Abstract
Metabolism was once relegated to the supply of energy and biosynthetic precursors, but it has now become clear that it is a specific mediator of nearly all physiological processes. In the context of microbial pathogenesis, metabolism has expanded outside its canonical role in bacterial replication. Among human pathogens, this expansion has emerged perhaps nowhere more visibly than for Mycobacterium tuberculosis, the causative agent of tuberculosis. Unlike most pathogens, M. tuberculosis has evolved within humans, which are both host and reservoir. This makes unrestrained replication and perpetual quiescence equally incompatible strategies for survival as a species. In this Review, we summarize recent work that illustrates the diversity of metabolic functions that not only enable M. tuberculosis to establish and maintain a state of chronic infection within the host but also facilitate its survival in the face of drug pressure and, ultimately, completion of its life cycle.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yaffe, M. B. Looking ahead to the past. Sci. Signal. 3, eg7 (2010).
Gutierrez, M. C. et al. Ancient origin and gene mosaicism of the progenitor of Mycobacterium tuberculosis. PLoS Pathog. 1, e5 (2005).
Pavelka, M. S., Chen, B., Kelley, C. L., Collins, F. M. & Jacobs, W. R. Jr. Vaccine efficacy of a lysine auxotroph of Mycobacterium tuberculosis. Infect. Immun. 71, 4190–4192 (2003).
Hondalus, M. K. et al. Attenuation of and protection induced by a leucine auxotroph of Mycobacterium tuberculosis. Infect. Immun. 68, 2888–2898 (2000).
Sambandamurthy, V. K. et al. A pantothenate auxotroph of Mycobacterium tuberculosis is highly attenuated and protects mice against tuberculosis. Nat. Med. 8, 1171–1174 (2002).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Micro 6, 613–624 (2008).
de Carvalho, L. P. S. et al. Metabolomics of Mycobacterium tuberculosis reveals compartmentalized co-catabolism of carbon substrates. Chem. Biol. 17, 1122–1131 (2010).
Puckett, S. et al. Inactivation of fructose-1,6-bisphosphate aldolase prevents optimal co-catabolism of glycolytic and gluconeogenic carbon substrates in Mycobacterium tuberculosis. PLoS Pathog. 10, e1004144 (2014).
Trujillo, C. et al. Triosephosphate isomerase is dispensable in vitro yet essential for Mycobacterium tuberculosis to establish infection. mBio 5, e00085–14 (2014).
Kayne, F. J. 11 Pyruvate Kinase. Enzymes 8, 353–382 (1973).
Zhong, W. et al. Allosteric pyruvate kinase-based “logic gate” synergistically senses energy and sugar levels in Mycobacterium tuberculosis. Nat. Commun. 8, 1986 (2017).
Noy, T. et al. Central role of pyruvate kinase in carbon co-catabolism of Mycobacterium tuberculosis. J. Biol. Chem. 291, 7060–7069 (2016).
Beste, D. J. V. et al. C-flux spectral analysis of host-pathogen metabolism reveals a mixed diet for intracellular Mycobacterium tuberculosis. Chem. Biol. 20, 1012–1021 (2013). This study shows that intracellular M. tuberculosis has access to glycolytic C3 substrates.
Somashekar, B. S. et al. Metabolic profiling of lung granuloma in Mycobacterium tuberculosis infected guinea pigs: ex vivo 1H magic angle spinning NMR studies. J. Proteome Res. 10, 4186–4195 (2011).
Zimmermann, M. et al. Integration of metabolomics and transcriptomics reveals a complex diet of Mycobacterium tuberculosis during early macrophage infection. mSystems 2, e00057–17 (2017).
Vanderven, B. C. et al. Novel inhibitors of cholesterol degradation in Mycobacterium tuberculosis reveal how the bacterium’s metabolism is constrained by the intracellular environment. PLoS Pathog. 11, e1004679 (2015). This work provides a framework to exploit carbon metabolism for TB drug development.
Johnson, R. M. et al. Chemical activation of adenylyl cyclase Rv1625c inhibits growth of Mycobacterium tuberculosis on cholesterol and modulates intramacrophage signaling. Mol. Microbiol. 105, 294–308 (2017).
Nazarova, E. V. et al. Rv3723/LucA coordinates fatty acid and cholesterol uptake in Mycobacterium tuberculosis. eLife 6, e26969 (2017).
Larrouy-Maumus, G. et al. Discovery of a glycerol 3-phosphate phosphatase reveals glycerophospholipid polar head recycling in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 110, 11320–11325 (2013).
Kalscheuer, R., Weinrick, B., Veeraraghavan, U., Besra, G. S. & Jacobs, W. R. Trehalose-recycling ABC transporter LpqY-SugA-SugB-SugC is essential for virulence of Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 107, 21761–21766 (2010).
Rustad, T. R., Sherrid, A. M., Minch, K. J. & Sherman, D. R. Hypoxia: a window into Mycobacterium tuberculosislatency. Cell. Microbiol. 11, 1151–1159 (2009).
Eoh, H. et al. Metabolic anticipation in Mycobacterium tuberculosis. Nat. Microbiol. 2, 17084 (2017).
Maksymiuk, C., Balakrishnan, A., Bryk, R., Rhee, K. Y. & Nathan, C. F. E1 of α-ketoglutarate dehydrogenase defends Mycobacterium tuberculosis against glutamate anaplerosis and nitroxidative stress. Proc. Natl Acad. Sci. USA 112, E5834–E5843 (2015).
Nandakumar, M., Nathan, C. & Rhee, K. Y. Isocitrate lyase mediates broad antibiotic tolerance in Mycobacterium tuberculosis. Nat. Commun. 5, 4306 (2014).
Venugopal, A. et al. Virulence of Mycobacterium tuberculosis depends on lipoamide dehydrogenase, a member of three multienzyme complexes. Cell Host Microbe 9, 21–31 (2011).
Bryk, R., Lima, C. D., Erdjument-Bromage, H., Tempst, P. & Nathan, C. Metabolic enzymes of mycobacteria linked to antioxidant defense by a thioredoxin-like protein. Science 295, 1073–1077 (2002).
Nathan, C. Taming tuberculosis: a challenge for science and society. Cell Host Microbe 5, 220–224 (2009).
Nathan, C. Fresh approaches to anti-infective therapies. Sci. Transl Med. 4, 140sr2 (2012).
McKinney, J. et al. Persistence of Mycobacterium tuberculosis in macrophages and mice requires the glyoxylate shunt enzyme isocitrate lyase. Nature 406, 735–738 (2000). This is a seminal study that identifies for the first time a gene required for persistence of M. tuberculosis in mice.
Marrero, J., Rhee, K. Y., Schnappinger, D., Pethe, K. & Ehrt, S. Gluconeogenic carbon flow of tricarboxylic acid cycle intermediates is critical for Mycobacterium tuberculosis to establish and maintain infection. Proc. Natl Acad. Sci. USA 107, 9819–9824 (2010).
Ehrt, S., Rhee, K. & Schnappinger, D. Mycobacterial genes essential for the pathogen’s survival in the host. Immunol. Rev. 264, 319–326 (2015).
Zhang, Y. J. et al. Tryptophan biosynthesis protects mycobacteria from CD4 T-cell-mediated killing. Cell 155, 1296–1308 (2013).
Sassetti, C. M. & Rubin, E. J. Genetic requirements for mycobacterial survival during infection. Proc. Natl Acad. Sci. USA 100, 12989–12994 (2003). This is the first study that screens a transposon mutant library to identify M. tuberculosis genes that are required for virulence in the mouse model.
Muñoz-Elías, E. & McKinney, J. Mycobacterium tuberculosis isocitrate lyases 1 and 2 are jointly required for in vivo growth and virulence. Nat. Med. 11, 638–644 (2005).
Blumenthal, A., Trujillo, C., Ehrt, S. & Schnappinger, D. Simultaneous analysis of multiple Mycobacterium tuberculosis knockdown mutants in vitro and in vivo. PLoS ONE 5, e15667 (2010).
Pandey, A. K. & Sassetti, C. M. Mycobacterial persistence requires the utilization of host cholesterol. Proc. Natl Acad. Sci. USA 105, 4376–4380 (2008). This work reveals that M. tuberculosis requires cholesterol for persistence in vivo.
Marrero, J., Trujillo, C., Rhee, K. Y. & Ehrt, S. Glucose phosphorylation is required for Mycobacterium tuberculosis persistence in mice. PLoS Pathog. 9, e1003116 (2013).
Puckett, S. et al. Glyoxylate detoxification is an essential function of malate synthase required for carbon assimilation in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 114, E2225–E2232 (2017).
Ruecker, N. et al. Fumarase deficiency causes protein and metabolite succination and intoxicates Mycobacterium tuberculosis. Cell Chem. Biol. 24, 306–315 (2017).
Rhee, K. Y. et al. Central carbon metabolism in Mycobacterium tuberculosis: an unexpected frontier. Trends Microbiol. 19, 307–314 (2011).
Dahl, J. L. et al. The role of Rel Mtb-mediated adaptation to stationary phase in long-term persistence of Mycobacterium tuberculosis in mice. Proc. Natl Acad. Sci. USA 100, 10026–10031 (2003).
Gould, T. A., van de Langemheen, H., Muñoz-Elías, E. J., Mckinney, J. D. & Sacchettini, J. C. Dual role of isocitrate lyase 1 in the glyoxylate and methylcitrate cycles in Mycobacterium tuberculosis. Mol. Microbiol. 61, 940–947 (2006).
Eoh, H. & Rhee, K. Y. Methylcitrate cycle defines the bactericidal essentiality of isocitrate lyase for survival of Mycobacterium tuberculosis on fatty acids. Proc. Natl Acad. Sci. USA 111, 4976–4981 (2014). This study dissects the specific biochemical mechanism underlying the essentiality of isocitrate lyase for growth and survival of M. tuberculosis with fatty acids as a carbon source.
Eoh, H. & Rhee, K. Y. Multifunctional essentiality of succinate metabolism in adaptation to hypoxia in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 110, 6554–6559 (2013).
Ganapathy, U. et al. Two enzymes with redundant fructose bisphosphatase activity sustain gluconeogenesis and virulence in Mycobacterium tuberculosis. Nat. Commun. 6, 7912 (2015).
Machová, I. et al. Mycobacterium tuberculosis phosphoenolpyruvate carboxykinase is regulated by redox mechanisms and interaction with thioredoxin. J. Biol. Chem. 289, 13066–13078 (2014).
Lee, W., VanderVen, B. C., Fahey, R. J. & Russell, D. G. Intracellular Mycobacterium tuberculosis exploits host-derived fatty acids to limit metabolic stress. J. Biol. Chem. 288, 6788–6800 (2013).
Thomas, S. T., VanderVen, B. C., Sherman, D. R., Russell, D. G. & Sampson, N. S. Pathway profiling in Mycobacterium tuberculosis: elucidation of cholesterol-derived catabolite and enzymes that catalyze its metabolism. J. Biol. Chem. 286, 43668–43678 (2011).
Kim, J.-H. et al. A genetic strategy to identify targets for the development of drugs that prevent bacterial persistence. Proc. Natl Acad. Sci. USA 110, 19095–19100 (2013).
Woong Park, S. et al. Evaluating the sensitivity of Mycobacterium tuberculosis to biotin deprivation using regulated gene expression. PLoS Pathog. 7, e1002264 (2011).
Dick, T., Manjunatha, U., Kappes, B. & Gengenbacher, M. Vitamin B6 biosynthesis is essential for survival and virulence of Mycobacterium tuberculosis. Mol. Microbiol. 78, 980–988 (2010).
Boshoff, H. I. M. et al. Biosynthesis and recycling of nicotinamide cofactors in Mycobacterium tuberculosis. An essential role for NAD in nonreplicating bacilli. J. Biol. Chem. 283, 19329–19341 (2008).
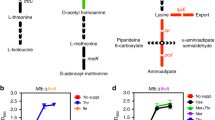
Berney, M. et al. Essential roles of methionine and S-adenosylmethionine in the autarkic lifestyle of Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 112, 10008–10013 (2015).
Bryk, R., Griffin, P. & Nathan, C. Peroxynitrite reductase activity of bacterial peroxiredoxins. Nature 407, 211–215 (2000).
Hartman, T. E., Gardete, S., Rhee, K. Y., Jansen, R. S. & Wang, Z. Metabolic perspectives on persistence. Microbiol. Spectr. 5, TBTB2-0026-2016 (2017).
Wayne, L. & Hayes, L. An in vitro model for sequential study of shiftdown of Mycobacterium tuberculosis through two stages of nonreplicating persistence. Infect. Immun. 64, 2062 (1996). This is a seminal study that establishes a model for non-replication persistence of M. tuberculosis.
Watanabe, S. et al. Fumarate reductase activity maintains an energized membrane in anaerobic Mycobacterium tuberculosis. PLoS Pathog. 7, e1002287 (2011).
Hartman, T. et al. Succinate dehydrogenase is the regulator of respiration in Mycobacterium tuberculosis. PLoS Pathog. 10, e1004510 (2014).
Dawson, R. et al. Efficiency and safety of the combination of moxifloxacin, pretomanid (PA-824), and pyrazinamide during the first 8 weeks of antituberculosis treatment: a phase 2b, open-label, partly randomised trial in patients with drug-susceptible or drug-resistant pulmonary tuberculosis. Lancet 385, 1738–1747 (2015).
Diacon, A. H. et al. The diarylquinoline TMC207 for multidrug-resistant tuberculosis. N. Engl. J. Med. 360, 2397–2405 (2009).
Gler, M. T. et al. Delamanid for multidrug-resistant pulmonary tuberculosis. N. Engl. J. Med. 366, 2151–2160 (2012).
Andries, K. A. Diarylquinoline drug active on the ATP synthase of Mycobacterium tuberculosis. Science 307, 223–227 (2005). This paper reports on bedaquiline, which in 2012 became the first new anti-TB drug with a novel mechanism of action since rifampicin was approved in 1974.
Singh, R. et al. PA-824 kills nonreplicating Mycobacterium tuberculosis by intracellular NO release. Science 322, 1392–1395 (2008).
Stamm, C. E., Collins, A. C. & Shiloh, M. U. Sensing of Mycobacterium tuberculosis and consequences to both host and bacillus. Immunol. Rev. 264, 204–219 (2015).
Ishikawa, E. et al. Direct recognition of the mycobacterial glycolipid, trehalose dimycolate, by C-type lectin Mincle. J. Exp. Med. 206, 2879–2888 (2009).
Neyrolles, O. & Guilhot, C. Recent advances in deciphering the contribution of Mycobacterium tuberculosis lipids to pathogenesis. Tuberculosis 91, 187–195 (2011). This is a comprehensive review of the cell envelope lipids of M. tuberculosis and their contribution to virulence.
Geisel, R. E., Sakamoto, K., Russell, D. G. & Rhoades, E. R. In vivo activity of released cell wall lipids of Mycobacterium bovis bacillus Calmette-Guérin is due principally to trehalose mycolates. J. Immunol. 174, 5007–5015 (2005).
Rhoades, E. et al. Identification and macrophage-activating activity of glycolipids released from intracellular Mycobacterium bovis BCG. Mol. Microbiol. 48, 875–888 (2003).
Galagan, J. E. et al. The Mycobacterium tuberculosis regulatory network and hypoxia. Nature 499, 178–183 (2013).
Glickman, M. S., Cox, J. S. & Jacobs, W. R. A novel mycolic acid cyclopropane synthetase is required for cording, persistence, and virulence of Mycobacterium tuberculosis. Mol. Cell 5, 717–727 (2000). This is a foundational study that describes the importance of cyclopropanation of mycolic acids for long-term persistence and virulence of M. tuberculosis.
Vilchèze, C. et al. Phosphorylation of KasB regulates virulence and acid-fastness in Mycobacterium tuberculosis. PLoS Pathog. 10, e1004115 (2014).
Seiler, P. et al. Cell-wall alterations as an attribute of Mycobacterium tuberculosis in latent infection. J. Infect. Dis. 188, 1326–1331 (2003).
Chancellor, A. et al. CD1b-restricted GEM T cell responses are modulated by Mycobacterium tuberculosis mycolic acid meromycolate chains. Proc. Natl Acad. Sci. USA 114, E10956–E10964 (2017).
Jain, M. et al. Lipidomics reveals control of Mycobacterium tuberculosis virulence lipids via metabolic coupling. Proc. Natl Acad. Sci. USA 104, 5133–5138 (2007).
Yang, X., Nesbitt, N. M., Dubnau, E., Smith, I. & Sampson, N. S. Cholesterol metabolism increases the metabolic pool of propionate in Mycobacterium tuberculosis. Biochemistry 48, 3819–3821 (2009).
Griffin, J. E. et al. Cholesterol catabolism by Mycobacterium tuberculosis requires transcriptional and metabolic adaptations. Chem. Biol. 19, 218–227 (2012).
Rousseau, C. et al. Production of phthiocerol dimycocerosates protects Mycobacterium tuberculosis from the cidal activity of reactive nitrogen intermediates produced by macrophages and modulates the early immune response to infection. Cell. Microbiol. 6, 277–287 (2004).
Astarie-Dequeker, C. et al. Phthiocerol dimycocerosates of M. tuberculosis participate in macrophage invasion by inducing changes in the organization of plasma membrane lipids. PLoS Pathog. 5, e1000289 (2009).
Cambier, C. J., Falkow, S. & Ramakrishnan, L. Host evasion and exploitation schemes of Mycobacterium tuberculosis. Cell 159, 1497–1509 (2014).
Quigley, J. et al. The cell wall lipid PDIM contributes to phagosomal escape and host cell exit of Mycobacterium tuberculosis. mBio 8, e00148–17 (2017).
Cox, J. S., Chen, B., McNeil, M. & Jacobs, W. R. Complex lipid determines tissue-specific replication of Mycobacterium tuberculosis in mice. Nature 402, 79–83 (1999).
Blanc, L. et al. Mycobacterium tuberculosis inhibits human innate immune responses via the production of TLR2 antagonist glycolipids. Proc. Natl Acad. Sci. USA 114, 11205–11210 (2017).
Baek, S.-H., Li, A. H. & Sassetti, C. M. Metabolic regulation of mycobacterial growth and antibiotic sensitivity. PLoS Biol. 9, e1001065 (2011).
Martinot, A. J. et al. Mycobacterial metabolic syndrome: LprG and Rv1410 regulate triacylglyceride levels, growth rate and virulence in Mycobacterium tuberculosis. PLoS Pathog 12, e1005351 (2016).
Daniel, J., Maamar, H., Deb, C., Sirakova, T. D. & Kolattukudy, P. E. Mycobacterium tuberculosis uses host triacylglycerol to accumulate lipid droplets and acquires a dormancy-like phenotype in lipid-loaded macrophages. PLoS Pathog. 7, e1002093 (2011).
Javid, B. et al. Mycobacterial mistranslation is necessary and sufficient for rifampicin phenotypic resistance. Proc. Natl Acad. Sci. USA 111, 1132–1137 (2014).
Su, H.-W. et al. The essential mycobacterial amidotransferase GatCAB is a modulator of specific translational fidelity. Nat. Microbiol. 1, 16147 (2016).
Liu, Y. et al. Immune activation of the host cell induces drug tolerance in Mycobacterium tuberculosis both in vitro and in vivo. J. Exp. Med. 213, 809–825 (2016).
Peterson, E. J. R., Ma, S., Sherman, D. R. & Baliga, N. S. Network analysis identifies Rv0324 and Rv0880 as regulators of bedaquiline tolerance in Mycobacterium tuberculosis. Nat. Microbiol. 1, 16078 (2016).
Koul, A. et al. Delayed bactericidal response of Mycobacterium tuberculosis to bedaquiline involves remodelling of bacterial metabolism. Nat. Commun. 5, 3369 (2014).
Dhillon, J., Andries, K., Phillips, P. P. J. & Mitchison, D. A. Bactericidal activity of the diarylquinoline TMC207 against Mycobacterium tuberculosis outside and within cells. Tuberculosis 90, 301–305 (2010).
Greenwood, D. in Antibiotic and Chemotherapy 9th edn (eds Finch, R., Greenwood, D., Whitley, R. & Norrby, S. R.) 2–9 (Saunders, 2010).
Bushby, S. R. & Hitchings, G. H. Trimethoprim, a sulphonamide potentiator. Br. J. Pharmacol. Chemother. 33, 72–90 (1968).
Volkamer, A., Kuhn, D., Grombacher, T., Rippmann, F. & Rarey, M. Combining global and local measures for structure-based druggability predictions. J. Chem. Inf. Model. 52, 360–372 (2012).
Wellington, S. et al. A small-molecule allosteric inhibitor of Mycobacterium tuberculosis tryptophan synthase. Nat. Chem. Biol. 13, 943–950 (2017). This study provides a thorough and elegant mechanistic analysis of novel TrpAB inhibitors.
Abrahams, K. A. et al. Inhibiting mycobacterial tryptophan synthase by targeting the inter-subunit interface. Sci. Rep. 7, 9430 (2017).
Bernstein, J., Lott, W. A., Steinberg, B. A. & Yale, H. L. Chemotherapy of experimental tuberculosis. V. Isonicotinic acid hydrazide (nydrazid) and related compounds. Am. Rev. Tuberc. 65, 357–364 (1952).
Vilchèze, C. & Jacobs, W. R. Resistance to isoniazid and ethionamide in Mycobacterium tuberculosis: genes, mutations, and causalities. Microbiol. Spectr. 2, MGM2-0014-2013 (2014).
Soutter, H. H. et al. Discovery of cofactor-specific, bactericidal Mycobacterium tuberculosis InhA inhibitors using DNA-encoded library technology. Proc. Natl Acad. Sci. USA 113, E7880–E7889 (2016).
Portevin, D. et al. A polyketide synthase catalyzes the last condensation step of mycolic acid biosynthesis in mycobacteria and related organisms. Proc. Natl Acad. Sci. USA 101, 314–319 (2004).
Aggarwal, A. et al. Development of a novel lead that targets M. tuberculosis polyketide synthase 13. Cell 170, 249–259.e25 (2017).
DeLaBarre, B., Hurov, J., Cianchetta, G., Murray, S. & Dang, L. Action at a distance: allostery and the development of drugs to target cancer cell metabolism. Chem. Biol. 21, 1143–1161 (2014).
Reaves, M. L. & Rabinowitz, J. D. Metabolomics in systems microbiology. Curr. Opin. Biotechnol. 22, 17–25 (2011).
Yus, E. et al. Impact of genome reduction on bacterial metabolism and its regulation. Science 326, 1263–1268 (2009).
Lofthouse, E. K. et al. Systems-based approaches to probing metabolic variation within the Mycobacterium tuberculosis complex. PLoS ONE 8, e75913 (2013).
Garay, C. D., Dreyfuss, J. M. & Galagan, J. E. Metabolic modeling predicts metabolite changes in Mycobacterium tuberculosis. BMC Syst. Biol. 9, 57 (2015).
World Health Organization. Global Tuberculosis Report 2016 (WHO, Geneva, 2016).
Lin, P. & Flynn, J. Understanding latent tuberculosis: a moving target. J. Immunol. 185, 15 (2010).
Ernst, J. D. The immunological life cycle of tuberculosis. Nat. Rev. Immunol. 12, 581–591 (2012).
Russell, D. G. Mycobacterium tuberculosis: here today, and here tomorrow. Nat. Rev. Mol. Cell Biol. 2, 569–577 (2001).
Barry, C. E. et al. The spectrum of latent tuberculosis: rethinking the biology and intervention strategies. Nat. Rev. Microbiol. 7, 845–855 (2009).
Philips, J. A. & Ernst, J. D. Tuberculosis Pathogenesis and Immunity. Annu. Rev. Pathol. Mech. Dis. 7, 353–384 (2012).
Stallings, C. L. & Glickman, M. S. Is Mycobacterium tuberculosis stressed out? A critical assessment of the genetic evidence. Microbes Infect. 12, 1091–1101 (2010).
Ehrt, S. & Schnappinger, D. Mycobacterial survival strategies in the phagosome: defence against host stresses. Cell. Microbiol. 11, 1170–1178 (2009).
van der Wel, N. et al. M. tuberculosis and M. leprae translocate from the phagolysosome to the cytosol in myeloid cells. Cell 129, 1287–1298 (2007).
Houben, E. N. G. et al. Composition of the type VII secretion system membrane complex. Mol. Microbiol. 86, 472–484 (2012).
Boshoff, H. & Barry, C. Tuberculosis — metabolism and respiration in the absence of growth. Nat. Rev. Microbiol. 3, 70–80 (2005).
Russell, D. G. Who puts the tubercle in tuberculosis? Nat. Rev. Microbiol. 5, 39–47 (2006).
Via, L. E. et al. Tuberculous granulomas are hypoxic in guinea pigs, rabbits, and nonhuman primates. Infect. Immun. 76, 2333–2340 (2008).
Elkington, P. T., D’Armiento, J. M. & Friedland, J. S. Tuberculosis immunopathology: the neglected role of extracellular matrix destruction. Sci. Transl Med. 3, 71ps6 (2011).
Garton, N. J. et al. Cytological and transcript analyses reveal fat and lazy persister-like bacilli in tuberculous sputum. PLoS Med. 5, e75 (2008).
Eum, S.-Y. et al. Neutrophils are the predominant infected phagocytic cells in the airways of patients with active pulmonary TB. Chest 137, 122–128 (2010).
Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G. & Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle bacilli in sputum. Am. J. Respir. Crit. Care Med. 181, 174–180 (2010).
Chengalroyen, M. D. et al. Detection and quantification of differentially culturable tubercle bacteria in sputum from patients with tuberculosis. Am. J. Respir. Crit. Care Med. 194, 1532–1540 (2016).
Acknowledgements
This work was supported by grants R01AI063446 (National Institute of Allergy and Infectious Diseases (NIAID)) and U19AI111143 (Tri-Institutional TB Research Unit, part of the NIAID Tuberculosis Research Units Network).
Reviewer information
Nature Reviews Microbiology thanks Johnjoe McFadden, Olivier Neyrolles and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, contributed substantially to discussion of the content, wrote the article and reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Caseation
-
The necrotic death of cells in the centre of a granuloma, resulting in an acellular mass that resembles a soft, crumbly cheese.
- Tricarboxylic acid cycle
-
(TCA cycle). A biochemical energy-generating pathway for the final steps of the oxidation of carbohydrates and fatty acids.
- Glyoxylate shunt
-
An anaplerotic pathway that converts isocitrate into malate and succinate, bypassing the two decarboxylation steps of the tricarboxylic acid cycle.
- Anaplerosis
-
The process of replenishing metabolite pools.
- Cataplerotic
-
Involving the process of extracting metabolites for biosynthetic reactions.
- Ketoacidosis
-
The accumulation of high concentrations of the ketones acetoacetate and β-hydroxybutyrate.
- Dienophile
-
A compound that readily reacts with an unsaturated hydrocarbon (diene).
- Cording
-
The aggregation of mycobacteria in a structure that resembles cords. This is due to the glycolipid trehalose dimycolate, also called cord factor.
- Acid-fastness
-
The physical property of the mycolic acid-containing mycobacterial cell envelope that is responsible for resistance to decolorization by acid and ethanol.
Rights and permissions
About this article
Cite this article
Ehrt, S., Schnappinger, D. & Rhee, K.Y. Metabolic principles of persistence and pathogenicity in Mycobacterium tuberculosis. Nat Rev Microbiol 16, 496–507 (2018). https://doi.org/10.1038/s41579-018-0013-4
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41579-018-0013-4
This article is cited by
-
Cryo- EM structure of the mycobacterial 70S ribosome in complex with ribosome hibernation promotion factor RafH
Nature Communications (2024)
-
Hypothetical protein CuvA (Rv1422) from Mycobacterium tuberculosis H37Rv interacts with uridine diphosphate N-acetylglucosamine as a key precursor of cell wall
Applied Biological Chemistry (2023)
-
The implication of Mycobacterium tuberculosis-mediated metabolism of targeted xenobiotics
Nature Reviews Chemistry (2023)
-
Mycobacterium tuberculosis and its clever approaches to escape the deadly macrophage
World Journal of Microbiology and Biotechnology (2023)
-
SynBioStrainFinder: A microbial strain database of manually curated CRISPR/Cas genetic manipulation system information for biomanufacturing
Microbial Cell Factories (2022)