, K. A. & Clegg, S. Role of Klebsiella pneumoniae type 1 and type 3 fimbriae in colonizing silicone tubes implanted into the bladders of mice as a model of catheter-associated urinary tract infections. Infect. Immun. 81, 3009–3017 (2013).
Bachman, M. A. et al. Klebsiella pneumoniae yersiniabactin promotes respiratory tract infection through evasion of lipocalin 2. Infect. Immun. 79, 3309–3316 (2011).
Russo, T. A., Olson, R., Macdonald, U., Beanan, J. & Davidson, B. A. Aerobactin, but not yersiniabactin, salmochelin, or enterobactin, enables the growth/survival of hypervirulent (hypermucoviscous) Klebsiella pneumoniae ex vivo and in vivo. Infect. Immun. 83, 3325–3333 (2015).
Russo, T. A. et al. Aerobactin mediates virulence and accounts for increased siderophore production under iron-limiting conditions by hypervirulent (hypermucoviscous) Klebsiella pneumoniae. Infect. Immun. 82, 2356–2367 (2014).
Fischbach, M. A. et al. The pathogen-associated iroA gene cluster mediates bacterial evasion of lipocalin 2. Proc. Natl Acad. Sci. USA 103, 16502–16507 (2006).
Lam, M. M. C. et al. Genetic diversity, mobilisation and spread of the yersiniabactin-encoding mobile element ICE Kp in Klebsiella pneumoniae populations. Microb. Genom. https://doi.org/10.1099/mgen.0.000196 (2018).
Lam, M. C. C. et al. Tracking key virulence loci encoding aerobactin and salmochelin siderophore synthesis in Klebsiella pneumoniae. Genome Med. 10, 77 (2018).
Holden, V. I., Bachman, M. A. & Holden, V. I. Diverging roles of bacterial siderophores during infection. Metallomics 7, 986–995 (2015).
Saha, P. et al. The bacterial siderophore enterobactin confers survival advantage to Salmonella in macrophages. Gut Microbes 10, 412–423 (2019).
Achard, M. E. S. et al. An antioxidant role for catecholate siderophores in Salmonella. Biochem. J. 454, 543–549 (2013).
Holden, V. I., Breen, P., Houle, S., Dozois, C. M. & Bachman, M. A. Klebsiella pneumoniae siderophores induce inflammation, bacterial dissemination, and HIF-1α stablization during pneumonia. mBio 7, e01397–e01416 (2016).
Lin, T.-L., Lee, C.-Z., Hsieh, P.-F., Tsai, S.-F. & Wang, J.-T. Characterization of integrative and conjugative element ICEKp1-associated genomic heterogeneity in a Klebsiella pneumoniae strain isolated from a primary liver abscess. J. Bacteriol. 190, 515–526 (2008).
Nassif, X., Fournier, J., Arondel, J. & Sansonetti, P. J. Mucoid phenotype of Klebsiella pneumoniae is a plasmid-encoded virulence factor. Infect. Immun. 57, 546–552 (1989).
Yang, X., Wai-Chi Chan, E., Zhang, R. & Chen, S. A conjugative plasmid that augments virulence in Klebsiella pneumoniae. Nat. Microbiol. 4, 2039–2043 (2019).
Nougayrède, J. P. et al. Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science 313, 848–851 (2006).
Lai, Y. C. et al. Genotoxic Klebsiella pneumoniae in Taiwan. PLoS One 9, e96292 (2014).
Lu, M.-C. et al. Colibactin contributes to the hypervirulence of pks + K1 CC23 Klebsiella pneumoniae in mouse meningitis infections. Front. Cell. Infect. Microbiol. 7, 1–14 (2017).
Wacharotayankun, R. et al. Enhancement of extracapsular polysaccharide synthesis in Klebsiella pneumoniae by RmpA2, which shows homology to NtrC and FixJ. Infect. Immun. 61, 3164–3174 (1993).
Arakawa, Y. et al. Biosynthesis of Klebsiella K2 capsular polysaccharide in Escherichia coli HB101 requires the functions of rmpA and the chromosomal cps gene cluster of the virulent strain Klebsiella pneumoniae Chedid (O1:K2). Infect. Immun. 59, 2043–2050 (1991).
Walker, K. A. et al. A Klebsiella pneumoniae regulatory mutant has reduced capsule expression but retains hypermucoviscosity. mBio 10, e00089–e00119 (2019). This study teases apart the roles of rmpA and a newly identified gene immediately downstream of it (rmpC) in hypermucoidy and capsule production.
Wu, C. C., Huang, Y. J., Fung, C. P. & Peng, H. L. Regulation of the Klebsiella pneumoniae Kpc fimbriae by the site-specific recombinase KpcI. Microbiology 156, 1983–1992 (2010).
Earle, S. G. et al. Identifying lineage effects when controlling for population structure improves power in bacterial association studies. Nat. Microbiol. 1, 16041 (2016).
Martin, R. M. et al. Identification of pathogenicity-associated loci in Klebsiella pneumoniae from hospitalized patients. mSystems 3, e00015–e00018 (2018). This genome-wide association study tests for K. pneumoniae genes associated with clinical isolates (bacteraemia or pneumonia) versus asymptomatic gut-colonizing isolates and demonstrates independent effects of the tellurite resistance operon ter and a psicose utilization locus.
Russo, T. A. et al. Identification of biomarkers for differentiation of hypervirulent Klebsiella pneumoniae from classical K. pneumoniae. J. Clin. Microbiol. 56, e00776-18 (2018). This study is the first systematic exploration of virulence markers and their relative effect on prediction of the hypervirulence phenotype, highlighting the importance of the aerobactin synthesis locus iuc and other plasmid co-located loci.
Liu, C. & Guo, J. Hypervirulent Klebsiella pneumoniae (hypermucoviscous and aerobactin positive) infection over 6 years in the elderly in China: antimicrobial resistance patterns, molecular epidemiology and risk factor. Ann. Clin. Microbiol. Antimicrob. 18, 4 (2019).
Ye, M. et al. Clinical and genomic analysis of liver abscess-causing Klebsiella pneumoniae identifies new liver abscess-associated virulence genes. Front. Cell Infect. Microbiol. 6, 1–12 (2016).
Wu, K. M. et al. Genome sequencing and comparative analysis of Klebsiella pneumoniae NTUH-K2044, a strain causing liver abscess and meningitis. J. Bacteriol. 191, 4492–4501 (2009).
Chen, Y., Chang, H., Lai, Y., Pan, C. & Tsai, S. Sequencing and analysis of the large virulence plasmid pLVPK of Klebsiella pneumoniae CG43. Gene 337, 189–198 (2004).
Lery, L. M. et al. Comparative analysis of Klebsiella pneumoniae genomes identifies a phospholipase D family protein as a novel virulence factor. BMC Biol. 12, 41 (2014).
Tu, Y. C. et al. Genetic requirements for Klebsiella pneumoniae-induced liver abscess in an oral infection model. Infect. Immun. 77, 2657–2671 (2009).
Bulger, J., MacDonald, U., Olson, R., Beanan, J. & Russo, T. A. Metabolite transporter PEG344 is required for full virulence of hypervirulent Klebsiella pneumoniae strain hvKP1 after pulmonary but not subcutaneous challenge. Infect. Immun. 85, e00093-17 (2017).
Xie, Y. et al. Emergence of the third-generation cephalosporin-resistant hypervirulent Klebsiella pneumoniae due to the acquisition of a self-transferable bla DHA-1-carrying plasmid by an ST23 strain. Virulence 9, 838–844 (2018).
Surgers, L., Boyd, A., Girard, P. M., Arlet, G. & Decré, D. ESBL-producing strain of hypervirulent Klebsiella pneumoniae K2, France. Emerg. Infect. Dis. 22, 1687–1688 (2016).
Shen, D. et al. Emergence of a multidrug-resistant hypervirulent Klebsiella pneumoniae of ST23 with a rare bla CTX-M-24-harboring virulence plasmid. Antimicrob. Agents Chemother. 63, e02273–e02318 (2019).
Dong, N., Lin, D., Zhang, R., Chan, E. W. C. & Chen, S. Carriage of bla KPC-2 by a virulence plasmid in hypervirulent Klebsiella pneumoniae. J. Antimicrob. Chemother. 73, 3317–3321 (2018).
Dong, N. et al. Genome analysis of clinical multilocus sequence Type 11 Klebsiella pneumoniae from China. Microb. Genomics 4, https://doi.org/10.1099/mgen.0.000149 (2018).
Weingarten, R. A. et al. Genomic analysis of hospital plumbing reveals diverse reservoir of bacterial plasmids conferring carbapenem resistance. mBio 9, e02011–e02017 (2018).
Sherry, N. L. et al. Genomics for molecular epidemiology and detecting transmission of carbapenemase-producing Enterobacterales in Victoria, Australia, 2012–2016. J. Clin. Microbiol. 57, e00573-19 (2019).
Brisse, S. & Verhoef, J. Phylogenetic diversity of Klebsiella pneumoniae and Klebsiella oxytoca clinical isolates revealed by randomly amplified polymorphic DNA, gyrA and parC genes sequencing and automated ribotyping. Int. J. Syst. Evol. Microbiol. 51, 915–924 (2001).
Brisse, S., Passet, V. & Grimont, P. A. D. Description of Klebsiella quasipneumoniae sp., isolated from human infections, with two subspecies, Klebsiella quasipneumoniae subsp. quasipneumoniae subsp. nov. and Klebsiella quasipneumoniae subsp. similipneumoniae subsp. nov., and. demonstration that Klebsiella singaporensis is a junior heterotypic synonym of Klebsiella variicola. Int. J. Syst. Evol. Microbiol. 64, 3146–3152 (2014).
Rosenblueth, M., Martínez, L., Silva, J. & Martínez-Romero, E. Klebsiella variicola, a novel species with clinical and plant-associated isolates. Syst. Appl. Microbiol. 27, 27–35 (2004).
Long, S. W. et al. Whole-genome sequencing of a human clinical isolate of the novel species Klebsiella quasivariicola sp. nov. Genome Announc. 5, e01057-17 (2017).
Blin, C., Passet, V., Touchon, M., Rocha, E. P. C. & Brisse, S. Metabolic diversity of the emerging pathogenic lineages of Klebsiella pneumoniae. Env. Microbiol. 19, 1881–1898 (2017). This study is the first exploration of metabolic variation among diverse K. pneumoniae lineages and other members of the species complex, showing that metabolic capabilities — in particular, carbon substrate utilization — can vary substantially between strains.
Potter, R. F. et al. Population structure, antibiotic resistance, and uropathogenicity of Klebsiella variicola. mBio 9, e02481-18 (2018).
Mathers, A. J. et al. Klebsiella quasipneumoniae provides a window into carbapenemase gene transfer, plasmid rearrangements, and patient interactions with the hospital environment. Antimicrob. Agents Chemother. 63, e02513–e02518 (2019).
Brinkac, L. M. et al. Emergence of New Delhi metallo-β-lactamase (NDM-5) in Klebsiella quasipneumoniae from neonates in a Nigerian hospital. mSphere 4, e00685-18 (2019).
Rodríguez-Medina, N., Barrios-Camacho, H., Duran-Bedolla, J. & Garza-Ramos, U. Klebsiella variicola: an emerging pathogen in humans. Emerg. Microbes Infect. 8, 973–988 (2019).
Breurec, S. et al. Liver abscess caused by infection with community-acquired Klebsiella quasipneumoniae subsp. quasipneumoniae. Emerg. Infect. Dis. 22, 529–531 (2016).
Brisse, S. & Van Duijkeren, E. Identification and antimicrobial susceptibility of 100 Klebsiella animal clinical isolates. Vet. Mirobiol. 105, 307–312 (2005).
Martínez-Romero, E. et al. Genome misclassification of Klebsiella variicola and Klebsiella quasipneumoniae isolated from plants, animals and humans. Salud Publica. Mex. 60, 52–62 (2018).
Ramirez, M. S., Iriarte, A., Reyes-Lamothe, R., Sherratt, D. J. & Tolmasky, M. E. Small Klebsiella pneumoniae plasmids: neglected contributors to antibiotic resistance. Front. Microbiol. 10, 2182 (2019).
Carattoli, A. et al. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
Robertson, J. & Nash, J. H. E. MOB-suite: software tools for clustering, reconstruction and typing of plasmids from draft assemblies. Microb. Genom. 4, https://doi.org/10.1099/mgen.0.000206 (2018).
Orlek, A. et al. Ordering the mob: insights into replicon and MOB typing schemes from analysis of a curated dataset of publicly available plasmids. Plasmid 91, 42–52 (2017).
Arredondo-Alonso, S., Willems, R. J., van Schaik, W. & Schürch, A. C. On the (im)possibility of reconstructing plasmids from whole-genome short-read sequencing data. Microb. Genom. 3, e000128 (2017).
De Maio, N. et al. Comparison of long-read sequencing technologies in the hybrid assembly of complex bacterial genomes. Microb. Genom. https://doi.org/10.1099/mgen.0.000294 (2019).
Villa, L. et al. Diversity, virulence and antimicrobial resistance of the KPC-producing Klebsiella pneumoniae ST307 clone. Microb. Genom. https://doi.org/10.1099/mgen.0.000110 (2017).
Yeh, K.-M. et al. Revisiting the importance of virulence determinant magA and its surrounding genes in Klebsiella pneumoniae causing pyogenic liver abscesses: exact role in serotype K1 capsule formation. J. Infect. Dis. 201, 1259–1267 (2010).
Yu, W.-L., Lee, M.-F., Tang, H.-J., Chang, M.-C. & Chuang, Y.-C. Low prevalence of rmpA and high tendency of rmpA mutation correspond to low virulence of extended spectrum β-lactamase-producing Klebsiella pneumoniae isolates. Virulence 6, 162–172 (2015).
Martinez, J., Martinez, L., Rosenblueth, M., Silva, J. & Martinez-Romero, E. How are gene sequence analyses modifying bacterial taxonomy? The case of Klebsiella. Int. Microbiol. 7, 261–268 (2004).
Ejaz, H. et al. Phylogenetic analysis of Klebsiella pneumoniae from hospitalized children, Pakistan. Emerg. Infect. Dis. 23, 1872–1875 (2017).
Cerqueira, G. C. et al. Multi-institute analysis of carbapenem resistance reveals remarkable diversity, unexplained mechanisms, and limited clonal outbreaks. Proc. Natl Acad. Sci. USA 114, 1135–1140 (2017).
Moradigaravand, D., Martin, V., Peacock, S. J. & Parkhill, J. Evolution and epidemiology of multidrug-resistant Klebsiella pneumoniae in the United Kingdom. mBio 8, e01976-16 (2017).
Liu, L. et al. Carbapenem-resistant isolates of the Klebsiella pneumoniae complex in western China: the common ST11 and the surprising hospital-specific types. Clin. Infect. Dis. 67 S263–S265 (2018).
Ocampo, A. M. et al. A two-year surveillance in five Colombian tertiary care hospitals reveals high frequency of non-CG258 clones of carbapenem-resistant Klebsiella pneumoniae with distinct clinical characteristics. Antimicrob. Agents Chemother. 60, 332–342 (2015).
Andrade, L. N. et al. Virulence genes, capsular and plasmid types of multidrug-resistant CTX-M(-2, -8, -15) and KPC-2-producing Klebsiella pneumoniae isolates from four major hospitals in Brazil. Diagn. Microbiol. Infect. Dis. 91, 164–168 (2018).
Deleo, F. R. et al. Molecular dissection of the evolution of carbapenem-resistant multilocus sequence type 258 Klebsiella pneumoniae. Proc. Natl Acad. Sci. USA 111, 4988–4993 (2014).
Lowe, M. et al. Klebsiella pneumoniae ST307 with bla OXA-181, South Africa, 2014–2016. Emerg. Infect. Dis. 25, 739–747 (2019).
Chung The, H. et al. A high-resolution genomic analysis of multidrug-resistant hospital outbreaks of Klebsiella pneumoniae. EMBO Mol. Med. 7, 227–239 (2015).
Acknowledgements
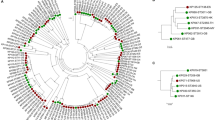
The authors are supported by Monash University and the Viertel Foundation of Australia. The authors thank Ryan Wick for assistance in creating the phylogenetic tree shown in Fig. 1.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Microbiology thanks J. Pitout, T. Russo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
BIGSdb: https://bigsdb.pasteur.fr/klebsiella/klebsiella.html
Kleborate: https://github.com/katholt/Kleborate
PathogenWatch: https://pathogen.watch/
Glossary
- Disability-adjusted life years
-
A measure of disease burden estimated as the number of years lost to ill-health, disability and/or death.
- Problem clones
-
Klebsiella pneumoniae clones that are over-represented among human infection isolates.
- Endophthalmitis
-
Inflammation of the eye, usually caused by bacterial or fungal infection.
- Necrotizing fasciitis
-
An infection resulting in rapid death of the skin and soft tissues.
- Hypervirulent
-
A term used to describe a clinical phenomenon of severe community-acquired Klebsiella pneumoniae disease, which is typically associated with the presence of a combination of multiple acquired virulence loci.
- Core genes
-
Genes that are present in all members of a given species (or nearly all, typically ≥95%).
- Accessory genes
-
Genes that are present in some members of a given species but not all (typically <95%).
- Core-genome multilocus sequence typing
-
(cgMLST). A classification scheme and nomenclature based on nucleotide sequence variation in core genes (usually hundreds of genes).
- Clones
-
A generic term used to describe subpopulations of Klebsiella pneumoniae strains with a recent common ancestor identified through allelic variation in core genes, either by multilocus sequence typing or core-genome multilocus sequence typing (specifically called ‘clonal groups’ or ‘CGs’) or by core-genome phylogenetics (specifically called ‘lineages’).
- Integrative conjugative elements
-
(ICEs). Mobile pieces of DNA that encode the machinery required for their own integration and excision from the bacterial host chromosome and transfer between bacterial cells.
- ESKAPE pathogens
-
The six most common causes of multidrug-resistant health-care-associated infection defined by the Infectious Diseases Society of America: Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter species.
- Simpson diversity
-
A measure of diversity that considers the total number of distinct entities (species, sequence types and so on) as well as their relative abundance.
- Minimum inhibitory concentration
-
(MIC). The lowest concentration of a compound (usually an antibiotic) that is able to inhibit growth of a given bacterial isolate.
- Pathogenicity factors
-
Features encoded by loci that are present in all Klebsiella pneumoniae (although there may be important allelic variants) and required for ‘classical’ opportunistic infections.
- K-locus
-
The capsule (K antigen) biosynthesis locus.
- Rhinoscleromatis
-
A rare chronic infection characterized by granulomas (structure formed from a collection of white blood cells) in the upper airways.
- O-loci
-
The outer lipopolysaccharide (O antigen) biosynthesis loci.
- Virulence factors
-
Features encoded by accessory loci which are entirely absent from most Klebsiella pneumoniae but whose presence increases either disease severity or propensity to cause disease.
- FIBK replicons
-
Plasmids of incompatibility type IncFIBK.
- Hypermucoidy
-
A phenotypic state characterized by ‘sticky’ growth and identified by production of a viscous filament (≥5 mm) when a colony is stretched by a culture loop.
- Psicose sugar
-
A monosaccharide molecule, also known as allulose.
- Classical infections
-
A term used to refer to opportunistic healthcare-associated Klebsiella pneumoniae infections.
Rights and permissions
About this article
Cite this article
Wyres, K.L., Lam, M.M.C. & Holt, K.E. Population genomics of Klebsiella pneumoniae. Nat Rev Microbiol 18, 344–359 (2020). https://doi.org/10.1038/s41579-019-0315-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-019-0315-1
This article is cited by
-
The overlap of accessory virulence factors and multidrug resistance among clinical and surveillance Klebsiella pneumoniae isolates from a neonatal intensive care unit in Nepal: a single-centre experience in a resource-limited setting
Tropical Medicine and Health (2024)
-
Carbapenem-resistant hypervirulent ST23 Klebsiella pneumoniae with a highly transmissible dual-carbapenemase plasmid in Chile
Biological Research (2024)
-
Respiratory carriage of hypervirulent Klebsiella pneumoniae by indigenous populations of Malaysia
BMC Genomics (2024)
-
Colistin monotherapy or combination for the treatment of bloodstream infection caused by Klebsiella pneumoniae: a systematic review and meta-analysis
BMC Infectious Diseases (2024)
-
Hepatitis E virus and Klebsiella pneumoniae co-infection detected by metagenomics next-generation sequencing in a patient with central nervous system and bloodstream Infection: a case report
BMC Infectious Diseases (2024)