Abstract
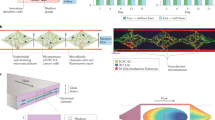
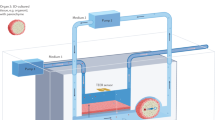
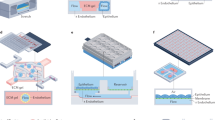
Predicting the effects of drugs before human clinical trials is at the heart of drug screening and discovery processes. The cost of drug discovery is steadily increasing owing to the limited predictability of 2D cell culture and animal models. The convergence of microfabrication and tissue engineering gave rise to organ-on-a-chip technologies, which offer an alternative to conventional preclinical models for drug screening. Organ-on-a-chip devices can replicate key aspects of human physiology crucial for the understanding of drug effects, improving preclinical safety and efficacy testing. In this Review, we discuss how organ-on-a-chip technologies can recreate functions of organs, focusing on tissue barrier properties, parenchymal tissue function and multi-organ interactions, which are three key aspects of human physiology. Specific organ-on-a-chip systems are examined in terms of cell sources, functional hallmarks and available disease models. Finally, we highlight the challenges that need to be overcome for the clinical translation of organ-on-a-chip devices regarding materials, cellular fidelity, multiplexing, sensing, scalability and validation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Shehab, N. et al. US emergency department visits for outpatient adverse drug events, 2013–2014. JAMA 316, 2115–2125 (2016).
Shahrbaf, F. G. & Assadi, F. Drug-induced renal disorders. J. Renal Injury Prevention 4, 57 (2015).
Kim, S. Y. & Moon, A. Drug-induced nephrotoxicity and its biomarkers. Biomolecules Ther. 20, 268 (2012).
Adams, C. P. & Brantner, V. V. Spending on new drug development1. Health Econom. 19, 130–141 (2010).
Kinch, M. S. & Merkel, J. An analysis of FDA-approved drugs for inflammation and autoimmune diseases. Drug Discov. Today 20, 920–923 (2015).
DiMasi, J. A., Grabowski, H. G. & Hansen, R. W. Innovation in the pharmaceutical industry: new estimates of R&D costs. J. Health Econom. 47, 20–33 (2016).
Paul, S. M. et al. How to improve R&D productivity: the pharmaceutical industry’s grand challenge. Nat. Rev. Drug Discov. 9, 203–214 (2010).
Ben-David, U. et al. Patient-derived xenografts undergo mouse-specific tumor evolution. Nat. Genet. 49, 1567–1575 (2017).
Rohr, S., Schölly, D. M. & Kleber, A. G. Patterned growth of neonatal rat heart cells in culture. Morphological and electrophysiological characterization. Circul. Res. 68, 114–130 (1991).
Fast, V. G., Rohr, S., Gillis, A. M. & Kléber, A. G. Activation of cardiac tissue by extracellular electrical shocks: formation of ‘secondary sources’ at intercellular clefts in monolayers of cultured myocytes. Circul. Res. 82, 375–385 (1998).
Kucera, J. P., Kléber, A. G. & Rohr, S. Slow conduction in cardiac tissue, II: effects of branching tissue geometry. Circul. Res. 83, 795–805 (1998).
Rohr, S., Kucera, J. P. & Kléber, A. G. Slow conduction in cardiac tissue, I: effects of a reduction of excitability versus a reduction of electrical coupling on microconduction. Circul. Res. 83, 781–794 (1998).
Rohr, S., Kucera, J. P., Fast, V. G. & Kléber, A. G. Paradoxical improvement of impulse conduction in cardiac tissue by partial cellular uncoupling. Science 275, 841–844 (1997).
Duffy, D. C., McDonald, J. C., Schueller, O. J. & Whitesides, G. M. Rapid prototyping of microfluidic systems in poly(dimethylsiloxane). Anal. Chem. 70, 4974–4984 (1998).
Xia, Y. & Whitesides, G. M. Soft lithography. Annu. Rev. Mater. Sci. 28, 153–184 (1998).
Whitesides, G. M. The origins and the future of microfluidics. Nature 442, 368 (2006).
Sin, A. et al. The design and fabrication of three-chamber microscale cell culture analog devices with integrated dissolved oxygen sensors. Biotechnol. Progress 20, 338–345 (2004).
Viravaidya, K., Sin, A. & Shuler, M. L. Development of a microscale cell culture analog to probe naphthalene toxicity. Biotechnol. Progress 20, 316–323 (2004).
Song, J. W. et al. Computer-controlled microcirculatory support system for endothelial cell culture and shearing. Anal. Chem. 77, 3993–3999 (2005).
Lam, M. T., Huang, Y.-C., Birla, R. K. & Takayama, S. Microfeature guided skeletal muscle tissue engineering for highly organized 3-dimensional free-standing constructs. Biomaterials 30, 1150–1155 (2009).
Jang, K., Sato, K., Igawa, K., Chung, U.-i. & Kitamori, T. Development of an osteoblast-based 3D continuous-perfusion microfluidic system for drug screening. Anal. Bioanal. Chem. 390, 825–832 (2008).
Huh, D. et al. Acoustically detectable cellular-level lung injury induced by fluid mechanical stresses in microfluidic airway systems. Proc. Natl Acad. Sci. USA 104, 18886–18891 (2007).
Carraro, A. et al. In vitro analysis of a hepatic device with intrinsic microvascular-based channels. Biomed. Microdevices 10, 795–805 (2008).
Lee, P. J., Hung, P. J. & Lee, L. P. An artificial liver sinusoid with a microfluidic endothelial-like barrier for primary hepatocyte culture. Biotechnol. Bioeng. 97, 1340–1346 (2007).
Harris, S. G. & Shuler, M. L. Growth of endothelial cells on microfabricated silicon nitride membranes for an in vitro model of the blood-brain barrier. Biotechnol. Bioprocess Eng. 8, 246–251 (2003).
Mahler, G. J., Esch, M. B., Glahn, R. P. & Shuler, M. L. Characterization of a gastrointestinal tract microscale cell culture analog used to predict drug toxicity. Biotechnol. Bioeng. 104, 193–205 (2009).
Kimura, H., Yamamoto, T., Sakai, H., Sakai, Y. & Fujii, T. An integrated microfluidic system for long-term perfusion culture and on-line monitoring of intestinal tissue models. Lab. Chip 8, 741–746 (2008).
Jang, K.-J. & Suh, K.-Y. A multi-layer microfluidic device for efficient culture and analysis of renal tubular cells. Lab. Chip 10, 36–42 (2010).
Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science 328, 1662–1668 (2010). This is a landmark study on recapitulating organ-level lung function in vitro with a microfluidic chip.
Ingber, D. E. Reverse engineering human pathophysiology with organs-on-chips. Cell 164, 1105–1109 (2016).
Langer, R. & Vacanti, J. P. Tissue engineering. Science 260, 920–926 (1993).
El-Ali, J., Sorger, P. K. & Jensen, K. F. Cells on chips. Nature 442, 403–411 (2006).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Park, I.-H. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008).
Reardon, S. Organs-on-chips’ go mainstream: drug companies put in vitro systems through their paces. Nature 523, 266–267 (2015).
Zhang, B. & Radisic, M. Organ-on-a-Chip devices advance to market. Lab. Chip 17, 2395–2420 (2017). This is a comprehensive review on organ-on-a-chip start-ups in the market space.
Zheng, Y., Chen, J. & López, J. A. Flow-driven assembly of VWF fibres and webs in in vitro microvessels. Nat. Commun. 6, 7858 (2015).
Zheng, Y. et al. In vitro microvessels for the study of angiogenesis and thrombosis. Proc. Natl Acad. Sci. USA 109, 9342–9347 (2012).
Jang, K.-J. et al. Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr. Biol. 5, 1119–1129 (2013).
Kim, H. J., Huh, D., Hamilton, G. & Ingber, D. E. Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab. Chip 12, 2165–2174 (2012).
Benam, K. H. et al. Small airway-on-a-chip enables analysis of human lung inflammation and drug responses in vitro. Nat. Methods 13, 151–157 (2016).
Musah, S. et al. Mature induced-pluripotent-stem-cell-derived human podocytes reconstitute kidney glomerular-capillary-wall function on a chip. Nat. Biomed. Eng. 1, 0069 (2017).
Adriani, G., Ma, D., Pavesi, A., Kamm, R. D. & Goh, E. L. A. 3D neurovascular microfluidic model consisting of neurons, astrocytes and cerebral endothelial cells as a blood–brain barrier. Lab. Chip (2017).
Jeon, J. S. et al. Human 3D vascularized organotypic microfluidic assays to study breast cancer cell extravasation. Proc. Natl Acad. Sci. USA 112, 214–219 (2015).
Trietsch, S. J. et al. Membrane-free culture and real-time barrier integrity assessment of perfused intestinal epithelium tubes. Nat. Commun. 8, 262 (2017).
Adler, M. et al. A quantitative approach to screen for nephrotoxic compounds in vitro. J. Am. Soc. Nephrol. 27, 1015–1028 (2016).
Homan, K. A. et al. Bioprinting of 3D convoluted renal proximal tubules on perfusable chips. Sci. Rep. 6, 34845 (2016).
Weber, E. J. et al. Development of a microphysiological model of human kidney proximal tubule function. Kidney Int. 90, 627–637 (2016).
Shlomai, A. et al. Modeling host interactions with hepatitis B virus using primary and induced pluripotent stem cell-derived hepatocellular systems. Proc. Natl Acad. Sci. USA 111, 12193–12198 (2014).
Khetani, S. R. & Bhatia, S. N. Microscale culture of human liver cells for drug development. Nat. Biotechnol. 26, 120–126 (2008).
Frey, O., Misun, P. M., Fluri, D. A., Hengstler, J. G. & Hierlemann, A. Reconfigurable microfluidic hanging drop network for multi-tissue interaction and analysis. Nat. Commun. 5, 4250 (2014).
Yazdi, S. R. et al. Adding the ‘heart’to hanging drop networks for microphysiological multi-tissue experiments. Lab. Chip 15, 4138–4147 (2015).
Nguyen, D. G. et al. Bioprinted 3D primary liver tissues allow assessment of organ-level response to clinical drug induced toxicity in vitro. PLoS One 11, e0158674 (2016).
Nunes, S. S. et al. Biowire: a platform for maturation of human pluripotent stem cell-derived cardiomyocytes. Nat. Methods 10, 781–787 (2013). This is a study on the maturation of engineered human cardiac tissues with electrical stimulation.
Mathur, A. et al. Human iPSC-based cardiac microphysiological system for drug screening applications. Sci. Rep. 5, 8883 (2015).
Legant, W. R. et al. Microfabricated tissue gauges to measure and manipulate forces from 3D microtissues. Proc. Natl Acad. Sci. USA 106, 10097–10102 (2009).
Jackman, C. P., Carlson, A. L. & Bursac, N. Dynamic culture yields engineered myocardium with near-adult functional output. Biomaterials 111, 66–79 (2016).
Lind, J. U. et al. Instrumented cardiac microphysiological devices via multimaterial three-dimensional printing. Nat. Mater. 16, 303–308 (2016). This study presents an entirely 3D-printed platform for the culture and analysis of engineered cardiac tissues.
Sidorov, V. Y. et al. I-Wire Heart-on-a-Chip I: three-dimensional cardiac tissue constructs for physiology and pharmacology. Acta Biomaterialia 48, 68–78 (2017).
Zimmermann, W.-H. et al. Engineered heart tissue grafts improve systolic and diastolic function in infarcted rat hearts. Nat. Med. 12, 452–458 (2006).
Wittmann, K. & Fischbach, C. Contextual control of adipose-derived stem cell function: implications for engineered tumor models. ACS Biomater. Sci. Eng. 3, 1483–1493 (2016).
Miller, P. G. & Shuler, M. L. Design and demonstration of a pumpless 14 compartment microphysiological system. Biotechnol. Bioeng. 113, 2213–2227 (2016). This study presents a body-on-a-chip platform integrated with 14 tissue compartments.
Oleaga, C. et al. Multi-Organ toxicity demonstration in a functional human in vitro system composed of four organs. Sci. Rep. 6, 20030 (2016).
Xiao, S. et al. A microfluidic culture model of the human reproductive tract and 28-day menstrual cycle. Nat. Commun. 8, 14584 (2017).
Coppeta, J. et al. A portable and reconfigurable multi-organ platform for drug development with onboard microfluidic flow control. Lab. Chip 17, 134–144 (2017).
Vernetti, L. et al. Functional coupling of human microphysiology systems: intestine, liver, kidney proximal tubule, blood-brain barrier and skeletal muscle. Sci. Rep. 7, 42296 (2017). This is a study on multi-organ interaction with multiple decentralized organ-on-a-chip systems located at different sites.
Maschmeyer, I. et al. A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin and kidney equivalents. Lab. Chip 15, 2688–2699 (2015).
Loskill, P., Marcus, S. G., Mathur, A., Reese, W. M. & Healy, K. E. μOrgano: A Lego®-like plug & play system for modular multi-organ-chips. PLoS One 10, e0139587 (2015).
Zhang, Y. S. et al. Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc. Natl Acad. Sci. USA 114, E2293–E2302 (2017).
Zhang, B. et al. Biodegradable scaffold with built-in vasculature for organ-on-a-chip engineering and direct surgical anastomosis. Nat. Mater. 15, 669–678 (2016). This study presents a biodegradable scaffold with a built-in blood vessel network that supports both organ-on-a-chip engineering and in vivo implantation.
Kolesky, D. B., Homan, K. A., Skylar-Scott, M. A. & Lewis, J. A. Three-dimensional bioprinting of thick vascularized tissues. Proc. Natl Acad. Sci. USA 113, 3179–3184 (2016).
Miller, J. S. et al. Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nat. Mater. 11, 768–774 (2012). This study discusses the rapid fabrication of microfluidic networks in hydrogel matrices by 3D printing of sacrificial materials.
Kang, H.-W. et al. A 3D bioprinting system to produce human-scale tissue constructs with structural integrity. Nat. Biotechnol. 34, 312–319 (2016).
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 526, 564–568 (2015).
Takebe, T., Zhang, B. & Radisic, M. Synergistic engineering: organoids meet organs-on-a-chip. Cell Stem Cell 21, 297–300 (2017). This is a review that promotes the integration of organoids and organs-on-a-chip as a future step in the field.
Berthier, E., Young, E. W. & Beebe, D. Engineers are from PDMS-land, biologists are from polystyrenia. Lab. Chip 12, 1224–1237 (2012).
Junaid, A., Mashaghi, A., Hankemeier, T. & Vulto, P. An end-user perspective on Organ-on-a-Chip: assays and usability aspects. Curr. Opin. Biomed. Engineer. 1, 15–22 (2017).
Maoz, B. M. et al. Organs-on-Chips with combined multi-electrode array and transepithelial electrical resistance measurement capabilities. Lab. Chip 17, 2294–2302 (2017).
Ingber, D. E. Cellular mechanotransduction: putting all the pieces together again. FASEB J. 20, 811–827 (2006).
Huh, D. et al. A human disease model of drug toxicity–induced pulmonary edema in a lung-on-a-chip microdevice. Sci. Transl Med. 4, 159ra147 (2012).
Huh, D. et al. Microfabrication of human organs-on-chips. Nat. Protoc. 8, 2135–2157 (2013).
Benam, K. H. et al. Matched-comparative modeling of normal and diseased human airway responses using a microengineered breathing lung chip. Cell Systems 3, 416–418 (2016).
Kim, H. J., Li, H., Collins, J. J. & Ingber, D. E. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc. Natl Acad. Sci. USA 113, E7–E15 (2016).
Villenave, R. et al. Human Gut-on-a-Chip supports polarized infection of coxsackie B1 virus in vitro. PLoS One 12, e0169412 (2017).
Kim, S. et al. Pharmacokinetic profile that reduces nephrotoxicity of gentamicin in a perfused kidney-on-a-chip. Biofabrication 8, 015021 (2016).
Tavana, H., Zamankhan, P., Christensen, P. J., Grotberg, J. B. & Takayama, S. Epithelium damage and protection during reopening of occluded airways in a physiologic microfluidic pulmonary airway model. Biomed. Microdevices 13, 731–742 (2011).
Tavana, H. et al. Dynamics of liquid plugs of buffer and surfactant solutions in a micro-engineered pulmonary airway model. Langmuir 26, 3744–3752 (2009).
Blundell, C. et al. A microphysiological model of the human placental barrier. Lab. Chip 16, 3065–3073 (2016).
Choi, Y. et al. A microengineered pathophysiological model of early-stage breast cancer. Lab. Chip 15, 3350–3357 (2015).
Heles, T. Emulate expands series B to $45m. Global University Venturing http://www.globaluniversityventuring.com/article.php/5521/emulate-expands-series-b-to-45m (2016).
Bouchie, A. & DeFrancesco, L. Nature Biotechnology’s academic spinouts of 2014. Nat. Biotechnol. 33, 247–255 (2015).
Timmerman, L. Organ-on-a-Chip startup, Emulate, grabs $45m to shake up drug discovery. Forbes https://www.forbes.com/sites/luketimmerman/2016/10/20/organ-on-a-chip-startup-emulate-grabs-45m-to-improve-drug-discovery/#3b20b0825fcb (2016).
Choi, N. W. et al. Microfluidic scaffolds for tissue engineering. Nat. Mater. 6, 908–915 (2007).
Leach, J. B. & Schmidt, C. E. Characterization of protein release from photocrosslinkable hyaluronic acid-polyethylene glycol hydrogel tissue engineering scaffolds. Biomaterials 26, 125–135 (2005).
Ligresti, G. et al. A novel three-dimensional human peritubular microvascular system. J. Am. Soc. Nephrol. 27, 2370–2381 (2015).
Tourovskaia, A., Fauver, M., Kramer, G., Simonson, S. & Neumann, T. Tissue-engineered microenvironment systems for modeling human vasculature. Exp. Biol. Med. 239, 1264–1271 (2014).
Stucki, A. O. et al. A lung-on-a-chip array with an integrated bio-inspired respiration mechanism. Lab. Chip 15, 1302–1310 (2015).
Gu, Z. Integrating intelligent materials into organ on a chip system (abstract W4B.1). Proceedings International Conference on Photonics and Imaging in Biology and Medicine (2017).
Moreno, E. L. et al. Differentiation of neuroepithelial stem cells into functional dopaminergic neurons in 3D microfluidic cell culture. Lab. Chip 15, 2419–2428 (2015).
Trietsch, S. J., Israëls, G. D., Joore, J., Hankemeier, T. & Vulto, P. Microfluidic titer plate for stratified 3D cell culture. Lab. Chip 13, 3548–3554 (2013).
Duinen, V. v. et al. 96 perfusable blood vessels to study vascular permeability in vitro. Sci. Rep. 7, 18071 (2017).
Jeon, N. L. In vitro formation and characterization of a perfusable three-dimensional tubular capillary network in microfluidic devices. Lab. Chip 12, 2815–2822 (2012).
Kim, S., Lee, H., Chung, M. & Jeon, N. L. Engineering of functional, perfusable 3D microvascular networks on a chip. Lab. Chip 13, 1489–1500 (2013).
Shin, Y. et al. Microfluidic assay for simultaneous culture of multiple cell types on surfaces or within hydrogels. Nat. Protoc. 7, 1247–1259 (2012).
Chung, S. et al. Cell migration into scaffolds under co-culture conditions in a microfluidic platform. Lab. Chip 9, 269–275 (2009).
Wang, X. et al. An on-chip microfluidic pressure regulator that facilitates reproducible loading of cells and hydrogels into microphysiological system platforms. Lab. Chip 16, 868–876 (2016).
Wang, X. et al. Engineering anastomosis between living capillary networks and endothelial cell-lined microfluidic channels. Lab. Chip 16, 282–290 (2016).
Moya, M. L., Hsu, Y.-H., Lee, A. P., Hughes, C. C. & George, S. C. In vitro perfused human capillary networks. Tissue Eng. Part C Methods 19, 730–737 (2013).
Hsu, Y.-H., Moya, M. L., Hughes, C. C., George, S. C. & Lee, A. P. A microfluidic platform for generating large-scale nearly identical human microphysiological vascularized tissue arrays. Lab. Chip 13, 2990–2998 (2013).
Wiedeman, M. P. Dimensions of blood vessels from distributing artery to collecting vein. Circul. Res. 12, 375–378 (1963).
Polacheck, W. J., German, A. E., Mammoto, A., Ingber, D. E. & Kamm, R. D. Mechanotransduction of fluid stresses governs 3D cell migration. Proc. Natl Acad. Sci. USA 111, 2447–2452 (2014).
Aref, A. R. et al. Screening therapeutic EMT blocking agents in a three-dimensional microenvironment. Integr. Biol. 5, 381–389 (2013).
Vickerman, V. & Kamm, R. D. Mechanism of a flow-gated angiogenesis switch: early signaling events at cell–matrix and cell–cell junctions. Integr. Biol. 4, 863–874 (2012).
Chung, M., Kim, S., Ahn, J., Lee, S. & Jeon, N. L. Interstitial flow regulates angiogenic response and phenotype of endothelial cells in a 3D culture model. Lab. Chip 16, 4189–4199 (2016).
Phan, D. T. et al. A vascularized and perfused organ-on-a-chip platform for large-scale drug screening applications. Lab. Chip 17, 511–520 (2017).
LiáJeon, N. Microfluidic vascularized bone tissue model with hydroxyapatite-incorporated extracellular matrix. Lab. Chip 15, 3984–3988 (2015).
Zervantonakis, I. K. et al. Three-dimensional microfluidic model for tumor cell intravasation and endothelial barrier function. Proc. Natl Acad. Sci. USA 109, 13515–13520 (2012).
Sobrino, A. et al. 3D microtumors in vitro supported by perfused vascular networks. Sci. Rep. 6, 31589 (2016).
Oh, S. et al. “Open-top” microfluidic device for in vitro three-dimensional capillary beds. Lab. Chip 17, 3405–3414 (2017).
Nashimoto, Y. et al. Integrating perfusable vascular networks with a three-dimensional tissue in a microfluidic device. Integr. Biol. 9, 506–518 (2017).
Kim, S., Chung, M. & Jeon, N. L. Three-dimensional biomimetic model to reconstitute sprouting lymphangiogenesis in vitro. Biomaterials 78, 115–128 (2016).
Wadman, M. Merck settles Vioxx lawsuits for $4.85 billion. Nature 450, 324 (2007).
Kattman, S. J. et al. Stage-specific optimization of activin/nodal and BMP signaling promotes cardiac differentiation of mouse and human pluripotent stem cell lines. Cell Stem Cell 8, 228–240 (2011).
Radisic, M. et al. Functional assembly of engineered myocardium by electrical stimulation of cardiac myocytes cultured on scaffolds. Proc. Natl Acad. Sci. USA 101, 18129–18134 (2004).
Tandon, N. et al. Electrical stimulation systems for cardiac tissue engineering. Nat. Protoc. 4, 155–173 (2009).
Stoehr, A. et al. Spontaneous formation of extensive vessel-like structures in murine engineered heart tissue. Tissue Eng. Part A 22, 326–335 (2016).
Mannhardt, I. et al. Human engineered heart tissue: analysis of contractile force. Stem Cell Rep. 7, 29–42 (2016).
Eder, A., Vollert, I., Hansen, A. & Eschenhagen, T. Human engineered heart tissue as a model system for drug testing. Adv. Drug Deliv. Rev. 96, 214–224 (2016).
Zimmermann, W.-H. et al. Tissue engineering of a differentiated cardiac muscle construct. Circul. Res. 90, 223–230 (2002).
Bian, W., Badie, N., Himel IV, H. D. & Bursac, N. Robust T-tubulation and maturation of cardiomyocytes using tissue-engineered epicardial mimetics. Biomaterials 35, 3819–3828 (2014).
Vunjak-Novakovic, G., Bhatia, S., Chen, C. & Hirschi, K. HeLiVa platform: integrated heart-liver-vascular systems for drug testing in human health and disease. Stem Cell Res. Ther. 4, S8 (2013).
Wilson, K., Das, M., Wahl, K. J., Colton, R. J. & Hickman, J. Measurement of contractile stress generated by cultured rat muscle on silicon cantilevers for toxin detection and muscle performance enhancement. PLoS One 5, e11042 (2010).
Schroer, A. K., Shotwell, M. S., Sidorov, V. Y., Wikswo, J. P. & Merryman, W. D. I-Wire Heart-on-a-Chip II: biomechanical analysis of contractile, three-dimensional cardiomyocyte tissue constructs. Acta Biomaterialia 48, 79–87 (2017).
Watkins, P. B. et al. The clinical liver safety assessment best practices workshop: rationale, goals, accomplishments and the future. Drug Safety 37, 1–7 (2014).
Lucena, M. I. et al. Trovafloxacin-induced acute hepatitis. Clin. Infect. Dis. 30, 400–401 (2000).
Bhatia, S., Balis, U., Yarmush, M. & Toner, M. Effect of cell–cell interactions in preservation of cellular phenotype: cocultivation of hepatocytes and nonparenchymal cells. FASEB J. 13, 1883–1900 (1999).
Hui, E. E. & Bhatia, S. N. Micromechanical control of cell–cell interactions. Proc. Natl Acad. Sci. USA 104, 5722–5726 (2007).
Stevens, K. et al. InVERT molding for scalable control of tissue microarchitecture. Nat. Commun. 4, 1847 (2013).
Ware, B. R., Berger, D. R. & Khetani, S. R. Prediction of drug-induced liver injury in micropatterned co-cultures containing iPSC-derived human hepatocytes. Toxicol. Sci. 145, 252–262 (2015).
Ploss, A. et al. Persistent hepatitis C virus infection in microscale primary human hepatocyte cultures. Proc. Natl Acad. Sci. USA 107, 3141–3145 (2010).
Khetani, S. R. et al. Use of micropatterned cocultures to detect compounds that cause drug-induced liver injury in humans. Toxicol. Sci. 132, 107–117 (2013).
Xu, J. J. et al. Cellular imaging predictions of clinical drug-induced liver injury. Toxicol. Sci. 105, 97–105 (2008).
Nguyen, T. V. et al. Establishment of a hepatocyte-kupffer cell coculture model for assessment of proinflammatory cytokine effects on metabolizing enzymes and drug transporters. Drug Metab. Dispos. 43, 774–785 (2015).
Stevens, K. R. et al. In situ expansion of engineered human liver tissue in a mouse model of chronic liver disease. Sci. Transl Med. 9, eaah5505 (2017).
Jakab, K., Neagu, A., Mironov, V., Markwald, R. R. & Forgacs, G. Engineering biological structures of prescribed shape using self-assembling multicellular systems. Proc. Natl Acad. Sci. USA 101, 2864–2869 (2004).
Norotte, C., Marga, F. S., Niklason, L. E. & Forgacs, G. Scaffold-free vascular tissue engineering using bioprinting. Biomaterials 30, 5910–5917 (2009).
Jakab, K. et al. Tissue engineering by self-assembly and bio-printing of living cells. Biofabrication 2, 022001 (2010).
Norona, L. M., Nguyen, D. G., Gerber, D. A., Presnell, S. C. & LeCluyse, E. L. Modeling compound-induced fibrogenesis in vitro using three-dimensional bioprinted human liver tissues. Toxicol. Sci. 154, 354–367 (2016).
Rodenhizer, D. et al. A three-dimensional engineered tumour for spatial snapshot analysis of cell metabolism and phenotype in hypoxic gradients. Nat. Mater. 15, 227–234 (2016).
Kelm, J. M., Timmins, N. E., Brown, C. J., Fussenegger, M. & Nielsen, L. K. Method for generation of homogeneous multicellular tumor spheroids applicable to a wide variety of cell types. Biotechnol. Bioeng. 83, 173–180 (2003).
Kelm, J. M. & Fussenegger, M. Microscale tissue engineering using gravity-enforced cell assembly. Trends Biotechnol. 22, 195–202 (2004).
Kim, J.-Y. et al. 3D spherical microtissues and microfluidic technology for multi-tissue experiments and analysis. J. Biotechnol. 205, 24–35 (2015).
Falkenberg, N. et al. Three-dimensional microtissues essentially contribute to preclinical validations of therapeutic targets in breast cancer. Cancer Med. 5, 703–710 (2016).
Horman, S. R. et al. High-content analysis of three-dimensional tumor spheroids: Investigating signaling pathways using small hairpin RNA. Nat. Methods 10, 1036 (2013).
Wagner, I. et al. A dynamic multi-organ-chip for long-term cultivation and substance testing proven by 3D human liver and skin tissue co-culture. Lab. Chip 13, 3538–3547 (2013).
Chung, K. et al. A microfluidic array for large-scale ordering and orientation of embryos. Nat. Methods 8, 171–176 (2011).
Levario, T. J., Zhan, M., Lim, B., Shvartsman, S. Y. & Lu, H. Microfluidic trap array for massively parallel imaging of Drosophila embryos. Nat. Protoc. 8, 721 (2013).
Jackson, E. & Lu, H. Three-dimensional models for studying development and disease: moving on from organisms to organs-on-a-chip and organoids. Integr. Biol. 8, 672–683 (2016).
Jackson-Holmes, E., McDevitt, T. C. & Lu, H. A microfluidic trap array for longitudinal monitoring and multi-modal phenotypic analysis of individual stem cell aggregates. Lab. Chip 17, 3634–3642 (2017).
Atienzar, F. A. et al. Predictivity of dog co-culture model, primary human hepatocytes and HepG2 cells for the detection of hepatotoxic drugs in humans. Toxicol. Appl. Pharmacol. 275, 44–61 (2014).
Novik, E., Maguire, T. J., Chao, P., Cheng, K. & Yarmush, M. L. A microfluidic hepatic coculture platform for cell-based drug metabolism studies. Biochem. Pharmacol. 79, 1036–1044 (2010).
Tatosian, D. A. & Shuler, M. L. A novel system for evaluation of drug mixtures for potential efficacy in treating multidrug resistant cancers. Biotechnol. Bioeng. 103, 187–198 (2009).
Kidambi, S. et al. Oxygen-mediated enhancement of primary hepatocyte metabolism, functional polarization, gene expression, and drug clearance. Proc. Natl Acad. Sci. USA 106, 15714–15719 (2009).
Chao, P., Maguire, T., Novik, E., Cheng, K.-C. & Yarmush, M. Evaluation of a microfluidic based cell culture platform with primary human hepatocytes for the prediction of hepatic clearance in human. Biochem. Pharmacol. 78, 625–632 (2009).
Wikswo, J. P. et al. Scaling and systems biology for integrating multiple organs-on-a-chip. Lab. Chip 13, 3496–3511 (2013).
Moraes, C. et al. On being the right size: scaling effects in designing a human-on-a-chip. Integr. Biol. 5, 1149–1161 (2013).
Esch, M. B., Ueno, H., Applegate, D. R. & Shuler, M. L. Modular, pumpless body-on-a-chip platform for the co-culture of GI tract epithelium and 3D primary liver tissue. Lab. Chip 16, 2719–2729 (2016).
Esch, M. B., Mahler, G. J., Stokol, T. & Shuler, M. L. Body-on-a-chip simulation with gastrointestinal tract and liver tissues suggests that ingested nanoparticles have the potential to cause liver injury. Lab. Chip 14, 3081–3092 (2014).
Sung, J. H. & Shuler, M. L. A micro cell culture analog (μCCA) with 3D hydrogel culture of multiple cell lines to assess metabolism-dependent cytotoxicity of anti-cancer drugs. Lab. Chip 9, 1385–1394 (2009).
Materne, E.-M. et al. The multi-organ chip-a microfluidic platform for long-term multi-tissue coculture. J. Vis. Exp. 98, e52526 (2015).
Bauer, S. et al. Functional coupling of human pancreatic islets and liver spheroids on-a-chip: Towards a novel human ex vivo type 2 diabetes model. Sci. Rep. 7, 14620 (2017).
Loskill, P. et al. WAT-on-a-chip: a physiologically relevant microfluidic system incorporating white adipose tissue. Lab. Chip 17, 1645–1654 (2017).
Yu, J. et al. Quantitative systems pharmacology approaches applied to microphysiological systems (MPS): data interpretation and multi-MPS integration. CPT Pharmacometrics Syst. Pharmacol. 4, 585–594 (2015).
Wikswo, J. P. & Porter, A. P. Biology coming full circle: joining the whole and the parts. Exp. Biol. Med. 240, 3–7 (2015).
Lai, B. F. L. et al. InVADE: integrated vasculature for assessing dynamic events. Adv. Funct. Mater. https://doi.org/10.1002/adfm.201703524 (2017). This study presents an organ-on-a-chip platform in the format of a multiwell plate that can model inter-organ cancer metastasis.
Yarmush, M. L., Freedman, R., Del Bufalo, A., Teissier, S. & Meunier, J.-R. Immune system modeling devices and methods. Patent application number US9535056B2 (2017).
Vargesson, N. Thalidomide-induced teratogenesis: history and mechanisms. Birth Defects Res. C Embryo Today 105, 140–156 (2015).
Therapontos, C., Erskine, L., Gardner, E. R., Figg, W. D. & Vargesson, N. Thalidomide induces limb defects by preventing angiogenic outgrowth during early limb formation. Proc. Natl Acad. Sci. USA 106, 8573–8578 (2009).
Nawroth, J., Rogal, J., Weiss, M., Brucker, S. Y. & Loskill, P. Organ-on-a-Chip systems for women’s health applications. Adv. Healthc. Mater. https://doi.org/10.1002/adhm.201700550 (2017).
Jo, B.-H., Van Lerberghe, L. M., Motsegood, K. M. & Beebe, D. J. Three-dimensional micro-channel fabrication in polydimethylsiloxane (PDMS) elastomer. J. Microelectromechan. Systems 9, 76–81 (2000).
Thorsen, T., Maerkl, S. J. & Quake, S. R. Microfluidic large-scale integration. Science 298, 580 (2002).
Unger, M. A., Chou, H. P., Thorsen, T., Scherer, A. & Quake, S. R. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113–116 (2000).
Woodruff, K. & Maerkl, S. J. A. High-throughput microfluidic platform for mammalian cell transfection and culturing. Sci. Rep. 6, 23937 (2016).
Fidalgo, L. M. & Maerkl, S. J. A software-programmable microfluidic device for automated biology. Lab. Chip 11, 1612–1619 (2011).
Piraino, F., Volpetti, F., Watson, C. & Maerkl, S. J. A. Digital–analog microfluidic platform for patient-centric multiplexed biomarker diagnostics of ultralow volume samples. ACS Nano 10, 1699–1710 (2016).
Miklas, J. W. et al. Bioreactor for modulation of cardiac microtissue phenotype by combined static stretch and electrical stimulation. Biofabrication 6, 024113–024113 (2014).
Borysiak, M. D. et al. Simple replica micromolding of biocompatible styrenic elastomers. Lab. Chip 13, 2773–2784 (2013).
Borysiak, M. D., Yuferova, E. & Posner, J. D. Simple, low-cost styrene-ethylene/butylene-styrene microdevices for electrokinetic applications. Anal. Chem. 85, 11700–11704 (2013).
Guillemette, M. D., Roy, E., Auger, F. A. & Veres, T. Rapid isothermal substrate microfabrication of a biocompatible thermoplastic elastomer for cellular contact guidance. Acta Biomaterialia 7, 2492–2498 (2011).
Domansky, K. et al. Clear castable polyurethane elastomer for fabrication of microfluidic devices. Lab. Chip 13, 3956–3964 (2013).
Sasaki, H., Onoe, H., Osaki, T., Kawano, R. & Takeuchi, S. Parylene-coating in PDMS microfluidic channels prevents the absorption of fluorescent dyes. Sensors Actuators B Chem. 150, 478–482 (2010).
Ren, K., Zhao, Y., Su, J., Ryan, D. & Wu, H. Convenient method for modifying poly (dimethylsiloxane) to be airtight and resistive against absorption of small molecules. Anal. Chem. 82, 5965–5971 (2010).
Tran, R. T. et al. Synthesis and characterization of a biodegradable elastomer featuring a dual crosslinking mechanism. Soft Matter 6, 2449–2461 (2010).
Davenport Huyer, L. et al. Highly elastic and moldable polyester biomaterial for cardiac tissue engineering applications. ACS Biomater. Sci. Eng. 2, 780–788 (2016).
Pan, C., Kumar, C., Bohl, S., Klingmueller, U. & Mann, M. Comparative proteomic phenotyping of cell lines and primary cells to assess preservation of cell type-specific functions. Mol. Cell. Proteom.: MCP 8, 443–450 (2009).
Zhao, Y., Korolj, A., Feric, N. & Radisic, M. Human pluripotent stem cell-derived cardiomyocyte based models for cardiotoxicity and drug discovery. Expert Opin. Drug Safety 15, 1455–1458 (2016).
Ahadian, S. et al. Organ-on-a-Chip platforms: a convergence of advanced materials, cells, and microscale technologies. Adv. Healthc Mater. https://doi.org/10.1002/adhm.201700506 (2017).
Unger, R. E., Krump-Konvalinkova, V., Peters, K. & Kirkpatrick, C. J. In vitro expression of the endothelial phenotype: comparative study of primary isolated cells and cell lines, including the novel cell line HPMEC-ST1.6R. Microvascular Res. 64, 384–397 (2002).
Baumann, K. Achieving pluripotency. Nat Rev. Mol. Cell Biol. 11, 677 (2010).
Saha, K. & Jaenisch, R. Technical challenges in using human induced pluripotent stem cells to model disease. Cell Stem Cell 5, 584–595.
Hanna, J. et al. Treatment of sickle cell anemia mouse model with iPS cells generated from autologous skin. Science 318, 1920 (2007).
Colman, A. & Dreesen, O. Pluripotent stem cells and disease modeling. Cell Stem Cell 5, 244–247 (2009).
Li, Y. Y. & Jones, S. J. M. Drug repositioning for personalized medicine. Genome Med. 4, 27–27 (2012).
Evans, W. E. & Relling, M. V. Pharmacogenomics: translating functional genomics into rational therapeutics. Science 286, 487 (1999).
van de Stolpe, A. & den Toonder, J. Workshop meeting report Organs-on-Chips: human disease models. Lab. Chip 13, 3449–3470 (2013).
Yazawa, M. et al. Using induced pluripotent stem cells to investigate cardiac phenotypes in Timothy syndrome. Nature 471, 230–234 (2011).
Moretti, A. et al. Patient-specific induced pluripotent stem-cell models for long-QT syndrome. N. Engl. J. Med. 363, 1397–1409 (2010).
Carvajal-Vergara, X. et al. Patient-specific induced pluripotent stem-cell-derived models of LEOPARD syndrome. Nature 465, 808–812 (2010).
Wang, G. et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat. Med. 20, 616–623 (2014).
Sun, N. et al. Patient-specific induced pluripotent stem cells as a model for familial dilated cardiomyopathy. Sci. Transl Med. 4, 130ra147 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Völkner, M. et al. Retinal organoids from pluripotent stem cells efficiently recapitulate retinogenesis. Stem Cell Rep. 6, 525–538 (2016).
Foster, J. W. et al. Cornea organoids from human induced pluripotent stem cells. Sci. Rep. 7, 41286 (2017).
Broutier, L. et al. Human primary liver cancer-derived organoid cultures for disease modeling and drug screening. Nat. Med. 23, 1424 (2017).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Boj, Sylvia, F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338.
Dedhia, P. H., Bertaux-Skeirik, N., Zavros, Y. & Spence, J. R. Organoid models of human gastrointestinal development and disease. Gastroenterology 150, 1098–1112 (2016).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373 (2013).
Chen, Y.-W. et al. A three-dimensional model of human lung development and disease from pluripotent stem cells. Nat. Cell Biol. 19, 542 (2017).
Lancaster, M. A. & Knoblich, J. A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125 (2014).
Byun, C. K., Abi-Samra, K., Cho, Y. K. & Takayama, S. Pumps for microfluidic cell culture. Electrophoresis 35, 245–257 (2014).
Li, X., Brooks, J. C., Hu, J., Ford, K. I. & Easley, C. J. 3D-templated, fully automated microfluidic input/output multiplexer for endocrine tissue culture and secretion sampling. Lab. Chip 17, 341–349 (2017).
Huh, D., Hamilton, G. A. & Ingber, D. E. From 3D cell culture to organs-on-chips. Trends Cell Biol. 21, 745–754 (2011).
Wevers, N. R. et al. High-throughput compound evaluation on 3D networks of neurons and glia in a microfluidic platform. Sci. Rep. 6, 38856 (2016).
Theberge, A. B. et al. Microfluidic multiculture assay to analyze biomolecular signaling in angiogenesis. Anal. Chem. 87, 3239–3246 (2015).
Kim, S., Chung, M., Ahn, J., Lee, S. & Jeon, N. L. Interstitial flow regulates the angiogenic response and phenotype of endothelial cells in a 3D culture model. Lab. Chip 16, 4189–4199 (2016).
Osaki, T., Sivathanu, V. & Kamm, R. D. Crosstalk between developing vasculature and optogenetically engineered skeletal muscle improves muscle contraction and angiogenesis. Biomaterials 156, 65–76 (2018).
Uzel, S. G. et al. Microfluidic device for the formation of optically excitable, three-dimensional, compartmentalized motor units. Sci. Adv. 2, e1501429 (2016).
Patra, B., Peng, C.-C., Liao, W.-H., Lee, C.-H. & Tung, Y.-C. Drug testing and flow cytometry analysis on a large number of uniform sized tumor spheroids using a microfluidic device. Sci. Rep. 6, 21061 (2016).
Gruber, P., Marques, M. P., Szita, N. & Mayr, T. Integration and application of optical chemical sensors in microbioreactors. Lab. Chip 17, 2693–2712 (2017).
Choi, J.-r., Song, H., Sung, J. H., Kim, D. & Kim, K. Microfluidic assay-based optical measurement techniques for cell analysis: a review of recent progress. Biosensors Bioelectron. 77, 227–236 (2016).
Xiao, F., Wang, L. & Duan, H. Nanomaterial based electrochemical sensors for in vitro detection of small molecule metabolites. Biotechnol. Adv. 34, 234–249 (2016).
Wang, L., Acosta, M. A., Leach, J. B. & Carrier, R. L. Spatially monitoring oxygen level in 3D microfabricated cell culture systems using optical oxygen sensing beads. Lab. Chip 13, 1586–1592 (2013).
Suzuki, H., Hirakawa, T., Watanabe, I. & Kikuchi, Y. Determination of blood pO 2 using a micromachined Clark-type oxygen electrode. Anal. Chim. Acta 431, 249–259 (2001).
Bellin, D. L. et al. Integrated circuit-based electrochemical sensor for spatially resolved detection of redox-active metabolites in biofilms. Nat. Commun. 5, 3256 (2014).
Wu, M.-H., Lin, J.-L., Wang, J., Cui, Z. & Cui, Z. Development of high throughput optical sensor array for on-line pH monitoring in micro-scale cell culture environment. Biomed. Microdevices 11, 265–273 (2009).
Eklund, S. E. et al. Modification of the Cytosensor™ Microphysiometer to simultaneously measure extracellular acidification and oxygen consumption rates. Anal. Chim. Acta 496, 93–101 (2003).
Lin, Z., Cherng-Wen, T., Roy, P. & Trau, D. In-situ measurement of cellular microenvironments in a microfluidic device. Lab. Chip 9, 257–262 (2009).
Lin, Y., Lu, F., Tu, Y. & Ren, Z. Glucose biosensors based on carbon nanotube nanoelectrode ensembles. Nano Lett. 4, 191–195 (2004).
Boero, C. et al. Design, development, and validation of an in-situ biosensor array for metabolite monitoring of cell cultures. Biosensors Bioelectron. 61, 251–259 (2014).
Sonker, M., Sahore, V. & Woolley, A. T. Recent advances in microfluidic sample preparation and separation techniques for molecular biomarker analysis: a critical review. Anal. Chim. Acta 986, 1–11 (2017).
Xiao, Y., Lubin, A. A., Heeger, A. J. & Plaxco, K. W. Label-free electronic detection of thrombin in blood serum by using an aptamer-based sensor. Angew. Chem. Int. Ed. 44, 5456–5459 (2005).
Shin, S. R. et al. Aptamer-based microfluidic electrochemical biosensor for monitoring cell-secreted trace cardiac biomarkers. Anal. Chem. 88, 10019–10027 (2016).
Shin, S. R. et al. Label-free and regenerative electrochemical microfluidic biosensors for continual monitoring of cell secretomes. Adv. Sci. 4, 1600522 (2017).
Xie, Y. et al. A novel electrochemical microfluidic chip combined with multiple biomarkers for early diagnosis of gastric cancer. Nanoscale Res. Lett. 10, 477 (2015).
Henry, O. et al. Organs-on-Chips with integrated electrodes for Trans-Epithelial Electrical Resistance (TEER) measurements of human epithelial barrier function. Lab. Chip 17, 2264–2271 (2017).
ávan der Meer, A. D., JungáKim, H., ávan der Helm, M. W. & den Berg, A. Measuring direct current trans-epithelial electrical resistance in organ-on-a-chip microsystems. Lab. Chip 15, 745–752 (2015).
Murphy, S. V. & Atala, A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 32, 773–785 (2014).
Waheed, S. et al. 3D printed microfluidic devices: enablers and barriers. Lab. Chip 16, 1993–2013 (2016).
Zhang, Y. S. et al. 3D bioprinting for tissue and organ fabrication. Ann. Biomed. Engineer. 45, 148–163 (2017).
Dehne, E.-M., Hasenberg, T. & Marx, U. The ascendance of microphysiological systems to solve the drug testing dilemma. Future Sci. OA 3, FSO0185 (2017).
Sager, P. T., Gintant, G., Turner, J. R., Pettit, S. & Stockbridge, N. Rechanneling the cardiac proarrhythmia safety paradigm: a meeting report from the Cardiac Safety Research Consortium. Am. Heart J. 167, 292–300 (2014).
US Food and Drug Administration. Paving the way for personalized medicine. (FDA, 2013).
Conant, G., Ahadian, S., Zhao, Y. & Radisic, M. Kinase inhibitor screening using artificial neural networks and engineered cardiac biowires. Sci. Rep. 7, 11807 (2017).
Sun, X. & Nunes, S. S. Biowire platform for maturation of human pluripotent stem cell-derived cardiomyocytes. Methods 101, 21–26 (2016).
Marga, F. et al. Toward engineering functional organ modules by additive manufacturing. Biofabrication 4, 022001 (2012).
Zhou, M. et al. Development of a functional glomerulus at the organ level on a chip to mimic hypertensive nephropathy. Sci. Rep. 6, 31771 (2016).
Wang, L. et al. A disease model of diabetic nephropathy in a glomerulus-on-a-chip microdevice. Lab. Chip 17, 1749–1760 (2017).
Lee, J. S. et al. Placenta-on-a-chip: a novel platform to study the biology of the human placenta. J. Maternal Fetal Neonatal Med. 29, 1046–1054 (2016).
Booth, R. & Kim, H. Characterization of a microfluidic in vitro model of the blood-brain barrier (μBBB). Lab. Chip 12, 1784–1792 (2012).
Brown, J. A. et al. Metabolic consequences of inflammatory disruption of the blood-brain barrier in an organ-on-chip model of the human neurovascular unit. J. Neuroinflamm. 13, 306 (2016).
Deosarkar, S. P. et al. A novel dynamic neonatal blood-brain barrier on a chip. PLoS One 10, e0142725 (2015).
Prabhakarpandian, B. et al. SyM-BBB: a microfluidic blood brain barrier model. Lab. Chip 13, 1093–1101 (2013).
Chung, M. et al. Wet-AMD on a chip: modeling outer blood-retinal barrier in vitro. Adv. Healthc. Mater. https://doi.org/10.1002/adhm.201700028 (2017).
Kolesky, D. B. et al. 3D bioprinting of vascularized, heterogeneous cell-laden tissue constructs. Adv. Mater. 26, 3124–3130 (2014).
Tsvirkun, D., Grichine, A., Duperray, A., Misbah, C. & Bureau, L. Microvasculature on a chip: study of the endothelial surface layer and the flow structure of red blood cells. Sci. Rep. 7, 45036 (2017).
Zhang, B., Peticone, C., Murthy, S. K. & Radisic, M. A standalone perfusion platform for drug testing and target validation in micro-vessel networks. Biomicrofluidics 7, 044125 (2013).
Ting, L., Feghhi, S., Karchin, A., Tooley, W. & White, N. J. Clot-on-a-chip: a microfluidic device to study platelet aggregation and contractility under shear. Blood 122, 2363–2363 (2013).
Lamberti, G. et al. Adhesive interaction of functionalized particles and endothelium in idealized microvascular networks. Microvascular Res. 89, 107–114 (2013).
Wang, L. et al. Patterning cells and shear flow conditions: convenient observation of endothelial cell remoulding, enhanced production of angiogenesis factors and drug response. Lab. Chip 11, 4235–4240 (2011).
Srigunapalan, S., Lam, C., Wheeler, A. R. & Simmons, C. A. A microfluidic membrane device to mimic critical components of the vascular microenvironment. Biomicrofluidics 5, 13409 (2011).
Zhang, Y. S. et al. Bioprinted thrombosis-on-a-chip. Lab. Chip 16, 4097–4105 (2016).
Zhang, W. et al. Elastomeric free-form blood vessels for interconnecting organs on chip systems. Lab. Chip 16, 1579–1586 (2016).
Yasotharan, S., Pinto, S., Sled, J. G., Bolz, S.-S. & Günther, A. Artery-on-a-chip platform for automated, multimodal assessment of cerebral blood vessel structure and function. Lab. Chip 15, 2660–2669 (2015).
Günther, A. et al. A microfluidic platform for probing small artery structure and function. Lab. Chip 10, 2341–2349 (2010).
Price, G. M., Chrobak, K. M. & Tien, J. Effect of cyclic AMP on barrier function of human lymphatic microvascular tubes. Microvascular Res. 76, 46–51 (2008).
Schaaf, S. et al. Human engineered heart tissue as a versatile tool in basic research and preclinical toxicology. PLoS One 6, e26397 (2011).
Yang, X., Pabon, L. & Murry, C. E. Engineering adolescence maturation of human pluripotent stem cell–derived cardiomyocytes. Circul. Res. 114, 511–523 (2014).
Bian, W., Jackman, C. P. & Bursac, N. Controlling the structural and functional anisotropy of engineered cardiac tissues. Biofabrication 6, 024109 (2014).
Nunes, S. S. et al. Human stem cell-derived cardiac model of chronic drug exposure. ACS Biomater. Sci. Eng. 3, 1911–1921 (2017).
Thavandiran, N. et al. Design and formulation of functional pluripotent stem cell-derived cardiac microtissues. Proc. Natl Acad. Sci. USA 110, E4698–E4707 (2013).
Guo, X., Gonzalez, M., Stancescu, M., Vandenburgh, H. H. & Hickman, J. J. Neuromuscular junction formation between human stem cell-derived motoneurons and human skeletal muscle in a defined system. Biomaterials 32, 9602–9611 (2011).
Davidson, M. D., Lehrer, M. & Khetani, S. R. Hormone and drug-mediated modulation of glucose metabolism in a microscale model of the human liver. Tissue Eng. Part C Methods 21, 716–725 (2015).
Chan, T. S., Yu, H., Moore, A., Khetani, S. R. & Tweedie, D. Meeting the challenge of predicting hepatic clearance of compounds slowly metabolized by cytochrome P450 using a novel hepatocyte model, HepatoPac. Drug Metab. Dispos. 41, 2024–2032 (2013).
Ballard, T. E. et al. Application of a micropatterned cocultured hepatocyte system to predict preclinical and human-specific drug metabolism. Drug Metab. Dispos. 44, 172–179 (2016).
Anastasov, N. et al. A 3D-microtissue-based phenotypic screening of radiation resistant tumor cells with synchronized chemotherapeutic treatment. BMC Cancer 15, 1 (2015).
Rimann, M. et al. An in vitro osteosarcoma 3D microtissue model for drug development. J. Biotechnol. 189, 129–135 (2014).
Huval, R. M. et al. Microengineered peripheral nerve-on-a-chip for preclinical physiological testing. Lab. Chip 15, 2221–2232 (2015).
Liazoghli, D., Roth, A. D., Thostrup, P. & Colman, D. R. Substrate micropatterning as a new in vitro cell culture system to study myelination. ACS Chem. Neurosci. 3, 90–95 (2011).
Magdesian, M. H. et al. Atomic force microscopy reveals important differences in axonal resistance to injury. Biophys. J. 103, 405–414 (2012).
Belkaid, W. et al. Cellular response to micropatterned growth promoting and inhibitory substrates. BMC Biotechnol. 13, 86 (2013).
Magdesian, M. H. et al. Rapid mechanically controlled rewiring of neuronal circuits. J. Neurosci. 36, 979–987 (2016).
Khoshakhlagh, P. & Moore, M. J. Photoreactive interpenetrating network of hyaluronic acid and Puramatrix as a selectively tunable scaffold for neurite growth. Acta Biomaterialia 16, 23–34 (2015).
Peyrin, J.-M. et al. Axon diodes for the reconstruction of oriented neuronal networks in microfluidic chambers. Lab. Chip 11, 3663–3673 (2011).
Deleglise, B. et al. β-Amyloid induces a dying-back process and remote trans-synaptic alterations in a microfluidic-based reconstructed neuronal network. Acta Neuropathol. Commun. 2, 145 (2014).
Taylor, A. M. et al. A microfluidic culture platform for CNS axonal injury, regeneration and transport. Nat. Methods 2, 599–605 (2005).
Acknowledgements
The authors thank M. Lecce and L. D. Huyer for helping with the editing of this manuscript. This work was made possible by the National Sciences and Engineering Research Council of Canada (NSERC) Postgraduate Scholarships-Doctoral Program awarded to B.F.L.L., Alexander Graham Bell Canada Graduate Scholarships-Doctoral Program awarded to A.K. and the Canadian Institutes of Health Research (CIHR) Banting Postdoctoral Fellowship to B.Z. This work was also funded by the CIHR Operating Grants (MOP-126027 and MOP-137107), NSERC Discovery Grant (RGPIN-2015-05952), NSERC Steacie Fellowship (SMFSU 4620), Heart and Stroke Foundation Grant-in-Aid (G-16-00012), NSERC-CIHR Collaborative Health Research Grant (CHRPJ 4937) and NSERC Strategic Grant (STPGP 5066) to M.R.
Author information
Authors and Affiliations
Contributions
B.Z., A.K. and B.F.L.L. wrote and edited the manuscript. M.R. supervised the work and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.Z. and M.R. hold equity in TARA Biosystems Inc.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
American Institute for Medical and Biological Engineering (AIMBE): http://aimbe.org
Comprehensive In Vitro Proarrhythmia Assay (CiPA): http://cipaproject.org
IQ Consortium: https://iqconsortium.org
Rights and permissions
About this article
Cite this article
Zhang, B., Korolj, A., Lai, B.F.L. et al. Advances in organ-on-a-chip engineering. Nat Rev Mater 3, 257–278 (2018). https://doi.org/10.1038/s41578-018-0034-7
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41578-018-0034-7
This article is cited by
-
Microfluidic high-throughput 3D cell culture
Nature Reviews Bioengineering (2024)
-
Vascularised cardiac spheroids-on-a-chip for testing the toxicity of therapeutics
Scientific Reports (2024)
-
Integration of multiple flexible electrodes for real-time detection of barrier formation with spatial resolution in a gut-on-chip system
Microsystems & Nanoengineering (2024)
-
Passive-Flow-Based MPS: Emerging Physiological Flow-Mimetic Platforms for Studying Effects of Flow on Single Tissues and Inter-tissue Interactions
BioChip Journal (2024)
-
Modular microfluidics for life sciences
Journal of Nanobiotechnology (2023)