Abstract
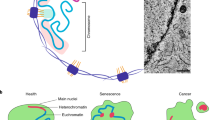
The genome is packaged and organized nonrandomly within the 3D space of the nucleus to promote efficient gene expression and to faithfully maintain silencing of heterochromatin. The genome is enclosed within the nucleus by the nuclear envelope membrane, which contains a set of proteins that actively participate in chromatin organization and gene regulation. Technological advances are providing views of genome organization at unprecedented resolution and are beginning to reveal the ways that cells co-opt the structures of the nuclear periphery for nuclear organization and gene regulation. These genome regulatory roles of proteins of the nuclear periphery have important influences on development, disease and ageing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Laubichler, M. D. & Davidson, E. H. Boveri’s long experiment: sea urchin merogones and the establishment of the role of nuclear chromosomes in development. Dev. Biol. 314, 1–11 (2008).
Cremer, T. & Cremer, C. Rise, fall and resurrection of chromosome territories: a historical perspective. Part I. The rise of chromosome territories. Eur. J. Histochem. 50, 161–176 (2006).
Croft, J. A. et al. Differences in the localization and morphology of chromosomes in the human nucleus. J. Cell Biol. 145, 1119–1131 (1999).
Sati, S. & Cavalli, G. Chromosome conformation capture technologies and their impact in understanding genome function. Chromosoma 126, 33–44 (2017).
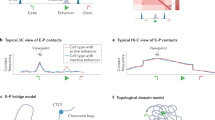
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009). This article provides the first demonstration of the HiC method for probing genome conformation indicating global properties of chromatin folding — that chromosomes occupy distinct territories and that chromatin separates into megabase-scale A and B compartments on the basis of chromatin activity and gene density.
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016). This study uses iterative FISH to demonstrate that regions of single chromosomes that partition into different compartments defined by genomic analyses also pack into distinct domains within single cells.
Wong, X., Luperchio, T. R. & Reddy, K. L. NET gains and losses: the role of changing nuclear envelope proteomes in genome regulation. Curr. Opin. Cell Biol. 28, 105–120 (2014).
Burke, B. & Stewart, C. L. The nuclear lamins: flexibility in function. Nat. Rev. Mol. Cell. Biol. 14, 13–24 (2013).
Beck, M. & Hurt, E. The nuclear pore complex: understanding its function through structural insight. Nat. Rev. Mol. Cell. Biol. 18, 73–89 (2017).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Ou, H. D. et al. ChromEMT: visualizing 3D chromatin structure and compaction in interphase and mitotic cells. Science 357, eaag0025 (2017).
Capelson, M. & Hetzer, M. W. The role of nuclear pores in gene regulation, development and disease. EMBO Rep. 10, 697–705 (2009).
Parada, L. A., McQueen, P. G. & Misteli, T. Tissue-specific spatial organization of genomes. Genome Biol. 5, R44 (2004).
Rego, A., Sinclair, P. B., Tao, W., Kireev, I. & Belmont, A. S. The facultative heterochromatin of the inactive X chromosome has a distinctive condensed ultrastructure. J. Cell Sci. 121, 1119–1127 (2008).
Chen, C.-K. et al. Xist recruits the X chromosome to the nuclear lamina to enable chromosome-wide silencing. Science 354, 468–472 (2016).
Kosak, S. T. et al. Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296, 158–162 (2002).
Blobel, G. Gene gating: a hypothesis. Proc. Natl Acad. Sci. 82, 8527–8529 (1985).
Wandke, C. & Kutay, U. Enclosing chromatin: reassembly of the nucleus after open mitosis. Cell 152, 1222–1225 (2013).
Gerlich, D. et al. Global chromosome positions are transmitted through mitosis in mammalian cells. Cell 112, 751–764 (2003).
Samwer, M. et al. DNA cross-bridging shapes a single nucleus from a set of mitotic chromosomes. Cell 170, 956–972.e23 (2017).
Anderson, D. J., Vargas, J. D., Hsiao, J. P. & Hetzer, M. W. Recruitment of functionally distinct membrane proteins to chromatin mediates nuclear envelope formation in vivo. J. Cell Biol. 186, 183–191 (2009).
Zierhut, C., Jenness, C., Kimura, H. & Funabiki, H. Nucleosomal regulation of chromatin composition and nuclear assembly revealed by histone depletion. Nat. Struct. Mol. Biol. 21, 617–625 (2014).
Vagnarelli, P. et al. Repo-man coordinates chromosomal reorganization with nuclear envelope reassembly during mitotic exit. Dev. Cell 21, 328–342 (2011).
Poleshko, A. et al. The human protein PRR14 tethers heterochromatin to the nuclear lamina during interphase and mitotic exit. Cell Rep. 5, 292–301 (2013).
Solovei, I. et al. LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152, 584–598 (2013). By comparing lamina composition with heterochromatin positioning across species and tissues, this study indicates that either LBR or lamin A/C is required in mammals to tether heterochromatin to the nuclear periphery.
Zullo, J. M. et al. DNA sequence-dependent compartmentalization and silencing of chromatin at the nuclear lamina. Cell 149, 1474–1487 (2012).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Walter, J., Schermelleh, L., Cremer, M., Tashiro, S. & Cremer, T. Chromosome order in HeLa cells changes during mitosis and early G1, but is stably maintained during subsequent interphase stages. J. Cell Biol. 160, 685–697 (2003).
Towbin, B. D. et al. Step-wise methylation of histone H3K9 positions heterochromatin at the nuclear periphery. Cell 150, 934–947 (2012).
Poleshko, A. et al. Genome-nuclear lamina interactions regulate cardiac stem cell lineage restriction. Cell 171, 573–587 (2017). This article presents a beautiful demonstration of a tissue context in which lamin-directed gene repression is essential for differentiation into a functional tissue.
Kumaran, R. I. & Spector, D. L. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J. Cell Biol. 180, 51–65 (2008).
Solovei, I. et al. Nuclear architecture of rod photoreceptor cells adapts to vision in mammalian evolution. Cell 137, 356–368 (2009).
Lottersberger, F., Karssemeijer, R. A., Dimitrova, N. & de Lange, T. 53BP1 and the LINC complex promote microtubule-dependent DSB mobility and DNA repair. Cell 163, 880–893 (2015).
Soutoglou, E. et al. Positional stability of single double-strand breaks in mammalian cells. Nat. Cell Biol. 9, 675–682 (2007).
Dion, V. & Gasser, S. M. Chromatin movement in the maintenance of genome stability. Cell 152, 1355–1364 (2013).
Lucas, J. S., Zhang, Y., Dudko, O. K. & Murre, C. 3D trajectories adopted by coding and regulatory DNA elements: first-passage times for genomic interactions. Cell 158, 339–352 (2014).
Chubb, J. R., Boyle, S., Perry, P. & Bickmore, W. A. Chromatin motion is constrained by association with nuclear compartments in human cells. Curr. Biol. 12, 439–445 (2002).
Therizols, P. et al. Chromatin decondensation is sufficient to alter nuclear organization in embryonic stem cells. Science 346, 1238–1242 (2014).
Canela, A. et al. Genome organization drives chromosome fragility. Cell 170, 507–521.e18 (2017).
Rabut, G., Doye, V. & Ellenberg, J. Mapping the dynamic organization of the nuclear pore complex inside single living cells. Nat. Cell Biol. 6, 1114–1121 (2004).
D’Angelo, M. A., Raices, M., Panowski, S. H. & Hetzer, M. W. Age-dependent deterioration of nuclear pore complexes causes a loss of nuclear integrity in postmitotic cells. Cell 136, 284–295 (2009).
Toyama, B. H. et al. Identification of long-lived proteins reveals exceptional stability of essential cellular structures. Cell 154, 971–982 (2013). This proteomic identification of long-lived proteins identifies several nuclear structures as being extremely long lived.
Lemke, E. A. The multiple faces of disordered nucleoporins. J. Mol. Biol. 428, 2011–2024 (2016).
Tang, S. & Presgraves, D. C. Evolution of the Drosophila nuclear pore complex results in multiple hybrid incompatibilities. Science 323, 779–782 (2009).
Lupu, F., Alves, A., Anderson, K., Doye, V. & Lacy, E. Nuclear pore composition regulates neural stem/progenitor cell differentiation in the mouse embryo. Dev. Cell 14, 831–842 (2008).
Jacinto, F. V., Benner, C. & Hetzer, M. W. The nucleoporin Nup153 regulates embryonic stem cell pluripotency through gene silencing. Genes Dev. 29, 1224–1238 (2015).
Toda, T. et al. Nup153 interacts with Sox2 to enable bimodal gene regulation and maintenance of neural progenitor cells. Cell Stem Cell 21, 618–634.e7 (2017).
Buchwalter, A. L., Liang, Y. & Hetzer, M. W. Nup50 is required for cell differentiation and exhibits transcription-dependent dynamics. Mol. Biol. Cell 25, 2472–2484 (2014).
D’Angelo, M. A., Gomez-Cavazos, J. S., Mei, A., Lackner, D. H. & Hetzer, M. W. A change in nuclear pore complex composition regulates cell differentiation. Dev. Cell 22, 446–458 (2012).
Casolari, J. M. et al. Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117, 427–439 (2004).
Schmid, M. et al. Nup-PI: the nucleopore-promoter interaction of genes in yeast. Mol. Cell 21, 379–391 (2006).
Brickner, J. H. & Walter, P. Gene recruitment of the activated INO1 locus to the nuclear membrane. PLOS Biol. 2, e342 (2004).
Taddei, A. et al. Nuclear pore association confers optimal expression levels for an inducible yeast gene. Nature 441, 774–778 (2006).
Van de Vosse, D. W. et al. A role for the nucleoporin Nup170p in chromatin structure and gene silencing. Cell 152, 969–983 (2013).
Light, W. H., Brickner, D. G., Brand, V. R. & Brickner, J. H. Interaction of a DNA zip code with the nuclear pore complex promotes H2A. Z incorporation and INO1 transcriptional memory. Mol. Cell 40, 112–125 (2010).
Brickner, J. Genetic and epigenetic control of the spatial organization of the genome. Mol. Biol. Cell 28, 364–369 (2017).
Raices, M. et al. Nuclear pores regulate muscle development and maintenance by assembling a localized Mef2C complex. Dev. Cell 41, 540–554 (2017).
Krull, S. et al. Protein Tpr is required for establishing nuclear pore-associated zones of heterochromatin exclusion. EMBO J. 29, 1659–1673 (2010).
Pascual-Garcia, P. et al. Metazoan nuclear pores provide a scaffold for poised genes and mediate induced enhancer-promoter contacts. Mol. Cell 66, 63–76 (2017).
Ibarra, A., Benner, C., Tyagi, S., Cool, J. & Hetzer, M. W. Nucleoporin-mediated regulation of cell identity genes. Genes Dev. 30, 2253–2258 (2016). This study demonstrates that NPCs bind and regulate cell type-specific super-enhancers, which are important regulatory structures in the human genome.
Capelson, M. et al. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 140, 372–383 (2010). This study provides the first evidence that NUPs regulate genes independently of the NPC in metazoans.
Liang, Y., Franks, T. M., Marchetto, M. C., Gage, F. H. & Hetzer, M. W. Dynamic association of NUP98 with the human genome. PLOS Genet. 9, e1003308 (2013).
Vaquerizas, J. M. et al. Nuclear pore proteins Nup153 and megator define transcriptionally active regions in the Drosophila genome. PLOS Genet. 6, e1000846 (2010).
Kalverda, B., Pickersgill, H., Shloma, V. V. & Fornerod, M. Nucleoporins directly stimulate expression of developmental and cell-cycle genes inside the nucleoplasm. Cell 140, 360–371 (2010).
Panda, D. et al. Nup98 promotes antiviral gene expression to restrict RNA viral infection in Drosophila. Proc. Natl Acad. Sci. 111, E3890–E3899 (2014).
Franks, T. M. et al. Nup98 recruits the Wdr82-Set1A/COMPASS complex to promoters to regulate H3K4 trimethylation in hematopoietic progenitor cells. Genes Dev. 31, 2222–2234 (2017).
Light, W. H. et al. A conserved role for human Nup98 in altering chromatin structure and promoting epigenetic transcriptional memory. PLOS Biol. 11, e1001524 (2013).
Kasper, L. H. et al. CREB binding protein interacts with nucleoporin-specific FG repeats that activate transcription and mediate NUP98-HOXA9 oncogenicity. Mol. Cell. Biol. 19, 764–776 (1999).
Pascual-Garcia, P., Jeong, J. & Capelson, M. Nucleoporin Nup98 associates with Trx/MLL and NSL histone-modifying complexes and regulates Hox gene expression. Cell Rep. 9, 433–442 (2014).
Akhtar, A. & Becker, P. B. Activation of transcription through histone H4 acetylation by MOF, an acetyltransferase essential for dosage compensation in Drosophila. Mol. Cell 5, 367–375 (2000).
Gough, S. M., Slape, C. I. & Aplan, P. D. NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood 118, 6247–6257 (2011).
Franks, T. M. & Hetzer, M. W. The role of Nup98 in transcription regulation in healthy and diseased cells. Trends Cell Biol. 23, 112–117 (2013).
Xu, H. et al. NUP98 fusion proteins interact with the NSL and MLL1 complexes to drive leukemogenesis. Cancer Cell 30, 863–878 (2016).
Franks, T. M. et al. Evolution of a transcriptional regulator from a transmembrane nucleoporin. Genes Dev. 30, 1155–1171 (2016). This paper defines a key step in the evolution of a NUP gene — losing functions related to nuclear transport and taking on functions in gene regulation.
Schoenberg, D. R. & Maquat, L. E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 13, 246–259 (2012).
García-Oliver, E., García-Molinero, V. & Rodriguez-Navarro, S. mRNA export and gene expression: the SAGA-TREX-2 connection. Biochim. Biophys. Acta 1819, 555–565 (2012).
Capitanio, J. S., Ben Montpetit & Wozniak, R. W. Human Nup98 regulates the localization and activity of DExH/D-box helicase DHX9. eLife 6, e18825 (2017).
Singer, S. et al. Nuclear pore component Nup98 is a potential tumor suppressor and regulates posttranscriptional expression of select p53 target genes. Mol. Cell 48, 799–810 (2012).
Makise, M. et al. The Nup153-Nup50 protein interface and its role in nuclear import. J. Biol. Chem. 287, 38515–38522 (2012).
Ren, Y., Seo, H.-S., Blobel, G. & Hoelz, A. Structural and functional analysis of the interaction between the nucleoporin Nup98 and the mRNA export factor Rae1. Proc. Natl Acad. Sci. 107, 10406–10411 (2010).
Kirby, T. J. & Lammerding, J. Emerging views of the nucleus as a cellular mechanosensor. Nat. Cell Biol. 103, 1 (2018).
Eckersley-Maslin, M. A., Bergmann, J. H., Lazar, Z. & Spector, D. L. Lamin A/C is expressed in pluripotent mouse embryonic stem cells. Nucleus 4, 53–60 (2013).
Rober, R. A., Weber, K. & Osborn, M. Differential timing of nuclear lamin A/C expression in the various organs of the mouse embryo and the young animal: a developmental study. Development 105, 365–378 (1989).
Coffinier, C. et al. Deficiencies in lamin B1 and lamin B2 cause neurodevelopmental defects and distinct nuclear shape abnormalities in neurons. Mol. Biol. Cell 22, 4683–4693 (2011).
Kim, Y. et al. Mouse B-type lamins are required for proper organogenesis but not by embryonic stem cells. Science 334, 1706–1710 (2011).
Kubben, N. et al. Post-natal myogenic and adipogenic developmental: defects and metabolic impairment upon loss of A-type lamins. Nucleus 2, 195–207 (2011).
Schreiber, K. H. & Kennedy, B. K. When lamins go bad: nuclear structure and disease. Cell 152, 1365–1375 (2013).
van Steensel, B., Delrow, J. & Henikoff, S. Chromatin profiling using targeted DNA adenine methyltransferase. Nat. Genet. 27, 304–308 (2001).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Meuleman, W. et al. Constitutive nuclear lamina-genome interactions are highly conserved and associated with A/T-rich sequence. Genome Res. 23, 270–280 (2013).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Kind, J. et al. Genome-wide maps of nuclear lamina interactions in single human cells. Cell 163, 134–147 (2015). This first analysis of genome association with the lamina in single cells indicates some consistent and some variable features of lamina association.
Harr, J. C. et al. Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J. Cell Biol. 208, 33–52 (2015).
Shevelyov, Y. Y. et al. The B-type lamin is required for somatic repression of testis-specific gene clusters. Proc. Natl Acad. Sci. USA 106, 3282–3287 (2009).
Reddy, K. L., Zullo, J. M., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008).
Akhtar, W. et al. Chromatin position effects assayed by thousands of reporters integrated in parallel. Cell 154, 914–927 (2013).
Pickersgill, H. et al. Characterization of the Drosophila melanogaster genome at the nuclear lamina. Nat. Genet. 38, 1005–1014 (2006).
Kind, J. & van Steensel, B. Stochastic genome-nuclear lamina interactions: modulating roles of Lamin A and BAF. Nucleus 5, 124–130 (2014).
Gesson, K. et al. A-type lamins bind both hetero- and euchromatin, the latter being regulated by lamina-associated polypeptide 2 alpha. Genome Res. 26, 462–473 (2016).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Luperchio, T. R. et al. Chromosome conformation paints reveal the role of lamina association in genome organization and regulation. Preprint at bioRxiv https://doi.org/10.1101/122226 (2017). Using a modified FISH technique, this study provides striking evidence that lamina-associated regions of chromatin are densely compacted at the lamina and that disrupting lamins or chromatin modifiers disrupts compaction and lamina tethering in single cells.
Yuan, J., Simos, G., Blobel, G. & Georgatos, S. D. Binding of lamin A to polynucleosomes. J. Biol. Chem. 266, 9211–9215 (1991).
Taniura, H., Glass, C. & Gerace, L. A chromatin binding site in the tail domain of nuclear lamins that interacts with core histones. J. Cell Biol. 131, 33–44 (1995).
Ye, Q. & Worman, H. J. Primary structure analysis and lamin B and DNA binding of human LBR, an integral protein of the nuclear envelope inner membrane. J. Biol. Chem. 269, 11306–11311 (1994).
Ye, Q. & Worman, H. J. Interaction between an integral protein of the nuclear envelope inner membrane and human chromodomain proteins homologous to Drosophila HP1. J. Biol. Chem. 271, 14653–14656 (1996).
Hirano, Y. et al. Lamin B receptor recognizes specific modifications of histone H4 in heterochromatin formation. J. Biol. Chem. 287, 42654–42663 (2012).
Clowney, E. J. et al. Nuclear aggregation of olfactory receptor genes governs their monogenic expression. Cell 151, 724–737 (2012).
Oldenburg, A. et al. A lipodystrophy-causing lamin A mutant alters conformation and epigenetic regulation of the anti-adipogenic MIR335 locus. J. Cell Biol. 216, 2731–2743 (2017).
Brachner, A. & Foisner, R. Evolvement of LEM proteins as chromatin tethers at the nuclear periphery. Biochem. Soc. Trans. 39, 1735–1741 (2011).
Frock, R. L. et al. Lamin A/C and emerin are critical for skeletal muscle satellite cell differentiation. Genes Dev. 20, 486–500 (2006).
Demmerle, J., Koch, A. J. & Holaska, J. M. Emerin and histone deacetylase 3 (HDAC3) cooperatively regulate expression and nuclear positions of MyoD, Myf5, and Pax7 genes during myogenesis. Chromosome Res. 21, 765–779 (2013).
Malik, P. et al. NET23/STING promotes chromatin compaction from the nuclear envelope. PLOS ONE 9, e111851 (2014).
Robson, M. I. et al. Tissue-specific gene repositioning by muscle nuclear membrane proteins enhances repression of critical developmental genes during myogenesis. Mol. Cell 62, 834–847 (2016).
Somech, R. et al. The nuclear-envelope protein and transcriptional repressor LAP2beta interacts with HDAC3 at the nuclear periphery, and induces histone H4 deacetylation. J. Cell Sci. 118, 4017–4025 (2005).
Yokochi, T. et al. G9a selectively represses a class of late-replicating genes at the nuclear periphery. Proc. Natl Acad. Sci. 106, 19363–19368 (2009).
Bian, Q., Khanna, N., Alvikas, J. & Belmont, A. S. β-Globin cis-elements determine differential nuclear targeting through epigenetic modifications. J. Cell Biol. 203, 767–783 (2013).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Lund, E. G., Duband-Goulet, I., Oldenburg, A., Buendia, B. & Collas, P. Distinct features of lamin A-interacting chromatin domains mapped by ChIP-sequencing from sonicated or micrococcal nuclease-digested chromatin. Nucleus 6, 30–39 (2015).
Naetar, N. et al. Loss of nucleoplasmic LAP2alpha-lamin A complexes causes erythroid and epidermal progenitor hyperproliferation. Nat. Cell Biol. 10, 1341–1348 (2008).
Kochin, V. et al. Interphase phosphorylation of lamin A. J. Cell Sci. 127, 2683–2696 (2014).
Bronshtein, I. et al. Loss of lamin A function increases chromatin dynamics in the nuclear interior. Nat. Commun. 6, 1–9 (2015).
Markiewicz, E. Remodelling of the nuclear lamina and nucleoskeleton is required for skeletal muscle differentiation in vitro. J. Cell Sci. 118, 409–420 (2005).
Vidak, S., Kubben, N., Dechat, T. & Foisner, R. Proliferation of progeria cells is enhanced by lamina-associated polypeptide 2α (LAP2α) through expression of extracellular matrix proteins. Genes Dev. 29, 2022–2036 (2015).
Shumaker, D. K. & Goldman, R. D. Mutant nuclear lamin A leads to progressive alterations of epigenetic control in premature aging. Proc. Natl Acad. Sci. 103, 8703–8708 (2006).
McCord, R. P. et al. Correlated alterations in genome organization, histone methylation, and DNA-lamin A/C interactions in Hutchinson-Gilford progeria syndrome. Genome Res. 23, 260–269 (2013).
Buchwalter, A. & Hetzer, M. W. Nucleolar expansion and elevated protein translation in premature aging. Nat. Commun. 8, 328 (2017).
McStay, B. & Grummt, I. The epigenetics of rRNA genes: from molecular to chromosome biology. Annu. Rev. Cell Dev. Biol. 24, 131–157 (2008).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Toyama, B. H. & Hetzer, M. W. Protein homeostasis: live long, won’t prosper. Nat. Rev. Mol. Cell. Biol. 14, 55–61 (2013).
Rodier, F. & Campisi, J. Four faces of cellular senescence. J. Cell Biol. 192, 547–556 (2011).
Shimi, T. et al. The role of nuclear lamin B1 in cell proliferation and senescence. Genes Dev. 25, 2579–2593 (2011).
Dou, Z. et al. Autophagy mediates degradation of nuclear lamina. Nature 527, 105–109 (2015).
Shah, P. P. et al. Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Genes Dev. 27, 1787–1799 (2013).
Lenain, C. et al. Massive reshaping of genome-nuclear lamina interactions during oncogene-induced senescence. Genome Res. 27, 1634–1644 (2017).
De Cecco, M. et al. Genomes of replicatively senescent cells undergo global epigenetic changes leading to gene silencing and activation of transposable elements. Aging Cell 12, 247–256 (2013).
Ivanov, A. et al. Lysosome-mediated processing of chromatin in senescence. J. Cell Biol. 202, 129–143 (2013).
Dou, Z. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 166, 822 (2017). This study indicates a link between weakening of the nuclear periphery, pro-inflammatory signalling and senescence that implicates declining nuclear function in ageing and cancer.
Bergmann, O. et al. Dynamics of cell generation and turnover in the human heart. Cell 161, 1566–1575 (2015).
Spalding, K. L., Bhardwaj, R. D., Buchholz, B. A., Druid, H. & Frisén, J. Retrospective birth dating of cells in humans. Cell 122, 133–143 (2005).
Mertens, J. et al. Directly reprogrammed human neurons retain aging-associated transcriptomic signatures and reveal age-related nucleocytoplasmic defects. Cell Stem Cell 17, 705–718 (2015).
Savas, J. N., Toyama, B. H., Xu, T., Yates, J. R. & Hetzer, M. W. Extremely long-lived nuclear pore proteins in the rat brain. Science 335, 942–942 (2012).
Gasset-Rosa, F. et al. Polyglutamine-expanded huntingtin exacerbates age-related disruption of nuclear integrity and nucleocytoplasmic transport. Neuron 94, 48–57.e4 (2017).
Zhang, K. et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature 525, 56–61 (2015). This article presents the first indication that NPCs are disrupted by toxic peptides that can cause neurodegenerative disease.
Lee, K.-H. et al. C9orf72 dipeptide repeats impair the assembly, dynamics, and function of membrane-less organelles. Cell 167, 774–788 (2016).
Shi, K. Y. et al. Toxic PRnpoly-dipeptides encoded by theC9orf72repeat expansion block nuclear import and export. Proc. Natl Acad. Sci. USA 114, E1111–E1117 (2017).
Devos, D. P., Gräf, R. & Field, M. C. Evolution of the nucleus. Curr. Opin. Cell Biol. 28, 8–15 (2014).
Goldman, R. D., Gruenbaum, Y., Moir, R. D., Shumaker, D. K. & Spann, T. P. Nuclear lamins: building blocks of nuclear architecture. Genes Dev. 16, 533–547 (2002).
Madhani, H. D. The frustrated gene: origins of eukaryotic gene expression. Cell 155, 744–749 (2013).
Amendola, M. & van Steensel, B. Nuclear lamins are not required for lamina-associated domain organization in mouse embryonic stem cells. EMBO Rep. 16, 610–617 (2015).
Zheng, X., Kim, Y. & Zheng, Y. Identification of lamin B-regulated chromatin regions based on chromatin landscapes. Mol. Biol. Cell 26, 2685–2697 (2015).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R. A. & Feinberg, A. P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat. Genet. 41, 246–250 (2009).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Arai, R. et al. Reduction in chromosome mobility accompanies nuclear organization during early embryogenesis in Caenorhabditis elegans. Sci. Rep. 7, 3631 (2017).
Pasque, V., Miyamoto, K. & Gurdon, J. B. Efficiencies and mechanisms of nuclear reprogramming. Cold Spring Harb. Symp. Quant. Biol. 75, 189–200 (2010).
Simonsson, S. & Gurdon, J. DNA demethylation is necessary for the epigenetic reprogramming of somatic cell nuclei. Nat. Cell Biol. 6, 984–990 (2004).
Sridharan, R. et al. Proteomic and genomic approaches reveal critical functions of H3K9 methylation and heterochromatin protein-1γ in reprogramming to pluripotency. Nat. Cell Biol. 15, 872–882 (2013).
Pasque, V. et al. Histone variant macroH2A marks embryonic differentiation in vivo and acts as an epigenetic barrier to induced pluripotency. J. Cell Sci. 125, 6094–6104 (2012).
Moreira, P. N., Robl, J. M. & Collas, P. Architectural defects in pronuclei of mouse nuclear transplant embryos. J. Cell Sci. 116, 3713–3720 (2003).
Abernathy, D. G. et al. MicroRNAs induce a permissive chromatin environment that enables neuronal subtype-specific reprogramming of adult human fibroblasts. Cell Stem Cell 21, 332–348.e9 (2017).
Ieda, M. et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 142, 375–386 (2010).
Acknowledgements
The authors thank members of the Hetzer laboratory for providing insightful critiques on this article and apologize to those whose work could not be cited owing to space restrictions. This work was supported by US National Institutes of Health grant R01NS096786, the Nomis Foundation and the Glenn Center for Aging Research.
Reviewer information
Nature Reviews Genetics thanks J. H. Brickner, R. Wozniak and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
A.B. and J.M.K. researched data for the article. All authors contributed equally to all other aspects of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- HiC
-
A technique used to study genome organization by identifying chromosomal interactions both in cis and in trans throughout the entire genome.
- Nuclear periphery
-
The outermost region of the nucleus, which includes the nuclear envelope and associated proteins, the nuclear lamina and the nuclear pore complexes.
- X chromosome inactivation
-
Transcriptional silencing of one X chromosome at random in XX female cells for dosage compensation between XX females and XY males.
- Telomeres
-
Repetitive sequences found at the ends of chromosomes for maintenance of genomic integrity.
- Chromatin immunoprecipitation followed by sequencing
-
(ChIP–seq). An assay that combines chromatin immunoprecipitation with DNA sequencing to identify protein–DNA interactions within the genome.
- DNA adenine methyltransferase identification
-
(DamID). An assay that fuses Escherichia coli DNA adenine methyltransferase with a protein of interest to induce adenine methylation in proximity to the fusion protein. Adenine-methylated DNA fragments represent protein-binding sites within the genome.
- Fluorescence in situ hybridization
-
(FISH). A fluorescence microscopy technique used to visualize specific genomic regions within the nucleus using fluorescently tagged DNA probes designed to hybridize to the region of interest.
- Transcriptional memory
-
The state in which genes are poised for rapid transcriptional reactivation after an initial stimulus.
- Satellite DNA
-
Highly repetitive non-coding sequences within heterochromatic regions of the genome.
- Nucleolus
-
A region within the nucleus where ribosomal RNA transcription, processing and assembly occur.
- Senescence
-
The cellular state in which cells permanently exit the cell cycle but do not undergo cell death.
- Autophagy
-
A process in which cellular material is recycled following degradation by the lysosome or vacuole.
Rights and permissions
About this article
Cite this article
Buchwalter, A., Kaneshiro, J.M. & Hetzer, M.W. Coaching from the sidelines: the nuclear periphery in genome regulation. Nat Rev Genet 20, 39–50 (2019). https://doi.org/10.1038/s41576-018-0063-5
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-018-0063-5
This article is cited by
-
Nuclear lamina component KAKU4 regulates chromatin states and transcriptional regulation in the Arabidopsis genome
BMC Biology (2024)
-
Histone FRET reports the spatial heterogeneity in nanoscale chromatin architecture that is imparted by the epigenetic landscape at the level of single foci in an intact cell nucleus
Chromosoma (2024)
-
Heterochromatin rewiring and domain disruption-mediated chromatin compaction during erythropoiesis
Nature Structural & Molecular Biology (2023)
-
Photoselective sequencing: microscopically guided genomic measurements with subcellular resolution
Nature Methods (2023)
-
Understanding the Ultra-Rare Disease Autosomal Dominant Leukodystrophy: an Updated Review on Morpho-Functional Alterations Found in Experimental Models
Molecular Neurobiology (2023)