Abstract
Primate genomics holds the key to understanding fundamental aspects of human evolution and disease. However, genetic diversity and functional genomics data sets are currently available for only a few of the more than 500 extant primate species. Concerted efforts are under way to characterize primate genomes, genetic polymorphism and divergence, and functional landscapes across the primate phylogeny. The resulting data sets will enable the connection of genotypes to phenotypes and provide new insight into aspects of the genetics of primate traits, including human diseases. In this Review, we describe the existing genome assemblies as well as genetic variation and functional genomic data sets. We highlight some of the challenges with sample acquisition. Finally, we explore how technological advances in single-cell functional genomics and induced pluripotent stem cell-derived organoids will facilitate our understanding of the molecular foundations of primate biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rogers, J. & Gibbs, R. A. Comparative primate genomics: emerging patterns of genome content and dynamics. Nat. Rev. Genet. 15, 347–359 (2014).
Kuderna, L. F., Esteller-Cucala, P. & Marques-Bonet, T. Branching out: what omics can tell us about primate evolution. Curr. Opin. Genet. Dev. 62, 65–71 (2020).
Johnson, M. E. et al. Positive selection of a gene family during the emergence of humans and African apes. Nature 413, 514–519 (2001).
Enard, W. et al. Intra- and interspecific variation in primate gene expression patterns. Science 296, 340–343 (2002).
Estrada, A. et al. Impending extinction crisis of the world’s primates: why primates matter. Sci. Adv. 3, e1600946 (2017).
Zoonomia Consortium. A comparative genomics multitool for scientific discovery and conservation. Nature 587, 240–245 (2020).
Church, D. M. et al. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 7, e1000112 (2009).
Rhie, A. et al. Towards complete and error-free genome assemblies of all vertebrate species. Nature 592, 737–746 (2021).
Meadows, J. R. S. & Lindblad-Toh, K. Dissecting evolution and disease using comparative vertebrate genomics. Nat. Rev. Genet. 18, 624–636 (2017).
Alkan, C., Coe, B. P. & Eichler, E. E. Genome structural variation discovery and genotyping. Nat. Rev. Genet. 12, 363–376 (2011).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Church, D. M. et al. Modernizing reference genome assemblies. PLoS Biol. 9, e1001091 (2011).
Schneider, V. A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Nurk, S. et al. The complete sequence of a human genome. Science 376, 44–53 (2022).
Miga, K. H. et al. Telomere-to-telomere assembly of a complete human X chromosome. Nature 585, 79–84 (2020).
Logsdon, G. A. et al. The structure, function and evolution of a complete human chromosome 8. Nature 593, 101–107 (2021).
Jarvis, E. D. et al. Semi-automated assembly of high-quality diploid human reference genomes. Nature https://doi.org/10.1038/s41586-022-05325-5 (2022).
Kronenberg, Z. N. et al. High-resolution comparative analysis of great ape genomes. Science 360, eaar6343 (2018). This study generated de novo high-quality long-read sequence assemblies of chimpanzee and orangutan genomes to recover human-specific structural variants.
He, Y. et al. Long-read assembly of the Chinese rhesus macaque genome and identification of ape-specific structural variants. Nat. Commun. 10, 4233 (2019).
Mao, Y. et al. A high-quality bonobo genome refines the analysis of hominid evolution. Nature 594, 77–81 (2021).
Dong, D., He, G., Zhang, S. & Zhang, Z. Evolution of olfactory receptor genes in primates dominated by birth-and-death process. Genome Biol. Evol. 1, 258–264 (2009).
Kazen, A. R. & Adams, E. J. Evolution of the V, D, and J gene segments used in the primate γδ T-cell receptor reveals a dichotomy of conservation and diversity. Proc. Natl Acad. Sci. USA 108, E332–E340 (2011).
Bruijnesteijn, J., de Groot, N. G. & Bontrop, R. E. The genetic mechanisms driving diversification of the KIR gene cluster in primates. Front. Immunol. 11, 582804 (2020).
Housman, G. & Gilad, Y. Prime time for primate functional genomics. Curr. Opin. Genet. Dev. 62, 1–7 (2020).
Armstrong, J., Fiddes, I. T., Diekhans, M. & Paten, B. Whole-genome alignment and comparative annotation. Annu. Rev. Anim. Biosci. 7, 41–64 (2019).
Benjamin, A. M., Nichols, M., Burke, T. W., Ginsburg, G. S. & Lucas, J. E. Comparing reference-based RNA-Seq mapping methods for non-human primate data. BMC Genomics 15, 570 (2014).
Zhu, Y., Li, M., Sousa, A. M. M. & Sestan, N. XSAnno: a framework for building ortholog models in cross-species transcriptome comparisons. BMC Genomics 15, 343 (2014).
Prado-Martinez, J. et al. Great ape genetic diversity and population history. Nature 499, 471–475 (2013). This work produced high-coverage whole-genome sequencing data for 89 individuals of six NHPs and seven subspecies, recovering an extensive catalogue of genetic polymorphisms and studying the recent evolution of great ape populations.
de Manuel, M. et al. Chimpanzee genomic diversity reveals ancient admixture with bonobos. Science 354, 477–481 (2016).
Xue, Y. et al. Mountain gorilla genomes reveal the impact of long-term population decline and inbreeding. Science 348, 242–245 (2015).
Nater, A. et al. Morphometric, behavioral, and genomic evidence for a new orangutan species. Curr. Biol. 27, 3487–3498.e10 (2017).
Rogers, J. et al. The comparative genomics and complex population history of Papio baboons. Sci. Adv. 5, eaau6947 (2019).
Warren, W. C. et al. Sequence diversity analyses of an improved rhesus macaque genome enhance its biomedical utility. Science 370, eabc6617 (2020).
Chiou, K. L. et al. Genomic signatures of high-altitude adaptation and chromosomal polymorphism in geladas. Nat. Ecol. Evol. 6, 630–643 (2022).
Kuhlwilm, M. et al. Evolution and demography of the great apes. Curr. Opin. Genet. Dev. 41, 124–129 (2016).
Svardal, H. et al. Ancient hybridization and strong adaptation to viruses across African vervet monkey populations. Nat. Genet. 49, 1705–1713 (2017).
Zhou, X. et al. Whole-genome sequencing of the snub-nosed monkey provides insights into folivory and evolutionary history. Nat. Genet. 46, 1303–1310 (2014).
Yu, L. et al. Genomic analysis of snub-nosed monkeys (Rhinopithecus) identifies genes and processes related to high-altitude adaptation. Nat. Genet. 48, 947–952 (2016).
van der Valk, T. et al. The genome of the endangered dryas monkey provides new insights into the evolutionary history of the vervets. Mol. Biol. Evol. 37, 183–194 (2020).
Skoglund, P. & Mathieson, I. Ancient genomics of modern humans: the first decade. Annu. Rev. Genomics Hum. Genet. 19, 381–404 (2018).
Gurdasani, D., Barroso, I., Zeggini, E. & Sandhu, M. S. Genomics of disease risk in globally diverse populations. Nat. Rev. Genet. 20, 520–535 (2019).
Anderson, J. A., Vilgalys, T. P. & Tung, J. Broadening primate genomics: new insights into the ecology and evolution of primate gene regulation. Curr. Opin. Genet. Dev. 62, 16–22 (2020).
Graham, S. E. et al. The power of genetic diversity in genome-wide association studies of lipids. Nature 600, 675–679 (2021).
Wojcik, G. L. et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature 570, 514–518 (2019).
Eilbeck, K., Quinlan, A. & Yandell, M. Settling the score: variant prioritization and Mendelian disease. Nat. Rev. Genet. 18, 599–612 (2017).
Kuhlwilm, M. & Boeckx, C. A catalog of single nucleotide changes distinguishing modern humans from archaic hominins. Sci. Rep. 9, 8463 (2019).
Sundaram, L. et al. Predicting the clinical impact of human mutation with deep neural networks. Nat. Genet. 50, 1161–1170 (2018). This study trained deep neural networks to predict pathogenic mutations in humans based on the common variants recovered from primate population genomic data.
Sudmant, P. H. et al. Evolution and diversity of copy number variation in the great ape lineage. Genome Res. 23, 1373–1382 (2013).
Spielmann, M., Lupiáñez, D. G. & Mundlos, S. Structural variation in the 3D genome. Nat. Rev. Genet. 19, 453–467 (2018).
Girirajan, S., Campbell, C. D. & Eichler, E. E. Human copy number variation and complex genetic disease. Annu. Rev. Genet. 45, 203–226 (2011).
Shaikh, T. H. Copy number variation disorders. Curr. Genet. Med. Rep. 5, 183–190 (2017).
Zhang, F. & Lupski, J. R. Non-coding genetic variants in human disease. Hum. Mol. Genet. 24, R102–R110 (2015).
Dennis, M. Y. & Eichler, E. E. Human adaptation and evolution by segmental duplication. Curr. Opin. Genet. Dev. 41, 44–52 (2016).
Hollox, E. J., Zuccherato, L. W. & Tucci, S. Genome structural variation in human evolution. Trends Genet. 38, 45–58 (2022).
Porubsky, D. et al. Recurrent inversion toggling and great ape genome evolution. Nat. Genet. 52, 849–858 (2020).
Axelsson, E. et al. The genomic signature of dog domestication reveals adaptation to a starch-rich diet. Nature 495, 360–364 (2013).
Serres-Armero, A. et al. Copy number variation underlies complex phenotypes in domestic dog breeds and other canids. Genome Res. 31, 762–774 (2021).
Rinker, D. C., Specian, N. K., Zhao, S. & Gibbons, J. G. Polar bear evolution is marked by rapid changes in gene copy number in response to dietary shift. Proc. Natl Acad. Sci. USA 116, 13446–13451 (2019).
Degenhardt, J. D. et al. Copy number variation of CCL3-like genes affects rate of progression to simian-AIDS in Rhesus Macaques (Macaca mulatta). PLoS Genet. 5, e1000346 (2009).
Guevara, E. E. et al. Comparative genomic analysis of sifakas (Propithecus) reveals selection for folivory and high heterozygosity despite endangered status. Sci. Adv. 7, eabd2274 (2021).
Ho, S. S., Urban, A. E. & Mills, R. E. Structural variation in the sequencing era. Nat. Rev. Genet. 21, 171–189 (2020).
Logsdon, G. A., Vollger, M. R. & Eichler, E. E. Long-read human genome sequencing and its applications. Nat. Rev. Genet. 21, 597–614 (2020).
De Coster, W., Weissensteiner, M. H. & Sedlazeck, F. J. Towards population-scale long-read sequencing. Nat. Rev. Genet. 22, 572–587 (2021).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Bernstein, B. E. et al. The NIH roadmap epigenomics mapping consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Mouse ENCODE Consortium et al. An encyclopedia of mouse DNA elements (Mouse ENCODE). Genome Biol. 13, 418 (2012).
Celniker, S. E. et al. Unlocking the secrets of the genome. Nature 459, 927–930 (2009).
Han, L. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature 604, 723–731 (2022).
Peng, X. et al. Tissue-specific transcriptome sequencing analysis expands the non-human primate reference transcriptome resource (NHPRTR). Nucleic Acids Res. 43, D737–D742 (2015). This resource contains the largest tissue-specific transcriptomic data set in primates to date.
Sato, K. et al. Resequencing of the common marmoset genome improves genome assemblies and gene-coding sequence analysis. Sci. Rep. 5, 16894 (2015).
Tan, T. et al. Improved Macaca fascicularis gene annotation reveals evolution of gene expression profiles in multiple tissues. BMC Genomics 19, 787 (2018).
Palesch, D. et al. Sooty mangabey genome sequence provides insight into AIDS resistance in a natural SIV host. Nature 553, 77–81 (2018).
Francescatto, M. et al. Transcription start site profiling of 15 anatomical regions of the Macaca mulatta central nervous system. Sci. Data 4, 170163 (2017).
Brawand, D. et al. The evolution of gene expression levels in mammalian organs. Nature 478, 343–348 (2011).
Cardoso-Moreira, M. et al. Gene expression across mammalian organ development. Nature 571, 505–509 (2019). This study produced and analyzed gene expression transcriptomic data along the development of different organs in six mammals, including human and rhesus macaque, recovering species-specific trajectories.
Villar, D. et al. Enhancer evolution across 20 mammalian species. Cell 160, 554–566 (2015).
García-Pérez, R. et al. Epigenomic profiling of primate lymphoblastoid cell lines reveals the evolutionary patterns of epigenetic activities in gene regulatory architectures. Nat. Commun. 12, 3116 (2021). This study performed extensive genomic, epigenomic and transcriptomic profiling of lymphoblastoid cell lines from human and four NHPs to provide the largest catalogue of gene regulatory architectures in primates to date and study the coordination of genetic and epigenetic evolution in primates.
Blake, L. E. et al. A comparison of gene expression and DNA methylation patterns across tissues and species. Genome Res. 30, 250–262 (2020).
Khan, Z. et al. Primate transcript and protein expression levels evolve under compensatory selection pressures. Science 342, 1100–1104 (2013).
Zhou, X. et al. Epigenetic modifications are associated with inter-species gene expression variation in primates. Genome Biol. 15, 547 (2014).
Reilly, S. K. et al. Evolutionary changes in promoter and enhancer activity during human corticogenesis. Science 347, 1155–1159 (2015).
Prescott, S. L. et al. Enhancer divergence and cis-regulatory evolution in the human and chimp neural crest. Cell 163, 68–83 (2015).
Hernando-Herraez, I. et al. The interplay between DNA methylation and sequence divergence in recent human evolution. Nucleic Acids Res. 43, 8204–8214 (2015).
Vermunt, M. W. et al. Epigenomic annotation of gene regulatory alterations during evolution of the primate brain. Nat. Neurosci. 19, 494–503 (2016).
Trizzino, M. et al. Transposable elements are the primary source of novelty in primate gene regulation. Genome Res. 27, 1623–1633 (2017).
Danko, C. G. et al. Dynamic evolution of regulatory element ensembles in primate CD4+ T cells. Nat. Ecol. Evol. 2, 537–548 (2018).
Li, M.-L. et al. 547 transcriptomes from 44 brain areas reveal features of the aging brain in non-human primates. Genome Biol. 20, 258 (2019).
Eres, I. E., Luo, K., Hsiao, C. J., Blake, L. E. & Gilad, Y. Reorganization of 3D genome structure may contribute to gene regulatory evolution in primates. PLoS Genet. 15, e1008278 (2019).
Gokhman, D. et al. Differential DNA methylation of vocal and facial anatomy genes in modern humans. Nat. Commun. 11, 1189 (2020).
Castelijns, B. et al. Recently evolved enhancers emerge with high interindividual variability and less frequently associate with disease. Cell Rep. 31, 107799 (2020).
Kozlenkov, A. et al. Evolution of regulatory signatures in primate cortical neurons at cell-type resolution. Proc. Natl Acad. Sci. USA 117, 28422–28432 (2020).
Jeong, H. et al. Evolution of DNA methylation in the human brain. Nat. Commun. 12, 2021 (2021).
Ferrandez-Peral, L. et al. Transcriptome innovations in primates revealed by single-molecule long-read sequencing. Genome Res. https://doi.org/10.1101/gr.276395.121 (2022).
Jasinska, A. J. et al. Genetic variation and gene expression across multiple tissues and developmental stages in a nonhuman primate. Nat. Genet. 49, 1714–1721 (2017). This study analyzes multi-tissue gene expression and genomic data in a vervet monkey pedigree to produce the first catalogue of eQTLs in an NHP, obtaining more significant eQTLs per sample than comparable human resources.
Tung, J., Zhou, X., Alberts, S. C., Stephens, M. & Gilad, Y. The genetic architecture of gene expression levels in wild baboons. eLife 4, e04729 (2015).
Fair, B. J. et al. Gene expression variability in human and chimpanzee populations share common determinants. eLife 9, e59929 (2020). This study compared inter-individual gene expression variability in human and chimpanzee primary heart samples and found that genetically controlled expression variability is conserved in humans and chimpanzees.
Umans, B. D., Battle, A. & Gilad, Y. Where are the disease-associated eQTLs? Trends Genet. 37, 109–124 (2021).
Hujoel, M. L. A., Gazal, S., Hormozdiari, F., van de Geijn, B. & Price, A. L. Disease heritability enrichment of regulatory elements is concentrated in elements with ancient sequence age and conserved function across species. Am. J. Hum. Genet. 104, 611–624 (2019).
Chan, E. T. et al. Conservation of core gene expression in vertebrate tissues. J. Biol. 8, 33 (2009).
Merkin, J., Russell, C., Chen, P. & Burge, C. B. Evolutionary dynamics of gene and isoform regulation in Mammalian tissues. Science 338, 1593–1599 (2012).
Berthelot, C., Villar, D., Horvath, J. E., Odom, D. T. & Flicek, P. Complexity and conservation of regulatory landscapes underlie evolutionary resilience of mammalian gene expression. Nat. Ecol. Evol. 2, 152–163 (2018).
Visel, A., Rubin, E. M. & Pennacchio, L. A. Genomic views of distant-acting enhancers. Nature 461, 199–205 (2009).
Mattioli, K. et al. Cis and trans effects differentially contribute to the evolution of promoters and enhancers. Genome Biol. 21, 210 (2020).
Wang, S. H., Hsiao, C. J., Khan, Z. & Pritchard, J. K. Post-translational buffering leads to convergent protein expression levels between primates. Genome Biol. 19, 83 (2018).
Wang, Z.-Y. et al. Transcriptome and translatome co-evolution in mammals. Nature 588, 642–647 (2020).
Prabhakar, S., Noonan, J. P., Pääbo, S. & Rubin, E. M. Accelerated evolution of conserved noncoding sequences in humans. Science 314, 786 (2006).
Doan, R. N. et al. Mutations in human accelerated regions disrupt cognition and social behavior. Cell 167, 341–354.e12 (2016).
Meyer, K. A., Marques-Bonet, T. & Sestan, N. Differential gene expression in the human brain is associated with conserved, but not accelerated, noncoding sequences. Mol. Biol. Evol. 34, 1217–1229 (2017).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Ryu, H. et al. Massively parallel dissection of human accelerated regions in human and chimpanzee neural progenitors. bioRxiv https://doi.org/10.1101/256313 (2018).
Uebbing, S. et al. Massively parallel discovery of human-specific substitutions that alter enhancer activity. Proc. Natl Acad. Sci. USA 118, e2007049118 (2021).
Johnson, M. B. et al. Functional and evolutionary insights into human brain development through global transcriptome analysis. Neuron 62, 494–509 (2009).
Somel, M. et al. Transcriptional neoteny in the human brain. Proc. Natl Acad. Sci. USA 106, 5743–5748 (2009).
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Liu, X. et al. Extension of cortical synaptic development distinguishes humans from chimpanzees and macaques. Genome Res. 22, 611–622 (2012).
Zeng, J. et al. Divergent whole-genome methylation maps of human and chimpanzee brains reveal epigenetic basis of human regulatory evolution. Am. J. Hum. Genet. 91, 455–465 (2012).
Shulha, H. P. et al. Human-specific histone methylation signatures at transcription start sites in prefrontal neurons. PLoS Biol. 10, e1001427 (2012).
Pletikos, M. et al. Temporal specification and bilaterality of human neocortical topographic gene expression. Neuron 81, 321–332 (2014).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, eaat8077 (2018). This paper provides a large-scale transcriptomic comparison between humans and rhesus macaques in multiple brain regions through prenatal and postnatal development.
Luo, X. et al. 3D Genome of macaque fetal brain reveals evolutionary innovations during primate corticogenesis. Cell 184, 723–740.e21 (2021).
Levchenko, A., Kanapin, A., Samsonova, A. & Gainetdinov, R. R. Human accelerated regions and other human-specific sequence variations in the context of evolution and their relevance for brain development. Genome Biol. Evol. 10, 166–188 (2018).
Girskis, K. M. et al. Rewiring of human neurodevelopmental gene regulatory programs by human accelerated regions. Neuron 109, 3239–3251.e7 (2021).
Pattabiraman, K., Muchnik, S. K. & Sestan, N. The evolution of the human brain and disease susceptibility. Curr. Opin. Genet. Dev. 65, 91–97 (2020).
Won, H., Huang, J., Opland, C. K., Hartl, C. L. & Geschwind, D. H. Human evolved regulatory elements modulate genes involved in cortical expansion and neurodevelopmental disease susceptibility. Nat. Commun. 10, 2396 (2019).
Phillips, K. A. et al. Why primate models matter. Am. J. Primatol. 76, 801–827 (2014).
Estes, J. D., Wong, S. W. & Brenchley, J. M. Nonhuman primate models of human viral infections. Nat. Rev. Immunol. 18, 390–404 (2018).
Ehrenberg, P. K. et al. A vaccine-induced gene expression signature correlates with protection against SIV and HIV in multiple trials. Sci. Transl. Med. 11, eaaw4236 (2019).
Yu, S. et al. Experimental treatment of SIV-infected macaques via autograft of CCR5-disrupted hematopoietic stem and progenitor cells. Mol. Ther. Methods Clin. Dev. 17, 520–531 (2020).
Ozirmak Lermi, N. et al. Comparative molecular genomic analyses of a spontaneous rhesus macaque model of mismatch repair-deficient colorectal cancer. PLoS Genet. 18, e1010163 (2022).
Sibal, L. R. & Samson, K. J. Nonhuman primates: a critical role in current disease research. ILAR J. 42, 74–84 (2001).
Yu, H. et al. Metabolism by conjugation appears to confer resistance to paracetamol (acetaminophen) hepatotoxicity in the cynomolgus monkey. Xenobiotica 45, 270–277 (2015).
Solis-Moruno, M. et al. Potential damaging mutation in LRP5 from genome sequencing of the first reported chimpanzee with the Chiari malformation. Sci. Rep. 7, 15224 (2017).
Fox, A. S. et al. Infant inhibited temperament in primates predicts adult behavior, is heritable, and is associated with anxiety-relevant genetic variation. Mol. Psychiatry 26, 6609–6618 (2021).
Huang, Y. S. et al. Sequencing strategies and characterization of 721 vervet monkey genomes for future genetic analyses of medically relevant traits. BMC Biol. 13, 41 (2015).
Bimber, B. N., Yan, M. Y., Peterson, S. M. & Ferguson, B. mGAP: the macaque genotype and phenotype resource, a framework for accessing and interpreting macaque variant data, and identifying new models of human disease. BMC Genomics 20, 176 (2019).
Aida, T. & Feng, G. The dawn of non-human primate models for neurodevelopmental disorders. Curr. Opin. Genet. Dev. 65, 160–168 (2020).
Feng, G. et al. Opportunities and limitations of genetically modified nonhuman primate models for neuroscience research. Proc. Natl Acad. Sci. USA 117, 24022–24031 (2020).
Chen, Y. et al. Modeling Rett syndrome using TALEN-Edited MECP2 mutant cynomolgus monkeys. Cell 169, 945–955.e10 (2017).
Yang, W. et al. CRISPR/Cas9-mediated PINK1 deletion leads to neurodegeneration in rhesus monkeys. Cell Res. 29, 334–336 (2019).
Vitale, A. & Borgi, M. In Evolution of Primate Social Cognition (eds Di Paolo, L. D., Di Vincenzo, F. & De Petrillo, F.) 143–161 (Springer International Publishing, 2018).
Aguilera, B., Perez Gomez, J. & DeGrazia, D. Should biomedical research with great apes be restricted? A systematic review of reasons. BMC Med. Ethics 22, 15 (2021).
Galán-Acedo, C., Arroyo-Rodríguez, V., Andresen, E. & Arasa-Gisbert, R. Ecological traits of the world’s primates. Sci. Data 6, 55 (2019).
Lüpold, S., Simmons, L. W. & Grueter, C. C. Sexual ornaments but not weapons trade off against testes size in primates. Proc. Biol. Sci. 286, 20182542 (2019).
Pitirri, M. K. & Richtsmeier, K. T. (eds) Evolutionary Cell Processes in Primates: Bone, Brains, and Muscle (Taylor & Francis, 2021).
Yuste, R. et al. A community-based transcriptomics classification and nomenclature of neocortical cell types. Nat. Neurosci. 23, 1456–1468 (2020).
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Cao, J. et al. A human cell atlas of fetal gene expression. Science 370, eaba7721 (2020).
Shafer, M. E. R. Cross-species analysis of single-cell transcriptomic data. Front. Cell Dev. Biol. 7, 175 (2019).
Lähnemann, D. et al. Eleven grand challenges in single-cell data science. Genome Biol. 21, 31 (2020).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. 20, 257–272 (2019).
Khrameeva, E. et al. Single-cell-resolution transcriptome map of human, chimpanzee, bonobo, and macaque brains. Genome Res. 30, 776–789 (2020).
Pollen, A. A. et al. Establishing cerebral organoids as models of human-specific brain evolution. Cell 176, 743–756.e17 (2019).
Ma, S. et al. Molecular and cellular evolution of the primate dorsolateral prefrontal cortex. Science 377, eabo7257 (2022).
Krienen, F. M. et al. Innovations present in the primate interneuron repertoire. Nature 586, 262–269 (2020).
Bakken, T. E. et al. Single-cell and single-nucleus RNA-seq uncovers shared and distinct axes of variation in dorsal LGN neurons in mice, non-human primates, and humans. eLife 10, e64875 (2021).
Schmitz, M. T. et al. The development and evolution of inhibitory neurons in primate cerebrum. Nature 603, 871–877 (2022).
Peng, Y.-R. et al. Molecular classification and comparative taxonomics of foveal and peripheral cells in primate retina. Cell 176, 1222–1237.e22 (2019).
Yi, W. et al. A single-cell transcriptome atlas of the aging human and macaque retina. Natl Sci. Rev. 8, nwaa179 (2021).
Yuan, W. et al. Temporally divergent regulatory mechanisms govern neuronal diversification and maturation in the mouse and marmoset neocortex. Nat. Neurosci. 25, 1049–1058 (2022).
Micali, N. et al. Molecular programs of regional specification and neural stem cell fate progression in developing macaque telencephalon. bioRxiv https://doi.org/10.1101/2022.10.18.512724 (2022).
Franjic, D. et al. Transcriptomic taxonomy and neurogenic trajectories of adult human, macaque, and pig hippocampal and entorhinal cells. Neuron 110, 452–469.e14 (2022).
Thomson, J. A. et al. Isolation of a primate embryonic stem cell line. Proc. Natl Acad. Sci. USA 92, 7844–7848 (1995).
Thomson, J. A. et al. Pluripotent cell lines derived from common marmoset (Callithrix jacchus) blastocysts. Biol. Reprod. 55, 254–259 (1996).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Suemori, H. et al. Establishment of embryonic stem cell lines from cynomolgus monkey blastocysts produced by IVF or ICSI. Dev. Dyn. 222, 273–279 (2001).
Simerly, C. R. et al. Establishment and characterization of baboon embryonic stem cell lines: an Old World Primate model for regeneration and transplantation research. Stem Cell Res. 2, 178–187 (2009).
Shimozawa, N., Nakamura, S., Takahashi, I., Hatori, M. & Sankai, T. Characterization of a novel embryonic stem cell line from an ICSI-derived blastocyst in the African green monkey. Reproduction 139, 565–573 (2010).
Shi, Y., Inoue, H., Wu, J. C. & Yamanaka, S. Induced pluripotent stem cell technology: a decade of progress. Nat. Rev. Drug Discov. 16, 115–130 (2017).
Wu, Y. et al. Nonhuman primate induced pluripotent stem cells in regenerative medicine. Stem Cell Int. 2012, 767195 (2012).
Liu, H. et al. Generation of induced pluripotent stem cells from adult rhesus monkey fibroblasts. Cell Stem Cell 3, 587–590 (2008).
Wu, Y., Zhang, Y., Mishra, A., Tardif, S. D. & Hornsby, P. J. Generation of induced pluripotent stem cells from newborn marmoset skin fibroblasts. Stem Cell Res. 4, 180–188 (2010).
Deleidi, M., Hargus, G., Hallett, P., Osborn, T. & Isacson, O. Development of histocompatible primate-induced pluripotent stem cells for neural transplantation. Stem Cell 29, 1052–1063 (2011).
Zhong, B. et al. Efficient generation of nonhuman primate induced pluripotent stem cells. Stem Cell Dev. 20, 795–807 (2011).
Mishra, A. et al. In Induced Pluripotent Stem (IPS) Cells: Methods and Protocols (eds Turksen, K. & Nagy, A.) 183–193 (Springer New York, 2016).
Ben-Nun, I. F. et al. Induced pluripotent stem cells from highly endangered species. Nat. Methods 8, 829–831 (2011).
Marchetto, M. C. N. et al. Differential L1 regulation in pluripotent stem cells of humans and apes. Nature 503, 525–529 (2013).
Wunderlich, S. et al. Primate iPS cells as tools for evolutionary analyses. Stem Cell Res. 12, 622–629 (2014).
Field, A. R. et al. Structurally conserved primate LncRNAs are transiently expressed during human cortical differentiation and influence cell-type-specific genes. Stem Cell Rep. 12, 245–257 (2019).
Geuder, J. et al. A non-invasive method to generate induced pluripotent stem cells from primate urine. Sci. Rep. 11, 3516 (2021).
Dannemann, M. & Romero, I. G. Harnessing pluripotent stem cells as models to decipher human evolution. FEBS J. 289, 2992–3010 (2022).
Fujie, Y. et al. New type of Sendai virus vector provides transgene-free iPS cells derived from chimpanzee blood. PLoS One 9, e113052 (2014).
Gallego Romero, I. et al. A panel of induced pluripotent stem cells from chimpanzees: a resource for comparative functional genomics. eLife 4, e07103 (2015).
Ward, M. C. et al. Silencing of transposable elements may not be a major driver of regulatory evolution in primate iPSCs. eLife 7, e33084 (2018).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Chen, C., Ji, W. & Niu, Y. Primate organoids and gene-editing technologies toward next-generation biomedical research. Trends Biotechnol. 39, 1332–1342 (2021).
Kelley, K. W. & Pașca, S. P. Human brain organogenesis: toward a cellular understanding of development and disease. Cell 185, 42–61 (2022).
Marchetto, M. C. et al. Species-specific maturation profiles of human, chimpanzee and bonobo neural cells. eLife 8, e37527 (2019).
Otani, T., Marchetto, M. C., Gage, F. H., Simons, B. D. & Livesey, F. J. 2D and 3D stem cell models of primate cortical development identify species-specific differences in progenitor behavior contributing to brain size. Cell Stem Cell 18, 467–480 (2016).
Cakir, B. et al. Engineering of human brain organoids with a functional vascular-like system. Nat. Methods 16, 1169–1175 (2019).
Andersen, J. et al. Generation of functional human 3D cortico-motor assembloids. Cell 183, 1913–1929.e26 (2020).
Agoglia, R. M. et al. Primate cell fusion disentangles gene regulatory divergence in neurodevelopment. Nature 592, 421–427 (2021).
Gokhman, D. et al. Human-chimpanzee fused cells reveal cis-regulatory divergence underlying skeletal evolution. Nat. Genet. 53, 467–476 (2021). Gokhman et al.197 and Agoglia et al.196 provide the first application of comparative functional genomics using allotetraploid organoids.
Song, J. H. T. et al. Genetic studies of human-chimpanzee divergence using stem cell fusions. Proc. Natl Acad. Sci. USA 118, e2117557118 (2021).
Pavlovic, B. J., Fox, D., Schaefer, N. K. & Pollen, A. A. Rethinking nomenclature for interspecies cell fusions. Nat. Rev. Genet. 23, 315–320 (2022).
Benito-Kwiecinski, S. et al. An early cell shape transition drives evolutionary expansion of the human forebrain. Cell 184, 2084–2102.e19 (2021).
Mora-Bermúdez, F. et al. Differences and similarities between human and chimpanzee neural progenitors during cerebral cortex development. eLife 5, e18683 (2016).
Muchnik, S. K., Lorente-Galdos, B., Santpere, G. & Sestan, N. Modeling the evolution of human brain development using organoids. Cell 179, 1250–1253 (2019).
Grassi, D. A. et al. Post-transcriptional mechanisms distinguish human and chimp forebrain progenitor cells. bioRxiv https://doi.org/10.1101/582197 (2019).
Kanton, S. et al. Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature 574, 418–422 (2019). This paper offers a paradigmatic functional comparative study in organoids leveraging single-cell multimodal data.
Singleton, M. Primate cranial diversity. Nat. Educ. Knowl. 4, 1 (2013).
Varki, N. et al. Heart disease is common in humans and chimpanzees, but is caused by different pathological processes. Evol. Appl. 2, 101–112 (2009).
Ward, M. C. & Gilad, Y. A generally conserved response to hypoxia in iPSC-derived cardiomyocytes from humans and chimpanzees. eLife 8, e42374 (2019).
Zhao, X. et al. Comparison of non-human primate versus human induced pluripotent stem cell-derived cardiomyocytes for treatment of myocardial infarction. Stem Cell Rep. 10, 422–435 (2018).
Jacobo Lopez, A. et al. Retinal organoids derived from rhesus macaque iPSCs undergo accelerated differentiation compared to human stem cells. Cell Prolif. 55, e13198 (2022).
Onozato, D. et al. Efficient generation of cynomolgus monkey induced pluripotent stem cell-derived intestinal organoids with pharmacokinetic functions. Stem Cell Dev. 27, 1033–1045 (2018).
Domingues, S. et al. Differentiation of nonhuman primate pluripotent stem cells into functional keratinocytes. Stem Cell Res. Ther. 8, 285 (2017).
Housman, G., Briscoe, E. & Gilad, Y. Evolutionary insights into primate skeletal gene regulation using a comparative cell culture model. PLoS Genet. 18, e1010073 (2022).
Cagan, A. et al. Natural selection in the great apes. Mol. Biol. Evol. 33, 3268–3283 (2016).
Blake, L. E. et al. A comparative study of endoderm differentiation in humans and chimpanzees. Genome Biol. 19, 162 (2018).
Moris, N. et al. An in vitro model of early anteroposterior organization during human development. Nature 582, 410–415 (2020).
Hockemeyer, D. & Jaenisch, R. Induced pluripotent stem cells meet genome editing. Cell Stem Cell 18, 573–586 (2016).
Acknowledgements
The authors thank William R. Bradley for help collecting metadata and all the authors with important contributions to primate comparative omics, many of which could not be included in this Review. T.M.-B. is supported by funding from the European Research Council (ERC) under the European Union Horizon 2020 Research and Innovation Programme (grant agreement No. 864203), PID2021-126004NB-100 (MINECO/FEDER, UE), “Unidad de Excelencia María de Maeztu”, funded by the AEI (CEX2018-000792-M), US National Institutes of Health (NIH) 1R01HG010898-01A1, and Secretaria d’Universitats i Recerca and CERCA Programme del Departament d’Economia i Coneixement de la Generalitat de Catalunya (GRC 2017 SGR 880). G.S. is supported by grant MS20/00064 from Instituto de Salud Carlos III (Spain) and the European Social Fund, and grant PID2019-104700GA-I00 funded by the AEI, Spain. G.S. is also supported by the NIH grant R01HG010898-01. J.L.K. is supported by National Science Foundation IOS 1931650 and María de Maeztu Mobility Fellowship.
Author information
Authors and Affiliations
Contributions
All the authors contributed equally to all aspects of the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Genetics thanks Katja Nowick, Christian Roos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Barcelona Cryozoo: https://www.zoobarcelona.cat/es/cryozoo
Frozen Zoo: https://science.sandiegozoo.org/resources/frozen-zoo®
Human Cell Atlas: https://humancellatlas.org
Macaque Genotype and Phenotype Resource: https://mgap.ohsu.edu
Non-human Primate Reference Transcriptome Resource: http://www.nhprtr.org
Species360: https://www.species360.org
The Frozen Ark: https://www.frozenark.org
The Nagoya Protocol Learning Portal: https://learnnagoya.com
Time Capsule Program for Environmental Specimens: https://www.nies.go.jp/timecaps1/summary/objectiveE.htm
TimeTree of Life: http://www.timetree.org
Supplementary information
Glossary
- Allotetraploid
-
Composite cell lines from different diploid species.
- Assembloids
-
Assemblies of organoids derived from different cell lineages.
- Balancing selection
-
The set of selective processes that maintain genetic variation longer than expected from genetic drift.
- Cellular primatology
-
Comparative studies focused on the cellular process underpinning the phenotypic variation among humans and non-human primates, including cell development, molecular features, tissue composition and organization.
- Composite cell lines
-
Cell lines obtained from the fusion of two diploid cell lines from the same or different species.
- Copy number variants
-
(CNVs). Variable regions in the genome in which sections of the genome are repeated; the number of repeats differs between individuals.
- CRISPR–Cas9
-
A widely used genome-editing technique consisting of a nuclease linked to a guide RNA cutting specific sites, which can be then repaired by homology-directed repair introducing the desired mutations.
- Deep learning
-
A type of machine learning that achieves great power to predict the output concepts or classes associated with the input cases. It works by learning a representation of many nested concepts connecting the input data to the desired outputs from the exposition to multiple cases.
- Epigenomic
-
Pertains to chemical modifications on DNA (for example, CpG-methylation) and histones (for example, methylation and acetylation) that correlate with functional features of the genome (for example, promoters, enhancers and gene transcription).
- Expression quantitative trait locus
-
(eQTL). Site in the genome where genetic variation is associated with altered gene expression.
- Genome-wide association studies
-
(GWAS). Studies designed to interrogate the contribution of millions of common variants to a trait or disease of interest.
- Gyrification
-
The process of the formation of the cortical folds during brain development.
- Induced pluripotent stem cells
-
(iPSCs). Pluripotent stem cells that are reprogrammed from somatic cells by inducing the expression of pluripotency factors.
- Long-read sequencing
-
A group of sequencing techniques that generate sequences of thousands to millions of nucleotides. They are especially suitable for resolving structural variants and repetitive sequences.
- Organoids
-
In vitro self-organizing 3D cell cultures resembling real organs in several aspects of structure, composition, development and function.
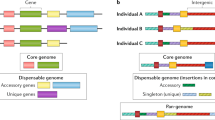
- Pangenome reference
-
A representation of the genome that captures the full genomic diversity for a given species.
- Reference genomes
-
Representative genome assemblies for given species, usually presented in haploid form, that facilitate the interpretation and analysis of resequenced data for organisms of the same or closely related species.
- Topologically associating domains
-
Regions of the genome with a high probability of self-interaction through the formation of chromatin loops.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Juan, D., Santpere, G., Kelley, J.L. et al. Current advances in primate genomics: novel approaches for understanding evolution and disease. Nat Rev Genet 24, 314–331 (2023). https://doi.org/10.1038/s41576-022-00554-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-022-00554-w
This article is cited by
-
Generation of chimpanzee induced pluripotent stem cell lines for cross-species comparisons
In Vitro Cellular & Developmental Biology - Animal (2024)
-
Identification of constrained sequence elements across 239 primate genomes
Nature (2024)