Abstract
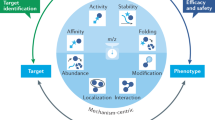
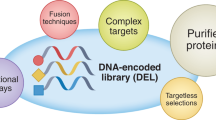
Affinity selection-mass spectrometry (AS-MS) is a high-throughput screening (HTS) technique for drug discovery that enables rapid screening of large collections of compounds to identify ligands for a specific biomolecular target. AS-MS is a binding assay that is insensitive to the functional effects a ligand might have, which is important because it lets us identify novel ligands irrespective of their binding site. This approach is gaining popularity, notably due to its role in the emergence of useful agents for targeted protein degradation. This Perspective highlights the use of AS-MS techniques to explore broad chemical space and identify small-molecule ligands for biological targets that have proven challenging to address with other screening paradigms. We present chemical structures of reported AS-MS hits to illustrate the potential of this screening approach to deliver high-quality hits for further optimization. AS-MS has, thus, evolved from being an infrequent alternative to traditional HTS or DNA-encoded library strategies to now firmly establishing itself as a HTS approach for drug discovery.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mignani, S., Huber, S., Tomás, H., Rodrigues, J. & Majoral, J.-P. Why and how have drug discovery strategies in pharma changed? What are the new mindsets? Drug Discov. Today 21, 239–249 (2016).
Erlanson, D. A., McDowell, R. S. & O’Brien, T. Fragment-based drug discovery. J. Med. Chem. 47, 3463–3482 (2004).
Erlanson, D. A., Fesik, S. W., Hubbard, R. E., Jahnke, W. & Jhoti, H. Twenty years on: the impact of fragments on drug discovery. Nat. Rev. Drug Discov. 15, 605–619 (2016).
Yuen, L. H. & Franzini, R. M. Achievements, challenges, and opportunities in DNA-encoded library research: an academic point of view. ChemBioChem 18, 829–836 (2017).
Annis, D. A. et al. An affinity selection–mass spectrometry method for the identification of small molecule ligands from self-encoded combinatorial libraries: Discovery of a novel antagonist of E. coli dihydrofolate reductase. Int. J. Mass. Spectrom. 238, 77–83 (2004).
Schreiber, S. L. A chemical biology view of bioactive small molecules and a binder-based approach to connect biology to precision medicines. Isr. J. Chem. 59, 52–59 (2018).
Toure, M. & Crews, C. M. Small-molecule PROTACS: new approaches to protein degradation. Angew. Chem. Int. Ed. Engl. 55, 1966–1973 (2016).
No Authors Listed. Retooling chemical probes. Nat. Chem. Biol. 6, 157 (2010).
Annis, D. A., Nickbarg, E., Yang, X., Ziebell, M. R. & Whitehurst, C. E. Affinity selection-mass spectrometry screening techniques for small molecule drug discovery. Curr. Opin. Chem. Biol. 11, 518–526 (2007).
Bergsdorf, C. & Ottl, J. Affinity-based screening techniques: their impact and benefit to increase the number of high quality leads. Expert Opin. Drug Discov. 5, 1095–1107 (2010).
Andrews, C. L., Ziebell, M. R., Nickbarg, E. & Yang, X. in Protein and Peptide Mass Spectrometry in Drug Discovery Ch. 10 (eds Gross, M. L., Chen G. & Pramanik, B. N.) 253–286 (Wiley, 2011).
Flusberg, D. A. et al. Identification of G-quadruplex-binding inhibitors of Myc expression through affinity selection–mass spectrometry. SLAS Discov. 24, 142–157 (2019).
Zehender, H., Le Goff, F., Lehmann, N., Filipuzzi, I. & Mayr, L. M. SpeedScreen: the “missing link” between genomics and lead discovery. J. Biomol. Screen. 9, 498–505 (2004).
Zehender, H. & Mayr, L. M. Application of high-throughput affinity-selection mass spectrometry for screening of chemical compound libraries in lead discovery. Expert Opin. Drug Discov. 2, 285–294 (2007).
Annis, A., Chuang, C.-C. & Nazef, N. in Mass Spectrometry in Medicinal Chemistry Ch. 3 (eds Wanner, K. T. & Höfner, G.) (Wiley, 2007).
Comess, K. M. et al. An ultraefficient affinity-based high-throughput screening process: application to bacterial cell wall biosynthesis enzyme MurF. J. Biomol. Screen. 11, 743–754 (2006).
Comess, K. M. et al. Kinase drug discovery by affinity selection/mass spectrometry (ASMS): application to DNA damage checkpoint kinase Chk1. J. Biomol. Screen. 11, 755–764 (2006).
Schriemer, D. C., Bundle, D. R., Li, L. & Hindsgaul, O. Micro-scale frontal affinity chromatography with mass spectrometric detection: a new method for the screening of compound libraries. Angew. Chem. Int. Ed. 37, 3383–3387 (1999).
Slon-Usakiewicz, J. J., Ng, W., Dai, J. R., Pasternak, A. & Redden, P. R. Frontal affinity chromatography with MS detection (FAC-MS) in drug discovery. Drug Discov. Today 10, 409–416 (2005).
Rush, M. D., Walker, E. M., Burton, T. & van Breemen, R. B. Magnetic microbead affinity selection screening (MagMASS) of botanical extracts for inhibitors of 15-lipoxygenase. J. Nat. Prod. 79, 2898–2902 (2016).
Rush, M. D., Walker, E. M., Prehna, G., Burton, T. & van Breemen, R. B. Development of a magnetic microbead affinity selection screen (MagMASS) using mass spectrometry for ligands to the retinoid X receptor-α. J. Am. Soc. Mass. Spectrom. 28, 479–485 (2017).
Lu, Y. et al. Accelerating the throughput of affinity mass spectrometry-based ligand screening toward a G protein-coupled receptor. Anal. Chem. 91, 8162–8169 (2019).
Qin, S. et al. High-throughput identification of G protein-coupled receptor modulators through affinity mass spectrometry screening. Chem. Sci. 9, 3192–3199 (2018).
Chen, X. et al. Identification of inhibitors of the antibiotic-resistance target New Delhi metallo-β-lactamase 1 by both nanoelectrospray ionization mass spectrometry and ultrafiltration liquid chromatography/mass spectrometry approaches. Anal. Chem. 85, 7957–7965 (2013).
Chen, X. et al. A ligand-observed mass spectrometry approach integrated into the fragment based lead discovery pipeline. Sci. Rep. 5, 8361 (2015).
VanderPorten, E. C., Scholle, M. D., Sherrill, J., Tran, J. C. & Liu, Y. Identification of small-molecule noncovalent binders utilizing SAMDI technology. SLAS Discov. 22, 1211–1217 (2017).
Qin, S. et al. Multiple ligand detection and affinity measurement by ultrafiltration and mass spectrometry analysis applied to fragment mixture screening. Anal. Chim. Acta 886, 98–106 (2015).
Fu, X. et al. Novel chemical ligands to Ebola virus and Marburg virus nucleoproteins identified by combining affinity mass spectrometry and metabolomics approaches. Sci. Rep. 6, 29680 (2016).
Siu, T. et al. Discovery of a novel cGAMP competitive ligand of the inactive form of STING. ACS Med. Chem. Lett. 10, 92–97 (2019).
Petrilli, W. L. et al. From screening to targeted degradation: strategies for the discovery and optimization of small molecule ligands for PCSK9. Cell Chem. Biol. 27, 32–40 (2020).
Valeur, E. et al. New modalities for challenging targets in drug discovery. Angew. Chem. Int. Ed. Engl. 56, 10294–10323 (2017).
Shoichet, B. K. Virtual screening of chemical libraries. Nature 432, 862–865 (2004).
Zhang, T. et al. Definitive metabolite identification coupled with automated ligand identification system (ALIS) technology: a novel approach to uncover structure–activity relationships and guide drug design in a factor IXa inhibitor program. J. Med. Chem. 59, 1818–1829 (2016).
Zhang, B. et al. A novel G protein-biased and subtype-selective agonist for a G protein-coupled receptor discovered from screening herbal extracts. ACS Cent. Sci. 6, 213–225 (2020).
Annis, D. A. et al. Inhibitors of the lipid phosphatase SHIP2 discovered by high throughput affinity selection-mass spectrometry screening of combinatorial libraries. Comb. Chem. High Throughput Screen. 12, 760–771 (2009).
Zhang, H. Acoustic dispensing-mass spectrometry: the next high throughput bioanalytical platform for early drug discovery. Bioanalysis 9, 1619–1621 (2017).
Jenkins, J. & Cook, M. Mosquito®: An accurate nanoliter dispensing technology. JALA 9, 257–261 (2004).
Makara, G. M., Nash, H., Zheng, Z., Orminati, J. P. A. & Wintner, E. A. A reagent-based strategy for the design of large combinatorial libraries: a preliminary experimental validation. Mol. Divers. 7, 3–14 (2003).
Gesmundo, N. J. et al. Nanoscale synthesis and affinity ranking. Nature 557, 228–232 (2018).
Kumar, K. & Waldmann, H. Synthesis of natural product inspired compound collections. Angew. Chem. Int. Ed. 48, 3224–3242 (2009).
Nelson, A. & Roche, D. Innovative approaches to the design and synthesis of small molecule libraries. Bioorg. Med. Chem. 23, 2613 (2015).
Oprea, T. I., Davis, A. M., Teague, S. J. & Leeson, P. D. Is there a difference between leads and drugs? A historical perspective. J. Chem. Inf. Comput. Sci. 41, 1308–1315 (2001).
Polinsky, A. in The Practice of Medicinal Chemistry 3rd edn (ed. Wermuth, C. G.) 244–254 (Elsevier, 2008).
MacArrón, R. & Luengo, J. I. Yin and Yang in medicinal chemistry: what does drug-likeness mean? Future Med. Chem. 3, 505–507 (2011).
Oprea, T. I. Current trends in lead discovery: are we looking for the appropriate properties? Mol. Divers. 5, 199–208 (2000).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–25 (2012).
Lipinski, C. A. Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov. Today Technol. 1, 337–341 (2004).
Kuenemann, M. A., Labbé, C. M., Cerdan, A. H. & Sperandio, O. Imbalance in chemical space: how to facilitate the identification of protein–protein interaction inhibitors. Sci. Rep. 6, 23815 (2016).
Ran, X. & Gestwicki, J. E. Inhibitors of protein–protein interactions (PPIs): an analysis of scaffold choices and buried surface area. Curr. Opin. Chem. Biol. 44, 75–86 (2018).
Doak, B. C., Zheng, J., Dobritzsch, D. & Kihlberg, J. How beyond rule of 5 drugs and clinical candidates bind to their targets. J. Med. Chem. 59, 2312–2327 (2016).
Wilson, A. J. Inhibition of protein–protein interactions using designed molecules. Chem. Soc. Rev. 38, 3289–3300 (2009).
Lovering, F., Bikker, J. & Humblet, C. Escape from flatland: Increasing saturation as an approach to improving clinical success. J. Med. Chem. 52, 6752–6756 (2009).
Quartararo, A. J. et al. Ultra-large chemical libraries for the discovery of high-affinity peptide binders. Nat. Commun. 11, 3183 (2020).
Lam, K. S. et al. A new type of synthetic peptide library for identifying ligand-binding activity. Nature 354, 82–84 (1991).
Furka, Á., Sebestyén, F., Asgedom, M. & Dibó, G. General method for rapid synthesis of multicomponent peptide mixtures. Int. J. Pept. Protein Res. 37, 487–493 (1991).
Fu, Y. et al. Affinity selection-based two-dimensional chromatography coupled with high-performance liquid chromatography-mass spectrometry for discovering xanthine oxidase inhibitors from Radix Salviae Miltiorrhizae. Anal. Bioanal. Chem. 406, 4987–4995 (2014).
Fei, F. et al. Rapid screening and identification of bioactive compounds specifically binding to beta 2-adrenoceptor from San-ao decoction using affinity magnetic fine particles coupled with high-performance liquid chromatography–mass spectrometry. Chin. Med. 13, 49 (2018).
Sun, Y. et al. Ultrafiltration tandem mass spectrometry of estrogens for characterization of structure and affinity for human estrogen receptors. J. Am. Soc. Mass. Spectrom. 16, 271–279 (2005).
Wang, Z. et al. Efficient ligand discovery from natural herbs by integrating virtual screening, affinity mass spectrometry and targeted metabolomics. Analyst 144, 2881–2890 (2019).
Malmqvist, M. BIACORE: an affinity biosensor system for characterization of biomolecular interactions. Biochem. Soc. Trans. 27, 335–340 (1999).
Rich, R. L. & Myszka, D. G. Advances in surface plasmon resonance biosensor analysis. Curr. Opin. Biotechnol. 11, 54–61 (2000).
Comess, K. M. et al. Discovery and characterization of non-ATP site inhibitors of the mitogen activated protein (MAP) kinases. ACS Chem. Biol. 6, 234–244 (2011).
Su, H.-P. et al. Structural characterization of nonactive site, TrkA-selective kinase inhibitors. Proc. Natl Acad. Sci. USA 114, E297–E306 (2017).
Song, X. S. et al. Identification of DGAT2 inhibitors using mass spectrometry. J. Biomol. Screen. 21, 117–126 (2016).
Walker, S. S. et al. Affinity selection–mass spectrometry identifies a novel antibacterial RNA polymerase inhibitor. ACS Chem. Biol. 12, 1346–1352 (2017).
Coburn, C. A. et al. Identification of a small molecule nonpeptide active site β-secretase inhibitor that displays a nontraditional binding mode for aspartyl proteases. J. Med. Chem. 47, 6117–6119 (2004).
Pantoliano, M. W. et al. Large increases in general stability for subtilisin BPN′ through incremental changes in the free energy of unfolding. Biochemistry 28, 7205–7213 (1989).
Brown, N. et al. A chemoinformatics analysis of hit lists obtained from high-throughput affinity-selection screening. J. Biomol. Screen. 11, 123–130 (2006).
Feng, B. Y. & Shoichet, B. K. A detergent-based assay for the detection of promiscuous inhibitors. Nat. Protoc. 1, 550–553 (2006).
Jadhav, A. et al. Quantitative analyses of aggregation, autofluorescence, and reactivity artifacts in a screen for inhibitors of a thiol protease. J. Med. Chem. 53, 37–51 (2010).
Whitehurst, C. E. et al. Application of affinity selection-mass spectrometry assays to purification and affinity-based screening of the chemokine receptor CXCR4. Comb. Chem. High Throughput Screen. 15, 473–485 (2012).
Whitehurst, C. E. et al. Discovery and characterization of orthosteric and allosteric muscarinic M2 acetylcholine receptor ligands by affinity selection–mass spectrometry. J. Biomol. Screen. 11, 194–207 (2006).
Gabriel, J., Höfner, G. & Wanner, K. T. A library screening strategy combining the concepts of MS binding assays and affinity selection mass spectrometry. Front. Chem. 7, 665 (2019).
Igonet, S. et al. Enabling STD-NMR fragment screening using stabilized native GPCR: a case study of adenosine receptor. Sci. Rep. 8, 8142 (2018).
Matsui, M. & Corey, D. R. Non-coding RNAs as drug targets. Nat. Rev. Drug Discov. 16, 167–179 (2017).
Rizvi, N. F. et al. Discovery of selective RNA-binding small molecules by affinity-selection mass spectrometry. ACS Chem. Biol. 13, 820–831 (2018).
Rizvi, N. F. et al. Targeting RNA with small molecules: identification of selective, RNA-binding small molecules occupying drug-like chemical space. SLAS Discov. 25, 384–396 (2020).
Petersen, D. N. et al. A small-molecule anti-secretagogue of PCSK9 targets the 80S ribosome to inhibit PCSK9 protein translation. Cell Chem. Biol. 23, 1362–1371 (2016).
Maria, J. P. S. et al. Linking high-throughput screens to identify MoAs and novel inhibitors of Mycobacterium tuberculosis dihydrofolate reductase. ACS Chem. Biol. 12, 2448–2456 (2017).
Yang, X.-X. et al. Development of a mitochondria-based centrifugal ultrafiltration/liquid chromatography/mass spectrometry method for screening mitochondria-targeted bioactive constituents from complex matrixes: herbal medicines as a case study. J. Chromatogr. A 1413, 33–46 (2015).
Tao, Y., Yan, J. & Cai, B. Label-free bio-affinity mass spectrometry for screening and locating bioactive molecules. Mass Spectrom. Rev. https://doi.org/10.1002/mas.21613 (2019).
Kutilek, V. D. et al. Integration of affinity selection–mass spectrometry and functional cell-based assays to rapidly triage druggable target space within the NF-κB pathway. J. Biomol. Screen. 21, 608–619 (2016).
Motoyaji, T. Revolution of small molecule drug discovery by affinity selection-mass spectrometry technology. Chem. Pharm. Bull. 68, 191–193 (2020).
Salcius, M. et al. SEC-TID: a label-free method for small-molecule target identification. J. Biomol. Screen. 19, 917–927 (2014).
McMillan, E. A. et al. Chemistry-first approach for nomination of personalized treatment in lung cancer. Cell. 173, 864–878 (2018).
Musetti, C. et al. High-throughput assessment of structural continuity in biologics. Anal. Chem. 90, 2970–2975 (2018).
Wei, J. N., Belanger, D., Adams, R. P. & Sculley, D. Rapid prediction of electron-ionization mass spectrometry using neural networks. ACS Cent. Sci. 5, 700–708 (2019).
Domingo-Almenara, X. et al. The METLIN small molecule dataset for machine learning-based retention time prediction. Nat. Commun. 10, 5811 (2019).
Boström, J., Brown, D. G., Young, R. J. & Keserü, G. M. Expanding the medicinal chemistry synthetic toolbox. Nat. Rev. Drug Discov. 10, 709–727 (2018).
Piper, D. E. et al. The crystal structure of PCSK9: a regulator of plasma LDL-cholesterol. Structure 15, 545–552 (2007).
Dai, J. et al. Structure of the intramolecular human telomeric G-quadruplex in potassium solution: a novel adenine triple formation. Nucleic Acids Res. 35, 2440–2450 (2007).
Howe, J. A. et al. Selective small-molecule inhibition of an RNA structural element. Nature 526, 672–677 (2015).
Klaholz, B. P. et al. Structure of the Escherichia coli ribosomal termination complex with release factor 2. Nature 421, 90–94 (2003).
Acknowledgements
The authors thank Calixar for their generous gift of purified native target to investigate the A2AR receptor.
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the preparation of this manuscript.
Corresponding author
Ethics declarations
Competing interests
J.-Y.O. and D.R. are cofounders, and R.P. is an employee of Edelris SAS, which has developed the commercial AS-MS service ‘EDEN platform’.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Prudent, R., Annis, D.A., Dandliker, P.J. et al. Exploring new targets and chemical space with affinity selection-mass spectrometry. Nat Rev Chem 5, 62–71 (2021). https://doi.org/10.1038/s41570-020-00229-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41570-020-00229-2
This article is cited by
-
Strategies for developing DNA-encoded libraries beyond binding assays
Nature Chemistry (2022)
-
Affinity Selection from Synthetic Peptide Libraries Enabled by De Novo MS/MS Sequencing
International Journal of Peptide Research and Therapeutics (2022)