Abstract
Identifying and validating molecular targets of interventions that extend the human health span and lifespan has been difficult, as most clinical biomarkers are not sufficiently representative of the fundamental mechanisms of ageing to serve as their indicators. In a recent breakthrough, biomarkers of ageing based on DNA methylation data have enabled accurate age estimates for any tissue across the entire life course. These ‘epigenetic clocks’ link developmental and maintenance processes to biological ageing, giving rise to a unified theory of life course. Epigenetic biomarkers may help to address long-standing questions in many fields, including the central question: why do we age?
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Baker, G. & Sprott, R. Biomarkers of aging. Exp. Gerontol. 23, 223–239 (1988).
Warner, H. R. The future of aging interventions. J. Gerontol. A Biol Sci. Med. Sci. 59, B692–B696 (2004).
Laird, P. W. Principles and challenges of genome-wide DNA methylation analysis. Nat. Rev. Genet. 11, 191–203 (2010).
Tibshirani, R. Regression shrinkage and selection via the Lasso. J. Royal Stat. Soc. B 58, 267–288 (1996).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. Royal Stat. Soc. B 67, 301–320 (2005).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
The Cancer Genome Atlas Research Netwok et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013). This paper presents the first multi-tissue DNAm age estimator that applies to all sources of DNA (except sperm) and to the entire life course (from prenatal samples to centenarians).
Horvath, S. et al. Accelerated epigenetic aging in Down syndrome. Aging Cell 14, 491–495 (2015). This is the first study to demonstrate that Down syndrome is accompanied by strong epigenetic age acceleration in brain and blood tissue.
Maierhofer, A. et al. Accelerated epigenetic aging in Werner syndrome. Aging 9, 1143–1152 (2017).
Horvath, S. et al. An epigenetic clock analysis of race/ethnicity, sex, and coronary heart disease. Genome Biol. 17, 171 (2016).
Levine, M. E. et al. Menopause accelerates biological aging. Proc. Natl Acad. Sci. USA 113, 9327–9332 (2016).
Quach, A. et al. Epigenetic clock analysis of diet, exercise, education, and lifestyle factors. Aging 9, 419–446 (2017). This is the largest study (n>4,500) to date to evaluate the effect of lifestyle factors (diet, education, exercise and clinical biomarkers) on epigenetic ageing rates.
Huh, C. J. et al. Maintenance of age in human neurons generated by microRNA-based neuronal conversion of fibroblasts. eLife 5, e18648 (2016). This is the first study to show that neurons resulting from direction conversion (transdifferentiation) of fibroblasts maintain the DNAm age of the fibroblast, which is in stark contrast to an iPS procedure.
Jylhävä, J., Pedersen, N. L. & Hägg, S. Biological age predictors. EBioMedicine 21, 29–36 (2017).
Lee, H. Y., Lee, S. D. & Shin, K.-J. Forensic DNA methylation profiling from evidence material for investigative leads. BMB Rep. 49, 359–369 (2016).
Berdyshev, G., Korotaev, G., Boiarskikh, G. & Vaniushin, B. Nucleotide composition of DNA and RNA from somatic tissues of humpback and its changes during spawning. Biokhimiia 31, 88–993 (1967).
Ahuja, N., Li, Q., Mohan, A. L., Baylin, S. B. & Issa, J. P. Aging and DNA methylation in colorectal mucosa and cancer. Cancer Res. 58, 5489–5494 (1998).
Rakyan, V. K. et al. Human aging-associated DNA hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res. 20, 434–439 (2010).
Teschendorff, A. E. et al. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 20, 440–446 (2010). This is the first study to show that one can define a signature of CpGs (near stem cell Polycomb group protein targets) for which the age-related gain in DNA methylation can be observed in multiple tissues.
Hernandez, D. et al. Distinct DNA methylation changes highly correlated with chronological age in the human brain. Hum. Mol. Genet. 20, 1164–1172 (2011).
Bell, J. T. et al. Epigenome-wide scans identify differentially methylated regions for age and age-related phenotypes in a healthy ageing population. PLoS Genet. 8, e1002629 (2012).
Christensen, B. et al. Aging and environmental exposures alter tissue-specific DNA methylation dependent upon CpG island context. PLoS Genet. 5, e1000602 (2009).
Maegawa, S. et al. Widespread and tissue specific age-related DNA methylation changes in mice. Genome Res. 20, 332–340 (2010).
Jung, M. & Pfeifer, G. P. Aging and DNA methylation. BMC Biol. 13, 7 (2015).
Fraga, M. F. & Esteller, M. Epigenetics and aging: the targets and the marks. Trends Genet. 23, 413–418 (2007).
Fraga, M. F., Agrelo, R. & Esteller, M. Cross-talk between aging and cancer. Ann. NY Acad. Sci. 1100, 60–74 (2007).
Bollati, V. et al. Decline in genomic DNA methylation through aging in a cohort of elderly subjects. Mech. Ageing Dev. 130, 234–239 (2009).
Mugatroyd, C., Wu, Y., Bockmühl, Y. & Spengler, D. The Janus face of DNA methylation in aging. Aging 2, 107–110 (2010).
Rodríguez-Rodero, S., Fernández-Morera, J., Fernandez, A., Menéndez-Torre, E. & Fraga, M. Epigenetic regulation of aging. Discov. Med. 10, 225–233 (2010).
Horvath, S. et al. Aging effects on DNA methylation modules in human brain and blood tissue. Genome Biol. 13, R97 (2012).
Zheng, S. C., Widschwendter, M. & Teschendorff, A. E. Epigenetic drift, epigenetic clocks and cancer risk. Epigenomics 8, 705–719 (2016).
Bocklandt, S. et al. Epigenetic predictor of age. PLoS ONE 6, e14821 (2011). This paper presents the first DNAm age estimator (that is, a mathematical algorithm for estimating the chronological age of a person on the basis of (saliva) methylation data).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013). This article describes a highly accurate and widely used DNAm age estimator for blood, which is highly correlated with age in many other tissues.
Simpkin, A. J. et al. Prenatal and early life influences on epigenetic age in children: a study of mother-offspring pairs from two cohort studies. Hum. Mol. Genet. 25, 191–201 (2016).
Simpkin, A. J. et al. The epigenetic clock and physical development during childhood and adolescence: longitudinal analysis from a UK birth cohort. Int. J. Epidemiol. 46, 549–558 (2017).
Marioni, R. et al. DNA methylation age of blood predicts all-cause mortality in later life. Genome Biol. 16, 25 (2015). This is the first study to demonstrate that epigenetic age acceleration in blood predicts lifespan even after adjusting for other risk factors.
Chen, B. H. et al. DNA methylation-based measures of biological age: meta-analysis predicting time to death. Aging 8, 1844–1865 (2016). This study extends the findings of reference 37 to several ethnic groups.
Koch, C. & Wagner, W. Epigenetic-aging-signature to determine age in different tissues. Aging 3, 1018–1027 (2011).
Garagnani, P. et al. Methylation of ELOVL2 gene as a new epigenetic marker of age. Aging Cell 11, 1132–1134 (2012). This article demonstrates that chronological age is highly correlated with a single CpG in the ELOVL2 gene (r = 0.92).
Weidner, C. I. et al. Aging of blood can be tracked by DNA methylation changes at just three CpG sites. Genome Biol. 15, R24 (2014).
Bekaert, B., Kamalandua, A., Zapico, S. C., Van de Voorde, W. & Decorte, R. Improved age determination of blood and teeth samples using a selected set of DNA methylation markers. Epigenetics 10, 922–930 (2015).
Hamano, Y. et al. Forensic age prediction for dead or living samples by use of methylation-sensitive high resolution melting. Leg. Med. 21, 5–10 (2016).
Zbiec-Piekarska, R. et al. Development of a forensically useful age prediction method based on DNA methylation analysis. Forensic Sci. Int. Genet. 17, 173–179 (2015).
Bormann, F. et al. Reduced DNA methylation patterning and transcriptional connectivity define human skin aging. Aging Cell 15, 563–571 (2016).
Florath, I., Butterbach, K., Muller, H., Bewerunge-Hudler, M. & Brenner, H. Cross-sectional and longitudinal changes in DNA methylation with age: an epigenome-wide analysis revealing over 60 novel age-associated CpG sites. Hum. Mol. Genet. 23, 1186–1201 (2014).
Lee, H. Y. et al. Epigenetic age signatures in the forensically relevant body fluid of semen: a preliminary study. Forensic Sci. Int. Genet. 19, 28–34 (2015).
Lin, Q. et al. DNA methylation levels at individual age-associated CpG sites can be indicative for life expectancy. Aging 8, 394–401 (2016).
Li, S. et al. Genetic and environmental causes of variation in the difference between biological age based on DNA methylation and chronological age for middle-aged women. Twin Res. Hum. Genet. 18, 720–726 (2015).
Bacalini, M. G. et al. Systemic age-associated DNA hypermethylation of ELOVL2 gene: in vivo and in vitro evidences of a cell replication process. J. Gerontol. A Biol Sci. Med. Sci. 72, 1015–1023 (2017). This study extends the findings from reference 40 from blood to other tissues.
Bernstein, B. E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Illingworth, R. et al. A novel CpG island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol. 6, e22 (2008).
Li, Y. et al. The DNA methylome of human peripheral blood mononuclear cells. PLoS Biol. 8, e1000533 (2010).
Thompson, R. F. et al. Tissue-specific dysregulation of DNA methylation in aging. Aging Cell 9, 506–518 (2010).
Vidaki, A., Daniel, B. & Court, D. S. Forensic DNA methylation profiling–potential opportunities and challenges. Forensic Sci. Int. Genet. 7, 499–507 (2013).
Horvath, S. et al. The cerebellum ages slowly according to the epigenetic clock. Aging 7, 294–306 (2015).
Sehl, M. E., Henry, J. E., Storniolo, A. M., Ganz, P. A. & Horvath, S. DNA methylation age is elevated in breast tissue of healthy women. Breast Cancer Res. Treat. 164, 209–219 (2017).
Jenkins, T. G., Aston, K. I. & Carrell, D. T. Germ line aging and regional epigenetic instability: age prediction using human sperm DNA methylation signatures. Preprint at bioRxiv https://doi.org/10.1101/220764 (2017).
Levine, M. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (US Albany) 2018. Preprint at bioRxiv https://doi.org/10.1101/276162 (2018).
Carroll, J. E. et al. Epigenetic aging and immune senescence in women with insomnia symptoms: findings from the Women’s Health Initiative study. Biol. Psychiatry 81, 136–144 (2017).
Marioni, R. E. et al. The epigenetic clock is correlated with physical and cognitive fitness in the Lothian Birth Cohort 1936. Int. J. Epidemiol. 44, 1388–1396 (2015).
Levine, M. E., Lu, A. T., Bennett, D. A. & Horvath, S. Epigenetic age of the pre-frontal cortex is associated with neuritic plaques, amyloid load, and Alzheimer’s disease related cognitive functioning. Aging 7, 1198–1211 (2015).
Horvath, S. & Ritz, B. R. Increased epigenetic age and granulocyte counts in the blood of Parkinson’s disease patients. Aging 7, 1130–1142 (2015).
Breitling, L. P. et al. Frailty is associated with the epigenetic clock but not with telomere length in a German cohort. Clin. Epigenet. 8, 21 (2016).
Horvath, S. et al. Decreased epigenetic age of PBMCs from Italian semi-supercentenarians and their offspring. Aging 7, 1159–1170 (2015).
Levine, M. E. et al. DNA methylation age of blood predicts future onset of lung cancer in the women’s health initiative. Aging 7, 690–700 (2015).
Zheng, Y. et al. Blood epigenetic age may predict cancer incidence and mortality. EBioMedicine 5, 68–73 (2016).
Ambatipudi, S. et al. DNA methylome analysis identifies accelerated epigenetic ageing associated with postmenopausal breast cancer susceptibility. Eur. J. Cancer 75, 299–307 (2017).
Durso, D. F. et al. Acceleration of leukocytes’ epigenetic age as an early tumor and sex-specific marker of breast and colorectal cancer. Oncotarget 8, 23237–23245 (2017).
Hodgson, K. et al. Epigenetic age acceleration assessed with human white-matter images. J. Neurosci. 37, 4735 (2017).
Raina, A. et al. Cerebral white matter hyperintensities on MRI and acceleration of epigenetic aging: the atherosclerosis risk in communities study. Clin. Epigenet. 9, 21 (2017).
Degerman, S. et al. Maintained memory in aging is associated with young epigenetic age. Neurobiol. Aging 55, 167–171 (2017).
Christiansen, L. et al. DNA methylation age is associated with mortality in a longitudinal Danish twin study. Aging Cell 15, 149–154 (2016).
Perna, L. et al. Epigenetic age acceleration predicts cancer, cardiovascular, and all-cause mortality in a German case cohort. Clin. Epigenet. 8, 64 (2016).
Dugue, P. A. et al. Association of DNA methylation-based biological age with health risk factors, and overall and cause-specific mortality. Am. J. Epidemiol. https://doi.org/10.1093/aje/kwx291 (2017).
Pourquie, O. The segmentation clock: converting embryonic time into spatial pattern. Science 301, 328–330 (2003).
Kruse, K. & Jülicher, F. Oscillations in cell biology. Curr. Opin. Cell Biol. 17, 20–26 (2005).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Oh, G. et al. Cytosine modifications exhibit circadian oscillations that are involved in epigenetic diversity and aging. Nat. Commun. 9, 644 (2018).
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
Yu, B. et al. Genome-wide, single-cell DNA methylomics reveals increased non-CpG methylation during human oocyte maturation. Stem Cell Rep. 9, 397–407 (2017).
Kelsey, G., Stegle, O. & Reik, W. Single-cell epigenomics: Recording the past and predicting the future. Science 358, 69 (2017).
Blattler, A. & Farnham, P. J. Cross-talk between site-specific transcription factors and DNA methylation states. J. Biol. Chem. 288, 34287–34294 (2013).
Yuan, T. et al. An integrative multi-scale analysis of the dynamic DNA methylation landscape in aging. PLoS Genet. 11, e1004996 (2015).
Horvath, S. et al. Obesity accelerates epigenetic aging of human liver. Proc. Natl Acad. Sci. USA 111, 15538–15543 (2014).
Lu, A. T. et al. Genetic variants near MLST8 and DHX57 affect the epigenetic age of the cerebellum. Nat. Commun. 7, 10561 (2016).
Lu, A. T. et al. Genetic architecture of epigenetic and neuronal ageing rates in human brain regions. Nat. Commun. 8, 15353 (2017).
Kananen, L. et al. The trajectory of the blood DNA methylome ageing rate is largely set before adulthood: evidence from two longitudinal studies. Age 38, 65 (2016).
Lu, A. T. et al. GWAS of epigenetic aging rates in blood reveals a critical role for TERT. Nat. Commun. 9, 387 (2018). This is the first GWAS of epigenetic ageing rates in blood to find genome-wide significant genetic variants, including a paradoxical role for variants in the TERT locus.
Mather, K. A., Jorm, A. F., Parslow, R. A. & Christensen, H. Is telomere length a biomarker of aging? A review. J. Gerontol. A Biol. Sci. Med. Sci 66A, 202–213 (2011).
Sanders, J. L. & Newman, A. B. Telomere length in epidemiology: a biomarker of aging, age-related disease, both, or neither? Epidemiol. Rev. 35, 112–131 (2013).
Marioni, R. E. et al. The epigenetic clock and telomere length are independently associated with chronological age and mortality. Int. J. Epidemiol. 45, 424–432 (2016).
Blackburn, E. H. Telomere states and cell fates. Nature 408, 53–56 (2000).
Chen, B. H. et al. Leukocyte telomere length, T cell composition and DNA methylation age. Aging 9, 1983–1995 (2017).
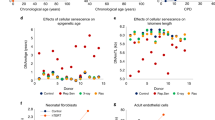
Lowe, D., Horvath, S. & Raj, K. Epigenetic clock analyses of cellular senescence and ageing. Oncotarget 7, 8524–8531 (2016).
Pucci, F., Gardano, L. & Harrington, L. Short telomeres in ESCs lead to unstable differentiation. Cell Stem Cell 12, 479–486 (2013).
Harrington, L. & Pucci, F. In medio stat virtus: unanticipated consequences of telomere dysequilibrium. Phil. Trans. R. Soc. B Biol Sci. 373, 20160444 (2018).
Yang, Z. et al. Correlation of an epigenetic mitotic clock with cancer risk. Genome Biol. 17, 205 (2016).
Horvath, S. Erratum to: DNA methylation age of human tissues and cell types. Genome Biol. 16, 96 (2015).
Beerman, I. et al. Proliferation-dependent alterations of the DNA methylation landscape underlie hematopoietic stem cell aging. Cell Stem Cell 12, 413–425 (2013).
Beerman, I. & Rossi, D. J. Epigenetic regulation of hematopoietic stem cell aging. Exp. Cell Res. 329, 192–199 (2014).
Issa, J.-P. Aging and epigenetic drift: a vicious cycle. J. Clin. Invest. 124, 24–29 (2014).
Collado, M., Blasco, M. A. & Serrano, M. Cellular senescence in cancer and aging. Cell 130, 223–233 (2007).
Drummond-Barbosa, D. Stem cells, their niches and the systemic environment: an aging network. Genetics 180, 1787–1797 (2008).
Melk, A. et al. Effects of donor age and cell senescence on kidney allograft survival. Am. J. Transplant 9, 114–123 (2009).
Halloran, P. F., Melk, A. & Barth, C. Rethinking chronic allograft nephropathy: the concept of accelerated senescence. J. Am. Soc. Nephrol. 10, 167–181 (1999).
Campisi, J. Aging, cellular senescence, and cancer. Annu. Rev. Physiol. 75, 685–705 (2013).
Campisi, J. & d’Adda di Fagagna, F. Cellular senescence: when bad things happen to good cells. Nat. Rev. Mol. Cell Biol. 8, 729–740 (2007).
Coppe, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Baker, D. J. et al. Opposing roles for p16Ink4a and p19Arf in senescence and ageing caused by BubR1 insufficiency. Nat. Cell Biol. 10, 825–836 (2008).
Chen, H. et al. PDGF signalling controls age-dependent proliferation in pancreatic beta-cells. Nature 478, 349–355 (2011).
Berent-Maoz, B., Montecino-Rodriguez, E., Signer, R. A. & Dorshkind, K. Fibroblast growth factor-7 partially reverses murine thymocyte progenitor aging by repression of Ink4a. Blood 119, 5715–5721 (2012).
Franzen, J. et al. Senescence-associated DNA methylation is stochastically acquired in subpopulations of mesenchymal stem cells. Aging Cell 16, 183–191 (2017).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Adams, P. D., Jasper, H. & Rudolph, K. L. Aging-induced stem cell mutations as drivers for disease and cancer. Cell Stem Cell 16, 601–612 (2015).
Grolleau-Julius, A., Ray, D. & Yung, R. L. The role of epigenetics in aging and autoimmunity. Clin. Rev. Allergy Immunol. 39, 42–50 (2010).
Neri, F. et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543, 72–77 (2017).
Jones, P. A. & Takai, D. The role of DNA methylation in mammalian epigenetics. Science 293, 1068–1070 (2001).
Knight, A. K. et al. An epigenetic clock for gestational age at birth based on blood methylation data. Genome Biol. 17, 206 (2016).
Spiers, H. et al. Methylomic trajectories across human fetal brain development. Genome Res. 25, 338–352 (2015).
Binder, A. M. et al. Faster ticking rate of the epigenetic clock is associated with faster pubertal development in girls. Epigenetics https://doi.org/10.1080/15592294.2017.1414127 (2018).
Bredesen, D. E. The non-existent aging program: how does it work? Aging Cell 3, 255–259 (2004).
Skulachev, V. P. Programmed death phenomena: from organelle to organism. Ann. NY Acad. Sci. 959, 214–237 (2002).
Mitteldorf, J. Chaotic population dynamics and the evolution of aging: proposing a demographic theory of senescence. Evol. Ecol. Res. 8, 561–574 (2006).
Takasugi, M. Progressive age-dependent DNA methylation changes start before adulthood in mouse tissues. Mech. Ageing Dev. 132, 65–71 (2011).
de Magalhaes, J. P. Programmatic features of aging originating in development: aging mechanisms beyond molecular damage? FASEB J. 26, 4821–4826 (2012).
Rando, T. A. & Chang, H. Y. Aging, Rejuvenation, and epigenetic reprogramming: resetting the aging clock. Cell 148, 46–57 (2012).
Johnson, A. A. et al. The role of DNA methylation in aging, rejuvenation, and age-related disease. Rejuven. Res. 15, 483–494 (2012).
Mitteldorf, J. J. How does the body know how old it is? Introducing the epigenetic clock hypothesis. Biochemistry 78, 1048–1053 (2013).
Mitteldorf, J. An epigenetic clock controls aging. Biogerontology 17, 257–265 (2016).
Blagosklonny, M. V. & Hall, M. N. Growth and aging: a common molecular mechanism. Aging 1, 357–362 (2009).
Walker, R. Why we age: Insight into the cause of growing old (Dove Medical Press, 2013).
Weidner, C. et al. Epigenetic aging upon allogeneic transplantation: the hematopoietic niche does not affect age-associated DNA methylation. Leukemia 29, 985 (2015). This is the first study to demonstrate that the chronological age of the donor of an allogeneic haematopoietic stem cell transplantation determines the DNAm age of the recipient (that is, the stem cell niche does not affect DNAm age).
Stolzel, F. et al. Dynamics of epigenetic age following hematopoietic stem cell transplantation. Haematologica 102, e321–e323 (2017). This study replicates and extends the results of reference 133.
Petkovich, D. A. et al. Using DNA Methylation profiling to evaluate biological age and longevity interventions. Cell Metab. 25, 954–960.e6 (2017).
Wang, T. et al. Epigenetic aging signatures in mice livers are slowed by dwarfism, calorie restriction and rapamycin treatment. Genome Biol. 18, 57 (2017).
Cole, J. J. et al. Diverse interventions that extend mouse lifespan suppress shared age-associated epigenetic changes at critical gene regulatory regions. Genome Biol. 18, 58 (2017).
Stubbs, T. M. et al. Multi-tissue DNA methylation age predictor in mouse. Genome Biol. 18, 68 (2017). References 135–138 demonstrate that DNAm age estimators in mice respond as expected to gold-standard anti-ageing interventions, for example, calorie restriction and growth hormone receptor knockout.
Polanowski, A. M., Robbins, J., Chandler, D. & Jarman, S. N. Epigenetic estimation of age in humpback whales. Mol. Ecol. Resources 14, 976–987 (2014).
Thompson, M. J., vonHoldt, B., Horvath, S. & Pellegrini, M. An epigenetic aging clock for dogs and wolves. Aging 9, 1055–1068 (2017).
Simpson, V. J., Johnson, T. E. & Hammen, R. F. Caenorhabditis elegans DNA does not contain 5-methylcytosine at any time during development or aging. Nucleic Acids Res. 14, 6711–6719 (1986).
Newman, A. B. Is the onset of obesity the same as aging? Proc. Natl Acad. Sci. USA 112, E7163–E7163 (2015).
Ward-Caviness, C. K. et al. Long-term exposure to air pollution is associated with biological aging. Oncotarget 7, 74510–74525 (2016).
Nwanaji-Enwerem, J. C. et al. Long-term ambient particle exposures and blood DNA methylation age: findings from the VA normative aging study. Environ. Epigenet. 2, dvw006 (2016).
Horvath, S. & Levine, A. J. HIV-1 infection accelerates age according to the epigenetic clock. J. Infect. Dis. 212, 1563–1573 (2015).
Gross, A. M. et al. Methylome-wide analysis of chronic HIV infection reveals five-year increase in biological age and epigenetic targeting of HLA. Mol. Cell 62, 157–168 (2016).
Kananen, L. et al. Cytomegalovirus infection accelerates epigenetic aging. Exp. Gerontol. 72, 227–229 (2015).
Gao, X., Zhang, Y. & Brenner, H. Associations of Helicobacter pylori infection and chronic atrophic gastritis with accelerated epigenetic ageing in older adults. Br. J. Cancer 117, 1211–1214 (2017).
Zannas, A. et al. Lifetime stress accelerates epigenetic aging in an urban, African American cohort: relevance of glucocorticoid signaling. Genome Biol. 16, 266 (2015).
Zannas, A. S. Editorial Perspective: Psychological stress and epigenetic aging - what can we learn and how can we prevent? J. Child Psychol. Psychiatry 57, 674–675 (2016).
Dugue, P. A. et al. DNA methylation-based biological aging and cancer risk and survival: pooled analysis of seven prospective studies. Int. J. Cancer 142, 1611–1619 (2018). This is the largest study to date to demonstrate that epigenetic age acceleration in blood is associated with increased cancer risk and shorter cancer survival independently of major health risk factors.
Horvath, S. et al. Huntington’s disease accelerates epigenetic aging of human brain and disrupts DNA methylation levels. Aging 8, 1485–1512 (2016).
Vidal, L. et al. Specific increase of methylation age in osteoarthritis cartilage. Osteoarthritis Cartilage 24, S63 (2016).
Acknowledgements
Although the authors have made every effort to be objective, it is proper to mention that S.H. co-authored many of the articles mentioned in this Review. The authors apologize for not being able to cover all publications owing to oversight or space limitations. To ensure accuracy, the authors highlighted articles that employed large sample sizes, rigorous study designs and validated biomarkers.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of this article.
Corresponding author
Ethics declarations
Competing interests
The Regents of the University of California is the sole owner of several patent applications directed at the invention of measures of epigenetic age estimation for which S.H. is a named inventor. K.R. declares no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related Links
NIH Wednesday Afternoon Lecture Series (presentation by S. H. from 15 June 2016): https://www.youtube.com/watch?v = 0zaCKAnFogQ
Supplementary information
Glossary
- Chronological age
-
The calendar time that has passed since birth. Zero is the time at birth. Negative numbers indicate prenatal ages, whereas positive numbers indicate postnatal ages.
- Biological age
-
Also known as physiological age, organismal age or phenotypic age. This ambiguous concept is held to be dependent on the biological state of the individual.
- Epigenetic age
-
The age estimate in years resulting from a mathematical algorithm based on the methylation state of specific CpGs in the genome. Negative numbers indicate prenatal ages.
- CpG dinucleotides
-
Regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5′ to 3′ direction. Cytosines in CpG dinucleotides can be methylated to form 5-methylcytosine.
- Epigenetic age estimators
-
Mathematical algorithms that use values assigned to the methylation state of specific CpGs in the genome to estimate the age of a person or biological sample. A multi-tissue age estimator allows one to estimate the age of any nucleated cell, tissue or organ.
- Quasi-programme theories of ageing
-
Several variations of a theory that posits that ageing is not the intended outcome of biological processes but that some programmed processes nevertheless result in ageing. Therefore, the process of ageing can be viewed as quasi-programmed.
Rights and permissions
About this article
Cite this article
Horvath, S., Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat Rev Genet 19, 371–384 (2018). https://doi.org/10.1038/s41576-018-0004-3
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-018-0004-3
This article is cited by
-
Meta-analysis of epigenetic aging in schizophrenia reveals multifaceted relationships with age, sex, illness duration, and polygenic risk
Clinical Epigenetics (2024)
-
Bazi Bushen ameliorates age-related energy metabolism dysregulation by targeting the IL-17/TNF inflammatory pathway associated with SASP
Chinese Medicine (2024)
-
The interaction between ageing and Alzheimer's disease: insights from the hallmarks of ageing
Translational Neurodegeneration (2024)
-
Consequences of heterogeneity in aging: parental age at death predicts midlife all-cause mortality and hospitalization in a Swedish national birth cohort
BMC Geriatrics (2024)
-
Causal association of obesity with epigenetic aging and telomere length: a bidirectional mendelian randomization study
Lipids in Health and Disease (2024)