Abstract
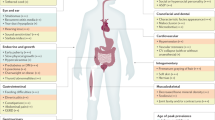
Trisomy 21, the presence of a supernumerary chromosome 21, results in a collection of clinical features commonly known as Down syndrome (DS). DS is among the most genetically complex of the conditions that are compatible with human survival post-term, and the most frequent survivable autosomal aneuploidy. Mouse models of DS, involving trisomy of all or part of human chromosome 21 or orthologous mouse genomic regions, are providing valuable insights into the contribution of triplicated genes or groups of genes to the many clinical manifestations in DS. This endeavour is challenging, as there are >200 protein-coding genes on chromosome 21 and they can have direct and indirect effects on homeostasis in cells, tissues, organs and systems. Although this complexity poses formidable challenges to understanding the underlying molecular basis for each of the many clinical features of DS, it also provides opportunities for improving understanding of genetic mechanisms underlying the development and function of many cell types, tissues, organs and systems. Since the first description of trisomy 21, we have learned much about intellectual disability and genetic risk factors for congenital heart disease. The lower occurrence of solid tumours in individuals with DS supports the identification of chromosome 21 genes that protect against cancer when overexpressed. The universal occurrence of the histopathology of Alzheimer disease and the high prevalence of dementia in DS are providing insights into the pathology and treatment of Alzheimer disease. Clinical trials to ameliorate intellectual disability in DS signal a new era in which therapeutic interventions based on knowledge of the molecular pathophysiology of DS can now be explored; these efforts provide reasonable hope for the future.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 1 digital issues and online access to articles
$99.00 per year
only $99.00 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Down, J. L. H. Observations on an ethnic classification of idiots. Lond. Hosp. Rep. 3, 259–262 (1866).
Hasle, H., Clemmensen, I. H. & Mikkelsen, M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet 355, 165–169 (2000).
LeJeune, J., Gautier, M. & Turpin, R. Study of somatic chromosomes from 9 mongoloid children [French]. C. R. Hebd. Seances. Acad. Sci. 248, 1721–1722 (1959). This study is the first description of the chromosomal abnormality in DS.
Davisson, M. T., Schmidt, C. & Akeson, E. C. Segmental trisomy of murine chromosome 16: a new model system for studying Down syndrome. Prog. Clin. Biol. Res. 360, 263–280 (1990).
Hattori, M. et al. The DNA sequence of human chromosome 21. Nature 405, 311–319 (2000). A landmark paper on the sequencing of the long arm of HSA21.
Antonarakis, S. E. Down syndrome and the complexity of genome dosage imbalance. Nat. Rev. Genet. 18, 147–163 (2017).
Herault, Y. et al. Rodent models in Down syndrome research: impact and future opportunities. Dis. Model. Mech. 10, 1165–1186 (2017).
Chen, X. Q. & Mobley, W. C. Exploring the pathogenesis of Alzheimer disease in basal forebrain cholinergic neurons: converging insights from alternative hypotheses. Front. Neurosci. 13, 446 (2019).
Reeves, R. H. et al. Paving the way for therapy: the Second International Conference of the Trisomy 21 Research Society. Mol. Syndromol. 9, 279–286 (2019).
de Graaf, G., Buckley, F. & Skotko, B. People living with Down syndrome in the USA: births and population. Down Syndrome Education International https://dsuri.net/us-population-factsheet (2019).
de Graaf, G., Buckley, F. & Skotko, B. G. Estimation of the number of people with Down syndrome in the United States. Genet. Med. 19, 439–447 (2017).
de Graaf, G., Buckley, F. & Skotko, B. G. Birth and population prevalence of Down syndrome in European countries. (Poster presented at the World Down Syndrome Congress 2018).
Bray, I., Wright, D. E., Davies, C. & Hook, E. B. Joint estimation of Down syndrome risk and ascertainment rates: a meta-analysis of nine published data sets. Prenat. Diagn. 18, 9–20 (1998).
Hecht, C. A. & Hook, E. B. Rates of Down syndrome at livebirth by one-year maternal age intervals in studies with apparent close to complete ascertainment in populations of European origin: a proposed revised rate schedule for use in genetic and prenatal screening. Am. J. Med. Genet. 62, 376–385 (1996).
Hassold, T. & Hunt, P. To err (meiotically) is human: the genesis of human aneuploidy. Nat. Rev. Genet. 2, 280–291 (2001).
Morris, J. K., Wald, N. J., Mutton, D. E. & Alberman, E. Comparison of models of maternal age-specific risk for Down syndrome live births. Prenat. Diagn. 23, 252–258 (2003).
de Graaf, G., Buckley, F. & Skotko, B. G. Estimates of the live births, natural losses, and elective terminations with Down syndrome in the United States. Am. J. Med. Genet. A 167, 756–767 (2015).
Savva, G. M., Morris, J. K., Mutton, D. E. & Alberman, E. Maternal age-specific fetal loss rates in Down syndrome pregnancies. Prenat. Diagn. 26, 499–504 (2006).
Bittles, A. H., Bower, C., Hussain, R. & Glasson, E. J. The four ages of Down syndrome. Eur. J. Public Health 17, 221–225 (2007).
Deng, C. et al. Recent trends in the birth prevalence of Down syndrome in China: impact of prenatal diagnosis and subsequent terminations. Prenat. Diagn. 35, 311–318 (2015).
Morris, J. K. & Alberman, E. Trends in Down’s syndrome live births and antenatal diagnoses in England and Wales from 1989 to 2008: analysis of data from the National Down Syndrome cytogenetic register. BMJ 339, b3794 (2009).
Tul, N., Verdenik, I., Srsen, T. P. & Antolic, Z. N. P31.05: incidence of Down syndrome in Slovenia in the last 15 years. Ultrasound Obstet. Gynecol. 30, 569–570 (2007).
Collins, V. R., Muggli, E. E., Riley, M., Palma, S. & Halliday, J. L. Is Down syndrome a disappearing birth defect? J. Pediatr. 152, 20–24.e1 (2008).
Dolk, H. et al. Trends and geographic inequalities in the prevalence of Down syndrome in Europe, 1980-1999. Rev. Epidemiol Sante Publique 53, 2S87–2S95 (2005).
de Graaf, G. et al. Estimates of live birth prevalence of children with Down syndrome in the period 1991–2015 in the Netherlands. J. Intellect. Disabil. Res. 61, 461–470 (2017).
Nagaoka, S. I., Hassold, T. J. & Hunt, P. A. Human aneuploidy: mechanisms and new insights into an age-old problem. Nat. Rev. Genet. 13, 493–504 (2012).
Gruhn, J. R. et al. Chromosome errors in human eggs shape natural fertility over reproductive life span. Science 365, 1466–1469 (2019).
Antonarakis, S. E. et al. The meiotic stage of nondisjunction in trisomy 21: determination by using DNA polymorphisms. Am. J. Hum. Genet. 50, 544–550 (1992).
Allen, E. G. et al. Maternal age and risk for trisomy 21 assessed by the origin of chromosome nondisjunction: a report from the Atlanta and National Down Syndrome Projects. Hum. Genet. 125, 41–52 (2009).
Yoon, P. W. et al. Advanced maternal age and the risk of Down syndrome characterized by the meiotic stage of chromosomal error: a population-based study. Am. J. Hum. Genet. 58, 628–633 (1996).
Freeman, S. B. et al. The National Down Syndrome Project: design and implementation. Public Health Rep. 122, 62–72 (2007).
Ghosh, S., Feingold, E. & Dey, S. K. Etiology of Down syndrome: evidence for consistent association among altered meiotic recombination, nondisjunction, and maternal age across populations. Am. J. Med. Genet. A 149, 1415–1420 (2009).
Oliver, T. R. et al. New insights into human nondisjunction of chromosome 21 in oocytes. PLoS Genet. 4, e1000033 (2008).
Chernus, J. M. et al. A candidate gene analysis and GWAS for genes associated with maternal nondisjunction of chromosome 21. PLoS Genet. 15, e1008414 (2019).
Coppede, F. Risk factors for Down syndrome. Arch. Toxicol. 90, 2917–2929 (2016).
Hunter, J. E. et al. The association of low socioeconomic status and the risk of having a child with Down syndrome: a report from the National Down Syndrome Project. Genet. Med. 15, 698–705 (2013).
Torfs, C. P. & Christianson, R. E. Socioeconomic effects on the risk of having a recognized pregnancy with Down syndrome. Birth Defects Res. A Clin. Mol. Teratol. 67, 522–528 (2003).
Christianson, R. E., Sherman, S. L. & Torfs, C. P. Maternal meiosis II nondisjunction in trisomy 21 is associated with maternal low socioeconomic status. Genet. Med. 6, 487–494 (2004).
Ghosh, S., Ghosh, P. & Dey, S. K. Altered incidence of meiotic errors and Down syndrome birth under extreme low socioeconomic exposure in the Sundarban area of India. J. Community Genet. 5, 119–124 (2014).
Keen, C. et al. The association between maternal occupation and Down syndrome: a report from the national Down syndrome project. Int. J. Hyg. Environ. Health 223, 207–213 (2020).
Sartain, C. V. & Hunt, P. A. An old culprit but a new story: bisphenol A and "NextGen" bisphenols. Fertil. Steril. 106, 820–826 (2016).
Horan, T. S. et al. Replacement bisphenols adversely affect mouse gametogenesis with consequences for subsequent generations. Curr. Biol. 28, 2948–2954.e3 (2018).
Grandjean, P. et al. Timescales of developmental toxicity impacting on research and needs for intervention. Basic Clin. Pharmacol. Toxicol. 125, 70–80 (2019).
Antonarakis, S. E., Avramopoulos, D., Blouin, J. L., Talbot, C. C. Jr. & Schinzel, A. A. Mitotic errors in somatic cells cause trisomy 21 in about 4.5% of cases and are not associated with advanced maternal age. Nat. Genet. 3, 146–150 (1993).
Antonarakis, S. E. Parental origin of the extra chromosome in trisomy 21 as indicated by analysis of DNA polymorphisms. Down Syndrome Collaborative Group. N. Engl. J. Med. 324, 872–876 (1991). This paper reports the use of DNA polymorphic markers to determine the parental origin of the supernumerary HSA21 in DS.
Antonarakis, S. E. 10 years of genomics, chromosome 21, and Down syndrome. Genomics 51, 1–16 (1998).
Morris, J. K., Alberman, E., Mutton, D. & Jacobs, P. Cytogenetic and epidemiological findings in Down syndrome: England and Wales 1989-2009. Am. J. Med. Genet. A 158, 1151–1157 (2012).
Lyle, R. et al. Genotype-phenotype correlations in Down syndrome identified by array CGH in 30 cases of partial trisomy and partial monosomy chromosome 21. Eur. J. Hum. Genet. 17, 454–466 (2009).
Barlow, G. M. et al. Down syndrome congenital heart disease: a narrowed region and a candidate gene. Genet. Med. 3, 91–101 (2001).
Gupta, M., Dhanasekaran, A. R. & Gardiner, K. J. Mouse models of Down syndrome: gene content and consequences. Mamm. Genome 27, 538–555 (2016).
Epstein, C. J. et al. Protocols to establish genotype-phenotype correlations in Down syndrome. Am. J. Hum. Genet. 49, 207–235 (1991).
Bray, I. C. & Wright, D. E. Estimating the spontaneous loss of Down syndrome fetuses between the times of chorionic villus sampling, amniocentesis and livebirth. Prenat. Diagn. 18, 1045–1054 (1998).
Hassold, T. J. & Jacobs, P. A. Trisomy in man. Ann.Rev.Genet. 18, 69–97 (1984).
Shapiro, B. L. Down syndrome – a disruption of homeostasis. Am. J. Med. Genet. 14, 241–269 (1983).
Pritchard, M. & Kola, I. The ‘gene dosage effect’ hypothesis versus the ‘amplified developmental instability’ hypothesis in Down syndrome. J. Neural Transm. 57, 293–303 (1999).
Stamoulis, G. et al. Single cell transcriptome in aneuploidies reveals mechanisms of gene dosage imbalance. Nat. Commun. 10, 4495 (2019).
Salehi, A. et al. Increased App expression in a mouse model of Down’s syndrome disrupts NGF transport and causes cholinergic neuron degeneration. Neuron 51, 29–42 (2006). A study that provided support for the hypothesis that neuronal degeneration in DS is due to increased APP expression.
Pelleri, M. C. et al. Integrated quantitative transcriptome maps of human trisomy 21 tissues and cells. Front. Genet. 9, 125 (2018).
Sullivan, K. D. et al. Trisomy 21 consistently activates the interferon response. eLife 5, e16220 (2016).
Gonzales, P. K. et al. Transcriptome analysis of genetically matched human induced pluripotent stem cells disomic or trisomic for chromosome 21. PLoS One 13, e0194581 (2018).
Letourneau, A. et al. Domains of genome-wide gene expression dysregulation in Down’s syndrome. Nature 508, 345–350 (2014).
Do, C., Xing, Z., Yu, Y. E. & Tycko, B. Trans-acting epigenetic effects of chromosomal aneuploidies: lessons from Down syndrome and mouse models. Epigenomics 9, 189–207 (2017).
Do, L. H., Mobley, W. C. & Singhal, N. Questioned validity of gene expression dysregulated domains in Down’s syndrome. F1000Res. 4, 269 (2015).
Ahlfors, H. et al. Gene expression dysregulation domains are not a specific feature of Down syndrome. Nat. Commun. 10, 2489 (2019).
Umlauf, D. & Mourad, R. The 3D genome: from fundamental principles to disease and cancer. Semin. Cell Dev. Biol. 90, 128–137 (2019).
Kemeny, S. et al. Spatial organization of chromosome territories in the interphase nucleus of trisomy 21 cells. Chromosoma 127, 247–259 (2018).
Mendioroz, M. et al. Trans effects of chromosome aneuploidies on DNA methylation patterns in human Down syndrome and mouse models. Genome Biol. 16, 263 (2015).
El Hajj, N. et al. Epigenetic dysregulation in the developing Down syndrome cortex. Epigenetics 11, 563–578 (2016).
Horvath, S. et al. Accelerated epigenetic aging in Down syndrome. Aging Cell 14, 491–495 (2015).
Scott, H. S. et al. Identification and characterization of two putative human arginine methyltransferases (HRMT1L1 and HRMT1L2). Genomics 48, 330–340 (1998).
Xiao, C. L. et al. N6-methyladenine DNA modification in the human genome. Mol. Cell 71, 306–318.e7 (2018).
Kim, I. S. et al. Roles of Mis18α in epigenetic regulation of centromeric chromatin and CENP-A loading. Mol. Cell 46, 260–273 (2012).
Veland, N. et al. DNMT3L facilitates DNA methylation partly by maintaining DNMT3A stability in mouse embryonic stem cells. Nucleic Acids Res. 47, 152–167 (2019).
Lu, J. et al. Global hypermethylation in fetal cortex of Down syndrome due to DNMT3L overexpression. Hum. Mol. Genet. 25, 1714–1727 (2016).
Sailani, M. R. et al. DNA-methylation patterns in trisomy 21 using cells from monozygotic twins. PLoS One 10, e0135555 (2015).
Guo, X., Williams, J. G., Schug, T. T. & Li, X. DYRK1A and DYRK3 promote cell survival through phosphorylation and activation of SIRT1. J. Biol. Chem. 285, 13223–13232 (2010).
Lepagnol-Bestel, A. M. et al. DYRK1A interacts with the REST/NRSF-SWI/SNF chromatin remodelling complex to deregulate gene clusters involved in the neuronal phenotypic traits of Down syndrome. Hum. Mol. Genet. 18, 1405–1414 (2009).
Liu, Y. et al. Systematic proteome and proteostasis profiling in human trisomy 21 fibroblast cells. Nat. Commun. 8, 1212 (2017). First quantitative study of the trisomy 21 proteome, using fibroblasts from individuals with DS.
Gillet, L. C. et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol. Cell Proteom. 11, O111.016717 (2012).
Sullivan, K. D. et al. Trisomy 21 causes changes in the circulating proteome indicative of chronic autoinflammation. Sci. Rep. 7, 14818 (2017).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One 5, e15004 (2010).
Izzo, A. et al. Mitochondrial dysfunction in Down syndrome: molecular mechanisms and therapeutic targets. Mol. Med. 24, 2 (2018).
Zamponi, E. & Helguera, P. R. The shape of mitochondrial dysfunction in Down syndrome. Dev. Neurobiol. 79, 613–621 (2019).
Valenti, D. et al. Mitochondria as pharmacological targets in Down syndrome. Free. Radic. Biol. Med. 114, 69–83 (2018).
Vilardell, M. et al. Meta-analysis of heterogeneous Down syndrome data reveals consistent genome-wide dosage effects related to neurological processes. BMC Genomics 12, 229 (2011).
Reeves, R. et al. A mouse model for Down syndrome exhibits learning and behaviour deficits. Nat. Genet. 11, 177–183 (1995). This paper describes the first complex model of trisomy 21, Ts65Dn, and lays out characterizations for comparing the effects of aneuploidy in mice and humans.
Sussan, T., Yang, A., Li, F., Ostrowski, M. & Reeves, R. H. Trisomy protects against ApcMin-mediated tumors in mouse models of Down syndrome. Nature 451, 73–75 (2008).
Garcia-Cerro, S., Rueda, N., Vidal, V., Lantigua, S. & Martinez-Cue, C. Normalizing the gene dosage of Dyrk1A in a mouse model of Down syndrome rescues several Alzheimer’s disease phenotypes. Neurobiol. Dis. 106, 76–88 (2017).
Aziz, N. M. et al. Lifespan analysis of brain development, gene expression and behavioral phenotypes in the Ts1Cje, Ts65Dn and Dp(16)1/Yey mouse models of Down syndrome. Disease Models Mech 11, dmm031013 (2018).
Gribble, S. M. et al. Massively parallel sequencing reveals the complex structure of an irradiated human chromosome on a mouse background in the Tc1 model of Down syndrome. PLoS One 8, e60482 (2013).
Kazuki, Y. et al. A non-mosaic humanized mouse model of Down syndrome, trisomy of a nearly complete long arm of human chromosome 21 in mouse chromosome background. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/862433v1 (2019).
Ramirez-Solis, R., Liu, P. & Bradley, A. Chromosome engineering in mice. Nature 378, 720–724 (1995).
Olson, L. E., Richtsmeier, J. T., Leszl, J. & Reeves, R. H. A chromosome 21 critical region does not cause specific Down syndrome phenotypes. Science 306, 687–690 (2004).
Yu, T. et al. A mouse model of Down syndrome trisomic for all human chromosome 21 syntenic regions. Hum. Mol. Genet. 19, 2780–2791 (2010). This paper reports a mouse model that is trisomic for all mouse chromosomal regions syntenic with HSA21.
Stefanidis, K. et al. Causes of infertility in men with Down syndrome. Andrologia 43, 353–357 (2011).
Williams, B. R. et al. Aneuploidy affects proliferation and spontaneous immortalization in mammalian cells. Science 322, 703–709 (2008).
Contestabile, A. et al. Cell cycle alteration and decreased cell proliferation in the hippocampal dentate gyrus and in the neocortical germinal matrix of fetuses with Down syndrome and in Ts65Dn mice. Hippocampus 17, 665–678 (2007).
Kleschevnikov, A. M. et al. Hippocampal long-term potentiation suppressed by increased inhibition in the Ts65Dn mouse, a genetic model of Down syndrome. J. Neurosci. 24, 8153–8160 (2004).
Siarey, R. J., Stoll, J., Rapoport, S. I. & Galdzicki, Z. Altered long-term potentiation in the young and old Ts65Dn mouse, a model for Down syndrome. Neuropharmacology 36, 1549–1554 (1997).
Ferencz, C. et al. Congenital cardiovascular malformations associated with chromosome abnormalities: an epidemiologic study. J. Pediatr. 114, 79–86 (1989).
Lana-Elola, E. et al. Genetic dissection of Down syndrome-associated congenital heart defects using a new mouse mapping panel. eLife 5, e11614 (2016).
Li, H. et al. Penetrance of congenital heart disease in a mouse model of Down syndrome depends on a trisomic potentiator of a disomic modifier. Genetics 203, 763–770 (2016).
Edie, S. et al. Survey of human chromosome 21 gene expression effects on early development in Danio rerio. G3 8, 2215–2223 (2018).
Yang, Q., Rasmussen, S. A. & Friedman, J. M. Mortality associated with Down’s syndrome in the USA from 1983 to 1997: a population-based study. Lancet 359, 1019–1025 (2002).
Escorihuela, R. M. et al. A behavioral assessment of Ts65Dn mice: a putative Down syndrome model. Neurosci. Lett. 199, 143–146 (1995).
Holtzman, D. M. et al. Developmental abnormalities and age-related neurodegeneration in a mouse model of Down syndrome. Proc. Natl Acad. Sci. USA 93, 13333–13338 (1996).
Hanson, J. E., Blank, M., Valenzuela, R. A., Garner, C. C. & Madison, D. V. The functional nature of synaptic circuitry is altered in area CA3 of the hippocampus in a mouse model of Down’s syndrome. J. Physiol. 579, 53–67 (2007). This study establishes the imbalance between excitatory and inhibitory inputs in the hippocampus, which has been the focus of efforts to antagonize signalling through GABA-A receptors, including a trial to improve cognitive ability in individuals with DS (NCT01436955).
Fernandez, F. et al. Pharmacotherapy for cognitive impairment in a mouse model of Down syndrome. Nat. Neurosci. 10, 411–413 (2007). One of the first mouse studies that raised the possibility of pharmacotherapy for the cognitive impairment in DS.
Braudeau, J. et al. Specific targeting of the GABA-A receptor α5 subtype by a selective inverse agonist restores cognitive deficits in Down syndrome mice. J Psychopharmacol (2011).
Duchon, A. et al. Long-lasting correction of in vivo LTP and cognitive deficits of mice modelling Down syndrome with an α5-selective GABAA inverse agonist. Br. J. Pharmacol. https://doi.org/10.1111/bph.14903 (2019).
Hart, S. J. et al. Pharmacological interventions to improve cognition and adaptive functioning in Down syndrome: strides to date. Am. J. Med. Genet. A 173, 3029–3041 (2017).
de la Torre, R. et al. Safety and efficacy of cognitive training plus epigallocatechin-3-gallate in young adults with Down’s syndrome (TESDAD): a double-blind, randomised, placebo-controlled, phase 2 trial. Lancet Neurol. 15, 801–810 (2016). A successful randomized trial to improve cognitive functioning in young adults with DS.
Jack, C. R. Jr. et al. A/T/N: An unbiased descriptive classification scheme for Alzheimer disease biomarkers. Neurology 87, 539–547 (2016).
Esparza, T. J. et al. Amyloid-β oligomerization in Alzheimer dementia versus high-pathology controls. Ann. Neurol. 73, 104–119 (2013).
Strydom, A. et al. Alzheimer’s disease in Down syndrome: an overlooked population for prevention trials. Alzheimers Dement. 4, 703–713 (2018).
Thonberg, H. et al. Mutation screening of patients with Alzheimer disease identifies APP locus duplication in a Swedish patient. BMC Res. Notes 4, 476 (2011).
Wallon, D. et al. The French series of autosomal dominant early onset Alzheimer’s disease cases: mutation spectrum and cerebrospinal fluid biomarkers. J. Alzheimers Dis. 30, 847–856 (2012).
Rovelet-Lecrux, A. et al. APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nat. Genet. 38, 24–26 (2006). This study provides strong evidence for a link between the triplication of APP in trisomy 21 and early-onset AD in individuals with DS.
Sleegers, K. et al. APP duplication is sufficient to cause early onset Alzheimer’s dementia with cerebral amyloid angiopathy. Brain 129, 2977–2983 (2006).
Wiseman, F. K. et al. A genetic cause of Alzheimer disease: mechanistic insights from Down syndrome. Nat. Rev. Neurosci. 16, 564–574 (2015).
Doran, E. et al. Down syndrome, partial trisomy 21, and absence of Alzheimer’s disease: the role of APP. J. Alzheimers Dis. 56, 459–470 (2017).
Ryoo, S. R. et al. Dual-specificity tyrosine(Y)-phosphorylation regulated kinase 1A-mediated phosphorylation of amyloid precursor protein: evidence for a functional link between Down syndrome and Alzheimer’s disease. J. Neurochem. 104, 1333–1344 (2008).
Wegiel, J. et al. Intraneuronal Aβ immunoreactivity is not a predictor of brain amyloidosis-β or neurofibrillary degeneration. Acta Neuropathol. 113, 389–402 (2007).
Vingtdeux, V. et al. Phosphorylation of amyloid precursor carboxy-terminal fragments enhances their processing by a gamma-secretase-dependent mechanism. Neurobiol. Dis. 20, 625–637 (2005).
Woods, Y. L. et al. The kinase DYRK phosphorylates protein-synthesis initiation factor eIF2Bε at Ser539 and the microtubule-associated protein tau at Thr212: potential role for DYRK as a glycogen synthase kinase 3-priming kinase. Biochem. J. 355, 609–615 (2001).
Lindwall, G. & Cole, R. D. Phosphorylation affects the ability of tau protein to promote microtubule assembly. J. Biol. Chem. 259, 5301–5305 (1984).
LaPointe, N. E. et al. The amino terminus of tau inhibits kinesin-dependent axonal transport: implications for filament toxicity. J. Neurosci. Res. 87, 440–451 (2009).
Goedert, M. & Jakes, R. Mutations causing neurodegenerative tauopathies. Biochim. Biophys. Acta 1739, 240–250 (2005).
Liu, F. et al. Overexpression of Dyrk1A contributes to neurofibrillary degeneration in Down syndrome. FASEB J. 22, 3224–3233 (2008).
Yin, X. et al. Dyrk1A overexpression leads to increase of 3R-tau expression and cognitive deficits in Ts65Dn Down syndrome mice. Sci. Rep. 7, 619 (2017).
Chen, X. Q., Sawa, M. & Mobley, W. C. Dysregulation of neurotrophin signaling in the pathogenesis of Alzheimer disease and of Alzheimer disease in Down syndrome. Free Radic. Biol. Med. 114, 52–61 (2018).
Stenmark, H. Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 10, 513–525 (2009).
Roberts, R. L. et al. Endosome fusion in living cells overexpressing GFP-rab5. J. Cell Sci. 112, 3667–3675 (1999).
Cataldo, A. et al. Endocytic disturbances distinguish among subtypes of Alzheimer’s disease and related disorders. Ann. Neurol. 50, 661–665 (2001).
Cataldo, A. M., Barnett, J. L., Pieroni, C. & Nixon, R. A. Increased neuronal endocytosis and protease delivery to early endosomes in sporadic Alzheimer’s disease: neuropathologic evidence for a mechanism of increased β-amyloidogenesis. J. Neurosci. 17, 6142–6151 (1997).
Cataldo, A. M. et al. Endocytic pathway abnormalities precede amyloid beta deposition in sporadic Alzheimer’s disease and Down syndrome: differential effects of APOE genotype and presenilin mutations. Am. J. Pathol. 157, 277–286 (2000).
Corlier, F. et al. Modifications of the endosomal compartment in peripheral blood mononuclear cells and fibroblasts from Alzheimer’s disease patients. Transl. Psychiatry 5, e595 (2015).
Saunders, A. M. et al. Association of apolipoprotein E allele epsilon 4 with late-onset familial and sporadic Alzheimer’s disease. Neurology 43, 1467–1472 (1993).
Rohn, T. T., McCarty, K. L., Love, J. E. & Head, E. Is apolipoprotein E4 an important risk factor for dementia in persons with Down syndrome? J. Parkinsons Dis. Alzheimers Dis. 1, 7 (2014).
Cataldo, A. M. et al. Down syndrome fibroblast model of Alzheimer-related endosome pathology: accelerated endocytosis promotes late endocytic defects. Am. J. Pathol. 173, 370–384 (2008).
Cossec, J. C. et al. Trisomy for synaptojanin1 in Down syndrome is functionally linked to the enlargement of early endosomes. Hum. Mol. Genet. 21, 3156–3172 (2012).
Nuriel, T. et al. The endosomal-lysosomal pathway is dysregulated by APOE4 expression in vivo. Front. Neurosci. 11, 702 (2017).
Xu, W. et al. Amyloid precursor protein-mediated endocytic pathway disruption induces axonal dysfunction and neurodegeneration. J. Clin. Invest. 126, 1815–1833 (2016).
Jiang, Y. et al. Alzheimer’s-related endosome dysfunction in Down syndrome is Aβ-independent but requires APP and is reversed by BACE-1 inhibition. Proc. Natl Acad. Sci. USA 107, 1630–1635 (2010).
Jiang, Y. et al. Lysosomal dysfunction in Down syndrome is APP-dependent and mediated by APP-βCTF (C99). J. Neurosci. 39, 5255–5268 (2019).
Kwart, D. et al. A large panel of isogenic APP and PSEN1 mutant human iPSC neurons reveals shared endosomal abnormalities mediated by APP β-CTFs, not Aβ. Neuron 104, 256–270.e1–e5 (2019).
Cuckle, H. & Maymon, R. Development of prenatal screening – a historical overview. Semin. Perinatol. 40, 12–22 (2016).
Bianchi, D. W., Crombleholme, T. M., D’Alton M. E. & Malone, F. D. Fetology: Diagnosis and Management of the Fetal Patient Ch. 131 (McGraw-Hill Medical, 2010).
Bianchi, D. W., Rava, R. P. & Sehnert, A. J. DNA sequencing versus standard prenatal aneuploidy screening. N. Engl. J. Med. 371, 577–578 (2014).
Norton, M. E. & Wapner, R. J. Cell-free DNA analysis for noninvasive examination of trisomy. N. Engl. J. Med. 373, 2581–2582 (2015).
Bianchi, D. W. & Chiu, R. W. K. Sequencing of circulating cell-free DNA during pregnancy. N. Engl. J. Med. 379, 464–473 (2018).
Liu, S. et al. Genomic analyses from non-invasive prenatal testing reveal genetic associations, patterns of viral infections, and Chinese population history. Cell 175, 347–359.e14 (2018).
Taylor-Phillips, S. et al. Accuracy of non-invasive prenatal testing using cell-free DNA for detection of Down, Edwards and Patau syndromes: a systematic review and meta-analysis. BMJ Open. 6, e010002 (2016).
Hill, M. et al. Has noninvasive prenatal testing impacted termination of pregnancy and live birth rates of infants with Down syndrome? Prenat. Diagn. 37, 1281–1290 (2017).
Ralston, S. J., Wertz, D., Chelmow, D., Craigo, S. D. & Bianchi, D. W. Pregnancy outcomes after prenatal diagnosis of aneuploidy. Obstet. Gynecol. 97, 729–733 (2001).
Alexander, M. et al. Morbidity and medication in a large population of individuals with Down syndrome compared to the general population. Dev. Med. Child. Neurol. 58, 246–254 (2016).
Bull, M. J. & Committee on Genetics,. Health supervision for children with Down syndrome. Pediatrics 128, 393–406 (2011).
Jensen, K. M. & Bulova, P. D. Managing the care of adults with Down’s syndrome. BMJ 349, g5596 (2014).
Capone, G. et al. Co-occurring medical conditions in adults with Down syndrome: a systematic review toward the development of health care guidelines. Am. J. Med. Genet. A 176, 116–133 (2018).
Roizen, N. J. & Patterson, D. Down’s syndrome. Lancet 361, 1281–1289 (2003).
Glasson, E. J., Dye, D. E. & Bittles, A. H. The triple challenges associated with age-related comorbidities in Down syndrome. J. Intellect. Disabil. Res. 58, 393–398 (2014).
Bergstrom, S. et al. Trends in congenital heart defects in infants with Down syndrome. Pediatrics 138, e20160123 (2016).
Morales-Demori, R. Congenital heart disease and cardiac procedural outcomes in patients with trisomy 21 and Turner syndrome. Congenit. Heart Dis. 12, 820–827 (2017).
Russell, M. W., Chung, W. K., Kaltman, J. R. & Miller, T. A. Advances in the understanding of the genetic determinants of congenital heart disease and their impact on clinical outcomes. J. Am. Heart Assoc. 7, e006906 (2018).
Churchill, S. S., Kieckhefer, G. M., Landis, C. A. & Ward, T. M. Sleep measurement and monitoring in children with Down syndrome: a review of the literature, 1960–2010. Sleep. Med. Rev. 16, 477–488 (2012).
Smith, D. S. Health care management of adults with Down syndrome. Am. Fam. Physician 64, 1031–1039 (2001).
Esbensen, A. J., Hoffman, E. K., Stansberry, E. & Shaffer, R. Convergent validity of actigraphy with polysomnography and parent reports when measuring sleep in children with Down syndrome. J. Intellect. Disabil. Res. 62, 281–291 (2018).
Hill, C. M. et al. Home oximetry to screen for obstructive sleep apnoea in Down syndrome. Arch. Dis. Child. 103, 962–967 (2018).
Venekamp, R. P. et al. Tonsillectomy or adenotonsillectomy versus non-surgical management for obstructive sleep-disordered breathing in children. Cochrane Database Syst. Rev. 10, CD011165 (2015).
Purdy, I. B., Singh, N., Brown, W. L., Vangala, S. & Devaskar, U. P. Revisiting early hypothyroidism screening in infants with Down syndrome. J. Perinatol. 34, 936–940 (2014).
Iughetti, L., Lucaccioni, L., Fugetto, F., Mason, A. & Predieri, B. Thyroid function in Down syndrome. Expert Rev. Endocrinol. Metab. 10, 525–532 (2015).
Sarici, D. et al. Thyroid functions of neonates with Down syndrome. Ital. J. Pediatr. 38, 44 (2012).
Hardy, O. et al. Hypothyroidism in Down syndrome: screening guidelines and testing methodology. Am. J. Med. Genet. A 124, 436–437 (2004).
Bittles, A. H. & Glasson, E. J. Clinical, social, and ethical implications of changing life expectancy in Down syndrome. Dev. Med. Child. Neurol. 46, 282–286 (2004).
Hithersay, R. et al. Association of dementia with mortality among adults with Down syndrome older than 35 years. JAMA Neurol. 76, 152–160 (2019). This paper identifies AD as the most prominent cause of mortality in adults with DS.
Strydom, A. et al. Dementia in older adults with intellectual disabilities—epidemiology, presentation, and diagnosis. J. Policy Pract. Intellect. Disabil. 7, 96–110 (2010).
Ballard, C., Mobley, W., Hardy, J., Williams, G. & Corbett, A. Dementia in Down’s syndrome. Lancet Neurol. 15, 622–636 (2016).
Bayen, E., Possin, K. L., Chen, Y., Cleret de Langavant, L. & Yaffe, K. Prevalence of aging, dementia, and multimorbidity in older adults with Down syndrome. JAMA Neurol. 75, 1399–1406 (2018).
Lott, I. T. & Head, E. Dementia in Down syndrome: unique insights for Alzheimer disease research. Nat. Rev. Neurol. 15, 135–147 (2019).
Startin, C. M. et al. Cognitive markers of preclinical and prodromal Alzheimer’s disease in Down syndrome. Alzheimers Dement. 15, 245–257 (2019).
Firth, N. C. et al. Aging related cognitive changes associated with Alzheimer’s disease in Down syndrome. Ann. Clin. Transl. Neurol. 5, 741–751 (2018).
De Simone, R., Puig, X. S., Gelisse, P., Crespel, A. & Genton, P. Senile myoclonic epilepsy: delineation of a common condition associated with Alzheimer’s disease in Down syndrome. Seizure 19, 383–389 (2010).
Royal College of Psychiatrists & The British Psychological Society. Dementia and People with Intellectual Disabilities: Guidance on the assessment, diagnosis, interventions and support of people with intellectual disabilities who develop dementia. https://www.dsrf.org/media/REP77%20final%20proof%20(3).pdf (The British Psychological Society, 2015).
Eady, N. et al. Impact of cholinesterase inhibitors or memantine on survival in adults with Down syndrome and dementia: clinical cohort study. Br. J. Psychiatry 212, 155–160 (2018).
Livingstone, N., Hanratty, J., McShane, R. & Macdonald, G. Pharmacological interventions for cognitive decline in people with Down syndrome. Cochrane Database Syst. Rev. 10, CD011546 (2015).
Dodd, K. et al. Consensus statement of the international summit on intellectual disability and Dementia related to post-diagnostic support. Aging Mental Health 22, 1406–1415 (2018).
Sanmaneechai, O. et al. Treatment outcomes of West syndrome in infants with Down syndrome. Pediatric Neurol. 48, 42–47 (2013).
Gholipour, T., Mitchell, S., Sarkis, R. A. & Chemali, Z. The clinical and neurobehavioral course of Down syndrome and dementia with or without new-onset epilepsy. Epilepsy Behav. 68, 11–16 (2017).
Austeng, M. E. et al. Otitis media with effusion in children with in Down syndrome. Int. J. Pediatr Otorhinolaryngol. 77, 1329–1332 (2013).
Fisher, P. G. Congenital hearing loss in Down syndrome. J. Pediatr. 166, 1–3 (2015).
Shott, S. R., Joseph, A. & Heithaus, D. Hearing loss in children with Down syndrome. Int. J. Pediatr. Otorhinolaryngol. 61, 199–205 (2001).
Krinsky-McHale, S. J. et al. Vision deficits in adults with Down syndrome. J. Appl. Res. Intellect. Disabil. 27, 247–263 (2014).
Li, E. Y., Chan, T. C., Lam, N. M. & Jhanji, V. Cataract surgery outcomes in adult patients with Down’s syndrome. Br. J. Ophthalmol. 98, 1273–1276 (2014).
Brockmeyer, D. Down syndrome and craniovertebral instability. Topic review and treatment recommendations. Pediatr. Neurosurg. 31, 71–77 (1999).
Capone, G., Goyal, P., Ares, W. & Lannigan, E. Neurobehavioral disorders in children, adolescents, and young adults with Down syndrome. Am. J. Med. Genet. C 142, 158–172 (2006).
Mantry, D. et al. The prevalence and incidence of mental ill-health in adults with Down syndrome. J. Intellect. Disabil. Res. 52, 141–155 (2008).
Tassé, M. J. et al. Psychiatric conditions prevalent among adults with Down syndrome. J. Policy Pract. Intellect. Disabil. 13, 173–180 (2016).
Mircher, C. et al. Acute regression in young people with Down syndrome. Brain Sci. 7, 57 (2017).
Spendelow, J. S. Assessment of mental health problems in people with Down syndrome: key considerations. Br. J. Learn. Disabil. 39, 306–313 (2011).
Nevill, R. E. & Benson, B. A. Risk factors for challenging behaviour and psychopathology in adults with Down syndrome. J.Intellect. Disabil. Res. 62, 941–951 (2018).
Liogier d’Ardhuy, X. et al. Assessment of cognitive scales to examine memory, executive function and language in individuals with Down syndrome: implications of a 6-month observational study. Front. Behav. Neurosci. 9, 300 (2015).
Dykens, E. M. Psychiatric and behavioral disorders in persons with Down syndrome. Ment. Retard. Dev. Disabil. Res. Rev. 13, 272–278 (2007).
Aman, M. G., Buican, B. & Arnold, L. E. Methylphenidate treatment in children with borderline IQ and mental retardation: analysis of three aggregated studies. J. Child. Adolesc. Psychopharmacol. 13, 29–40 (2003).
Guedj, F., Bianchi, D. W. & Delabar, J. M. Prenatal treatment of Down syndrome: a reality? Curr. Opin. Obstet. Gynecol. 26, 92–103 (2014).
Bianchi, D. W. From prenatal genomic diagnosis to fetal personalized medicine: progress and challenges. Nat. Med. 18, 1041–1051 (2012).
Stagni, F., Giacomini, A., Emili, M., Guidi, S. & Bartesaghi, R. Neurogenesis impairment: an early developmental defect in Down syndrome. Free. Radic. Biol. Med. 114, 15–32 (2018).
Guidi, S. et al. Neurogenesis impairment and increased cell death reduce total neuron number in the hippocampal region of fetuses with Down syndrome. Brain Pathol. 18, 180–197 (2008).
Guidi, S., Ciani, E., Bonasoni, P., Santini, D. & Bartesaghi, R. Widespread proliferation impairment and hypocellularity in the cerebellum of fetuses with Down syndrome. Brain Pathol. 21, 361–373 (2011).
Dierssen, M. & de Lagran, M. M. DYRK1A (dual-specificity tyrosine-phosphorylated and -regulated kinase 1A): a gene with dosage effect during development and neurogenesis. ScientificWorldJournal 6, 1911–1922 (2006).
Tarui, T. et al. Quantitative MRI analyses of regional brain growth in living fetuses with Down syndrome. Cereb. Cortex. https://doi.org/10.1093/cercor/bhz094 (2019).
Stagni, F., Giacomini, A., Guidi, S., Ciani, E. & Bartesaghi, R. Timing of therapies for Down syndrome: the sooner, the better. Front. Behav. Neurosci. 9, 265 (2015).
de Wert, G., Dondorp, W. & Bianchi, D. W. Fetal therapy for Down syndrome: an ethical exploration. Prenat. Diagn. 37, 222–228 (2017).
Incerti, M. et al. Prenatal treatment prevents learning deficit in Down syndrome model. PLoS One 7, e50724 (2012).
Kelley, C. M. et al. Effects of maternal choline supplementation on the septohippocampal cholinergic system in the Ts65Dn mouse model of Down syndrome. Curr. Alzheimer Res. 13, 84–96 (2016).
McElyea, S. D. et al. Influence of prenatal EGCG treatment and Dyrk1a dosage reduction on craniofacial features associated with Down syndrome. Hum. Mol. Genet. 25, 4856–4869 (2016).
Nakano-Kobayashi, A. et al. Prenatal neurogenesis induction therapy normalizes brain structure and function in Down syndrome mice. Proc. Natl Acad. Sci. USA 114, 10268–10273 (2017).
Izzo, A. et al. Metformin restores the mitochondrial network and reverses mitochondrial dysfunction in Down syndrome cells. Hum. Mol. Genet. 26, 1056–1069 (2017).
Guidi, S. et al. Prenatal pharmacotherapy rescues brain development in a Down’s syndrome mouse model. Brain 137, 380–401 (2014).
Gao, S. Y. et al. Fluoxetine and congenital malformations: a systematic review and meta-analysis of cohort studies. Br. J. Clin. Pharmacol. 83, 2134–2147 (2017).
Chan, M. et al. The burden of respiratory syncytial virus (RSV) associated acute lower respiratory infections in children with Down syndrome: a systematic review and meta-analysis. J. Glob. Health 7, 020413 (2017).
Grut, V., Söderström, L. & Naumburg, E. National cohort study showed that infants with Down’s syndrome faced a high risk of hospitalisation for the respiratory syncytial virus. Acta Paediatr. 106, 1519–1524 (2017).
Bloemers, B. L. P. et al. Increased risk of respiratory tract infections in children with Down syndrome: the consequence of an altered immune system. Microbes Infect. 12, 799–808 (2010).
Giménez-Barcons, M. et al. Autoimmune predisposition in Down syndrome may result from a partial central tolerance failure due to insufficient intrathymic expression of AIRE and peripheral antigens. J. Immunol. 193, 3872–3879 (2014).
Swigonski, N. L., Kuhlenschmidt, H. L., Bull, M. J., Corkins, M. R. & Downs, S. M. Screening for celiac disease in asymptomatic children with Down syndrome: cost-effectiveness of preventing lymphoma. Pediatrics 118, 594–602 (2006).
Tunstall, O. et al. Guidelines for the investigation and management of transient leukaemia of Down syndrome. Br. J. Haematol. 182, 200–211 (2018).
Mateos, M. K., Barbaric, D., Byatt, S.-A., Sutton, R. & Marshall, G. M. Down syndrome and leukemia: insights into leukemogenesis and translational targets. Transl. Pediatr. 4, 76–92 (2015).
Gamis, A. S. & Smith, F. O. Transient myeloproliferative disorder in children with Down syndrome: clarity to this enigmatic disorder. Br. J. Haematol. 159, 277–287 (2012).
Skotko, B. G., Levine, S. P., Macklin, E. A. & Goldstein, R. D. Family perspectives about Down syndrome. Am. J. Med. Genet. A 170, 930–941 (2016).
Van Herwegen, J., Ashworth, M. & Palikara, O. Parental views on special educational needs provision: cross-syndrome comparisons in Williams syndrome, Down syndrome, and autism spectrum disorders. Res. Dev. Disabil. 80, 102–111 (2018).
Buckley, S., Bird, G., Sacks, B. & Archer, T. A comparison of mainstream and special education for teenagers with Down syndrome: implications for parents and teachers. Syndrome Res. Pract. 9, 54–67 (2006).
Kumin, L. & Schoenbrodt, L. Employment in adults with Down syndrome in the United States: results from a national survey. Appl. Res. Intellect. Disabil. 29, 330–345 (2015).
Mihaila, I. et al. Leisure activity and caregiver involvement in middle-aged and older adults with Down syndrome. Intellect. Dev. Disabil. 55, 97–109 (2017).
Xanthopoulos, M. S. et al. Caregiver-reported quality of life in youth with Down syndrome. J. Pediatr. 189, 98–104.e1 (2017).
Diaz, K. M. Physical inactivity among parents of children with and without Down syndrome: the National Health Interview Survey. J. Intellect. Disabil. Res. 64, 38–44 (2020).
Hardee, J. P. & Fetters, L. The effect of exercise intervention on daily life activities and social participation in individuals with Down syndrome: a systematic review. Res. Dev. Disabil. 62, 81–103 (2017).
O’Roak, B. J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012).
Becker, W., Soppa, U. & Tejedor, F. J. DYRK1A: a potential drug target for multiple Down syndrome neuropathologies. CNS Neurol. Disord. Drug Targets 13, 26–33 (2014).
Mowery, C. T. et al. Trisomy of a Down syndrome critical region globally amplifies transcription via HMGN1 overexpression. Cell Rep. 25, 1898–1911.e5 (2018).
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 541, 331–338 (2017).
Annus, T. et al. The Down syndrome brain in the presence and absence of fibrillar β-amyloidosis. Neurobiol. Aging 53, 11–19 (2017). Describes the age of onset of amyloid deposition in the brain of individuals with DS using PET imaging.
Rafii, M. S. Tau PET imaging for staging of Alzheimer’s disease in Down syndrome. Dev. Neurobiol. 79, 711–715 (2019).
Fortea, J. et al. Plasma and CSF biomarkers for the diagnosis of Alzheimer’s disease in adults with Down syndrome: a cross-sectional study. Lancet Neurol. 17, 860–869 (2018). Explores age-related changes in fluid biomarkers associated with AD in individuals with DS to show similarities with sporadic AD.
Strydom, A. et al. Neurofilament light as a blood biomarker for neurodegeneration in Down syndrome. Alzheimer’s Res. Ther. 10, 39 (2018).
Xiao, M. F. et al. NPTX2 and cognitive dysfunction in Alzheimer’s disease. eLife 6, e23798 (2017).
Rafii, M. S. et al. Plasma neurofilament light and Alzheimer’s disease biomarkers in Down syndrome: results from the Down syndrome biomarker initiative (DSBI). J. Alzheimers Dis. 70, 131–138 (2019).
Das, I. & Reeves, R. H. The use of mouse models to understand and improve cognitive deficits in Down syndrome. Dis. Model.Mech. 4, 596–606 (2011).
Duchon, A. et al. Identification of the translocation breakpoints in the Ts65Dn and Ts1Cje mouse lines: relevance for modeling Down syndrome. Mamm. Genome 22, 674–684 (2011).
Reinholdt, L. et al. Molecular characterization of the translocation breakpoints in the Down syndrome mouse model Ts65Dn. Mamm. Genome 22, 685–691 (2011).
Kanekiyo, T., Xu, H. & Bu, G. ApoE and Aβ in Alzheimer’s disease: accidental encounters or partners? Neuron 81, 740–754 (2014).
Startin, C. M. et al. The LonDownS adult cognitive assessment to study cognitive abilities and decline in Down syndrome. Wellcome Open Res. 1, 11 (2016).
Sinai, A. et al. Predictors of age of diagnosis and survival of Alzheimer’s disease in Down syndrome. J. Alzheimers Dis. 61, 717–728 (2018).
Acknowledgements
The authors thank the members of the London Down Syndrome (LonDownS) Consortium, G. de Graaf of the Dutch Down Syndrome Foundation and F. Buckley of Down Syndrome Education International for their review of the epidemiology section of this article. Work in the authors’ laboratories and clinics was supported by grants from the SNF, EU, ERC, Jerome Lejeune, and ChildCare Foundations to S.E.A.; a Wellcome Trust Strategic Award (grant number 098330/Z/12/Z) conferred upon the LonDownS Consortium, an MRC project grant (LonDownsPREVENT MR/S011277/1), and grants from the EU Joint Programme - Neurodegenerative Disease Research (MR/R024901/1, as part of the HEROES consortium), Network of Centres of Excellence in Neurodegeneration (COEN) (MR/S005145/1), Lumind Foundation and Jerome Lejeune Foundation to A.S.; the Jerome Lejeune Foundation USA, Anna and John Sie Foundation, US National Institutes of Health (NIH; HD42053-10, UL1TR001064, ZIA HG200399-04) to D.W.B.; NIH and Lumind Foundation to S.L.S.; HD038384-20, HD098540 and the Lumind Foundation to R.H.R.; and the Alzheimer’s Society and BRC to S.P. The authors thank all past and present members of their laboratories, their collaborators and the patients and their families for their inspiration and support.
Author information
Authors and Affiliations
Contributions
Introduction (S.E.A.); Epidemiology (B.G.S.); Mechanisms/pathophysiology (S.E.A., M.S.R., S.L.S. and R.H.R.); Diagnosis, screening and prevention (S.E.A. and D.W.B.); Management (A.S. and S.E.P.); Quality of life (A.S. and S.E.P.); Outlook (S.E.A.).
Corresponding author
Ethics declarations
Competing interests
S.E.A. is the co-founder and CEO of MediGenome, a clinical and laboratory diagnostic company. B.G.S. occasionally consults on the topic of Down syndrome through Gerson Lehrman Group. B.G.S. receives remuneration from Down syndrome non-profit organizations for speaking engagements and associated travel expenses. B.G.S. receives annual royalties from Woodbine House, Inc., for the publication of his book, Fasten Your Seatbelt: A Crash Course on Down Syndrome for Brothers and Sisters. Within the past 2 years, B.G.S. has received research funding from F. Hoffmann-La Roche, Inc., and LuMind IDSC Down Syndrome Foundation to conduct clinical trials for people with Down syndrome. B.G.S. is occasionally asked to serve as an expert witness in legal cases where Down syndrome is discussed. B.G.S. serves in a non-paid capacity on the Honorary Board of Directors for the Massachusetts Down Syndrome Congress and the Professional Advisory Committee for the National Center for Prenatal and Postnatal Down Syndrome Resources. B.G.S. has a sister with Down syndrome. M.S.R. is a consultant to AC Immune SA. A.S. has consulted for Roche Pharmaceuticals, ONO Pharma, Aelis Farma and AC Immune, and he serves on the Scientific Advisory Board of ProMIS Neurosciences. A.S. and S.E.P. provide clinical services within the UK National Health Service to individuals with Down syndrome. The remaining authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Down’s Syndrome Association and the Down Syndrome Medical Interest Group fact sheet: http://www.perinatalservicesbc.ca/Documents/Guidelines-Standards/Maternal/DownSyndromePracticeResource.pdf
Protein-coding genes on HSA21: https://www.ensembl.org/Homo_sapiens/Location/Chromosome?r=21
Rights and permissions
About this article
Cite this article
Antonarakis, S.E., Skotko, B.G., Rafii, M.S. et al. Down syndrome. Nat Rev Dis Primers 6, 9 (2020). https://doi.org/10.1038/s41572-019-0143-7
Accepted:
Published:
DOI: https://doi.org/10.1038/s41572-019-0143-7
This article is cited by
-
The detailed profile of congenital heart diseases in 254 children with Down syndrome in Saudi Arabia
The Cardiothoracic Surgeon (2024)
-
Walking capacity and its association with quality of life among children with down syndrome in Saudi Arabia
BMC Pediatrics (2024)
-
Nationwide hospitalizations of patients with down syndrome and congenital heart disease over a 15-year period
European Journal of Pediatrics (2024)
-
Trisomy 21-driven metabolite alterations are linked to cellular injuries in Down syndrome
Cellular and Molecular Life Sciences (2024)
-
The Ethical, Legal, and Social Implications of Genomics and Disability: Findings from a Scoping Review and Their Human Rights Implications
Advances in Neurodevelopmental Disorders (2024)