Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has already infected more than 500 million people globally (as of May 2022), creating the coronavirus disease 2019 (COVID-19) pandemic. Nanotechnology has played a pivotal role in the fight against SARS-CoV-2 in various aspects, with the successful development of the two highly effective nanotechnology-based messenger RNA vaccines being the most profound. Despite the remarkable efficacy of mRNA vaccines against the original SARS-CoV-2 strain, hopes for quickly ending this pandemic have been dampened by the emerging SARS-CoV-2 variants, which have brought several new pandemic waves. Thus, novel strategies should be proposed to tackle the crisis presented by existing and emerging SARS-CoV-2 variants. Here, we discuss the SARS-CoV-2 variants from biological and immunological perspectives, and the rational design and development of novel and potential nanotechnology-based strategies to combat existing and possible future SARS-CoV-2 variants. The lessons learnt and design strategies developed from this battle against SARS-CoV-2 variants could also inspire innovation in the development of nanotechnology-based strategies for tackling other global infectious diseases and their future variants.
Similar content being viewed by others
Main
The coronavirus disease 2019 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has already infected millions of people globally. To cope with this unprecedented crisis, scientists from different disciplines have worked collaboratively to develop various strategies. Among these, vaccines are the most effective strategies to prevent infection by SARS-CoV-2, because our own immune system is the most important line of defence against the infection of new viral strains1,2,3. Until now, several vaccines based on different technologies, including messenger RNA, protein subunit, adenoviral-vectored and whole-cell inactivated virus vaccines, have been deployed worldwide4. While most have a protective efficacy of 50–80%, the two lipid nanoparticle (LNP)-based mRNA vaccines from Moderna5 and Pfizer–BioNTech6 (mRNA-1273 and BNT162b2, respectively) have shown much greater protective efficacy, of 94.1% and 95%, respectively. These two mRNA vaccines are currently the most widely used, demonstrating the pivotal role of nanotechnology in the response to the COVID-19 pandemic7.
With the large-scale rollout of the COVID-19 vaccines in many regions, the number of cases of COVID-19 has remarkably declined over time. However, new waves of COVID-19 caused by emerging SARS-CoV-2 variants have posed new risks to global public health. So far, the World Health Organization (WHO) has designated five SARS-CoV-2 variants of concern: B.1.1.7 (Alpha), B.1.351 (Beta), P.1 (Gamma), B.1.617.2 (Delta) and B.1.1.529 (Omicron)8. Accumulating evidence indicates that these new variants have increased transmissibility and virulence along with reduced neutralization, making them even more troublesome and dangerous9. Clinical data have shown that several currently deployed vaccines demonstrate significantly lower protective efficacy against these new variants. Therefore, the crisis arising from the existing and emerging SARS-CoV-2 variants increases the demands for novel strategies.
The SARS-CoV-2 variants
The emergence of SARS-CoV-2 variants of concern
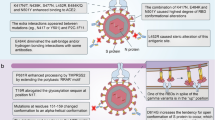
SARS-CoV-2 is an ~29-kilobase single-stranded positive RNA virus with a mutation rate of two single-letter mutations per month, which is relatively slow compared with other RNA viruses owing to its proofreading capabilities (Fig. 1a)10,11. However, due to its rapid spread, more than 4,000 variants have been reported12. Currently, the major concern regarding certain variants is their capability to hamper the immunity created by vaccines or previous infection13. The WHO ranks variants as follows, in order of increasing demand for attention and action: variants under monitoring, variants of interest (VOI) and variants of concern (VOC). VOI and VOC are categorized on the basis of genetic changes predicted or known to affect transmissibility, disease severity, immune escape, and therapeutic and diagnostic escape. Once these predictions manifest on a global scale, variants are reclassified as VOC. Currently, five variants are considered VOC and two are considered VOI (Table 1).
a, The structure of SARS-CoV-2, showing the major components, including the S protein, nucleocapsid protein, envelope protein, membrane protein and viral RNA. The locations of the RBD and NTD in the S protein are highlighted. b, SARS-CoV-2 VOC. The key S-protein mutations in each variant are presented. Protein Data Bank ID: 7DWZ.
While most variant strains carry several mutations, the most studied and worrying mutations are located in the SARS-CoV-2 spike (S) protein14. The S protein comprises >1,200 amino acids, yet only a small 25-amino acid stretch mediates the interactions between its receptor-binding domain (RBD) and the angiotensin-converting enzyme 2 (ACE2) receptor of the host cells. Mutations in and around this interaction area have been noted in all VOC (Fig. 1b)15. Furthermore, RBD mutations have the greatest effect on the ability of antibodies to neutralize the virus16. In the early emerging variants, that is, before Omicron, <3% of the residues in the S protein have mutated, yet this handful of mutations is suspected of severely reducing the function of the neutralizing antibodies17,18.
SARS-CoV-2 variants are exacerbating the pandemic
To determine how variants are affecting the pandemic, institutes and governments are required to piece together epidemiological data on variant spread and laboratory data on how efficient sera samples from vaccinated and previously infected individuals are at neutralizing the specific variant12. While in vitro assays generally yield similar trends across studies, they can be inconsistent, because different studies use different cell and pseudovirus systems14,19,20. Furthermore, while T-cell responses are proposed to play a crucial protective role, they are rarely considered in studies addressing viral immune evasion. In contrast to epitopes responsible for antibody production, T-cell epitopes are located along the full length of the S protein. This suggests a limited effect of viral mutations on cellular immunity11.
How the immunity provided by the vaccines and previous infection is stacking up against the new VOCs is of high interest. Notable mutations include D614G, N501Y, K417N, E484K, L452R and P681H. Among these, the L452R and E484K mutations can dramatically reduce the neutralizing capacity of antibodies and help evade previous immunity from infected or vaccinated individuals’ sera16,17,21. Although the RBD region is studied the most, the N-terminal domain (NTD) also harbours potential targets for antibody neutralization15. This can perhaps explain why the Beta and Gamma strains demonstrate a different profile when tested against immunized sera, even though they have the same RBD mutations22. Furthermore, it is important to consider that some common mutations in the same variant can synergistically induce a more severe effect16.
Currently, sera with high titres of neutralizing antibodies post-infection or -vaccination are capable of neutralizing all current VOCs20,23,24. If, as suggested, high titres of neutralizing antibodies elicited by current vaccines or previous infections protect from infection against all variants, the next important topic to address is waning immunity. Vaccines have induced high titres of virus-specific neutralizing antibodies that, as expected, decline over time13,16. For example, data on the duration of BNT162b2 mRNA vaccine protection demonstrate that neutralizing antibody levels rapidly decreased in the first 3 months after vaccination, followed by a more gradual decrease. It was concluded that, although the humoral response substantially decreased 6 months post-vaccination with BNT162b2, it remains quite potent at preventing severe disease even facing the Delta variants (clinical trial number NCT04368728)25,26,27.
The B.1.1.529 Omicron variant, designated a VOC on 26 November 2021, was the last variant to make the headlines. Omicron gained attention because it has over 30 S-protein mutations, 15 of which are located in the RBD, and spreads rapidly28,29,30,31. Furthermore, Omicron possesses mutations in its RNA-dependent RNA polymerase and its main protease, both of which are targets of antiviral intervention31. While most of these mutations had already been recorded separately in previous strains and confer enhanced viral transmission and immune evasion, their simultaneous appearance in a single strain accompanied by new mutations is concerning. Accordingly, Omicron demonstrates an enhanced ability to escape previous immunity established by vaccines or natural infections, and can evade a large number of monoclonal antibody therapies29,32,33,34.
Nanotechnology solutions for the SARS-CoV-2 variant challenge
SARS-CoV-2 infects through the binding of its S protein to the ACE2 receptor expressed on host cells. The increase in transmission and decrease in antibody neutralization of SARS-CoV-2 variants are closely related to the mutations in their S protein. Thus, targeting the S protein of the variants to inhibit the interaction with ACE2 receptors could be the most straightforward and promising strategy. Nanotechnology offers various solutions, including nanoparticle (NP) vaccine-elicited neutralizing antibodies, engineered neutralizing antibodies and ACE2-based nanodecoys.
Eliciting neutralizing antibodies using NP-based vaccines
Mounting evidence indicates that vaccines are the most efficient solution to mitigate the transmission of SARS-CoV-2. Compared with the mature and widely employed live-attenuated or inactivated vaccines, NP-based vaccines, in particular mRNA-LNP vaccines35,36, have key advantages, including modularity, rapid manufacturing and high efficacy37. Indeed, Moderna launched the phase I clinical trial of an LNP-based mRNA vaccine (mRNA-1273) in a record time of 63 days after sequence selection.
Much of the discussion surrounding vaccine efficacy revolves around T follicular helper (TFH) cells. These are a unique subset of CD4+ T cells that are required for the development of germinal centre (GC) responses and facilitate immunoglobulin class switch, affinity maturation and long-term B-cell memory persistence38. Therefore, antigen-specific TFH cells are crucial for the development of a sustained, broadly protective antibody response. Nucleoside-modified mRNA-LNP vaccines are known to induce high levels of antigen-specific TFH and GC B cells39. Interestingly, mRNA-LNP vaccines outperform inactivated virus vaccines, adjuvanted protein vaccines and live pathogen infection in terms of TFH cell abundance and induction of long-lived plasma and memory B cells40. It is suggested that this is driven by the robust and sustained antigen production from encoded mRNA, as well as the adjuvant effect elicited by both the mRNA and LNPs themselves41. The ionizable lipid in the LNP formulation drives early cytokine production in the draining lymph nodes after intramuscular administration, most importantly through interleukin-6 signalling. This rapid spike in interleukin-6 concentration is critical for the induction of the downstream effectors that drive the TFH and GC responses41. Although the mRNA in the LNP has been modified to reduce sensing by pathogen recognition receptors, it yields cytokine responses that support, to some extent, the generation of TFH cells. The general scheme of TFH-cell differentiation unfolds as follows: (1) priming of naïve CD4+ T cells in the spleen or draining lymph nodes by antigen-presenting dendritic cells, (2) TFH differentiation following expression of costimulatory molecules and migration to the T cell/B cell border, (3) antigen-dependent T and B cells relocate to the follicle to form GCs and (4) GC TFH cells provide essential signals to B cells and drive the selection of high-affinity B-cell clones and the generation of memory B cells38. The initial dendritic cell presentation of SARS-CoV-2 antigen can be derived either from the presentation of processed live viral antigen or from antigen encoded in the mRNA of the vaccine. Regarding SARS-CoV-2 infections, elevated numbers of activated TFH cells are detected in the blood of non-severely affected patients and in the lymph nodes of rhesus macaques following SARS-CoV-2 infection. Importantly, a correlation exists between elevated activated circulating TFH levels and low disease severity. These antigen-specific circulating TFH cells are reported to persist for at least 6 months post-infection38.
Despite the high efficacy of these mRNA vaccines against the original SARS-CoV-2 strain, their efficacy against the newly emerging SARS-CoV-2 variants is compromised due to multiple mutations in the S protein of the virus. Thus, several strategies have been developed or proposed to update current mRNA-LNP vaccines or create new mRNA-LNP vaccines (Fig. 2a), aiming to induce higher levels of neutralizing antibodies that can more efficiently bind to the mutated S protein and subsequently neutralize SARS-CoV-2 variants. After injection, mRNA-LNPs are internalized by antigen-presenting cells, where mRNA is translated into the S-protein antigen by the ribosome (Fig. 2b)42. The resulting S-protein antigen is subsequently degraded to antigen fragments to activate CD8+ T cells; in the meantime, endocytosis of the secreted S-protein antigen activates CD4+ T cells and B cells for the production of neutralizing antibodies. Given that waning vaccine-elicited immunity is one of the major reasons for decreased vaccine efficacy against SARS-CoV-2 variants, enhancing vaccine-elicited immunity by a third booster dose of the current vaccines is the most straightforward and widely proposed strategy to address this issue. Consequently, the Food and Drug Administration authorized a single booster dose of the Pfizer or Moderna mRNA vaccine for individuals 18 years of age and older. Recent studies showed that administration of a third booster dose of Pfizer’s BNT162b2 or Moderna’s mRNA-1273 vaccine to solid organ transplant recipients substantially elevated the immunogenicity of the vaccine in terms of SARS-CoV-2 antibodies43,44, suggesting the robust potential of a booster dose against SARS-CoV-2 variants. Indeed, in another study, a third booster dose of the BNT162b2 vaccine greatly increased the magnitude and breadth of the neutralization of SARS-CoV-2, as well as the Beta and Delta variants45. Despite the current prevalence of the highly contagious, mutated Omicron variant, a third or even more booster doses of the mRNA-1273 or BNT162b2 vaccine might still induce potent neutralizing antibodies against Omicron and provide protection against it, despite reduced efficacy46,47,48.
a, Original NP-based mRNA vaccine and its derivatives, including the variant-modified mRNA vaccine, the chimeric spike mRNA vaccine and the circular RNA vaccine. b, Eliciting immunity through mRNA and protein vaccines. Upon administration, mRNA-LNPs are internalized by antigen-presenting cells, where mRNA is translated into the S-protein antigen by ribosome. The resulting S-protein antigen is processed by proteasome, yielding antigen fragments that are presented on the cell surface through interaction with MHC class I proteins to activate CD8+ T cells. Meanwhile, the secreted S-protein antigen is endocytosed, followed by degradation in endosomes and presentation on the cell surface by interaction with MHC class II proteins to activate CD4+ T cells. Both CD4+ T cells and secreted antigens can stimulate B cells to produce neutralizing antibodies. Similarly, protein vaccines can also elicit immunity by introducing antigens to the cells directly. TCR, T-cell receptor; ER, endoplasmic reticulum; BCR, B-cell receptor. c, Two types of protein NP vaccine contain 24 and 20 RBD units. The RBD-24-mer NP vaccine uses ferritin NPs as the scaffold, and the RBD-20-mer NP vaccine uses helper proteins as building blocks.
Another strategy proposed by Moderna to cope with the SARS-CoV-2 variants is to develop updated mRNA vaccines49. To this end, a modified version of the prototype mRNA-1273 vaccine, named mRNA-1273.351, containing the genetic sequence of the S protein of the Beta variant has been developed (Fig. 2a)50. While a third dose boosting with both mRNA-1273 and mRNA-1273.351 elicited increasing neutralization titres against the variants, the latter appeared to yield numerically higher neutralizing antibody titres against the Beta variant. The S protein of SARS-CoV-2 contains the major immunogenic domains, including the RBD, NTD and subunit 2 (S2)51,52. Instead of encoding the full length of the S protein, a chimeric S mRNA vaccine encoding RBD, NTD and S2 from both SARS-CoV and SARS-CoV-2 produced efficient neutralizing antibodies against newly emergent SARS-CoV-2 variants53. One concern regarding the two current mRNA vaccines is the requirement for an extremely low storage temperature54, which greatly limits the wider deployment of the vaccines. One attempt to address this issue is the development of an LNP-based circular RNA vaccine (Fig. 2a)55. Administration of the circular RNA vaccine containing the RBDs of SARS-CoV-2 variants (Delta and Omicron) to mice and monkeys showed effective protection against the variants, including Delta and Omicron. Compared with the linear mRNA-based vaccines, the circular RNA vaccines exhibit higher stability because of their special circular structure, which cannot be easily degraded by the nuclease. Lyophilization of mRNA-LNPs is another possible solution, although some consider that process too expensive and time consuming37. Furthermore, it has recently been suggested that nucleic acid vaccines may be concealing ‘hitchhiker’ proteins encoded on overlapping open reading frames (ORFs) within the sequences. For example, it has been reported that there are other ORFs that overlap the SARS-CoV-2 S sequence. Therefore, the sequences encoded need to be screened to exclude the translation of undesired peptides that could alter clinical outcomes. This complication of nucleic acid vaccines can be circumvented by the use of traditional vaccine platforms or protein-based vaccines56.
Aside from LNP-based vaccines, novel protein NP vaccines have also been studied for combating SARS-CoV-2 variants (Fig. 2b). An RBD-24-mer NP vaccine was developed by attaching 24 sortase A-tagged RBDs to the surface of ferritin (Fig. 2c)57. Sortase A was used as a linker to conjugate the RBD and ferritin scaffold through a sortase A reaction58. Ferritin was employed as the scaffold due to its ability to self-assemble into octahedral particles that contain three-fold axes on the particle surface, allowing for the orderly display of various viral antigens59. Given that RBD antibodies have been demonstrated to cross-neutralize SARS-CoV, SARS-CoV-2 and bat coronaviruses60, RBD-based vaccines may elicit cross-neutralizing antibodies that target the RBD of SARS-CoV-2 and its variants. The immunization of macaques with RBD-24-mer NP vaccines induced extremely high titres of neutralizing antibodies against more transmissible or neutralization-resistant SARS-CoV-2 variants, including Alpha, Beta and Gamma. Similarly, another protein NP vaccine (denoted the RBD-20-mer NP vaccine) consisting of 20 SARS-CoV-2 RBD subunits was prepared using a computationally designed self-assembling protein NP (Fig. 2c)61,62. The administration of RBD-20-mer NP vaccines to non-human primates produced potent neutralizing antibodies targeting a panel of the human immunodeficiency virus and vesicular stomatitis virus pseudotyped SARS-CoV-2 variants. The two protein NP vaccines described above use RBDs as immunogens, while in another study63, a rationally designed highly immunogenic heptad repeat 2-deleted glycine-capped (S2GΔHR2) S2 was employed in the construction of a protein NP vaccine. The protein vaccine (denoted SApNP) was constructed through the self-assembly of S2GΔHR2 S and other protein-based building blocks63,64. Immunization of mice with the SApNP vaccine elicited potent neutralizing antibodies that were able to neutralize the original SARS-CoV-2 and its variants (Alpha, Beta, Gamma and Delta) with the same potency. Furthermore, several particle-based antigen display strategies for the design of modular vaccine platforms have been described65,66,67. Of these, the Tag/Catcher platform represents a versatile and efficient technology. This is a split-protein conjugation system for the covalent anchoring of vaccine antigens onto virus or capsid-like particles upon mixing in solution. This system has been reported to achieve complete and even decoration of the NP surface and yield high antibody titres against displayed antigens65.
Targeting the S protein of variants with engineered neutralizing antibody
In addition to targeting the S protein of variants with vaccine-induced neutralizing antibodies, variant S proteins can also be directly targeted with engineered neutralizing antibodies. While producing vaccine-induced neutralizing antibodies in immunocompromised individuals might prove inefficient, the application of engineered neutralizing antibodies could solve the problem. However, most highly effective conventional antibodies for SARS-CoV-2 show significantly less efficacy against emerging SARS-CoV-2 variants. One solution might be to develop highly efficient nanobodies against the variants. Nanobodies, antigen binding domains derived from camelid single-chain antibodies, are smaller than conventional antibodies (15 kDa versus 50 kDa), enabling them to bind to virus epitopes not usually accessible to conventional antibodies68. Nanobodies can be manufactured at scale through inexpensive but efficient microbial production. Meanwhile, the high stability of nanobodies enables them to survive the nebulization process, allowing them to be administered by inhalation, a highly attractive route of administration for tackling various respiratory viruses69,70. In particular, monomeric nanobodies can be conveniently engineered to multivalent nanobody conjugates that normally display improved efficacy71,72.
To target the S protein of the variants, three classes of nanobodies (I, II and III) were produced (Fig. 3a)73. A mechanistic study revealed that these nanobodies neutralize the SARS-CoV-2 variants through multiple mechanisms. Class I nanobodies, ultrapotent neutralizers for SARS-CoV-2, target ACE2 binding sites and disrupt the binding of the virus to the ACE2 receptor. Class II nanobodies are especially interesting, because they bind to the highly conserved epitopes of the S protein, retaining potent binding activity against the VOCs. Class III nanobodies use a different neutralizing mechanism from other classes by recognizing unique epitopes usually inaccessible to conventional antibodies. The efficacy of monomeric nanobodies can be further improved by engineering them to multivalent nanobodies. A llama nanobody hexamer (linked homotrimers), a protein conjugate that consists of six nanobodies, displayed ultrapotent neutralization activity against SARS-CoV-2 variants, while the monomeric form of the nanobody was unable to neutralize some of the variants (Fig. 3a)74. Further analysis indicated that the multivalent llama nanobody hexamer potently neutralized the SARS-CoV-2 variants probably by promoting the avidity for the ACE2 binding domain, simultaneous binding to multiple S, blocking the binding of ACE2 to RBD or recognizing the conserved epitopes that are normally difficult to access by conventional antibodies. Similarly, a nanobody dimer, consisting of nanobody V and E targeting two independent epitopes of the RBD, largely suppressed the escape of SARS-CoV-2 variants from neutralization (Fig. 3a)75,76. Mechanistically, the nanobody dimer induced and stabilized the RBD trimers in an active conformation with all RBDs in the ‘up’ conformation, preventing their binding to ACE2 receptors. More importantly, this active conformation of the RBD further triggered their premature cleavage and irreversibly inactivated the cell fusion ability of the S protein. These studies are particularly interesting because they demonstrate that displaying the ineffective neutralizing nanobodies for SARS-CoV-2 variants on a well-designed nanostructure may create a multivalent nanobody NP that is ultrapotent against the variants.
a, Engineered neutralizing antibodies: a single nanobody, a nanobody hexamer containing an IgG Fc, a nanobody dimer consisting of two nanobodies (V and E) targeting two independent epitopes, a nanobody trimer containing linkers and a trimerization domain, and an IgM pentamer containing a J chain. b, ACE2-based nanodecoys: an ACE-2 microbody consisting of two ACE2 receptors, EV-based nanodecoys and nanodecoys based on different cell membranes, such as 293T, LSC, 293T/THP-1 and glycan THP-1. In some cases, a poly(lactic-co-glycolic acid) (PLGA) core can be included. c, SARS-CoV-2 variants infect host cells through ACE2 receptors. d, Engineered neutralizing antibodies and ACE2-based nanodecoys inhibit the binding of the virus to ACE2, protecting host cells from infection.
Given the trimeric structures of the S proteins of SARS-CoV-2 and its variants, it is reasonable to speculate that a nanobody trimer that matches the geometry of the S protein may have an extremely high binding affinity to the S protein. To explore this possibility, nanobody trimers were prepared by conjugating three nanobodies to immunoneutral collagen XVIII using proper immunoneutral linkers (Fig. 3a)77. As expected, the nanobody trimer developed with Re6B06 was 10,000 times more potent than the monomeric nanobody in neutralizing SARS-CoV-2. Meanwhile, the nanobody trimer prepared with Re9F06 was able to neutralize the Beta variant at a concentration as low as 5.8 pM. In addition to the nanobody strategy, another solution to address the limited efficiency of conventional antibodies (IgGs) against the variants is to develop potent immunoglobulin M (IgM) neutralizing antibodies. A rationally designed IgM pentamer (IgM-14), a protein conjugate consisting of five immunoglobulin dimers (ten binding sites), assembled in the presence of a joining chain (J chain), potently neutralized three VOCs: Alpha, Gamma and Beta (Fig. 3a)78. In mice, single-dose intranasal administration of IgM pentamers protected against Gamma and Beta variants, while IgG-14, an IgG1 monoclonal antibody (CoV2-14), failed.
Targeting the S protein of variants with ACE2-based nanodecoys
One extensively investigated nano-based strategy to combat SARS-CoV-2 and its variants is the preparation of ACE2 receptor-modified decoy NPs. Despite their continuing evolution, all SARS-CoV-2 variants enter cells through the interactions of their S protein with the ACE2 receptor expressed on cells. Therefore, strong interactions between the ACE2 receptors of decoy NPs and the S proteins of the SARS-CoV-2 variants can ensure that the sequestration of viruses by the ACE2-based nanodecoys is impervious to virus mutations. An ACE2 microbody consisting of two ACE2 fused to an IgG Fc domain inhibited the entry of SARS-CoV-2 pseudotyped virus in vitro and protected K18-hACE2 transgenic mice from SARS-CoV-2 infection-induced weight loss after intranasal co-administration of microbodies and the SARS-CoV-2 virus (Fig. 3b)79. In addition to engineered ACE2-based nanodecoys, circulating ACE2-expressing extracellular vesicles (EVs) isolated directly from the plasma of patients with COVID-19 also potently inhibited the infection of SARS-CoV-2 variants in vitro (Fig. 3b)80. In a well-established human ACE2 (hACE2) transgenic COVID-19 mouse model, intranasal co-administration of EV nanodecoys and SARS-CoV-2 improved the survival rates of infected mice when monitored on day 6. The robust antiviral efficacy of EV nanodecoys could be ascribed to two mechanisms: first, proteins presented on the surface of EVs might further hinder the access of SARS-CoV-2 to the host cell surface, and second, synergistic binding of more than one ACE2 of the EV to the SARS-CoV-2 S protein could promote the binding affinity of EV nanodecoys to viruses.
Another widely used method for the preparation of ACE2-based nanodecoys is the construction of membrane NPs derived from ACE2-rich cells. For instance, 293T membrane-based ACE2 nanodecoys showed excellent sequestration ability against both SARS-CoV-2 and D614G variant pseudotyped viruses (Fig. 3b)81. Importantly, in a hACE2-expressing mouse model, the inhalation of a formulation containing 293T nanodecoys and hyaluronic acids 4 h and 8 h before virus exposure effectively inhibited the infection of the SARS-CoV-2 pseudotyped virus. One major advantage of this inhalable formulation, consisting of 293T nanodecoys and mucoadhesive excipient hyaluronic acid, is the notably enhanced retention time of 293T nanodecoys in the lung, the main target organ of SARS-CoV-2. A similar 293T nanodecoy has also been described as blocking the infection of SARS-CoV-2 and its variants82.
As mentioned above, ACE2 is the viral receptor of human cells regardless of the mutations of the virus. Thus, the ACE2 nanodecoys that have shown efficacy against SARS-CoV-283,84 are considered to have similar efficacy against SARS-CoV-2 variants. To construct a cell membrane-based ACE2 nanodecoy, a high ACE2-expressing human lung spheroid cell (LSC) line was employed (Fig. 3b)85. In mice, the inhalation of LSC nanodecoys 1 day after SARS-CoV-2 mimic virus infection showed obvious accumulation in the lungs, yielding accelerated clearance of SARS-CoV-2 mimics from the lungs from day 2 to day 6. Furthermore, in a non-human primate model challenged with SARS-CoV-2 on day 0, the inhalation of four doses of LSC nanodecoys on days 2, 3, 4 and 5 greatly enhanced the clearance of the virus and mitigated lung injury when analysed on day 8. Several mechanisms might be responsible for these post-infection therapeutic effects of LSC nanodecoys. As not all the viruses were internalized by the host cells 1 or 2 days after viral exposure, nanodecoys could prevent the remaining viruses from entering the cells. Furthermore, the endocytosized nanodecoys could also bind to intracellular viruses, reducing further infection. To further strengthen the ability of nanodecoys to combat SARS-CoV-2, hybrid cell membranes can be used. For example, a 293T/THP-1 nanodecoy has been developed by fusing cell membranes from genetically engineered ACE2-expressing 293T cells and cytokine receptor-expressing human myeloid mononuclear THP-1 cells (Fig. 3b)86. In this design, the ACE2 and cytokine receptors of nanodecoys concurrently sequestered SARS-CoV-2 and the related inflammatory cytokines, including interleukin-6 and granulocyte-macrophage colony-stimulating factor. All the cell membrane nanodecoys described above have a hollow spherical structure. However, a polymer core can also be introduced to facilitate the formation of nanodecoys and the display of ACE2 receptors. For instance, a THP-1 nanodecoy prepared by coating ACE2-rich THP-1 cell membranes onto polymeric cores made from poly(lactic-co-glycolic acid) exhibited excellent sequestration ability against SARS-CoV-2 and protected cells from viral infection87. In addition, the sequestration ability of the THP-1 nanodecoys can be further enhanced by introducing heparin (denoted glycan THP-1 nanodecoys) at their surface (Fig. 3b)88.
Besides these cell membrane-based nanodecoys, mRNA NPs encoded by ACE2 receptor mimics capable of generating soluble ACE2 decoys in situ efficiently inhibited the binding of SARS-CoV-2 to ACE2 receptors89,90. Despite the fact that several soluble ACE2 (recombinant) decoys have entered clinical trials and shown sequestering efficacy against SARS-CoV-291,92, a short half-life might limit further improvement in their efficacy93. Apart from serving as a receptor for SARS-CoV-2, ACE2 also plays a key role in the renin–angiotensin system94. Therefore, one concern regarding the exogenous administration of soluble ACE2 decoys is the possibility of reduced blood pressure and inflammation owing to a downregulated renin–angiotensin system95,96. Although several ACE2-based nanodecoys have proven quite capable in fighting against SARS-CoV-2 and its variants, their performance can be further improved using engineered ACE2 receptors that possess a higher binding affinity to the virus97,98,99,100,101. One major limitation of these ACE2-based nanodecoys is the requirement for repeated clinical administration owing to their short half-lives. In addition, in many studies, the antiviral efficacy of nanodecoys has been evaluated by co-administering them with SARS-CoV-2, or by administering the nanodecoys immediately before virus exposure. However, this is not the scenario in real clinical use, because it is impossible to predict when an individual will become infected. In contrast, one study showed that the administration of LSC nanodecoys85 1 or 2 days after virus exposure can still efficiently sequester the virus and mitigate lung injury, demonstrating its robust potential as a post-infection therapeutic in the clinic. All SARS-CoV-2 variants infect host cells in the same way as the original SARS-CoV-2 strain, by binding the ACE2 receptors (Fig. 3c). Both the above-mentioned strategies (engineered neutralizing antibodies and ACE2-based nanodecoys) have demonstrated efficacy in inhibiting the infection of SARS-CoV-2 variants by strongly binding to their S proteins, making them attractive strategies for combating SARS-CoV-2 variants (Fig. 3d).
Potential nanotechnology strategies against SARS-CoV-2 variants
Nanotechnology-based non-specific physical inactivation of variants
Antiviral nanomaterials with intrinsic virucidal abilities are highly valuable in combating the ongoing COVID-19 pandemic and future ones. Like most respiratory viruses, SARS-CoV-2 is covered by a phospholipid bilayer membrane, which is vital for its structural integrity and cellular entry. Virucidal nanomaterials, such as polymer surfactants and pore-forming peptides, can directly disrupt the membrane and thus kill the virus102,103,104. For example, NanoViricides is developing a topical nanoviricide using ligand-decorated polymeric micelles, which can bind to viral glycoproteins and then disrupt its lipid envelope. Alternatively, Starpharma’s Viraleze, an antiviral nasal spray containing a negatively charged naphthalenedisulfonate-modified poly(L-lisine)-based SPL7013 dendrimer, can establish a physical barrier to trap and inactivate viruses. In principle, SPL7013 acts as a heparan sulfate mimetic that strongly binds to the heparan sulfate proteoglycan binding motifs on the S protein, thereby blocking the virus–host interaction. In a preclinical study, Viraleze was shown to inactivate more than 99.9% of SARS-CoV-2 when used before or after exposure105. As these nanomaterials act by physically damaging or dampening the virus, they are effective against a very broad spectrum of viruses, including SARS-CoV-2 and its variants.
Nanotechnology-based spatial capture of variants
As mentioned above, ACE2-based nanodecoys have been proven to effectively bind and immobilize SARS-CoV-2. As nanoarchitecture with a topography matching the virus can induce multivalent interactions and maximize binding106, the combination of geometry-matching topography and ACE2-enriched membrane coating could be a promising strategy to design next-generation nanodecoys against variants (Fig. 4a). As an alternative to decoys, virus-specific binders such as DNA aptamers and antibodies can be modified to yield customized nanoarchitectures with virus-matching geometries to achieve multivalent interactions (Fig. 4b,c)107,108. Exploiting the precise construction of geometrically defined three-dimensional objects by DNA origami technology109, DNA-based nanostructures have been designed to trap viruses. For example, by mimicking the symmetrical structure found in viral capsids, a self-assembling icosahedral DNA shell with its interior functionalized with virus-specific binders has been built to entrap the entire virus and prevent infection108. Although this nanoshell has been demonstrated to trap other infectious viruses, it can be swiftly customized to combat SARS-CoV-2 and its variants by simply switching the virus binders103. However, a structural mismatch between the icosahedral geometry and the enveloped virus could reduce multivalent interactions. To address this issue, a nanobody trimer binder that matches the geometry of the S protein could be used to increase the binding affinity of the DNA nanoshell to SARS-CoV-2 variants (Fig. 3a).
a, A spiky nanostructure can be coated with an ACE2-enriched membrane to generate a potent decoy for SARS-CoV-2 variants. b, A DNA icosahedral shell can be modified with neutralizing antibodies to yield a virus trap. c, A star-shaped DNA architecture can be modified with neutralizing antibodies to prepare a SARS-CoV-2 variant inhibitor. d, Instead of simply combining different neutralizing antibodies (for example, B1-182.1 and A19-46.1), multivalent display of these antibodies on the surface of NPs can substantially enhance the neutralizing efficacy of the antibodies against SARS-CoV-2 variants.
Nanotechnology-based immune neutralization of the variants
SARS-CoV-2 neutralizing monoclonal antibodies (mAbs) can provide effective therapeutic protection in patients with COVID-19, but several authorized mAbs have recently been found to have reduced neutralizing potency towards the circulating variants, especially the Omicron variant110. Nanotechnology-based cryoelectron microscopy is a crucial tool for understanding the neutralizing mechanisms and binding epitopes of mAbs, which is helpful to discover broad-spectrum mAbs targeting highly conserved epitopes of SARS-CoV-2 and ultrapotent mAbs neutralizing the SARS-CoV-2 variants111,112. To prevent viral escape and increase coverage against the variants, cocktails of two or more mAbs targeting multiple sites of vulnerability on the S glycoprotein are highly preferred over a single antibody113. Therefore, rational assembly of those mAbs as multispecific antibodies that simultaneously target two or more non-overlapping epitopes is a promising means to induce broad cross-protection against current and future variants (Fig. 4d)112,114 due to complementary neutralization and reduced risk of mutational escape. In particular, nanotechnology can play an active role in the non-covalent assembly or chemical conjugation of mAbs115,116. For example, as an alternative to displaying multivalent antigens, self-assembling ferritin can be modified to display antibody fragments with a high degree of valency for broader neutralization and stronger affinity.
Nanotechnology-based therapeutic drugs for variants
Anti-inflammatory drugs and broad-spectrum antivirals (for example, nucleoside analogues and protease inhibitors) remain promising treatment options against variants in terms of reducing morbidity and mortality, as exemplified by dexamethasone, remdesivir and Paxlovid (nirmatrelvir and ritonavir), currently used to treat patients with COVID-19117,118,119. The repurposing of drugs that are licensed or in clinical trials for other viruses provides a cost- and time-effective therapeutic solution120,121. However, high-dose, repeated administration of small-molecule drugs is necessary to maintain clinically effective concentrations, which can cause severe adverse effects118. NPs are promising drug delivery vehicles for these therapeutic drugs, and formulated nanomedicines are expected to achieve passive/active targeting, prolong circulation time and reduce side effects122,123,124. For example, some NPs preferentially accumulate in macrophages, providing the possibility of targeting dexamethasone to these hyperactivated immune cells to suppress the aberrant inflammatory responses related to COVID-19125. Apart from small-molecule therapeutic drugs, NP-enabled nucleic acid drugs that can precisely interfere with either pro-inflammatory signalling or viral replication represent another powerful tool to combat COVID-19126. Notably, effectiveness against SARS-CoV-2 and its variants can be further improved by integrating multiple treatment modalities into one nanosystem. For instance, the aforementioned virucidal nanomaterials could be loaded with broad-spectrum antiviral drugs to achieve a synergistic effect103.
Outlook
While the original SARS-CoV-2 initiated the current pandemic, the emerging SARS-CoV-2 variants generated by the continuing mutations of the S protein of SARS-CoV-2 have exacerbated and prolonged it. Despite its enhanced viral transmission and immune evasion, the currently dominant Omicron variant has resulted in lower rates of hospitalization and death compared with previous COVID-19 waves28,127,128. However, conclusions regarding severity should be assessed carefully, because the global population has already experienced several COVID-19 waves and many countries have high vaccination rates, especially among vulnerable elderly populations. Further complicating matters, in some countries it is difficult to ascertain what percentage of the population has already been infected129. Currently, it seems that a boost of the original mRNA vaccines can enhance neutralizing antibodies and might prevent Omicron infection shortly after administration46,47. Specifically, individuals recovering from infection or having had a recent mRNA vaccine dose had a substantial gain in neutralizing activity29,130. However, some claim that variant-specific vaccines are warranted. To this end, both current mRNA vaccine manufacturers (Moderna131 and Pfizer–BioNTech132) and Johnson & Johnson133 have announced plans to develop new Omicron variant-specific vaccines. Accordingly, the Food and Drug Administration has stated that it is setting rules for the accelerated review of updated vaccines against specific variants12.
Targeting the mutated S protein of the SARS-CoV-2 variants with novel nanotechnology-based strategies holds great promise for combating SARS-CoV-2 variants. Eliciting neutralizing antibodies using vaccines is the most effective method to target the S protein of the variants, inhibiting viral infection. Various technologies have been employed to develop a large variety of vaccines, and several of them demonstrating high protective efficacy have been deployed worldwide. However, when faced with the emerging SARS-CoV-2 variants, the protective efficacy of these vaccines has decreased. Compared with conventional vaccines, nanotechnology-based vaccines are still able to protect individuals from the SARS-CoV-2 variants due to their original high efficacy; in addition, they can be conveniently and rapidly updated to improve their efficacy against the SARS-CoV-2 variants. We anticipate that their extremely high efficacy and short manufacturing cycle time will prove crucial in the implementation of next-generation vaccines against SARS-CoV-2 variants.
Engineered neutralizing antibodies provide another method to target the S protein of variants, especially when vaccine-elicited neutralizing antibodies are insufficient because of a compromised immune response. While a large number of engineered monomeric neutralizing antibodies that neutralize SARS-CoV-2 fail to neutralize the emerging SARS-CoV-2 variants, multivalent display of these antibodies on a nanoplatform can dramatically promote their neutralizing activity. Future efforts could focus on the development of multivalent antibody nanoplatforms that are able to bind to different sites of the S protein simultaneously, aiming to effectively target different variants without the need to adjust the platforms. The development of ACE2-based nanodecoys is considered an ideal strategy to target the S protein of the variants, due to the fact that all the variants have high binding affinity to the ACE2 receptors regardless of their continuing mutations. Thus, an effective ACE2-based nanodecoy for SARS-CoV-2 should also be effective for current and future SARS-CoV-2 variants, holding great potential to become a universal platform for all variants. Notably, although the latest Omicron variant harbours several times more mutations than other circulating variants, it still infects host cells through ACE2 receptors. Potentially, a variety of nanostructures (for example, DNA origami) that directly capture variants or precisely display neutralizing antibodies or ACE2 receptors on their surfaces can be adapted to prepare novel nanotechnology-based therapeutics to combat SARS-CoV-2 variants. Despite the ongoing nature of the COVID-19 pandemic, owing to the rapid spread of SARS-CoV-2 variants globally, we expect the advances and innovation offered by nanotechnology can provide diverse approaches to hasten the end of the pandemic.
References
Florindo, H. F. et al. Immune-mediated approaches against COVID-19. Nat. Nanotechnol. 15, 630–645 (2020).
Tang, Z. et al. A materials science perspective on tackling COVID-19. Nat. Rev. Mater. 5, 847–860 (2020).
Tang, Z. et al. Insights from nanotechnology in COVID-19 treatment. Nano Today 36, 101019 (2021).
Sadarangani, M., Marchant, A. & Kollmann, T. R. Immunological mechanisms of vaccine-induced protection against COVID-19 in humans. Nat. Rev. Immunol. 21, 475–484 (2021).
Baden, L. R. et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N. Engl. J. Med. 384, 403–416 (2021).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA COVID-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Kirtane, A. R. et al. Nanotechnology approaches for global infectious diseases. Nat. Nanotechnol. 16, 369–384 (2021).
Tracking SARS-CoV-2 variants. WHO https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/ (2022).
Krause, P. R. et al. SARS-CoV-2 variants and vaccines. N. Engl. J. Med. 385, 179–186 (2021).
Callaway, E. The coronavirus is mutating—does it matter? Nature 585, 174–178 (2020).
Williams, T. C. & Burgers, W. A. SARS-CoV-2 evolution and vaccines: cause for concern? Lancet Respir. Med. 9, 333–335 (2021).
Bian, L. et al. Effects of SARS-CoV-2 variants on vaccine efficacy and response strategies. Expert Rev. Vaccines 20, 365–373 (2021).
Cohn, B. A., Cirillo, P. M., Murphy, C. C., Krigbaum, N. Y. & Wallace, A. W. SARS-CoV-2 vaccine protection and deaths among US veterans during 2021. Science 375, 331–336 (2022).
Hoffmann, M. et al. SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 184, 2384–2393.e2312 (2021).
Liu, C. et al. Reduced neutralization of SARS-CoV-2 B.1.617 by vaccine and convalescent serum. Cell 184, 4220–4236.e4213 (2021).
Lucas, C. et al. Impact of circulating SARS-CoV-2 variants on mRNA vaccine-induced immunity. Nature 600, 523–529 (2021).
Greaney, A. J. et al. Mapping mutations to the SARS-CoV-2 RBD that escape binding by different classes of antibodies. Nat. Commun. 12, 4196 (2021).
COG-UK Mutation Explorer (COG-UK, 2021); https://sars2.cvr.gla.ac.uk/cog-uk/
Planas, D. et al. Sensitivity of infectious SARS-CoV-2 B.1.1.7 and B.1.351 variants to neutralizing antibodies. Nat. Med. 27, 917–924 (2021).
Choi, A. et al. Serum neutralizing activity of mRNA-1273 against SARS-CoV-2 variants. J. Virol. 95, e01313–01321 (2021).
Jangra, S. et al. SARS-CoV-2 spike E484K mutation reduces antibody neutralisation. Lancet Microbe 2, e283–e284 (2021).
Dejnirattisai, W. et al. Antibody evasion by the P.1 strain of SARS-CoV-2. Cell 184, 2939–2954.e2939 (2021).
Liu, J. et al. BNT162b2-elicited neutralization of B.1.617 and other SARS-CoV-2 variants. Nature 596, 273–275 (2021).
Stamatatos, L. et al. mRNA vaccination boosts cross-variant neutralizing antibodies elicited by SARS-CoV-2 infection. Science 372, 1413–1418 (2021).
Levin, E. G. et al. Waning immune humoral response to BNT162b2 COVID-19 vaccine over 6 months. N. Engl. J. Med. 385, e84 (2021).
Thomas, S. J. et al. Safety and efficacy of the BNT162b2 mRNA COVID-19 vaccine through 6 months. N. Engl. J. Med. 385, 1761–1773 (2021).
Scott, J., Richterman, A. & Cevik, M. COVID-19 vaccination: evidence of waning immunity is overstated. Brit. Med. J. 374, n2320 (2021).
Karim, S. S. A. & Karim, Q. A. Omicron SARS-CoV-2 variant: a new chapter in the COVID-19 pandemic. Lancet 398, 2126–2128 (2021).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature 602, 664–670 (2022).
Schmidt, F. et al. Plasma neutralization of the SARS-CoV-2 Omicron variant. N. Engl. J. Med. 386, 599–601 (2021).
Takashita, E. et al. Efficacy of antibodies and antiviral drugs against COVID-19 Omicron variant. N. Engl. J. Med. 386, 995–998 (2022).
Rössler, A., Riepler, L., Bante, D., von Laer, D. & Kimpel, J. SARS-CoV-2 Omicron variant neutralization in serum from vaccinated and convalescent persons. N. Engl. J. Med. 386, 698–700 (2022).
Altarawneh, H. N. et al. Protection against the Omicron variant from previous SARS-CoV-2 infection. N. Engl. J. Med. 386, 1288–1290 (2022).
VanBlargan, L. A. et al. An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by therapeutic monoclonal antibodies. Nat. Med. 28, 490–495 (2022).
Elia, U. et al. Design of SARS-CoV-2 hFc-conjugated receptor-binding domain mRNA vaccine delivered via lipid nanoparticles. ACS Nano 15, 9627–9637 (2021).
Elia, U. et al. Lipid nanoparticle RBD-hFc mRNA vaccine protects hACE2 transgenic mice against a lethal SARS-CoV-2 infection. Nano Lett. 21, 4774–4779 (2021).
Kon, E., Elia, U. & Peer, D. Principles for designing an optimal mRNA lipid nanoparticle vaccine. Curr. Opin. Biotechnol. 73, 329–336 (2022).
Baumjohann, D. & Fazilleau, N. Antigen-dependent multistep differentiation of T follicular helper cells and its role in SARS-CoV-2 infection and vaccination. Eur. J. Immunol. 51, 1325–1333 (2021).
Pardi, N. et al. Nucleoside-modified mRNA vaccines induce potent T follicular helper and germinal center B cell responses. J. Exp. Med. 215, 1571–1588 (2018). This study demonstrates how modified mRNA-LNP vaccines induce highly potent and durable neutralizing antibody responses.
Lederer, K. et al. SARS-CoV-2 mRNA vaccines foster potent antigen-specific germinal center responses associated with neutralizing antibody generation. Immunity 53, 1281–1295.e1285 (2020).
Alameh, M.-G. et al. Lipid nanoparticles enhance the efficacy of mRNA and protein subunit vaccines by inducing robust T follicular helper cell and humoral responses. Immunity 54, 2877–2892.e2877 (2021).
Teijaro, J. R. & Farber, D. L. COVID-19 vaccines: modes of immune activation and future challenges. Nat. Rev. Immunol. 21, 195–197 (2021).
Kamar, N. et al. Three doses of an mRNA COVID-19 vaccine in solid-organ transplant recipients. N. Engl. J. Med. 385, 661–662 (2021).
Hall, V. G. et al. Randomized trial of a third dose of mRNA-1273 vaccine in transplant recipients. N. Engl. J. Med. 385, 1244–1246 (2021).
Falsey, A. R. et al. SARS-CoV-2 neutralization with BNT162b2 vaccine dose 3. N. Engl. J. Med. 385, 1627–1629 (2021).
Nemet, I. et al. Third BNT162b2 vaccination neutralization of SARS-CoV-2 Omicron infection. N. Engl. J. Med. 386, 492–494 (2021). This study shows that a third dose does elicit Omicron neutralizing antibodies shortly after administration.
Pajon, R. et al. SARS-CoV-2 Omicron variant neutralization after mRNA-1273 booster vaccination. N. Engl. J. Med. 386, 1088–1091 (2022).
Wu, M. et al. Three-dose vaccination elicits neutralising antibodies against Omicron. Lancet 399, 715–717 (2022).
Tanne, J. H. COVID-19: Moderna plans booster doses to counter variants. Brit. Med. J. 372, n232 (2021).
Choi, A. et al. Safety and immunogenicity of SARS-CoV-2 variant mRNA vaccine boosters in healthy adults: an interim analysis. Nat. Med. 27, 2025–2031 (2021).
Yang, Y. & Du, L. SARS-CoV-2 spike protein: a key target for eliciting persistent neutralizing antibodies. Signal Transduct. Target. Ther. 6, 95 (2021).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Martinez, D. R. et al. Chimeric spike mRNA vaccines protect against Sarbecovirus challenge in mice. Science 373, 991–998 (2021).
Xiao, Y. et al. Emerging mRNA technologies: Delivery strategies and biomedical applications. Chem. Soc. Rev. 51, 3828–3845 (2022).
Qu, L. et al. Circular RNA vaccines against SARS-CoV-2 and emerging variants. Cell 185, 1–17 (2022). This report describes circular RNA vaccines for SARS-CoV-2 and its variants.
Beaudoin, C. A., Bartas, M., Volná, A., Pečinka, P. & Blundell, T. L. Are there hidden genes in DNA/RNA vaccines? Front. Immunol. 13, 801915 (2022).
Saunders, K. O. et al. Neutralizing antibody vaccine for pandemic and pre-emergent coronaviruses. Nature 594, 553–559 (2021). This study provides a promising protein nanoparticle platform for developing pancoronavirus vaccines.
Saunders, K. O. et al. Targeted selection of HIV-specific antibody mutations by engineering B cell maturation. Science 366, eaay7199 (2019).
Houser, K. V. et al. Safety and immunogenicity of a ferritin nanoparticle H2 influenza vaccine in healthy adults: a phase 1 trial. Nat. Med. 28, 383–391 (2022).
Li, D. et al. In vitro and in vivo functions of SARS-CoV-2 infection-enhancing and neutralizing antibodies. Cell 184, 4203–4219.e4232 (2021).
Walls, A. C. et al. Elicitation of broadly protective sarbecovirus immunity by receptor-binding domain nanoparticle vaccines. Cell 184, 5432–5447 (2021).
Walls, A. C. et al. Elicitation of potent neutralizing antibody responses by designed protein nanoparticle vaccines for SARS-CoV-2. Cell 183, 1367–1382.e1317 (2020).
He, L. et al. Single-component, self-assembling, protein nanoparticles presenting the receptor binding domain and stabilized spike as SARS-CoV-2 vaccine candidates. Sci. Adv. 7, eabf1591 (2021).
Zhang, Y.-N. et al. Mechanism of a COVID-19 nanoparticle vaccine candidate that elicits a broadly neutralizing antibody response to SARS-CoV-2 variants. Sci. Adv. 7, eabj3107 (2021).
Aves, K.-L., Goksøyr, L. & Sander, A. F. Advantages and prospects of Tag/Catcher mediated antigen display on capsid-like particle-based vaccines. Viruses 12, 185 (2020).
Janitzek, C. M. et al. A proof-of-concept study for the design of a VLP-based combinatorial HPV and placental malaria vaccine. Sci. Rep. 9, 5260 (2019).
Brune, K. D. & Howarth, M. New routes and opportunities for modular construction of particulate vaccines: stick, click, and glue. Front. Immunol. 9, 1432 (2018).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Vanlandschoot, P. et al. Nanobodies®: new ammunition to battle viruses. Antiviral Res. 92, 389–407 (2011).
Detalle, L. et al. Generation and characterization of ALX-0171, a potent novel therapeutic nanobody for the treatment of respiratory syncytial virus infection. Antimicrob. Agents Chemother. 60, 6–13 (2016).
Xiang, Y. et al. Versatile and multivalent nanobodies efficiently neutralize SARS-CoV-2. Science 370, 1479–1484 (2020).
Schoof, M. et al. An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive Spike. Science 370, 1473–1479 (2020).
Sun, D. et al. Potent neutralizing nanobodies resist convergent circulating variants of SARS-CoV-2 by targeting diverse and conserved epitopes. Nat. Commun. 12, 4676 (2021).
Xu, J. et al. Nanobodies from camelid mice and llamas neutralize SARS-CoV-2 variants. Nature 595, 278–282 (2021).
Koenig, P.-A. et al. Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape. Science 371, eabe6230 (2021).
Saelens, X. & Schepens, B. Single-domain antibodies make a difference. Science 371, 681–682 (2021).
Güttler, T. et al. Neutralization of SARS-CoV-2 by highly potent, hyperthermostable, and mutation-tolerant nanobodies. EMBO J. 40, e107985 (2021).
Ku, Z. et al. Nasal delivery of an IgM offers broad protection from SARS-CoV-2 variants. Nature 595, 718–723 (2021).
Tada, T. et al. An ACE2 microbody containing a single immunoglobulin Fc domain is a potent inhibitor of SARS-CoV-2. Cell Rep. 33, 108528 (2020).
El-Shennawy, L. et al. Circulating ACE2-expressing extracellular vesicles block broad strains of SARS-CoV-2. Nat. Commun. 13, 405 (2022).
Zhang, H. et al. Inhalable nanocatchers for SARS-CoV-2 inhibition. Proc. Natl Acad. Sci. USA 118, e2102957118 (2021).
Wang, C. et al. Membrane nanoparticles derived from ACE2-rich cells block SARS-CoV-2 infection. ACS Nano 15, 6340–6351 (2021).
Wang, Z. et al. Inhaled ACE2-engineered microfluidic microsphere for intratracheal neutralization of COVID-19 and calming of the cytokine storm. Matter 5, 336–362 (2021).
Xie, F. et al. Engineering extracellular vesicles enriched with palmitoylated ACE2 as COVID-19 therapy. Adv. Mater. 33, 2103471 (2021).
Li, Z. et al. Cell-mimicking nanodecoys neutralize SARS-CoV-2 and mitigate lung injury in a non-human primate model of COVID-19. Nat. Nanotechnol. 16, 942–951 (2021). This study describes a nanodecoy that exhibits post-infection therapeutic effects for SARS-CoV-2.
Rao, L. et al. Decoy nanoparticles protect against COVID-19 by concurrently adsorbing viruses and inflammatory cytokines. Proc. Natl Acad. Sci. USA 117, 27141–27147 (2020).
Zhang, Q. et al. Cellular nanosponges inhibit SARS-CoV-2 infectivity. Nano Lett. 20, 5570–5574 (2020).
Ai, X. et al. Surface glycan modification of cellular nanosponges to promote SARS-CoV-2 inhibition. J. Am. Chem. Soc. 143, 17615–17621 (2021).
Li, M. et al. Secreted expression of mRNA-encoded truncated ACE2 variants for SARS-CoV-2 via lipid-like nanoassemblies. Adv. Mater. 33, 2101707 (2021).
Kim, J., Mukherjee, A., Nelson, D., Jozic, A. & Sahay, G. Rapid generation of circulating and mucosal decoy ACE2 using mRNA nanotherapeutics for the potential treatment of SARS-CoV-2. Preprint at bioRxiv https://doi.org/10.1101/2020.07.24.205583 (2020).
Zoufaly, A. et al. Human recombinant soluble ACE2 in severe COVID-19. Lancet Respir. Med. 8, 1154–1158 (2020).
Monteil, V. et al. Inhibition of SARS-CoV-2 infections in engineered human tissues using clinical-grade soluble human ACE2. Cell 181, 905–913.e907 (2020).
Haschke, M. et al. Pharmacokinetics and pharmacodynamics of recombinant human angiotensin-converting enzyme 2 in healthy human subjects. Clin. Pharmacokinet. 52, 783–792 (2013).
Romero, C. A., Orias, M. & Weir, M. R. Novel RAAS agonists and antagonists: clinical applications and controversies. Nat. Rev. Endocrinol. 11, 242–252 (2015).
South, A. M. et al. Fetal programming and the angiotensin-(1-7) axis: a review of the experimental and clinical data. Clin. Sci. 133, 55–74 (2019).
Warner, F. J., Rajapaksha, H., Shackel, N. & Herath, C. B. ACE2: from protection of liver disease to propagation of COVID-19. Clin. Sci. 134, 3137–3158 (2020).
Chan, K. K. et al. Engineering human ACE2 to optimize binding to the spike protein of SARS coronavirus 2. Science 369, 1261–1265 (2020).
Glasgow, A. et al. Engineered ACE2 receptor traps potently neutralize SARS-CoV-2. Proc. Natl Acad. Sci. USA 117, 28046–28055 (2020).
Chan, K. K., Tan, T. J. C., Narayanan, K. K. & Procko, E. An engineered decoy receptor for SARS-CoV-2 broadly binds protein S sequence variants. Sci. Adv. 7, eabf1738 (2021).
Higuchi, Y. et al. Engineered ACE2 receptor therapy overcomes mutational escape of SARS-CoV-2. Nat. Commun. 12, 3802 (2021).
Zhang, L. et al. Engineered ACE2 decoy mitigates lung injury and death induced by SARS-CoV-2 variants. Nat. Chem. Biol. 18, 342–351 (2022).
Jackman, J. A. et al. Therapeutic treatment of Zika virus infection using a brain-penetrating antiviral peptide. Nat. Mater. 17, 971–977 (2018).
Peplow, M. Nanotechnology offers alternative ways to fight COVID-19 pandemic with antivirals. Nat. Biotechnol. 39, 1172–1174 (2021).
Yoon, B. K., Jeon, W.-Y., Sut, T. N., Cho, N.-J. & Jackman, J. A. Stopping membrane-enveloped viruses with nanotechnology strategies: toward antiviral drug development and pandemic preparedness. ACS Nano 15, 125–148 (2021).
Paull, J. R. A. et al. Protective effects of astodrimer sodium 1% nasal spray formulation against SARS-CoV-2 nasal challenge in K18-hACE2 mice. Viruses 13, 1656 (2021).
Nie, C. et al. Spiky nanostructures with geometry-matching topography for virus inhibition. Nano Lett. 20, 5367–5375 (2020).
Kwon, P. S. et al. Designer DNA architecture offers precise and multivalent spatial pattern-recognition for viral sensing and inhibition. Nat. Chem. 12, 26–35 (2020).
Sigl, C. et al. Programmable icosahedral shell system for virus trapping. Nat. Mater. 20, 1281–1289 (2021). This study demonstrates the possibility of using DNA architecture to capture virus.
Saccà, B. & Niemeyer, C. M. DNA origami: the art of folding DNA. Angew. Chem. Int. Ed. 51, 58–66 (2012).
Eli Lilly, Regeneron antibody therapies lose out against Omicron. The Irish Times (14 December 2021); https://www.irishtimes.com/business/health-pharma/eli-lilly-regeneron-antibody-therapies-lose-out-against-omicron-1.4755091
Du, S. et al. Structures of SARS-CoV-2 B.1.351 neutralizing antibodies provide insights into cocktail design against concerning variants. Cell Res. 31, 1130–1133 (2021).
Wang, L. et al. Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants. Science 373, eabh1766 (2021).
Dussupt, V. et al. Low-dose in vivo protection and neutralization across SARS-CoV-2 variants by monoclonal antibody combinations. Nat. Immunol. 22, 1503–1514 (2021).
De Gasparo, R. et al. Bispecific IgG neutralizes SARS-CoV-2 variants and prevents escape in mice. Nature 593, 424–428 (2021).
Szijj, P. & Chudasama, V. The renaissance of chemically generated bispecific antibodies. Nat. Rev. Chem. 5, 78–92 (2021).
Shatz, W. et al. Ferritin as a natural protein scaffold: building a multivalent ferritin–Fab conjugate. LCGC Suppl. 37, 30–35 (2019).
Beigel, J. H. et al. Remdesivir for the treatment of COVID-19—final report. N. Engl. J. Med. 383, 1813–1826 (2020).
The RECOVERY Collaborative Group Dexamethasone in hospitalized patients with COVID-19. N. Engl. J. Med. 384, 693–704 (2020).
Mahase, E. COVID-19: Pfizer’s paxlovid is 89% effective in patients at risk of serious illness, company reports. Brit. Med. J. 375, n2713 (2021).
Saul, S. & Einav, S. Old drugs for a new virus: repurposed approaches for combating COVID-19. ACS Infect. Dis. 6, 2304–2318 (2020).
Cao, Y. The impact of the hypoxia-VEGF-vascular permeability on COVID-19-infected patients. Exploration 1, 20210051 (2021).
Anselmo, A. C. & Mitragotri, S. Nanoparticles in the clinic: an update post COVID-19 vaccines. Bioeng. Transl. Med. 6, e10246 (2021).
Zhao, Z. et al. Glycyrrhizic acid nanoparticles as antiviral and anti-inflammatory agents for COVID-19 treatment. ACS Appl. Mater. Interfaces 13, 20995–21006 (2021).
Liu, J., Wan, M., Lyon, C. J. & Hu, T. Y. Nanomedicine therapies modulating macrophage dysfunction: a potential strategy to attenuate cytokine storms in severe infections. Theranostics 10, 9591–9600 (2020).
Lammers, T. et al. Dexamethasone nanomedicines for COVID-19. Nat. Nanotechnol. 15, 622–624 (2020).
Han, X., Mitchell, M. J. & Nie, G. Nanomaterials for therapeutic RNA delivery. Matter 3, 1948–1975 (2020).
Bhattacharyya, R. P. & Hanage, W. P. Challenges in inferring intrinsic severity of the SARS-CoV-2 Omicron variant. N. Engl. J. Med. 386, e14 (2022).
Ulloa, A. C., Buchan, S. A., Daneman, N. & Brown, K.A. Estimates of SARS-CoV-2 Omicron variant severity in Ontario, Canada. JAMA 327, 1286–1288 (2022).
Nealon, J. & Cowling, B. J. Omicron severity: milder but not mild. Lancet 399, 412–413 (2022).
Pulliam, J. R. C. et al. Increased risk of SARS-CoV-2 reinfection associated with emergence of Omicron in South Africa. Science 376, eabn4947 (2022).
Moderna announces preliminary booster data and updates strategy to address Omicron variant. Business Wire https://www.businesswire.com/news/home/20211220005253/en/Moderna-Announces-Preliminary-Booster-Data-and-Updates-Strategy-to-Address-Omicron-Variant (2021).
Pfizer and BioNTech provide update on Omicron variant. Pfizer https://www.pfizer.com/news/press-release/press-release-detail/pfizer-and-biontech-provide-update-omicron-variant (2021).
Johnson & Johnson to evaluate its COVID-19 vaccine against new Omicron COVID-19 variant. Johnson & Johnson https://www.jnj.com/johnson-johnson-to-evaluate-its-covid-19-vaccine-against-new-omicron-covid-19-variant (2021).
Altmann, D. M., Boyton, R. J. & Beale, R. Immunity to SARS-CoV-2 variants of concern. Science 371, 1103–1104 (2021).
Tregoning, J. S., Flight, K. E., Higham, S. L., Wang, Z. & Pierce, B. F. Progress of the COVID-19 vaccine effort: viruses, vaccines and variants versus efficacy, effectiveness and escape. Nat. Rev. Immunol. 21, 626–636 (2021).
Wang, P. et al. Increased resistance of SARS-CoV-2 variant P.1 to antibody neutralization. Cell Host Microbe 29, 747–751.e744 (2021).
Mlcochova, P. et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 599, 114–119 (2021).
Wilhelm, A. et al. Reduced neutralization of SARS-CoV-2 Omicron variant by vaccine sera and monoclonal antibodies. Preprint at MedRxiv https://www.medrxiv.org/content/10.1101/2021.12.07.21267432v4 (2021).
Andrews, N. et al. COVID-19 vaccine effectiveness against the Omicron (B.1.1.529) variant. N. Engl. J. Med. 386, 1532–1546 (2022).
Enhancing Readiness for Omicron (B.1.1.529): Technical Brief and Priority Actions for Member States (WHO, 2021); https://www.who.int/docs/default-source/coronaviruse/2022-01-21-global-technical-brief-and-priority-action-on-omicron-sars-cov-2-variant.pdf?sfvrsn=f3ac8bc3_9&download=true © Springer Nature.
Mahase, E. COVID-19 booster vaccines: what we know and who’s doing what. Brit. Med. J. 374, n2082 (2021).
Kitchin, D. et al. Ad26.COV2.S breakthrough infections induce high titers of neutralizing antibodies against Omicron and other SARS-CoV-2 variants of concern. Cell Rep. Med. 3, 100535 (2022).
Wink, P. L. et al. First identification of SARS-CoV-2 Lambda (C.37) variant in Southern Brazil. Infect. Control Hosp. Epidemiol. 1–2 (2021).
Messali, S. et al. A cluster of the new SARS-CoV-2 B.1.621 lineage in Italy and sensitivity of the viral isolate to the BNT162b2 vaccine. J. Med. Virol. 93, 6468–6470 (2021).
Acknowledgements
The authors are supported by a US METAvivor Early Career Investigator Award (No. 2018A020560, W.T.), a Department Basic Scientist Grant (No. 2420 BPA075, W.T.), a Khoury Innovation Award (No. 2020A003219, W.T.), a Gillian Reny Stepping Strong Center for Trauma Innovation Breakthrough Innovator Award (No. 113548, W.T.), the Nanotechnology Foundation (No. 2022A002721, W.T.), the Farokhzad Family Distinguished Chair Foundation (No. 018129, W.T.), an AHA Collaborative Sciences Award (No. 2018A004190, W.T.), the Innovation Authority in Israel (Kamin-Corona; D.P.), a US National Institutes of Health (NIH) Director’s New Innovator Award (DP2 TR002776, M.J.M.), a Burroughs Wellcome Fund Career Award at the Scientific Interface (CASI; M.J.M.), the National Institutes of Health (NCI R01 CA241661, NCI R37 CA244911 and NIDDK R01 DK123049, M.J.M.), a US National Science Foundation CAREER Award (CBET-2145491, M.J.M.) and the Yoran Institute for Human Genome Research (E.K.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
D.P. declares the following competing financial interest(s): D.P. receives licensing fees (to patents on which he was an inventor) from, invested in, consults (or on scientific advisory boards or boards of directors) for, lectured (and received a fee) or conducts sponsored research at TAU for the following entities: ART Biosciences, BioNtech SE, EPM Inc., Earli Inc., Kernal Biologics, Merck, Newphase Ltd., NeoVac Ltd., RiboX Therapeutics, Roche, SirTLabs Corporation, Teva Pharmaceuticals Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Tony Hu and Pere Santamaria for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, X., Kon, E., Han, X. et al. Nanotechnology-based strategies against SARS-CoV-2 variants. Nat. Nanotechnol. 17, 1027–1037 (2022). https://doi.org/10.1038/s41565-022-01174-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-022-01174-5
This article is cited by
-
Inhalation of ACE2-expressing lung exosomes provides prophylactic protection against SARS-CoV-2
Nature Communications (2024)
-
Research Progress of Nanomaterials for Prevention, Diagnosis, and Treatment of SARS-CoV-2
BioNanoScience (2024)
-
Nanotechnology-based theranostic and prophylactic approaches against SARS-CoV-2
Immunologic Research (2024)
-
Nanoparticle-based DNA vaccine protects against SARS-CoV-2 variants in female preclinical models
Nature Communications (2024)
-
Emerging non-viral vectors for gene delivery
Journal of Nanobiotechnology (2023)