Abstract
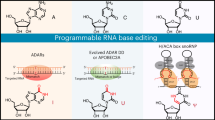
RNA-guided programmable nucleases from CRISPR systems generate precise breaks in DNA or RNA at specified positions. In cells, this activity can lead to changes in DNA sequence or RNA transcript abundance. Base editing is a newer genome-editing approach that uses components from CRISPR systems together with other enzymes to directly install point mutations into cellular DNA or RNA without making double-stranded DNA breaks. DNA base editors comprise a catalytically disabled nuclease fused to a nucleobase deaminase enzyme and, in some cases, a DNA glycosylase inhibitor. RNA base editors achieve analogous changes using components that target RNA. Base editors directly convert one base or base pair into another, enabling the efficient installation of point mutations in non-dividing cells without generating excess undesired editing by-products. In this Review, we summarize base-editing strategies to generate specific and precise point mutations in genomic DNA and RNA, highlight recent developments that expand the scope, specificity, precision and in vivo delivery of base editors and discuss limitations and future directions of base editing for research and therapeutic applications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
19 October 2018
The originally published article contained errors in reference numbering throughout table 1 (DNA base editors and their approximate editing windows) due to the unintended propagation of reference numbering from an earlier version of the table. The article has now been corrected online. The editors apologize for this error.
References
Cohen, S. N., Chang, A. C., Boyer, H. W., Helling, R. B. Construction of biologically functional bacterial plasmids in vitro. Proc. Natl Acad. Sci. USA 70, 3240–3244 (1973).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012). This paper describes the isolation and purification of SpCas9 and the development of Cas9 nickases.
Jansen, R., Embden, J. D., Gaastra, W. & Schouls, L. M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 43, 1565–1575 (2002).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Cho, S. W., Kim, S., Kim, J. M. & Kim, J. S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Sternberg, S. H. & Doudna, J. A. Expanding the biologist’s toolkit with CRISPR-Cas9. Mol. Cell 58, 568–574 (2015).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Bikard, D. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 41, 7429–7437 (2013).
Cheng, A. W. et al. Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell Res. 23, 1163–1171 (2013).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012).
Jeggo, P. A. DNA breakage and repair. Adv. Genet. 38, 185–218 (1998).
Rouet, P., Smih, F. & Jasin, M. Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease. Mol. Cell. Biol. 14, 8096–8106 (1994).
Lukacsovich, T., Yang, D. & Waldman, A. S. Repair of a specific double-strand break generated within a mammalian chromosome by yeast endonuclease I-SceI. Nucleic Acids Res. 22, 5649–5657 (1994).
Rudin, N., Sugarman, E. & Haber, J. E. Genetic and physical analysis of double-strand break repair and recombination in Saccharomyces cerevisiae. Genetics 122, 519–534 (1989).
Rouet, P., Smih, F. & Jasin, M. Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc. Natl Acad. Sci. USA 91, 6064–6068 (1994).
Chapman, J. R., Taylor, M. R. & Boulton, S. J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015). This review highlights the methods and challenges associated with therapeutic genome editing; it was published before the advent of base editing.
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129 (2016).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR-Cas9 leads to large deletions and complex rearrangements. Nat. Biotechnol. 38, 765–771 (2018).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Shin, H. Y. et al. CRISPR/Cas9 targeting events cause complex deletions and insertions at 17 sites in the mouse genome. Nat. Commun. 8, 15464 (2017).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Zhang, L. et al. Large genomic fragment deletions and insertions in mouse using CRISPR/Cas9. PLOS ONE 10, e0120396 (2015).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016). This paper describes the development of the first DNA base editors, the cytosine base editors BE1, BE2 and BE3.
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017). This paper reports the evolution and engineering of the first DNA adenine base editor.
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017). This paper describes improvements to our understanding of the mechanism of cytosine base editing in human cells and describes cytosine base editors with improved efficiency and product purity.
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Li, X. et al. Base editing with a Cpf1-cytidine deaminase fusion. Nat. Biotechnol. 36, 324–327 (2018).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017). This paper describes the development of the first Cas enzyme-guided RNA base editor, which can generate A-to-I mutations in RNA transcripts.
Montiel-Gonzalez, M. F., Vallecillo-Viejo, I., Yudowski, G. A. & Rosenthal, J. J. Correction of mutations within the cystic fibrosis transmembrane conductance regulator by site-directed RNA editing. Proc. Natl Acad. Sci. USA 110, 18285–18290 (2013). This paper describes a genetically encoded RNA base editor capable of generating A-to-I mutations in mRNA of mammalian cells.
Montiel-Gonzalez, M. F., Vallecillo-Viejo, I. C. & Rosenthal, J. J. An efficient system for selectively altering genetic information within mRNAs. Nucleic Acids Res. 44, e157 (2016).
Fukuda, M. et al. Construction of a guide-RNA for site-directed RNA mutagenesis utilising intracellular A-to-I RNA editing. Sci. Rep. 7, 41478 (2017).
Wettengel, J., Reautschnig, P., Geisler, S., Kahle, P. J. & Stafforst, T. Harnessing human ADAR2 for RNA repair — recoding a PINK1 mutation rescues mitophagy. Nucleic Acids Res. 45, 2797–2808 (2017).
Vogel, P., Hanswillemenke, A. & Stafforst, T. Switching protein localization by site-directed RNA editing under control of light. ACS Synth. Biol. 6, 1642–1649 (2017).
Hanswillemenke, A., Kuzdere, T., Vogel, P., Jekely, G. & Stafforst, T. Site-directed RNA editing in vivo can be triggered by the light-driven assembly of an artificial riboprotein. J. Am. Chem. Soc. 137, 15875–15881 (2015).
Vogel, P. & Stafforst, T. Site-directed RNA editing with antagomir deaminases — a tool to study protein and RNA function. ChemMedChem 9, 2021–2025 (2014).
Vogel, P., Schneider, M. F., Wettengel, J. & Stafforst, T. Improving site-directed RNA editing in vitro and in cell culture by chemical modification of the guideRNA. Angew. Chem. Int. Ed. 53, 6267–6271 (2014).
Stafforst, T. & Schneider, M. F. An RNA-deaminase conjugate selectively repairs point mutations. Angew. Chem. Int. Ed. 51, 11166–11169 (2012).
Vogel, P. et al. Efficient and precise editing of endogenous transcripts with SNAP-tagged ADARs. Nat. Methods 15, 535–538 (2018). In this paper, the authors address several of the limitations associated with RNA editing and generate a multiplexable, high-precision A-to-I RNA editor.
Harris, R. S., Petersen-Mahrt, S. K. & Neuberger, M. S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002).
Kunz, C., Saito, Y. & Schar, P. DNA repair in mammalian cells — mismatched repair: variations on a theme. Cell. Mol. Life Sci. 66, 1021–1038 (2009).
Pearl, L. H. Structure and function in the uracil-DNA glycosylase superfamily. Mutat. Res. 460, 165–181 (2000).
Mol, C. D. et al. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: protein mimicry of DNA. Cell 82, 701–708 (1995).
Tang, W. & Liu, D. R. Rewritable multi-event analog recording in bacterial and mammalian cells. Science 360, eaap8992 (2018).
Krokan, H. E., Drablos, F. & Slupphaug, G. Uracil in DNA — occurrence, consequences and repair. Oncogene 21, 8935–8948 (2002).
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Yasui, M. et al. Miscoding properties of 2΄-deoxyinosine, a nitric oxide-derived DNA Adduct, during translesion synthesis catalyzed by human DNA polymerases. J. Mol. Biol. 377, 1015–1023 (2008).
Losey, H. C., Ruthenburg, A. J. & Verdine, G. L. Crystal structure of Staphylococcus aureus tRNA adenosine deaminase TadA in complex with RNA. Nat. Struct. Mol. Biol. 13, 153–159 (2006).
Vogel, P. & Stafforst, T. Critical review on engineering deaminases for site-directed RNA editing. Curr. Opin. Biotechnol. 55, 74–80 (2018).
Bass, B. L. & Weintraub, H. An unwinding activity that covalently modifies its double-stranded RNA substrate. Cell 55, 1089–1098 (1988).
Matthews, M. M. et al. Structures of human ADAR2 bound to dsRNA reveal base-flipping mechanism and basis for site selectivity. Nat. Struct. Mol. Biol. 23, 426–433 (2016).
Yu, Y. T. et al. Internal modification of U2 small nuclear (sn)RNA occurs in nucleoli of Xenopus oocytes. J. Cell Biol. 152, 1279–1288 (2001).
Bass, B. L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Bazak, L. et al. A-To-I RNA editing occurs at over a hundred million genomic sites, located in a majority of human genes. Genome Res. 24, 365–376 (2014).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Schneider, M. F., Wettengel, J., Hoffmann, P. C. & Stafforst, T. Optimal guideRNAs for re-directing deaminase activity of hADAR1 and hADAR2 in trans. Nucleic Acids Res. 42, e87 (2014).
Herbert, A. & Rich, A. The role of binding domains for dsRNA and Z-DNA in the in vivo editing of minimal substrates by ADAR1. Proc. Natl Acad. Sci. USA 98, 12132–12137 (2001).
Lehmann, K. A. & Bass, B. L. Double-stranded RNA adenosine deaminases ADAR1 and ADAR2 have overlapping specificities. Biochemistry 39, 12875–12884 (2000).
Kuttan, A. & Bass, B. L. Mechanistic insights into editing-site specificity of ADARs. Proc. Natl Acad. Sci. USA 109, E3295–E3304 (2012).
Crick, F. H. Codon—anticodon pairing: the wobble hypothesis. J. Mol. Biol. 19, 548–555 (1966).
Kume, H., Hino, K., Galipon, J. & Ui-Tei, K. A-To-I editing in the miRNA seed region regulates target mRNA selection and silencing efficiency. Nucleic Acids Res. 42, 10050–10060 (2014).
Seeburg, P. H. & Hartner, J. Regulation of ion channel/neurotransmitter receptor function by RNA editing. Curr. Opin. Neurobiol. 13, 279–283 (2003).
Yang, J. H., Sklar, P., Axel, R. & Maniatis, T. Editing of glutamate receptor subunit B pre-mRNA in vitro by site-specific deamination of adenosine. Nature 374, 77–81 (1995).
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLOS Biol. 2, e391 (2004).
Hess, G. T. et al. Directed evolution using dCas9-targeted somatic hypermutation in mammalian cells. Nat. Methods 13, 1036–1042 (2016).
Kim, K. et al. Highly efficient RNA-guided base editing in mouse embryos. Nat. Biotechnol. 35, 435–437 (2017).
Ma, Y. et al. Targeted AID-mediated mutagenesis (TAM) enables efficient genomic diversification in mammalian cells. Nat. Methods 13, 1029–1035 (2016).
Wang, L. et al. Enhanced base editing by co-expression of free uracil DNA glycosylase inhibitor. Cell Res. 27, 1289–1292 (2017).
Di Noia, J. & Neuberger, M. S. Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature 419, 43–48 (2002).
Radany, E. H. et al. Increased spontaneous mutation frequency in human cells expressing the phage PBS2-encoded inhibitor of uracil-DNA glycosylase. Mutat. Res. 461, 41–58 (2000).
Hess, G. T., Tycko, J., Yao, D. & Bassik, M. C. Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell 68, 26–43 (2017).
Ryu, S. M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Hua, K., Tao, X., Yuan, F., Wang, D. & Zhu, J. K. Precise A·T to G·C base editing in the rice genome. Mol. Plant 11, 627–630 (2018).
Yan, F. et al. Highly efficient A·T to G·C base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol. Plant 11, 631–634 (2018).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Lau, A. Y., Wyatt, M. D., Glassner, B. J., Samson, L. D. & Ellenberger, T. Molecular basis for discriminating between normal and damaged bases by the human alkyladenine glycosylase, AAG. Proc. Natl Acad. Sci. USA 97, 13573–13578 (2000).
Kouzminova, E. A. & Kuzminov, A. Patterns of chromosomal fragmentation due to uracil-DNA incorporation reveal a novel mechanism of replication-dependent double-stranded breaks. Mol. Microbiol. 68, 202–215 (2008).
Liu, Z. et al. Highly efficient RNA-guided base editing in rabbit. Nat. Commun. 9, 2717 (2018).
Yeh, W. H., Chiang, H., Rees, H. A., Edge, A. S. B. & Liu, D. R. In vivo base editing of post-mitotic sensory cells. Nat. Commun. 9, 2184 (2018).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Kang, B. C. et al. Precision genome engineering through adenine base editing in plants. Nat. Plants 4, 427–431 (2018).
Rees, H. A. et al. Improving the DNA specificity and applicability of base editing through protein engineering and protein delivery. Nat. Commun. 8, 15790 (2017).
Kim, D. et al. Genome-wide target specificities of CRISPR RNA-guided programmable deaminases. Nat. Biotechnol. 35, 475–480 (2017).
Kleinstiver, B. P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Lee, J. K. et al. Directed evolution of CRISPR-Cas9 to increase its specificity. Nat. Commun. 9, 3048 (2018).
Yamanaka, S. et al. Apolipoprotein B mRNA-editing protein induces hepatocellular carcinoma and dysplasia in transgenic animals. Proc. Natl Acad. Sci. USA 92, 8483–8487 (1995).
Okazaki, I. M. et al. Constitutive expression of AID leads to tumorigenesis. J. Exp. Med. 197, 1173–1181 (2003).
Burns, M. B. et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494, 366–370 (2013).
Liu, Z. et al. Efficient generation of mouse models of human diseases via ABE- and BE-mediated base editing. Nat. Commun. 9, 2338 (2018).
Gehrke, J. M. et al. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. https://doi.org/10.1038/nbt.4199 (2018).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Kim, E. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 8, 14500 (2017).
Muller, M. et al. Streptococcus thermophilus CRISPR-Cas9 systems enable specific editing of the human genome. Mol. Ther. 24, 636–644 (2016).
Lee, C. M., Cradick, T. J. & Bao, G. The Neisseria meningitidis CRISPR-Cas9 system enables specific genome editing in mammalian cells. Mol. Ther. 24, 645–654 (2016).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Jiang, W. et al. BE-PLUS: a new base editing tool with broadened editing window and enhanced fidelity. Cell Res. 28, 855–861 (2018).
Kim, S., Bae, T., Hwang, J. & Kim, J. S. Rescue of high-specificity Cas9 variants using sgRNAs with matched 5’ nucleotides. Genome Biol. 18, 218 (2017).
Nishimasu, H. et al. Engineered CRISPR-Cas9 nuclease with expanded targeting space. Science https://doi.org/10.1126/science.aas9129 (2018).
Hua, K., Tao, X. & Zhu, J. K. Expanding the base editing scope in rice by using Cas9 variants. Plant Biotechnol. J. https://doi.org/10.1111/pbi.12993 (2018).
Yang, L. et al. Increasing targeting scope of adenosine base editors in mouse and rat embryos through fusion of TadA deaminase with Cas9 variants. Protein Cell 9, 814–819 (2018).
Wang, X. et al. Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion. Nat. Biotechnol. https://doi.org/10.1038/nbt.4198 (2018).
Pinello, L. et al. Analyzing CRISPR genome-editing experiments with CRISPResso. Nat. Biotechnol. 34, 695–697 (2016).
Clement, K. et al. Analysis and comparison of genome editing using CRISPResso2. Preprint at https://www.biorxiv.org/content/early/2018/08/15/392217 (2018).
Suzuki, K. et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Wheeler, L. C., Lim, S. A., Marqusee, S. & Harms, M. J. The thermostability and specificity of ancient proteins. Curr. Opin. Struct. Biol. 38, 37–43 (2016).
Zafra, M. P. et al. Optimized base editors enable efficient editing in cells, organoids and mice. Nat. Biotechnol. 36, (888–893 (2018).
Midoux, P., Pichon, C., Yaouanc, J. J. & Jaffres, P. A. Chemical vectors for gene delivery: a current review on polymers, peptides and lipids containing histidine or imidazole as nucleic acids carriers. Br. J. Pharmacol. 157, 166–178 (2009).
Bodles-Brakhop, A. M., Heller, R. & Draghia-Akli, R. Electroporation for the delivery of DNA-based vaccines and immunotherapeutics: current clinical developments. Mol. Ther. 17, 585–592 (2009).
Thomas, C. E., Ehrhardt, A. & Kay, M. A. Progress and problems with the use of viral vectors for gene therapy. Nat. Rev. Genet. 4, 346–358 (2003).
Zuris, J. A. et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat. Biotechnol. 33, 73–80 (2015).
Wang, M. et al. Efficient delivery of genome-editing proteins using bioreducible lipid nanoparticles. Proc. Natl Acad. Sci. USA 113, 2868–2873 (2016).
Kim, S., Kim, D., Cho, S. W., Kim, J. & Kim, J. S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Miao, C. H. et al. Nonrandom transduction of recombinant adeno-associated virus vectors in mouse hepatocytes in vivo: cell cycling does not influence hepatocyte transduction. J. Virol. 74, 3793–3803 (2000).
Dong, J. Y., Fan, P. D. & Frizzell, R. A. Quantitative analysis of the packaging capacity of recombinant adeno-associated virus. Hum. Gene Ther. 7, 2101–2112 (1996).
Lai, Y. et al. Efficient in vivo gene expression by trans-splicing adeno-associated viral vectors. Nat. Biotechnol. 23, 1435–1439 (2005).
Gao, X. et al. Treatment of autosomal dominant hearing loss by in vivo delivery of genome editing agents. Nature 553, 217–221 (2018).
Park, D. S. et al. Targeted base editing via RNA-guided cytidine deaminases in Xenopus laevis embryos. Mol. Cells 40, 823–827 (2017).
Liang, P. et al. Effective gene editing by high-fidelity base editor 2 in mouse zygotes. Protein Cell 8, 601–611 (2017).
Li, G. et al. Highly efficient and precise base editing in discarded human tripronuclear embryos. Protein Cell 8, 776–779 (2017).
Liang, P. et al. Correction of beta-thalassemia mutant by base editor in human embryos. Protein Cell 8, 811–822 (2017).
Ma, Y. et al. Highly efficient and precise base editing by engineered dCas9-guide tRNA adenosine deaminase in rats. Cell Discov. 4, 39 (2018).
Zhang, Y. et al. Programmable base editing of zebrafish genome using a modified CRISPR-Cas9 system. Nat. Commun. 8, 118 (2017).
Tanaka, S. et al. In vivo targeted single-nucleotide editing in zebrafish. Sci. Rep. 8, 11423 (2018).
Schenk, B. et al. MPDU1 mutations underlie a novel human congenital disorder of glycosylation, designated type If. J. Clin. Invest. 108, 1687–1695 (2001).
Zeng, Y. et al. Correction of the Marfan syndrome pathogenic FBN1 mutation by base editing in human cells and heterozygous embryos. Mol. Ther. https://doi.org/10.1016/j.ymthe.2018.08.007 (2018).
Chadwick, A. C., Wang, X. & Musunuru, K. In vivo base editing of PCSK9 (proprotein convertase subtilisin/kexin type 9) as a therapeutic alternative to genome editing. Arterioscler. Thromb. Vasc. Biol. 37, 1741–1747 (2017).
Roccio, M., Hahnewald, S., Perny, M. & Senn, P. Cell cycle reactivation of cochlear progenitor cells in neonatal FUCCI mice by a GSK3 small molecule inhibitor. Sci. Rep. 5, 17886 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Kuscu, C. et al. CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations. Nat. Methods 14, 710–712 (2017).
Billon, P. et al. CRISPR-mediated base editing enables efficient disruption of eukaryotic genes through induction of STOP codons. Mol. Cell 67, 1068–1079 (2017).
Haapaniemi, E., Botla, S., Persson, J., Schmierer, B. & Taipale, J. CRISPR-Cas9 genome editing induces a p53-mediated DNA damage response. Nat. Med. 24, 927–930 (2018).
Ihry, R. J. et al. p53 inhibits CRISPR-Cas9 engineering in human pluripotent stem cells. Nat. Med. 24, 939–946 (2018).
Aguirre, A. J. et al. Genomic copy number dictates a gene-independent cell response to CRISPR/Cas9 targeting. Cancer Discov. 6, 914–929 (2016).
Roukos, V. & Misteli, T. The biogenesis of chromosome translocations. Nat. Cell Biol. 16, 293–300 (2014).
Gapinske, M. et al. CRISPR-SKIP: programmable gene splicing with single base editors. Genome Biol. 19, 107 (2018).
Hur, J. K. et al. Targeted mutagenesis in mice by electroporation of Cpf1 ribonucleoproteins. Nat. Biotechnol. 34, 807–808 (2016).
Sung, Y. H. et al. Knockout mice created by TALEN-mediated gene targeting. Nat. Biotechnol. 31, 23–24 (2013).
Sung, Y. H. et al. Highly efficient gene knockout in mice and zebrafish with RNA-guided endonucleases. Genome Res. 24, 125–131 (2014).
Farzadfard, F. & Lu, T. K. Emerging applications for DNA writers and molecular recorders. Science 361, 870–875 (2018).
Tang, W., Hu, J. H. & Liu, D. R. Aptazyme-embedded guide RNAs enable ligand-responsive genome editing and transcriptional activation. Nat. Commun. 8, 15939 (2017).
Farzadfard, F. et al. Single-nucleotide-resolution computing and memory in living cells. Preprint at https://www.biorxiv.org/content/early/2018/02/15/263657 (2018).
Yin, K., Gao, C. & Qiu, J. L. Progress and prospects in plant genome editing. Nat. Plants 3, 17107 (2017).
Voytas, D. F. & Gao, C. Precision genome engineering and agriculture: opportunities and regulatory challenges. PLOS Biol. 12, e1001877 (2014).
Svitashev, S. et al. Targeted mutagenesis, precise gene editing, and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol. 169, 931–945 (2015).
Endo, M., Mikami, M. & Toki, S. Biallelic gene targeting in rice. Plant Physiol. 170, 667–677 (2016).
Zong, Y. et al. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 35, 438–440 (2017).
Shimatani, Z. et al. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 35, 441–443 (2017).
Li, C. et al. Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol. 19, 59 (2018).
Acknowledgements
D.R.L. acknowledges support from Defense Advanced Research Projects Agency (DARPA) HR0011-17-2-0049; the Ono Pharma Foundation; US National Institutes of Health (NIH) RM1 HG009490, R01 EB022376, U01 AI142756 and R35 GM118062; and Howard Hughes Medical Institute (HHMI). H.A.R. is supported by the Kilpatrick Educational fund from the Chemistry and Chemical Biology Department, Harvard University. The authors thank J.K. Joung, F. Zhang, A. Raguram, W.-H. Yeh, T. Huang, K. Zhao and W. Tang for their helpful comments.
Author information
Authors and Affiliations
Contributions
Both authors contributed to all aspects of the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.R.L. is a consultant and co-founder of Beam Therapeutics, Editas Medicine and Pairwise Plants, which are all companies that use genome-editing technologies. H.A.R. declares no competing financial interests. Both authors have no competing non-financial interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Guide RNA
-
Short RNA sequence comprising a scaffold for binding to the necessary CRISPR-associated (Cas) enzyme and a variable spacer region that defines the target site for the enzyme. In natural CRISPR systems, the guide RNA is often made of two molecules of RNA with complementarity. Engineered ‘single-guide’ RNAs connecting the two natural guide RNA components are often accepted by Cas enzymes.
- Protospacer adjacent motif
-
(PAM). A small region of nucleotides in the target DNA sequence adjacent to the sequence specified by a guide RNA. The PAM is not specified in the guide RNA, but CRISPR-associated (Cas) enzymes do not bind or cleave a sequence unless they are next to the appropriate PAM.
- Cas9 nickase
-
A catalytically disabled mutant of a Cas9 enzyme that is able to create a single-stranded DNA break but not a double-stranded DNA break.
- Activity window
-
The region of DNA or RNA, typically defined by the number of nucleotides from the protospacer adjacent motif (PAM), in which a particular base editor acts to induce efficient point mutations. The activity window for most base editors is approximately four to five nucleotides wide.
- Protospacer
-
A region in a guide RNA of 15–25 nucleotides in length that specifies the target RNA or DNA locus.
- Proximal off-target editing
-
Unwanted editing of bases that occurs outside of the activity window but is found nearby (for example, 100 nucleotides upstream or downstream of) the target site.
- Distal off-target editing
-
Unwanted editing of bases residing in locations of the genome or transcriptome unrelated to (for example, >100 nucleotides away from) the target site of the base editor.
- Cas13b
-
A class 2, type VI RNA-guided RNase from the CRISPR system. Variants from several species have been characterized. It catalyses site-specific cleavage of single-stranded RNA.
- Wobble position
-
The third nucleotide in a codon.
- Base-editing product purity
-
The term used to describe the spectrum of mutations induced by a particular base-editing technology. Low product purity occurs when a target base is mutated to bases other than the desired point mutation or when small insertions or deletions are generated in addition to the desired edit; for example, C-to-G or C-to-A edits, rather than the desired C-to-T edit, from a cytosine base editor.
- SpCas9
-
An RNA-guided endonuclease variant isolated from the CRISPR system of Streptococcus pyogenes. It catalyses site-specific cleavage of double-stranded DNA at sites with an NGG protospacer adjacent motif (PAM).
- SaCas9
-
An RNA-guided endonuclease variant isolated from the CRISPR system of Staphylococcus aureus. It catalyses site-specific cleavage of double-stranded DNA at sites with an NNGRRT protospacer adjacent motif (PAM).
- Bystander editing
-
Editing of a non-target base that resides in the activity window of a particular base editor and guide RNA. Bystander editing occurs in addition to editing of the target base.
- Cas12a
-
A class 2, type V RNA-guided endonuclease from the CRISPR system. Variants from several species have been characterized. It catalyses site-specific cleavage of double-stranded DNA at sites with a TTTV protospacer adjacent motif (where V is A, C or G).
- Mosaicism
-
A state in which two or more cell populations with distinct genotypes are present in the same organism and derived from a single fertilized egg.
Rights and permissions
About this article
Cite this article
Rees, H.A., Liu, D.R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat Rev Genet 19, 770–788 (2018). https://doi.org/10.1038/s41576-018-0059-1
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-018-0059-1
This article is cited by
-
Technologies of gene editing and related clinical trials for the treatment of genetic and acquired diseases: a systematic review
Egyptian Journal of Medical Human Genetics (2024)
-
Application of genome editing techniques to regulate gene expression in crops
BMC Plant Biology (2024)
-
RNA editing enzymes: structure, biological functions and applications
Cell & Bioscience (2024)
-
Adenine transversion editors enable precise, efficient A•T-to-C•G base editing in mammalian cells and embryos
Nature Biotechnology (2024)
-
Efficient prime editing in mouse brain, liver and heart with dual AAVs
Nature Biotechnology (2024)