Abstract
Transposable elements are abundant in the human genome, and great strides have been made in pinpointing variations in these repetitive sequences using whole-genome sequencing. Now, the focus is shifting to understanding their expression and regulation, and the functional consequences of their insertion and retention in the genome over time. Whereas transposable element insertions have been known to cause human genetic disease since the 1980s, the scope of their contributions to heritable phenotypes is now starting to be uncovered. Here, we review the many ways human retrotransposons contribute to genome function, their dysregulation in diseases including cancer and how they affect genetic disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0. Institute for Systems Biology http://www.repeatmasker.org (2013–2015).
Boissinot, S., Davis, J., Entezam, A., Petrov, D. & Furano, A. V. Fitness cost of LINE-1 (L1) activity in humans. Proc. Natl Acad. Sci. USA 103, 9590–9594 (2006).
Rishishwar, L. et al. Evidence for positive selection on recent human transposable element insertions. Gene 675, 69–79 (2018).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory activities of transposable elements: from conflicts to benefits. Nat. Rev. Genet. 18, 71–86 (2017).
Lowe, C. B. & Haussler, D. 29 mammalian genomes reveal novel exaptations of mobile elements for likely regulatory functions in the human genome. PLOS ONE 7, e43128 (2012).
Flemr, M. et al. A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell 155, 807–816 (2013).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 351, 1083–1087 (2016). This recent report demonstrates a ‘plug-and-play’ model whereby TEs provide co-opted regulatory sequences that wire a gene network.
Fuentes, D. R., Swigut, T. & Wysocka, J. Systematic perturbation of retroviral LTRs reveals widespread long-range effects on human gene regulation. eLife 7, e35989 (2018).
Attig, J. et al. Splicing repression allows the gradual emergence of new Alu-exons in primate evolution. eLife 5, e19545 (2016).
Aktas, T. et al. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 544, 115–119 (2017).
Zarnack, K. et al. Direct competition between hnRNP C and U2AF65 protects the transcriptome from the exonization of Alu elements. Cell 152, 453–466 (2013). This study describes the identification of the mechanism that suppresses Alu exonization.
Bailey, J. A., Liu, G. & Eichler, E. E. An Alu transposition model for the origin and expansion of human segmental duplications. Am. J. Hum. Genet. 73, 823–834 (2003).
Ewing, A. D. & Kazazian, H. H. Jr. High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res 20, 1262–1270 (2010).
Huang, C. R. et al. Mobile interspersed repeats are major structural variants in the human genome. Cell 141, 1171–1182 (2010).
Iskow, R. C. et al. Natural mutagenesis of human genomes by endogenous retrotransposons. Cell 141, 1253–1261 (2010).
Witherspoon, D. J. et al. Mobile element scanning (ME-Scan) by targeted high-throughput sequencing. BMC Genomics 11, 410 (2010).
Stewart, C. et al. A comprehensive map of mobile element insertion polymorphisms in humans. PLOS Genet. 7, e1002236 (2011).
Sudmant, P. H. et al. An integrated map of structural variation in 2,504 human genomes. Nature 526, 75–81 (2015). This study reports the mapping of TEs in whole-genome data and provides the best current catalogue of structural variants resulting from mobile element activity.
Chaisson, M. J. et al. Resolving the complexity of the human genome using single-molecule sequencing. Nature 517, 608–611 (2015).
Konkel, M. K., Walker, J. A. & Batzer, M. A. LINEs and SINEs of primate evolution. Evol. Anthropol. 19, 236–249 (2010).
Brouha, B. et al. Hot L1s account for the bulk of retrotransposition in the human population. Proc. Natl Acad. Sci. USA 100, 5280–5285 (2003).
Beck, C. R. et al. LINE-1 retrotransposition activity in human genomes. Cell 141, 1159–1170 (2010).
Martin, S. L. et al. LINE-1 retrotransposition requires the nucleic acid chaperone activity of the ORF1 protein. J. Mol. Biol. 348, 549–561 (2005).
Khazina, E. et al. Trimeric structure and flexibility of the L1ORF1 protein in human L1 retrotransposition. Nat. Struct. Mol. Biol. 18, 1006–1014 (2011).
Feng, Q., Moran, J. V., Kazazian, H. H. Jr. & Boeke, J. D. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell 87, 905–916 (1996).
Weichenrieder, O., Repanas, K. & Perrakis, A. Crystal structure of the targeting endonuclease of the human LINE-1 retrotransposon. Structure 12, 975–986 (2004).
Mathias, S. L., Scott, A. F., Kazazian, H. H. Jr., Boeke, J. D. & Gabriel, A. Reverse transcriptase encoded by a human transposable element. Science 254, 1808–1810 (1991).
Luan, D. D., Korman, M. H., Jakubczak, J. L. & Eickbush, T. H. Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell 72, 595–605 (1993).
Ostertag, E. M. & Kazazian, H. H. Jr. Twin priming: a proposed mechanism for the creation of inversions in L1 retrotransposition. Genome Res. 11, 2059–2065 (2001).
Kulpa, D. A. & Moran, J. V. Cis-preferential LINE-1 reverse transcriptase activity in ribonucleoprotein particles. Nat. Struct. Mol. Biol. 13, 655–660 (2006).
Ullu, E. & Tschudi, C. Alu sequences are processed 7SL RNA genes. Nature 312, 171–172 (1984).
Dewannieux, M., Esnault, C. & Heidmann, T. LINE-mediated retrotransposition of marked Alu sequences. Nat. Genet. 35, 41–48 (2003).
Ahl, V., Keller, H., Schmidt, S. & Weichenrieder, O. Retrotransposition and crystal structure of an Alu RNP in the ribosome-stalling conformation. Mol. Cell 60, 715–727 (2015).
Hancks, D. C., Goodier, J. L., Mandal, P. K., Cheung, L. E. & Kazazian, H. H. Jr. Retrotransposition of marked SVA elements by human L1s in cultured cells. Hum. Mol. Genet. 20, 3386–3400 (2011).
Buzdin, A. et al. A new family of chimeric retrotranscripts formed by a full copy of U6 small nuclear RNA fused to the 3′ terminus of L1. Genomics 80, 402–406 (2002).
Doucet, A. J., Droc, G., Siol, O., Audoux, J. & Gilbert, N. U6 snRNA pseudogenes: markers of retrotransposition dynamics in mammals. Mol. Biol. Evol. 32, 1815–1832 (2015).
Esnault, C., Maestre, J. & Heidmann, T. Human LINE retrotransposons generate processed pseudogenes. Nat. Genet. 24, 363–367 (2000).
Gagnier, L., Belancio, V. P. & Mager, D. L. Mouse germ line mutations due to retrotransposon insertions. Mob. DNA 10, 15 (2019).
Dewannieux, M. et al. Identification of an infectious progenitor for the multiple-copy HERV-K human endogenous retroelements. Genome Res. 16, 1548–1556 (2006).
Mager, D. L. & Stoye, J. P. Mammalian endogenous retroviruses. Microbiol. Spectr. 3, MDNA3-0009-2014 (2015).
Belshaw, R. et al. Genomewide screening reveals high levels of insertional polymorphism in the human endogenous retrovirus family HERV-K(HML2): implications for present-day activity. J. Virol. 79, 12507–12514 (2005).
Wildschutte, J. H. et al. Discovery of unfixed endogenous retrovirus insertions in diverse human populations. Proc. Natl Acad. Sci. USA 113, E2326–E2334 (2016).
Thomas, J., Perron, H. & Feschotte, C. Variation in proviral content among human genomes mediated by LTR recombination. Mob. DNA 9, 36 (2018).
Buzdin, A., Kovalskaya-Alexandrova, E., Gogvadze, E. & Sverdlov, E. At least 50% of human-specific HERV-K (HML-2) long terminal repeats serve in vivo as active promoters for host nonrepetitive DNA transcription. J. Virol. 80, 10752–10762 (2006).
Babaian, A. & Mager, D. L. Endogenous retroviral promoter exaptation in human cancer. Mob. DNA 7, 24 (2016).
Skowronski, J., Fanning, T. G. & Singer, M. F. Unit-length line-1 transcripts in human teratocarcinoma cells. Mol. Cell. Biol. 8, 1385–1397 (1988).
Deininger, P. et al. A comprehensive approach to expression of L1 loci. Nucleic Acids Res 45, e31 (2017).
Jacobs, F. M. et al. An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons. Nature 516, 242–245 (2014).
Imbeault, M., Helleboid, P. Y. & Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 543, 550–554 (2017). This report presents the large-scale mapping of KZFPs to transposable elements, demonstrating the potential for regulatory repurposing.
Quenneville, S. et al. The KRAB-ZFP/KAP1 system contributes to the early embryonic establishment of site-specific DNA methylation patterns maintained during development. Cell Rep. 2, 766–773 (2012).
Rowe, H. M. et al. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463, 237–240 (2010).
Molaro, A. & Malik, H. S. Hide and seek: how chromatin-based pathways silence retroelements in the mammalian germline. Curr. Opin. Genet. Dev. 37, 51–58 (2016).
Slotkin, R. K. & Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nat. Rev. Genet. 8, 272–285 (2007).
Kapusta, A. et al. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLOS Genet. 9, e1003470 (2013).
Crow, M. K. Long interspersed nuclear elements (LINE-1): potential triggers of systemic autoimmune disease. Autoimmunity 43, 7–16 (2010).
Poduri, A., Evrony, G. D., Cai, X. & Walsh, C. A. Somatic mutation, genomic variation, and neurological disease. Science 341, 1237758 (2013).
De Cecco, M. et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 566, 73–78 (2019).
Scott, E. C. et al. A hot L1 retrotransposon evades somatic repression and initiates human colorectal cancer. Genome Res. 26, 745–755 (2016).
Rodic, N. et al. Long interspersed element-1 protein expression is a hallmark of many human cancers. Am. J. Pathol. 184, 1280–1286 (2014).
Lee, E. et al. Landscape of somatic retrotransposition in human cancers. Science 337, 967–971 (2012).
Shukla, R. et al. Endogenous retrotransposition activates oncogenic pathways in hepatocellular carcinoma. Cell 153, 101–111 (2013).
Helman, E. et al. Somatic retrotransposition in human cancer revealed by whole-genome and exome sequencing. Genome Res. 24, 1053–1063 (2014).
Rodic, N. et al. Retrotransposon insertions in the clonal evolution of pancreatic ductal adenocarcinoma. Nat. Med. 21, 1060–1064 (2015).
Tang, Z. et al. Human transposon insertion profiling: analysis, visualization and identification of somatic LINE-1 insertions in ovarian cancer. Proc. Natl Acad. Sci. USA 114, E733–E740 (2017).
Nguyen, T. H. M. et al. L1 retrotransposon heterogeneity in ovarian tumor cell evolution. Cell Rep. 23, 3730–3740 (2018).
Burns, K. H. Transposable elements in cancer. Nat. Rev. Cancer 17, 415–424 (2017).
Miki, Y. et al. Disruption of the APC gene by a retrotransposal insertion of L1 sequence in a colon cancer. Cancer Res. 52, 643–645 (1992).
Jang, H. S. et al. Transposable elements drive widespread expression of oncogenes in human cancers. Nat. Genet. 51, 611–617 (2019).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Leonova, K. I. et al. p53 cooperates with DNA methylation and a suicidal interferon response to maintain epigenetic silencing of repeats and noncoding RNAs. Proc. Natl Acad. Sci. USA 110, E89–E98 (2013).
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961–973 (2015).
Jones, P. A., Ohtani, H., Chakravarthy, A. & De Carvalho, D. D. Epigenetic therapy in immune-oncology. Nat. Rev. Cancer 19, 151–161 (2019).
Gannon, H. S. et al. Identification of ADAR1 adenosine deaminase dependency in a subset of cancer cells. Nat. Commun. 9, 5450 (2018).
Hancks, D. C. & Kazazian, H. H. Jr. Roles for retrotransposon insertions in human disease. Mob. DNA 7, 9 (2016).
Kazazian, H. H. Jr. et al. Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 332, 164–166 (1988). This classic report is the first to report a disease-causing TE insertion.
Apoil, P. A., Kuhlein, E., Robert, A., Rubie, H. & Blancher, A. HIGM syndrome caused by insertion of an AluYb8 element in exon 1 of the CD40LG gene. Immunogenetics 59, 17–23 (2007).
Nakamura, Y. et al. SVA retrotransposition in exon 6 of the coagulation factor IX gene causing severe hemophilia B. Int. J. Hematol. 102, 134–139 (2015).
Taskesen, M. et al. Novel Alu retrotransposon insertion leading to Alstrom syndrome. Hum. Genet. 131, 407–413 (2012).
Claverie-Martin, F., Flores, C., Anton-Gamero, M., Gonzalez-Acosta, H. & Garcia-Nieto, V. The Alu insertion in the CLCN5 gene of a patient with Dent’s disease leads to exon 11 skipping. J. Hum. Genet. 50, 370–374 (2005).
Narita, N. et al. Insertion of a 5′ truncated L1 element into the 3′ end of exon 44 of the dystrophin gene resulted in skipping of the exon during splicing in a case of Duchenne muscular dystrophy. J. Clin. Invest. 91, 1862–1867 (1993).
Wallace, M. R. et al. A de novo Alu insertion results in neurofibromatosis type 1. Nature 353, 864–866 (1991).
Gallus, G. N. et al. Alu-element insertion in an OPA1 intron sequence associated with autosomal dominant optic atrophy. Mol. Vis. 16, 178–183 (2010).
Meischl, C., Boer, M., Ahlin, A. & Roos, D. A new exon created by intronic insertion of a rearranged LINE-1 element as the cause of chronic granulomatous disease. Eur. J. Hum. Genet. 8, 697–703 (2000).
Rodriguez-Martin, C. et al. Familial retinoblastoma due to intronic LINE-1 insertion causes aberrant and noncanonical mRNA splicing of the RB1 gene. J. Hum. Genet. 61, 463–466 (2016).
Lev-Maor, G. et al. Intronic Alus influence alternative splicing. PLOS Genet. 4, e1000204 (2008).
Hancks, D. C., Ewing, A. D., Chen, J. E., Tokunaga, K. & Kazazian, H. H. Jr. Exon-trapping mediated by the human retrotransposon SVA. Genome Res. 19, 1983–1991 (2009).
Sela, N., Mersch, B., Hotz-Wagenblatt, A. & Ast, G. Characteristics of transposable element exonization within human and mouse. PLOS ONE 5, e10907 (2010).
van der Klift, H. M., Tops, C. M., Hes, F. J., Devilee, P. & Wijnen, J. T. Insertion of an SVA element, a nonautonomous retrotransposon, in PMS2 intron 7 as a novel cause of Lynch syndrome. Hum. Mutat. 33, 1051–1055 (2012).
de Boer, M. et al. Primary immunodeficiency caused by an exonized retroposed gene copy inserted in the CYBB gene. Hum. Mutat. 35, 486–496 (2014).
Vogt, J. et al. SVA retrotransposon insertion-associated deletion represents a novel mutational mechanism underlying large genomic copy number changes with non-recurrent breakpoints. Genome Biol. 15, R80 (2014).
Mine, M. et al. A large genomic deletion in the PDHX gene caused by the retrotranspositional insertion of a full-length LINE-1 element. Hum. Mutat. 28, 137–142 (2007).
Peixoto, A. et al. Genomic characterization of two large Alu-mediated rearrangements of the BRCA1 gene. J. Hum. Genet. 58, 78–83 (2013).
Morrish, T. A. et al. DNA repair mediated by endonuclease-independent LINE-1 retrotransposition. Nat. Genet. 31, 159–165 (2002).
Morisada, N. et al. Branchio-oto-renal syndrome caused by partial EYA1 deletion due to LINE-1 insertion. Pediatr. Nephrol. 25, 1343–1348 (2010).
Kazazian, H. H. Jr. & Moran, J. V. Mobile DNA in health and disease. N. Engl. J. Med. 377, 361–370 (2017).
Wimmer, K., Callens, T., Wernstedt, A. & Messiaen, L. The NF1 gene contains hotspots for L1 endonuclease-dependent de novo insertion. PLOS Genet. 7, e1002371 (2011). This analysis focuses on the NF1 locus in patients with neurofibromatosis-identified frequent TPRT insertions; similar analyses at loci for other monogenic disease genes will likely find more de novo TE insertions.
Kobayashi, K. et al. An ancient retrotransposal insertion causes Fukuyama-type congenital muscular dystrophy. Nature 394, 388–392 (1998).
Taniguchi-Ikeda, M. et al. Pathogenic exon-trapping by SVA retrotransposon and rescue in Fukuyama muscular dystrophy. Nature 478, 127–131 (2011).
Kagawa, T. et al. Recessive inheritance of population-specific intronic LINE-1 insertion causes a Rotor syndrome phenotype. Hum. Mutat. 36, 327–332 (2015).
Tucker, B. A. et al. Exome sequencing and analysis of induced pluripotent stem cells identify the cilia-related gene male germ cell-associated kinase (MAK) as a cause of retinitis pigmentosa. Proc. Natl Acad. Sci. USA 108, E569–E576 (2011).
Makino, S. et al. Reduced neuron-specific expression of the TAF1 gene is associated with X-linked dystonia-parkinsonism. Am. J. Hum. Genet. 80, 393–406 (2007).
Aneichyk, T. et al. Dissecting the causal mechanism of X-linked dystonia-parkinsonism by integrating genome and transcriptome assembly. Cell 172, 897–909.e21 (2018).
Bragg, D. C. et al. Disease onset in X-linked dystonia-parkinsonism correlates with expansion of a hexameric repeat within an SVA retrotransposon in TAF1. Proc. Natl Acad. Sci. USA 114, E11020–E11028 (2017).
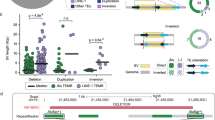
Payer, L. M. et al. Structural variants caused by Alu insertions are associated with risks for many human diseases. Proc. Natl Acad. Sci. USA 114, E3984–E3992 (2017). This study demonstrates that polymorphic TEs are potential causative variants in common diseases studied by GWAS.
Wang, L., Norris, E. T. & Jordan, I. K. Human retrotransposon insertion polymorphisms are associated with health and disease via gene regulatory phenotypes. Front. Microbiol. 8, 1418 (2017).
Payer, L. M. et al. Alu insertion variants alter mRNA splicing. Nucleic Acids Res. 47, 421–431 (2019). This study reports a mechanism by which common insertion variants contribute to disease risk by inducing splicing quantitative trait loci.
De Jager, P. L. et al. The role of the CD58 locus in multiple sclerosis. Proc. Natl Acad. Sci. USA 106, 5264–5269 (2009).
Wang, L., Rishishwar, L., Marino-Ramirez, L. & Jordan, I. K. Human population-specific gene expression and transcriptional network modification with polymorphic transposable elements. Nucleic Acids Res. 45, 2318–2328 (2017).
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10, 691–703 (2009).
Su, L. K. et al. Genomic rearrangements of the APC tumor-suppressor gene in familial adenomatous polyposis. Hum. Genet. 106, 101–107 (2000).
Garland, J. et al. Identification of an Alu element-mediated deletion in the promoter region of GNE in siblings with GNE myopathy. Mol. Genet. Genomic Med. 5, 410–417 (2017).
Rickman, K. A. et al. Deficiency of UBE2T, the E2 ubiquitin ligase necessary for FANCD2 and FANCI ubiquitination, causes FA-T subtype of fanconi anemia. Cell Rep. 12, 35–41 (2015).
Brooks, E. M., Branda, R. F., Nicklas, J. A. & O’Neill, J. P. Molecular description of three macro-deletions and an Alu–Alu recombination-mediated duplication in the HPRT gene in four patients with Lesch–Nyhan disease. Mutat. Res. 476, 43–54 (2001).
Gu, S. et al. Alu-mediated diverse and complex pathogenic copy-number variants within human chromosome 17 at p13.3. Hum. Mol. Genet. 24, 4061–4077 (2015).
Nazaryan-Petersen, L. et al. Germline chromothripsis driven by l1-mediated retrotransposition and Alu/Alu homologous recombination. Hum. Mutat. 37, 385–395 (2016).
Burwinkel, B. & Kilimann, M. W. Unequal homologous recombination between LINE-1 elements as a mutational mechanism in human genetic disease. J. Mol. Biol. 277, 513–517 (1998).
Temtamy, S. A. et al. Long interspersed nuclear element-1 (LINE1)-mediated deletion of EVC, EVC2, C4orf6, and STK32B in Ellis–van Creveld syndrome with borderline intelligence. Hum. Mutat. 29, 931–938 (2008).
Sun, C. et al. Deletion of azoospermia factor a (AZFa) region of human Y chromosome caused by recombination between HERV15 proviruses. Hum. Mol. Genet. 9, 2291–2296 (2000).
Segal, Y. et al. LINE-1 elements at the sites of molecular rearrangements in Alport syndrome–diffuse leiomyomatosis. Am. J. Hum. Genet. 64, 62–69 (1999).
Jacob, F. Evolution and tinkering. Science 196, 1161–1166 (1977).
Ecco, G. et al. Transposable elements and their KRAB-ZFP controllers regulate gene expression in adult tissues. Dev. Cell 36, 611–623 (2016).
Chuong, E. B., Rumi, M. A., Soares, M. J. & Baker, J. C. Endogenous retroviruses function as species-specific enhancer elements in the placenta. Nat. Genet. 45, 325–329 (2013).
Dunn-Fletcher, C. E. et al. Anthropoid primate-specific retroviral element THE1B controls expression of CRH in placenta and alters gestation length. PLOS Biol. 16, e2006337 (2018).
Sorek, R., Ast, G. & Graur, D. Alu-containing exons are alternatively spliced. Genome Res. 12, 1060–1067 (2002).
Lev-Maor, G., Sorek, R., Shomron, N. & Ast, G. The birth of an alternatively spliced exon: 3′ splice-site selection in Alu exons. Science 300, 1288–1291 (2003).
Agrawal, A., Eastman, Q. M. & Schatz, D. G. Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system. Nature 394, 744–751 (1998).
Hiom, K., Melek, M. & Gellert, M. DNA transposition by the RAG1 and RAG2 proteins: a possible source of oncogenic translocations. Cell 94, 463–470 (1998).
Kapitonov, V. V. & Jurka, J. RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLOS Biol. 3, e181 (2005).
Hencken, C. G., Li, X. & Craig, N. L. Functional characterization of an active Rag-like transposase. Nat. Struct. Mol. Biol. 19, 834–836 (2012).
Huang, S. et al. Discovery of an active RAG transposon illuminates the origins of V(D)J recombination. Cell 166, 102–114 (2016).
Zhang, Y. et al. Transposon molecular domestication and the evolution of the RAG recombinase. Nature 569, 79–84 (2019).
Sinzelle, L., Izsvak, Z. & Ivics, Z. Molecular domestication of transposable elements: from detrimental parasites to useful host genes. Cell Mol. Life Sci. 66, 1073–1093 (2009).
Smit, A. F. & Riggs, A. D. Tiggers and DNA transposon fossils in the human genome. Proc. Natl Acad. Sci. USA 93, 1443–1448 (1996).
Sarkar, A. et al. Molecular evolutionary analysis of the widespread piggyBac transposon family and related ‘domesticated’ sequences. Mol. Genet. Genomics 270, 173–180 (2003).
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
Stessman, H. A. F. et al. Disruption of POGZ is associated with intellectual disability and autism spectrum disorders. Am. J. Hum. Genet. 98, 541–552 (2016).
Henssen, A. G. et al. Genomic DNA transposition induced by human PGBD5. eLife 4, e10565 (2015).
Henssen, A. G. et al. PGBD5 promotes site-specific oncogenic mutations in human tumors. Nat. Genet. 49, 1005–1014 (2017).
Henssen, A. G. et al. Therapeutic targeting of PGBD5-induced DNA repair dependency in pediatric solid tumors. Sci. Transl Med. 9, eaam9078 (2017).
Blaise, S., de Parseval, N., Benit, L. & Heidmann, T. Genomewide screening for fusogenic human endogenous retrovirus envelopes identifies syncytin 2, a gene conserved on primate evolution. Proc. Natl Acad. Sci. USA 100, 13013–13018 (2003).
Blond, J. L. et al. An envelope glycoprotein of the human endogenous retrovirus HERV-W is expressed in the human placenta and fuses cells expressing the type D mammalian retrovirus receptor. J. Virol. 74, 3321–3329 (2000).
Mi, S. et al. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature 403, 785–789 (2000).
Cornelis, G. et al. An endogenous retroviral envelope syncytin and its cognate receptor identified in the viviparous placental Mabuya lizard. Proc. Natl Acad. Sci. USA 114, E10991–E11000 (2017).
Emerson, R. O. & Thomas, J. H. Gypsy and the birth of the SCAN domain. J. Virol. 85, 12043–12052 (2011).
Yang, W. R., Ardeljan, D., Pacyna, C. N., Payer, L. M. & Burns, K. H. SQuIRE reveals locus-specific regulation of interspersed repeat expression. Nucleic Acids Res 47, e27 (2019).
Jin, Y., Tam, O. H., Paniagua, E. & Hammell, M. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics 31, 3593–3599 (2015).
Philippe, C. et al. Activation of individual L1 retrotransposon instances is restricted to cell-type dependent permissive loci. eLife 5, e13926 (2016).
Goerner-Potvin, P. & Bourque, G. Computational tools to unmask transposable elements. Nat. Rev. Genet. 19, 688–704 (2018).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLOS Biol. 2, e391 (2004).
Chen, L. L., DeCerbo, J. N. & Carmichael, G. G. Alu element-mediated gene silencing. EMBO J. 27, 1694–1705 (2008).
Kawahara, Y. & Nishikura, K. Extensive adenosine-to-inosine editing detected in Alu repeats of antisense RNAs reveals scarcity of sense-antisense duplex formation. FEBS Lett. 580, 2301–2305 (2006).
Gong, C. & Maquat, L. E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–288 (2011).
Spengler, R. M., Oakley, C. K. & Davidson, B. L. Functional microRNAs and target sites are created by lineage-specific transposition. Hum. Mol. Genet. 23, 1783–1793 (2014).
Wang, L. & Jordan, I. K. Transposable element activity, genome regulation and human health. Curr. Opin. Genet. Dev. 49, 25–33 (2018).
Gardner, E. J. et al. The Mobile Element Locator Tool (MELT): population-scale mobile element discovery and biology. Genome Res. 27, 1916–1929 (2017).
1000 Genomes Project Consortium. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Acknowledgements
The authors thank many members of the transposable element research community for engaging conversations that have shaped their ideas, and they apologize to these colleagues for unreferenced work. The authors thank M. Gorbounov for technical assistance with L1 ORF1p staining.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
K.H.B. and the Johns Hopkins University School of Medicine have licenced antibodies for L1 ORF1p to EMD Millipore. L.M.P. declares no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks G. Faulkner, I. K. Jordan and D. Mager for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
dbvar: www.ncbi.nlm.nih.gov/dbvar
Online Mendelian Inheritance in Man: https://www.omim.org
Glossary
- Transposable elements
-
(TEs). DNA sequences that have the ability to move (transpose) to new locations in the genome.
- Retrotransposons
-
DNA sequences that proliferate in the genome using an RNA intermediate and a ‘copy-and-paste’ retrotransposition mechanism.
- Complex diseases
-
Common diseases caused by interactions of genetics, behaviour and the environment.
- Long interspersed element 1
-
(LINE-1 or L1). An autonomous (protein-coding) retrotransposon; currently, Homo sapiens-specific L1 (L1Hs) are active.
- Target primed reverse transcription
-
(TPRT). The form of retrotransposition used by non-long terminal repeat elements; in humans, this requires L1 ORF2p-encoded endonuclease and reverse transcriptase activities.
- Alu
-
A short interspersed element derived from 7SL RNA, a non-autonomous retrotransposon that relies on L1-encoded ORF2p.
- SVA
-
A composite retrotransposon made of short interspersed element, variable number tandem repeat and Alu sequences; also uses L1 protein to transpose.
- Processed pseudogenes
-
cDNA copy of a gene transcript inserted into the genome.
- Endogenous retroviruses
-
(ERVs). Autonomous (protein-coding) retrotransposons recently active in humans; also known as LTR elements for their long terminal repeats.
- Polymorphic TEs
-
Also known as retrotransposon insertion polymorphisms or polymorphic mobile element insertions. A transposable element (TE) insertion that is a structural variant in a population, present or absence at a locus.
- Minor allele frequency
-
For biallelic loci, the allele frequency for the second most common allele, as opposed to the major allele frequency; the two sum to 1 (p + q = 1).
- Synthetic lethalities
-
Cell deaths in response to a combination of two attributes, most often genetic lesions, when either one of which would be well tolerated.
- A-to-I editing
-
Conversion of adenosine (A) to inosine (I) in double-stranded RNA by adenosine deaminase acting on RNA 1 (ADAR1); relaxes strand annealing. Unedited hybrids in contact with cytoplasmic double-stranded RNA sensors can prompt interferon responses.
- Exonization
-
Incorporation of a new exon into a processed transcript; in this review, the incorporation of some transposable element sequences into a spliced mRNA.
- Endonuclease-independent retrotransposition
-
A process whereby a retrotransposon inserts at a pre-existing DNA break.
- Haplotype
-
An interval of DNA wherein a set of alleles is inherited as a group because of linkage disequilibrium.
- Linkage disequilibrium
-
The non-random association of alleles on the same DNA strand.
- Expression quantitative trait loci
-
(eQTLs). Sequence variants that are associated with alterations in mRNA levels.
- Splicing quantitative trait loci
-
Sequence variants that are associated with alterations in mRNA splicing.
- Chromothripsis
-
Chromosomal shattering. A large number of rearrangements occurring in a single event over limited genomic regions.
- GWAS trait-associated SNP
-
A single-nucleotide polymorphism (SNP) identified by a genome-wide association study (GWAS) as being associated with a disease or phenotypic trait.
Rights and permissions
About this article
Cite this article
Payer, L.M., Burns, K.H. Transposable elements in human genetic disease. Nat Rev Genet 20, 760–772 (2019). https://doi.org/10.1038/s41576-019-0165-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-019-0165-8
This article is cited by
-
Autophagy degrades immunogenic endogenous retroelements induced by 5-azacytidine in acute myeloid leukemia
Leukemia (2024)
-
Epigenomic insights into common human disease pathology
Cellular and Molecular Life Sciences (2024)
-
Transposable Elements: Emerging Therapeutic Targets in Neurodegenerative Diseases
Neurotoxicity Research (2024)
-
Assessing the efficacy of target adaptive sampling long-read sequencing through hereditary cancer patient genomes
npj Genomic Medicine (2024)
-
Taming transposable elements in livestock and poultry: a review of their roles and applications
Genetics Selection Evolution (2023)