Abstract
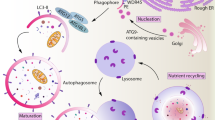
Across all branches of the immune system, the process of autophagy is fundamentally important in cellular development, function and homeostasis. Strikingly, this evolutionarily ancient pathway for intracellular recycling has been adapted to enable a high degree of functional complexity and specialization. However, although the requirement for autophagy in normal immune cell function is clear, the mechanisms involved are much less so and encompass control of metabolism, selective degradation of substrates and organelles and participation in cell survival decisions. We review here the crucial functions of autophagy in controlling the differentiation and homeostasis of multiple immune cell types and discuss the potential mechanisms involved.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dikic, I. & Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell. Biol. 19, 349–364 (2018).
Loukil, A. et al. High-resolution live-cell imaging reveals novel cyclin A2 degradation foci involving autophagy. J. Cell Sci. 127, 2145–2150 (2014).
Riffelmacher, T., Richter, F. C. & Simon, A. K. Autophagy dictates metabolism and differentiation of inflammatory immune cells. Autophagy 14, 199–206 (2018).
Puleston, D. J. et al. Autophagy is a critical regulator of memory CD8+ T cell formation. eLife 3, 2516–2521 (2014).
Xu, X. et al. Autophagy is essential for effector CD8+ T cell survival and memory formation. Nat. Immunol. 15, 1152–1161 (2014). References 4 and 5 show that T cell memory depends on autophagy and can be improved by the induction of autophagy, which suggests the possibility of improving vaccine responses in this manner.
Chen, M. et al. Essential role for autophagy in the maintenance of immunological memory against influenza infection. Nat. Med. 20, 503–510 (2014). This study shows the importance of autophagy in B cell memory, which opens up the possibility of new therapeutic approaches.
Wei, J. et al. Autophagy enforces functional integrity of regulatory T cells by coupling environmental cues and metabolic homeostasis. Nat. Immunol. 17, 277–285 (2016).
Riffelmacher, T. et al. Autophagy-dependent generation of free fatty acids is critical for normal neutrophil differentiation. Immunity 47, 466–480 (2017). This study is a key step towards understanding what autophagy provides for immune cell differentiation, rather than what it degrades.
Clarke, A. J., Riffelmacher, T., Braas, D., Cornall, R. J. & Simon, A. K. B1a B cells require autophagy for metabolic homeostasis and self-renewal. J. Exp. Med. 215, 399–413 (2018). This study examines why the B1 B cell subset is specifically dependent on autophagy owing to its microenvironment and metabolism.
Mortensen, M. et al. Loss of autophagy in erythroid cells leads to defective removal of mitochondria and severe anemia in vivo. Proc. Natl Acad. Sci. USA 107, 832–837 (2010).
Sandoval, H. et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature 454, 232–235 (2008).
Liu, F. et al. FIP200 is required for the cell-autonomous maintenance of fetal hematopoietic stem cells. Blood 116, 4806–4814 (2010).
Gómez-Puerto, M. C. et al. Activation of autophagy by FOXO3 regulates redox homeostasis during osteogenic differentiation. Autophagy 12, 1804–1816 (2017).
Liu, F. et al. Suppression of autophagy by FIP200 deletion leads to osteopenia in mice through the inhibition of osteoblast terminal differentiation. J. Bone Miner. Res. 28, 2414–2430 (2013).
Zhang, Y. et al. Adipose-specific deletion of autophagy-related gene 7 (atg7) in mice reveals a role in adipogenesis. Proc. Natl Acad. Sci. USA 106, 19860–19865 (2009).
Deretic, V. & Levine, B. Autophagy balances inflammation in innate immunity. Autophagy 14, 243–251 (2018).
Gomes, L. C. & Dikic, I. Autophagy in antimicrobial immunity. Mol. Cell 54, 224–233 (2014).
Münz, C. Autophagy beyond intracellular MHC class II antigen presentation. Trends Immunol. 37, 755–763 (2016).
Kraft, C., Peter, M. & Hofmann, K. Selective autophagy: ubiquitin-mediated recognition and beyond. Nat. Cell Biol. 12, 836–841 (2010).
Haynes, C. M., Fiorese, C. J. & Lin, Y.-F. Evaluating and responding to mitochondrial dysfunction: the mitochondrial unfolded-protein response and beyond. Trends Cell Biol. 23, 311–318 (2013).
Schweers, R. L. et al. NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc. Natl Acad. Sci. USA 104, 19500–19505 (2007).
Rawi, Al,S. et al. Postfertilization autophagy of sperm organelles prevents paternal mitochondrial DNA transmission. Science 334, 1144–1147 (2011).
Harper, J. W., Ordureau, A. & Heo, J.-M. Building and decoding ubiquitin chains for mitophagy. Nat. Rev. Mol. Cell Biol. 19, 93–108 (2018).
Valente, E. M. et al. Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 304, 1158–1160 (2004).
Kitada, T. et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392, 605–608 (1998).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009).
Murera, D. et al. CD4 T cell autophagy is integral to memory maintenance. Sci. Rep. 8, 5951 (2018).
Ouimet, M. et al. Autophagy regulates cholesterol efflux from macrophage foam cells via lysosomal acid lipase. Cell. Metab. 13, 655–667 (2011).
Walther, T. C. & Farese, R. V. Jr. Lipid droplets and cellular lipid metabolism. Annu. Rev. Biochem. 81, 687–714 (2012).
Zechner, R., Madeo, F. & Kratky, D. Cytosolic lipolysis and lipophagy: two sides of the same coin. Nat. Rev. Mol. Cell. Biol. 18, 671–684 (2017).
Khaminets, A. et al. Regulation of endoplasmic reticulum turnover by selective autophagy. Nature 522, 354–358 (2015).
Dou, Z. et al. Autophagy mediates degradation of nuclear lamina. Nature 527, 105–109 (2015).
Wyant, G. A. et al. NUFIP1 is a ribosome receptor for starvation-induced ribophagy. Science 360, 751–758 (2018).
Ktistakis, N. T. & Tooze, S. A. Digesting the expanding mechanisms of autophagy. Trends Cell Biol. 26, 624–635 (2016).
Galluzzi, L. et al. Molecular definitions of autophagy and related processes. EMBO J. 36, 1811–1836 (2017).
Martinez, J. et al. Molecular characterization of LC3-associated phagocytosis reveals distinct roles for Rubicon, NOX2 and autophagy proteins. Nat. Cell Biol. 17, 893–906 (2015).
Sanjuan, M. A. et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 450, 1253–1257 (2007).
Martinez-Martin, N. et al. A switch from canonical to noncanonical autophagy shapes B cell responses. Science 355, 641–647 (2017).
Afzal, S. et al. Autophagy-independent functions of UVRAG are essential for peripheral naive T cell homeostasis. Proc. Natl Acad. Sci. USA 112, 1119–1124 (2015).
Botbol, Y., Patel, B. & Macian, F. Common γ-chain cytokine signaling is required for macroautophagy induction during CD4+ T cell activation. Autophagy 11, 1864–1877 (2015).
Watanabe, K., Ichinose, S., Hayashizaki, K. & Tsubata, T. Induction of autophagy by B cell antigen receptor stimulation and its inhibition by costimulation. Biochem. Biophys. Res. Commun. 374, 274–281 (2008).
Andrade, R. M., Wessendarp, M., Gubbels, M.-J., Striepen, B. & Subauste, C. S. CD40 induces macrophage anti-Toxoplasma gondii activity by triggering autophagy-dependent fusion of pathogen-containing vacuoles and lysosomes. J. Clin. Invest. 116, 2366–2377 (2006).
Gutierrez, M. G. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Matsuzawa, T. et al. IFN-γ elicits macrophage autophagy via the p38 MAPK signaling pathway. J. Immunol. 189, 813–818 (2012).
Cooney, R. et al. NOD2 stimulation induces autophagy in dendritic cells influencing bacterial handling and antigen presentation. Nat. Med. 16, 90–97 (2010).
Delgado, M. A., Elmaoued, R. A., Davis, A. S., Kyei, G. & Deretic, V. Toll-like receptors control autophagy. EMBO J. 27, 1110–1121 (2008).
Kim, J., Kundu, M., Viollet, B. & Guan, K.-L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Lee, J. M. et al. Nutrient-sensing nuclear receptors coordinate autophagy. Nature 516, 112–115 (2014).
Seok, S. et al. Transcriptional regulation of autophagy by an FXR–CREB axis. Nature 516, 108–111 (2014).
Mammucari, C. et al. FoxO3 controls autophagy in skeletal muscle in vivo. Cell. Metab. 6, 458–471 (2007).
Lin, S. Y. et al. GSK3-TIP60-ULK1 signaling pathway links growth factor deprivation to autophagy. Science 336, 477–481 (2012).
Pattingre, S. et al. Bcl-2 antiapoptotic proteins inhibit beclin 1-dependent autophagy. Cell 122, 927–939 (2005).
Maiuri, M. C. et al. Functional and physical interaction between Bcl-XL and a BH3-like domain in Beclin-1. EMBO J. 26, 2527–2539 (2007).
Nazio, F. et al. mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat. Cell Biol. 15, 1–13 (2013).
Suda, T., Takubo, K. & Semenza, G. L. Metabolic regulation of hematopoietic stem cells in the hypoxic niche. Stem Cell 9, 298–310 (2011).
Mouttie, L. L.-E. et al. Autophagy is required for stem cell mobilization by G-CSF. Blood 125, 2933–2936 (2015).
Rožman, S. et al. The generation of neutrophils in the bone marrow is controlled by autophagy. Cell Death Differ. 22, 445–456 (2015).
Zhang, Y., Morgan, M. J., Chen, K., Choksi, S. & Liu, Z. G. Induction of autophagy is essential for monocyte-macrophage differentiation. Blood 119, 2895–2905 (2012).
Jacquel, A. et al. Autophagy is required for CSF-1-induced macrophagic differentiation and acquisition of phagocytic functions. Blood 119, 4527–4531 (2012).
Obba, S. et al. The PRKAA1/AMPKα1 pathway triggers autophagy during CSF1-induced human monocyte differentiation and is a potential target in CMML. Autophagy 11, 1114–1129 (2015).
Huang, S. C.-C. et al. Cell-intrinsic lysosomal lipolysis is essential for alternative activation of macrophages. Nat. Immunol. 15, 846–855 (2014).
Stranks, A. J. et al. Autophagy controls acquisition of aging features in macrophages. J. Innate Immun. 7, 375–391 (2015).
Martinez, J. et al. Noncanonical autophagy inhibits the autoinflammatory, lupus-like response to dying cells. Nature 533, 115–119 (2016). This study provides a link between homeostatic autophagy as a means of degrading engulfed cells and the autoimmune disease SLE.
Cunha, L. D. et al. LC3-associated phagocytosis in myeloid cells promotes tumor immune tolerance. Cell 175, 1–13 (2018).
Sieweke, M. H. & Allen, J. E. Beyond stem cells: self-renewal of differentiated macrophages. Science 342, 1242974–1242974 (2013).
Plaza-Zabala, A., Sierra-Torre, V. & Sierra, A. Autophagy and microglia: novel partners in neurodegeneration and aging. Int. J. Mol. Sci. 18, 598–528 (2017).
Ribeiro, C. M. S. et al. Receptor usage dictates HIV-1 restriction by human TRIM5a in dendritic cell subsets. Nature 540, 448–452 (2016).
Hubbard-Lucey, V. M. et al. Autophagy gene Atg16l1 prevents lethal T cell alloreactivity mediated by dendritic cells. Immunity 41, 579–591 (2014).
Weindel, C. G., Richey, L. J., Mehta, A. J., Shah, M. & Huber, B. T. Autophagy in dendritic cells and B cells is critical for the inflammatory state of TLR7-mediated autoimmunity. J. Immunol. 198, 1081–1092 (2017).
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015).
O’Sullivan, T. E., Johnson, L. R., Kang, H. H. & Sun, J. C. BNIP3- and BNIP3L-mediated mitophagy promotes the generation of natural killer cell memory. Immunity 43, 331–342 (2015).
O’Sullivan, T. E. et al. Atg5 is essential for the development and survival of innate lymphocytes. Cell Rep. 15, 1910–1919 (2016). This study elucidates the type of selective autophagy (mitophagy) that is involved in immune cell differentiation using genetic deletion of the cargo receptor BNIP3 and its ligand.
Starr, T. K., Jameson, S. C. & Hogquist, K. A. Positive and negative selection of T cells. Ann. Rev. Immunol. 21, 139–176 (2003).
Schmid, D., Pypaert, M. & Münz, C. Antigen-loading compartments for major histocompatibility complex class II molecules continuously receive input from autophagosomes. Immunity 26, 79–92 (2007). This work shows that autophagosomes provide a route for cross-presentation of endogenous antigens on MHC class II molecules.
Dengjel, J. et al. Autophagy promotes MHC class II presentation of peptides from intracellular source proteins. Proc. Natl Acad. Sci. USA 102, 7922–7927 (2005).
Aichinger, M., Wu, C., Nedjic, J. & Klein, L. Macroautophagy substrates are loaded onto MHC class II of medullary thymic epithelial cells for central tolerance. J. Exp. Med. 210, 287–300 (2013).
Nedjic, J., Aichinger, M., Emmerich, J., Mizushima, N. & Klein, L. Autophagy in thymic epithelium shapes the T cell repertoire and is essential for tolerance. Nature 455, 396–400 (2008).
Paludan, C. et al. Endogenous MHC class II processing of a viral nuclear antigen after autophagy. Science 307, 593–596 (2005).
Sukseree, S. et al. Autophagy in the thymic epithelium is dispensable for the development of self-tolerance in a novel mouse model. PLOS ONE 7, e38933 (2012).
Gateva, V. et al. A large-scale replication study identifies TNIP1, PRDM1, JAZF1, UHRF1BP1 and IL10 as risk loci for systemic lupus erythematosus. Nat. Genet. 41, 1228–1233 (2009).
Todd, J. A. et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat. Genet. 39, 857–864 (2007).
The International Multiple Sclerosis Genetics Consortium & The Wellcome Trust Case Control Consortium 2. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011).
Schuster, C. et al. The autoimmunity-associated gene CLEC16A modulates thymic epithelial cell autophagy and alters T cell selection. Immunity 42, 942–952 (2015).
Stephenson, L. M. et al. Identification of Atg5-dependent transcriptional changes and increases in mitochondrial mass in Atg5-deficient T lymphocytes. Autophagy 5, 625–635 (2009).
Arsov, I. et al. A role for autophagic protein beclin 1 early in lymphocyte development. J. Immunol. 186, 2201–2209 (2011).
Pua, H. H., Dzhagalov, I., Chuck, M., Mizushima, N. & He, Y.-W. A critical role for the autophagy gene Atg5 in T cell survival and proliferation. J. Exp. Med. 204, 25–31 (2007).
Miller, B. C. et al. The autophagy gene ATG5 plays an essential role in B lymphocyte development. Autophagy 4, 309–314 (2008).
Clarke, A. J. et al. Autophagy is activated in systemic lupus erythematosus and required for plasmablast development. Ann. Rheum. Dis. 74, 912–920 (2015).
Pua, H. H., Guo, J., Komatsu, M. & He, Y.-W. Autophagy is essential for mitochondrial clearance in mature T lymphocytes. J. Immunol. 182, 4046–4055 (2009).
Puleston, D. J. & Simon, A. K. Autophagy in the immune system. Immunology 141, 1–8 (2013).
Godfrey, D. I., Stankovic, S. & Baxter, A. G. Raising the NKT cell family. Nat. Immunol. 11, 197–206 (2010).
Salio, M. et al. Essential role for autophagy during invariant NKT cell development. Proc. Natl Acad. Sci. USA 111, E5678–E5687 (2014).
Pei, B. et al. Invariant NKT cells require autophagy to coordinate proliferation and survival signals during differentiation. J. Immunol. 194, 5872–5884 (2015).
Kovacs, J. R. et al. Autophagy promotes T cell survival through degradation of proteins of the cell death machinery. Cell Death Differ. 19, 144–152 (2012).
Vargas, T. R. et al. Selective degradation of PU.1 during autophagy represses the differentiation and antitumour activity of TH9 cells. Nat. Commun. 2017, 1–15 (2017).
Kabat, A. M. et al. The autophagy gene Atg16l1 differentially regulates Treg and TH2 cells to control intestinal inflammation. eLife 5, e12444 (2016).
Jia, W., Pua, H. H., Li, Q. J. & He, Y.-W. Autophagy regulates endoplasmic reticulum homeostasis and calcium mobilization in T lymphocytes. J. Immunol. 186, 1564–1574 (2011).
Parekh, V. V. et al. Impaired autophagy, defective T cell homeostasis, and a wasting syndrome in mice with a T cell-specific deletion of Vps34. J. Immunol. 190, 5086–5101 (2013).
McLeod, I. X., Zhou, X., Li, Q. J., Wang, F. & He, Y.-W. The class III kinase Vps34 promotes T lymphocyte survival through regulating IL-7R surface expression. J. Immunol. 187, 5051–5061 (2011).
Goldrath, A. W., Bogatzki, L. Y. & Bevan, M. J. Naive T cells transiently acquire a memory-like phenotype during homeostasis-driven proliferation. J. Exp. Med. 192, 557–564 (2000).
Jia, W. & He, Y.-W. Temporal regulation of intracellular organelle homeostasis in T lymphocytes by autophagy. J. Immunol. 186, 5313–5322 (2011).
Le Texier, L. et al. Autophagy-dependent regulatory T cells are critical for the control of graft-versus-host disease. JCI Insight 1, e86850 (2016).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Shi, L. Z. et al. HIF1α-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
Schlie, K. et al. Survival of effector CD8+ T cells during influenza infection is dependent on autophagy. J. Immunol. 194, 4277–4286 (2015).
Jia, W. et al. Autophagy regulates T lymphocyte proliferation through selective degradation of the cell-cycle inhibitor CDKN1B/p27Kip1. Autophagy 11, 2335–2345 (2015).
Pearce, E. L. et al. Enhancing CD8 T cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Arsov, I. et al. BAC-mediated transgenic expression of fluorescent autophagic protein Beclin 1 reveals a role for Beclin 1 in lymphocyte development. Cell Death Differ. 15, 1385–1395 (2008).
Arnold, J. et al. Autophagy is dispensable for B cell development but essential for humoral autoimmune responses. Cell Death Differ. 23, 853–864 (2016).
Baumgarth, N. The double life of a B-1 cell: self-reactivity selects for protective effector functions. Nat. Rev. Immunol. 11, 34–46 (2010).
Chen, M., Kodali, S., Jang, A., Kuai, L. & Wang, J. Requirement for autophagy in the long-term persistence but not initial formation of memory B cells. J. Immunol. 194, 2607–2615 (2015).
Raso, F. et al. αv integrins regulate germinal center B cell responses through noncanonical autophagy. J. Clin. Invest. 128, 4163–4178 (2018).
Acharya, M. et al. αv integrins combine with LC3 and atg5 to regulate Toll-like receptor signalling in B cells. Nat. Commun. 7, 10917 (2016).
Pengo, N. et al. Plasma cells require autophagy for sustainable immunoglobulin production. Nat. Immunol. 14, 298–305 (2013). This is an important study that shows the role of autophagy in plasma cell homeostasis as a system that complements the unfolded protein response.
Weindel, C. G. et al. B cell autophagy mediates TLR7-dependent autoimmunity and inflammation. Autophagy 11, 1010–1024 (2015).
Gros, F. et al. Macroautophagy is deregulated in murine and human lupus T lymphocytes. Autophagy 8, 1054–1053 (2012).
van Loosdregt, J. et al. Hydroxychloroquine preferentially induces apoptosis of CD45RO+ effector T cells by inhibiting autophagy: a possible mechanism for therapeutic modulation of T cells. J. Allergy Clin. Immunol. 131, 1443–1446 (2013).
Zhang, H., Puleston, D. J. & Simon, A. K. Autophagy and immune senescence. Trends Mol. Med. 22, 671–686 (2016).
Lamb, C. A., Yoshimori, T. & Tooze, S. A. The autophagosome: origins unknown, biogenesis complex. Nat. Rev. Mol. Cell. Biol. 14, 759–774 (2013).
Karanasios, E. et al. Autophagy initiation by ULK complex assembly on ER tubulovesicular regions marked by ATG9 vesicles. Nat. Commun. 7, 12420 (2016).
Ge, L., Zhang, M. & Schekman, R. Phosphatidylinositol 3-kinase and COPII generate LC3 lipidation vesicles from the ER-Golgi intermediate compartment. eL ife 3, 839–813 (2014).
Young, A. R. J. Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J. Cell Sci. 119, 3888–3900 (2006).
Yu, L., Chen, Y. & Tooze, S. A. Autophagy pathway: cellular and molecular mechanisms. Autophagy 14, 207–215 (2018).
Mizushima, N. et al. A protein conjugation system essential for autophagy. Nature 395, 395–398 (1998).
Mizushima, N. et al. Mouse Apg16L, a novel WD-repeat protein, targets to the autophagic isolation membrane with the Apg12-Apg5 conjugate. J. Cell Sci. 116, 1679–1688 (2003).
Hanada, T. et al. The Atg12-Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J. Biol. Chem. 282, 37298–37302 (2007).
Nakatogawa, H., Ichimura, Y. & Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130, 165–178 (2007).
Settembre, C. et al. TFEB links autophagy to lysosomal biogenesis. Science 332, 1429–1433 (2011).
Riffelmacher, T. & Simon, A. K. Mechanistic roles of autophagy in hematopoietic differentiation. FEBS J. 284, 1008–1020 (2016).
Warr, M. R. et al. FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494, 323–327 (2013).
Watson, A. S. et al. Autophagy limits proliferation and glycolytic metabolism in acute myeloid leukemia. Cell Death Discov. 1, 15008–15010 (2015).
Mortensen, M. et al. The autophagy protein Atg7 is essential for hematopoietic stem cell maintenance. J. Exp. Med. 208, 455–467 (2011).
Cao, Y. et al. Hierarchal autophagic divergence of hematopoietic system. J. Biol. Chem. 290, 23050–23063 (2015).
Ito, K. et al. Self-renewal of a purified Tie2+ hematopoietic stem cell population relies on mitochondrial clearance. Science 354, 1156–1160 (2016).
Acknowledgements
A.J.C. and A.K.S. are both funded by the Wellcome Trust (104549/Z/14/Z to A.J.C. and 103830/Z/14/Z to A.K.S.).
Reviewer information
Nature Reviews Immunology thanks F. Gros, J. Martinez and other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Both authors wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Oxidative phosphorylation
-
(OXPHOS). The production of ATP through the oxidation of nutrients. The electron transport chain in mitochondria produces an electrochemical gradient that is used to make ATP.
- Tricarboxylic acid cycle
-
(TCA cycle). A metabolic pathway that oxidizes acetyl-CoA derived from pyruvate to release stored energy in the form of NADH, which then enters oxidative phosphorylation. The TCA cycle is crucial for carbohydrate, protein and lipid metabolism.
- Autophagy genes
-
Genes related to autophagy. The core machinery of autophagy is encoded by ~30 genes. The most commonly deleted genes in experimental settings include Ulk1, Ulk2, Atg3, Atg5, Becn1, Atg7 and Atg16l1, which are all essential for autophagy.
- β-Oxidation
-
The catabolism of fatty acid molecules to produce acetyl-CoA.
- LC3-associated phagocytosis
-
(LAP). A form of non-canonical autophagy in which LC3 is conjugated to the phagosomal membrane using some components of the autophagy pathway.
- Unc51-like autophagy-activating kinase 1 complex
-
(ULK1 complex). The major complex controlling the initiation of autophagy, which is targeted by mechanistic target of rapamycin complex 1 (mTORC1) and 5′-AMP-activated protein kinase (AMPK).
- Mechanistic target of rapamycin complex 1
-
(mTORC1). A complex — which consists of mTOR together with the protein raptor — that senses nutrient status and is a master regulator of protein synthesis and cell growth.
- M1 macrophages and M2 macrophages
-
‘M1’ and ‘M2’ are classifications historically used to define macrophages activated in vitro as pro-inflammatory (when ‘classically’ activated with IFNγ and lipopolysaccharide) or anti-inflammatory (when ‘alternatively’ activated with IL-4 or IL-10), respectively. However, in vivo macrophages are highly specialized, transcriptomically dynamic and extremely heterogeneous with regard to their phenotypes and functions, which are continuously shaped by their tissue microenvironment. Therefore, the M1 or M2 classification is too simplistic to explain the true nature of in vivo macrophages, although these terms are still often used to indicate whether the macrophages in question are more pro- or anti-inflammatory.
- Senescence
-
A process that typically occurs in aged cells. It involves the acquisition of progressive and diverse cellular phenotypes including growth arrest, telomere attrition, damaged macromolecules and the secretion of cytokines, chemokines and proteases with pro-inflammatory properties (the senescence-associated secretory phenotype), which together lead to tissue dysfunction.
- Non-classical cross-presentation
-
The presentation of endogenous proteins, which enter the endosomal pathway, on MHC class II molecules by antigen-presenting cells.
- Antibody affinity maturation
-
The progressive increase in antibody affinity that occurs during the immune response as the germinal centre reaction selects for B cells producing higher-affinity immunoglobulin.
- Unfolded protein response
-
The stress response pathway that is activated by the accumulation of unfolded or defective proteins in the endoplasmic reticulum.
Rights and permissions
About this article
Cite this article
Clarke, A.J., Simon, A.K. Autophagy in the renewal, differentiation and homeostasis of immune cells. Nat Rev Immunol 19, 170–183 (2019). https://doi.org/10.1038/s41577-018-0095-2
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41577-018-0095-2
This article is cited by
-
The application of nanoparticles-based ferroptosis, pyroptosis and autophagy in cancer immunotherapy
Journal of Nanobiotechnology (2024)
-
Cancer cell metabolism and antitumour immunity
Nature Reviews Immunology (2024)
-
The aging mouse CNS is protected by an autophagy-dependent microglia population promoted by IL-34
Nature Communications (2024)
-
The role of H3K27me3 methylation in cancer development
Genome Instability & Disease (2024)
-
Targeting and regulation of autophagy in hepatocellular carcinoma: revisiting the molecular interactions and mechanisms for new therapy approaches
Cell Communication and Signaling (2023)