Abstract
Self-organization is a prerequisite of biological complexity. At the population level, it amounts to spontaneously sorting different individuals through space and time. Here, we reveal a simple mechanism by which different populations of motile cells can self-organize through a reciprocal control of their motilities. We first show how the reciprocal activation of motility between two populations of engineered Escherichia coli makes an initially mixed population of cells segregate, leading to out-of-phase population oscillations without the need of any preexisting positional or orientational cues. By redesigning the interaction, the original segregation between the two populations can be turned into co-localization. We account for this self-organization using a theoretical model that shows the reciprocal control of motility to be a robust and versatile self-organization pathway in multi-component systems. We finally show how our theoretical and experimental results can be generalized to three interacting bacterial populations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data that support the plots within this paper and other findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
The algorithms used to produce our numerical results are available from the corresponding author upon reasonable request.
Change history
11 April 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41567-024-02500-5
References
Koch, A. & Meinhardt, H. Biological pattern formation: from basic mechanisms to complex structures. Rev. Mod. Phys. 66, 1481 (1994).
Steinberg, M. Differential adhesion in morphogenesis: a modern view. Curr. Opin. Genet. Dev. 17, 281–286 (2007).
Friedl, P. & Gilmour, D. Collective cell migration in morphogenesis, regeneration and cancer. Nat. Rev. Mol. Cell Biol. 10, 445–457 (2009).
Nakamasu, A., Takahashi, G., Kanbe, A. & Kondo, S. Interactions between zebrafish pigment cells responsible for the generation of Turing patterns. Proc. Natl Acad. Sci. USA 106, 8429–8434 (2009).
Theveneau, E. et al. Chase-and-run between adjacent cell populations promotes directional collective migration. Nat. Cell Biol. 15, 763–772 (2013).
Yamanaka, H. & Kondo, S. In vitro analysis suggests that difference in cell movement during direct interaction can generate various pigment patterns in vivo. Proc. Natl Acad. Sci. USA 111, 1867–1872 (2014).
Scheffer, M. & van Nes, E. H. Self-organized similarity, the evolutionary emergence of groups of similar species. Proc. Natl Acad. Sci. USA 103, 6230–6235 (2006).
Hallatschek, O., Hersen, P., Ramanathan, S. & Nelson, D. R. Genetic drift at expanding frontiers promotes gene segregation. Proc. Natl Acad. Sci. USA 104, 19926–19930 (2007).
Reichenbach, T., Mobilia, M. & Frey, E. Mobility promotes and jeopardizes biodiversity in rock–paper–scissors games. Nature 448, 1046–1049 (2007).
Barbier, M., Arnoldi, J.-F., Bunin, G. & Loreau, M. Generic assembly patterns in complex ecological communities. Proc. Natl Acad. Sci. USA 115, 2156–2161 (2018).
Murray, C. J. L. Mathematical Biology (Springer, 2003).
Budrene, E. O. & Berg, H. C. Complex patterns formed by motile cells of Escherichia coli. Nature 349, 630–633 (1991).
Vicsek, T. & Zafeiris, A. Collective motion. Phys. Rep. 517, 71–140 (2012).
Marchetti, M. C. et al. Hydrodynamics of soft active matter. Rev. Mod. Phys. 85, 1143 (2013).
Cates, M. E. & Tailleur, J. Motility-induced phase separation. Annu. Rev. Condens. Matter Phys. 6, 219–244 (2015).
Basu, S., Gerchman, Y., Collins, C. H., H., A. F. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Tabor, J. J. et al. A synthetic genetic edge detection program. Cell 137, 1272–1281 (2009).
Payne, S. et al. Temporal control of self-organized pattern formation without morphogen gradients in bacteria. Mol. Syst. Biol. 9, 697 (2013).
Cao, Y. et al. Collective space-sensing coordinates pattern scaling in engineered bacteria. Cell 165, 620–630 (2016).
Kong, W., Blanchard, A. E., Liao, C. & Lu, T. Engineering robust and tunable spatial structures with synthetic gene circuits. Nucleic Acids Res. 45, 1005–1014 (2017).
Wolfe, A. J. & Berg, H. C. Migration of bacteria in semisolid agar. Proc. Natl Acad. Sci. USA 86, 6973–6977 (1989).
Sanna, M. G. & Simon, M. I. In vivo and in vitro characterization of Escherichia coli protein CheZ gain-and loss-of-function mutants. J. Bacteriol. 178, 6275–6280 (1996).
Silversmith, R. E., Levin, M. D., Schilling, E. & Bourret, R. B. Kinetic characterization of catalysis by the chemotaxis phosphatase CheZ—modulation of activity by the phosphorylated CheY substrate. J. Biol. Chem. 283, 756–765 (2008).
Liu, C. et al. Sequential establishment of stripe patterns in an expanding cell population. Science 334, 238–241 (2011).
You, L., Cox, R. S., Weiss, R. & Arnold, F. H. Programmed population control by cell–cell communication and regulated killing. Nature 428, 868–871 (2004).
Balagaddé, F. K. et al. A synthetic Escherichia coli predator–prey ecosystem. Mol. Syst. Biol. 4, 187 (2008).
Fuqua, C. W., Winans, S. C. & Greenberg, E. P. Quorum sensing in bacteria: the LuxR–LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 176, 269–275 (1994).
Gray, K. M., Passador, L., Iglewski, B. H. & Greenberg, E. P. Interchangeability and specificity of components from the quorum-sensing regulatory systems of Vibrio fischeri and Pseudomonas aeruginosa. J. Bacteriol. 176, 3076–3080 (1994).
Hallatschek, O., Hersen, P., Ramanathan, S. & Nelson, D. R. Genetic drift at expanding frontiers promotes gene segregation. Proc. Natl Acad. Sci. USA 104, 19926–19930 (2007).
Cates, M. E., Marenduzzo, D., Pagonabarraga, I. & Tailleur, J. Arrested phase separation in reproducing bacteria creates a generic route to pattern formation. Proc. Natl Acad. Sci. USA 107, 11715–11729 (2010).
Van Saarloos, W. Front propagation into unstable states. Phys. Rep. 386, 29–222 (2003).
Cates, M. & Tailleur, J. When are active Brownian particles and run-and-tumble particles equivalent? Consequences for motility-induced phase separation. Eur. Phys. Lett. 101, 20010 (2013).
Grant, P. K. et al. Orthogonal intercellular signaling for programmed spatial behavior. Mol. Syst. Biol. 12, 849 (2016).
Xue, X., Xue, C. & Tang, M. The role of intracellular signaling in the stripe formation in engineered Escherichia coli populations. PLoS Comput. Biol. 14, e1006178 (2018).
Acknowledgements
We thank M. Cates, H. Chaté, W. Huang, X. Fu and T. Hwa for discussions. A.I.C. acknowledges a doctoral fellowship from DIM ISC. A.I.C., A.D., J.T., J.H. and Y.Z. acknowledge support from ANR/RGC grants Bactterns and A-HKU712/14. J.H. acknowledges support from the Shenzhen Peacock project (KQTD2015033117210153), the Shenzhen Science and Technology Innovation Committee Basic Science Research Grant (JCYJ20170413154523577) and CAS Key Laboratory of Quantitative Engineering Biology, Shenzhen Institutes of Advanced Technology.
Author information
Authors and Affiliations
Contributions
A.I.C., N.Z., Y.Z., A.D., J.H. and J.T. conceived the project. N.Z. and J.H. conceived and realized the biology experiments. A.I.C. and J.T. conceived and realized the theoretical work. Y.Z. and A.D. built the set-up and carried out the measurements. A.I.C., N.Z., Y.Z., A.D., J.H. and J.T. wrote the manuscript. C.L. was involved in applications for grant support of the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Information and Figs. 1–15.
Supplementary Video 1
This video displays an independent replicate using the activator strains, leading to the formation of out-of-phase fluorescent stripe patterns.
Supplementary Video 2
This video displays an independent replicate using the inhibitor strains, leading to the formation of in-phase fluorescent stripe patterns.
Supplementary Data 1
This data file contains the simulation results of the density fields rho_A and rho_B shown in Supplementary Fig. 14, at different time points, as well as their sum at the final time point.
Supplementary Data 2
This data file contains the simulation results of the density fields rho_A and rho_B shown in Supplementary Fig. 15, at different time points, as well as their sum at the final time point.
Source data
Source Data Fig. 1
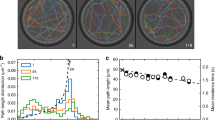
Relative intensity of fluorescence/dark-field signals in Fig. 1c.
Source Data Fig. 2
Relative intensity of fluorescence/dark-field signals in Fig. 2c.
Source Data Fig. 3
Simulation results of density fields in Fig. 3a,b.
Source Data Fig. 4
Relative intensity of fluorescence signals in Fig. 4c,f.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Curatolo, A.I., Zhou, N., Zhao, Y. et al. Cooperative pattern formation in multi-component bacterial systems through reciprocal motility regulation. Nat. Phys. 16, 1152–1157 (2020). https://doi.org/10.1038/s41567-020-0964-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-020-0964-z
This article is cited by
-
Photochromism from wavelength-selective colloidal phase segregation
Nature (2023)
-
Engineered bacterial swarm patterns as spatial records of environmental inputs
Nature Chemical Biology (2023)
-
Non-reciprocity across scales in active mixtures
Nature Communications (2023)
-
Reversible thermal regulation for bifunctional dynamic control of gene expression in Escherichia coli
Nature Communications (2021)