Abstract
The COVID-19 pandemic has infected millions of people with no clear signs of abatement owing to the high prevalence, long incubation period and lack of established treatments or vaccines. Vaccines are the most promising solution to mitigate new viral strains. The genome sequence and protein structure of the 2019-novel coronavirus (nCoV or SARS-CoV-2) were made available in record time, allowing the development of inactivated or attenuated viral vaccines along with subunit vaccines for prophylaxis and treatment. Nanotechnology benefits modern vaccine design since nanomaterials are ideal for antigen delivery, as adjuvants, and as mimics of viral structures. In fact, the first vaccine candidate launched into clinical trials is an mRNA vaccine delivered via lipid nanoparticles. To eradicate pandemics, present and future, a successful vaccine platform must enable rapid discovery, scalable manufacturing and global distribution. Here, we review current approaches to COVID-19 vaccine development and highlight the role of nanotechnology and advanced manufacturing.
Similar content being viewed by others
Main
In December 2019, a novel coronavirus (nCoV or SARS-CoV-2) belonging to the betacoronavirus family emerged1,2. All human betacoronaviruses are unique from one another, however, they do share a certain degree of genetic and structural homology. SARS-CoV-2 genome sequence homology with SARS-CoV and MERS-CoV is 77% and 50%, respectively3. In contrast to the relatively smaller outbreaks of SARS-CoV in 2002 and MERS-CoV in 2012, SARS-CoV-2 is exhibiting an unprecedented scale of infection, resulting in a global pandemic declaration of Coronavirus Infectious Disease (COVID-19) on 11 March 2020 by the World Health Organization (WHO). On 1 June 2020, the World Health Organization reported >6 million confirmed cases and 371 thousand deaths globally. Of note, during the 1918 influenza pandemic, more death was observed in the second phase of outbreak4. Similar to influenza, COVID-19 harbours the potential to become a seasonal disease5. The high infection rate, long incubation period, along with mild-to-moderate symptoms experienced by many, make COVID-19 a troubling disease. A vaccine is crucial, in particular because data indicate asymptomatic transmission of COVID-196,7,8. More than 10 years ago, scientists predicted the pandemic potential of the coronaviruses9. And for the past 30 years, a once-per-decade novel coronavirus has pushed our public health system to the limit, with SARS-CoV-2 being the most severe. Despite the repeated warnings and discussion, the world was not prepared for this pandemic. The rapid development, distribution and administration of a vaccine to the global population is the most effective approach to quell this pandemic and the only one that will lead to a complete lifting of restrictions. Challenges include the vaccine design itself, but also its manufacture and global distribution; cold chain requirements present logistical and fiscal barriers to the availability of important, life-saving vaccines in resource-poor areas of the world. Innovating vaccine delivery platforms and devices to break cold chain limitations are therefore an efficient solution to safeguard potent vaccination for both wealthy and lower-income countries.
The vaccine strategies
When designing a vaccine, principally, one needs to define the antigen, the adjuvant, the manufacturing system and the delivery strategy (Box 1). The rapid development of vaccines is possible because the genome and structural information of SARS-CoV-2 was made available in record time10,11,12,13,14. These data, along with expedited communication of bioinformatic predictions and epitope mapping15,16,17,18, has provided crucial knowledge enabling vaccine design beyond development of live-attenuated and inactivated vaccines19,20,21,22,23. Also, information available from prior development of SARS/MERS vaccine candidates aids in the development of SARS-CoV-2 vaccine candidates24,25. Nanotechnology platforms offer great utility in modern vaccine design and have helped catalyse novel candidate vaccines toward clinical testing at unprecedented speed. Along with inactivated vaccines, emerging nanotechnologies such as mRNA vaccines delivered by lipid nanoparticles and viral vector vaccines have already reached Phase II and III clinical trials (Fig. 1 and Table 1).
a, As of 1 June 2020, 157 vaccine candidates are undergoing development by academic labs and industry (and partnerships thereof). COVID-19 vaccine candidates include live-attenuated vaccines, inactivated vaccines, subunit vaccines, virus-like particles (VLPs), viral vectors (replicating and non-replicating), DNA and RNA vaccines. b, 16 vaccine candidates have entered clinical testing (Table 1). The vaccine candidate data was compiled from searching vaccine trackers resources: Milken Institute (https://milkeninstitute.org/covid-19-tracker), Regulatory Affairs Professionals Society104, BioCentury (https://www.biocentury.com/preclinical-vaccines-and-therapies), and World Health Organization (http://www.who.int/blueprint/priority-diseases/key-action/novel-coronavirus/en/).
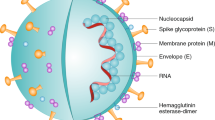
SARS-CoV-2 is an enveloped ssRNA virus with spike-like glycoproteins protruding from its exterior membrane surface forming a ‘corona’. The four major structural proteins of betacoronaviruses are spike (S) protein, envelope (E) protein, membrane (M) protein, and nucleocapsid (N) protein26. The S protein is an attractive target for vaccine design because it facilitates viral entry into the host cell during the infection process. The two spike protein subdomains, S1 and S2, are responsible for host cell angiotensin-converting enzyme 2 (ACE2) receptor binding and host cell membrane fusion, respectively27. S1 contains the receptor-binding domain (RBD) and S2 the fusion machinery enabling virus entry (Fig. 2). While the S1 domain is divergent across the coronaviruses, the S2 domain is more conserved12. Combining SARS-CoV-2 structural information and knowledge gained from SARS/MERS vaccine candidates28, researchers projected the full length S protein, as well as S1, RBD, and S2 subunit derivatives, to contain the prime target epitopes for the induction of neutralizing antibodies. Indeed, recent clinical data from a cohort of convalescent SARS-CoV-2 patients validates this approach—analysed patient sera indicates neutralizing antibodies targeting the different domains of S protein: S1, RBD and S229.
Assembled form three identical chains (depicted in orange, grey and white colours), the S protein is functionally subdivided in S1 (red) and S2 (green) domain; S1 contains the receptor binding domain (RBD). The SARS-CoV-2 structure was reproduced and adapted from the CDC Public Health Image Library. The S protein structure was prepared on the PyMol molecular graphics system (2.3.4) using the PDB file ‘6VSB’.
SARS-CoV-2 S protein is also extensively glycosylated30, and computational analysis indicates glycosylated SARS-CoV-2 S protein to have a more organized conformation versus its non-glycosylated counterpart31. Therefore, glycosylation of SARS-CoV-2 S vaccine candidates should be considered: DNA/RNA vaccines are produced in situ inside the patient and thus would carry native glycosylation; for recombinant subunit vaccines glycoengineering protocols may be applied. The hotspot of SARS-CoV-2, however, which is critical in binding to the ACE2 receptor, appears to be free from glycosylation32; therefore, it may not be required to glycosylate corresponding peptide epitopes.
Contemporary vaccines
Active immunization against viruses have traditionally relied on the usage of whole pathogen in a weakened or killed form through chemical or physical processes, and this has resulted in clinically-approved treatments. Further, several mammalian viral vectors such as oncolytic herpes simplex virus have been repurposed into clinically-approved treatments. Contemporary vaccines hold merit, and here, we highlight how live attenuated, inactivated, and viral vector vaccine development efforts are making the reality of a vaccine against SARS-CoV-2 palpable.
Live attenuated and inactivated vaccines
Live-attenuated vaccines (LAVs) are live, reproducing but avirulent viruses. LAV design intends single-dose immunity without illness. Because LAV technology is mature, LAVs are likely to emerge as one of the frontrunner vaccine candidates for the ongoing COVID-19 pandemic. Codagenix Incorporation’s proprietary deoptimized SARS-CoV-2 vaccine candidate currently leads the charge. However, LAVs bear risks of transfer of the virus and/or reversion to the pathogenic form, reactivation in immune-compromised individuals or recombination with related viruses circulating in the population—especially for novel diseases where pathophysiology is not yet fully understood. LAVs generally require cold chain distribution. Furthermore, loss of efficacy and reproductive potential of progeny viruses during vaccine production poses a significant challenge. Preliminary studies of silent codon change indicate some positive effect on mitigating reversion events, however, these are not general to all viruses33. New technologies such as genetic code expansion are being applied to create highly reproductive but genetically stable LAVs34. More recently, synthetic genomics approaches have enabled the synthesis of recombinant SARS-CoV-2 viruses from fragments of viral DNAs35,36. These strategies could be employed towards rapid generation of SARS-CoV-2 LAVs.
Inactivated vaccines (IVs) are heat or chemically inactivated pathogens or fractions thereof. These vaccine formulations are incapable of replication and safer than LAVs, but their inactivation results in lowered immunogenicity and requirement for multiple-dose regimens to establish long-lasting immunity; also, these vaccine formulations often require adjuvants to immunize the aging population due to immune senescence37. While IVs have better stability profiles compared to LAVs, they still require a cold chain. Several COVID-19 IVs are in development, with the first clinical trial approved recently for Sinovac.
Viral vectors
Mammalian viruses have been engineered and repurposed for several vaccine applications. For COVID-19, there are several vaccine candidates in development that utilize non-replicating adenoviral vectors. Leading adenoviral vectors in SARS-CoV-2 clinical trials are adenovirus type 5 vector (Ad5-nCoV) as of 16 March 2020 and chimpanzee adenovirus vaccine vector (ChAdOx1) as of 31 March 2020 by CanSino Biological and the University of Oxford, respectively (Table 1). Advantages of adenoviral vectors include their broad tissue tropism, inherent adjuvant qualities and scalability. A challenge for adenoviral vector platforms is pre-existing immunity in humans, which may dampen efficacy of the adenoviral vector. While pre-existing immunity against Ad5 is reportedly widespread, its clinical application continues and moreover, alternative ChAdOx1 with low human seroprevalence have been derived for use as vaccine platforms38,39.
Next-generation vaccines enabled through advances in nanotechnology
Viruses are nanoscale objects and therefore can be regarded as naturally occurring nanomaterials; per that definition, LAVs, IVs and viral vectors are nanotechnologies. Nanoparticles and viruses operate at the same length scale—this is what makes nanotechnology approaches in vaccine development and immunoengineering so powerful. Nanoparticles, natural or synthetic, mimic the structural features of viruses whereas chemical biology, biotechnology and nanochemistry enables the development of next-generation designer vaccine technologies. From a vaccine technology development point of view, this is an exciting time and novel technologies and approaches are poised to make a clinical impact for the first time.
Nucleic acid-based vaccines
Delivering the genetic code for in situ production of viral proteins is a promising alternative to conventional vaccine approaches. Both DNA and mRNA vaccines fall under this category and are being pursued in the context of the COVID-19 pandemic. While these platforms are attractive in terms of safety, speed, stability and scalability, they carry a significantly higher risk of failure in clinical development as seen previously with other novel technologies40. To date there is no licensed DNA or RNA vaccine. Nevertheless, a particular advantage of these vaccines is that in addition to antibody and CD4+ T cell responses, DNA or RNA vaccine elicit CD8+ cytotoxic T cell responses, which plays a key role for virus eradication41,42. For the DNA vaccines, the frontrunner in this space is Inovio Pharmaceuticals with their Phase I clinical trial having commenced 6 April 2020. Another rising company on track for a Phase I clinical trial is Entos Pharmaceuticals, Inc. a company based in Alberta, Canada. mRNA vaccines can be produced through in vitro transcription, which precludes the need for cells and their associated regulatory hurdles43. Moderna’s mRNA-based technology was the fastest to Phase I clinical trial in the US, which began on 16 March 2020 (see Table 1). Additionally, BioNTech-Pfizer recently announced regulatory approval in Germany for Phase I/II clinical trials to test four lead mRNA vaccine candidates44.
While DNA vaccines offer higher stability over mRNA vaccines, the mRNA is non-integrating and therefore poses no risk of insertional mutagenesis. Additionally, the half-life, stability and immunogenicity of mRNA can be tuned through established modifications45. For example, researchers at the Imperial College of London and Arcturus Therapeutics are incorporating self-amplifying RNA technology to prolong the otherwise short half-life of the RNA and thereby boost S protein expression levels46 (http://www.imperial.ac.uk/a-z-research/future-vaccine-hub/workstreams/rna-vaccine-manufacture). Nanotechnology-based approaches offer enabling solutions to the delivery challenge by trafficking the vaccine to appropriate cellular populations and subcellular locations. While synthetic nanocarriers including cationic liposomes and polymeric nanoparticles have been used for the delivery of DNA vaccines across cell membranes, targeted formulations could further enhance nuclear translocation of the plasmid DNA47. Moderna’s mRNA vaccine is based on a lipid nanoparticle platform, but there are many other emerging nanotechnologies for delivery of nucleic acid vaccines (several structures are shown in Fig. 3). Nanotechnology platforms including cationic nanoemulsions, liposomes, dendrimers or polysaccharide particles have been employed for improving the stability and delivery of mRNA based vaccines41,45.
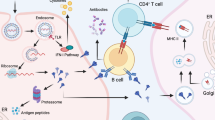
a, Protein nanoparticles and their size; sizes for the synthetic nanocarriers vary between 10–1000 nm. The protein nanoparticles were prepared using Chimera software using the PDB files (3IYI, 1FHA, 1NY7 for P22, ferritin and CPMV, respectively). b, Components of nanoparticle-based vaccines. c, Key steps involved in nanoparticles-based vaccine processing by APCs. The antigenic cargo is processed by the APC and epitopes are presented by MHC-I and MHC-II leading to production of CD8+ cytotoxic T cells or CD4+ T helper cells required for antiviral antibody production (or a combination thereof).
Subunit vaccines
Subunit vaccine candidates constitute minimal structural components of SARS-CoV-2 that can prime protective immune responses in the host when administered with molecular adjuvants for enhanced immunogenicity. For example, contemporary SARS-CoV-2 subunit vaccine candidates are formulations of full-length S protein or S1/S2 subunits with adjuvants. The frontrunner amongst developers is Novavax who initiated a Phase I/II trial on 25 May 2020. Also, Sanofi Pasteur/GSK, Vaxine, Johnson & Johnson and the University of Pittsburgh have announced that they expect to begin Phase I clinical trials within the next few months. Others, including Clover Biopharmaceuticals and the University of Queensland, are independently developing subunit vaccines engineered to present the prefusion trimer confirmation of S protein using the molecular clamp technology48 and the Trimer-tag technology49, respectively. Further, other groups are exploring subunit vaccines using only the RBD of the S protein50.
Alternatively, subunit vaccines can also take the form of protein nanoparticles or virus-like particles (VLPs). VLP vaccines can be produced by recombinant expression and allows for genetic engineering to incorporate ligands, immunomodulators and targeting moieties. Both self-assembled protein nanoparticles and VLPs offer highly ordered, stable and monodisperse vaccine formulations as well as scalable production through fermentation or molecular farming (Table 2). For example, Medicago and iBio are using Nicotiana benthamiana to produce VLPs using the S protein, and AdaptVac/ExpreS2ion is using insect cell expression system to make VLPs from the S2 protein; clinical trials are expected as early as July 2020 (https://www.medicago.com/en/newsroom/; https://ir.ibioinc.com/press-releases; https://news.cision.com/expres2ion-biotechnologies). Besides generating protein nanoparticles from antigenic subunits, their expression and/or display on proteinaceous biomaterial scaffolds such as ferritin, encapsulin51 and bacteriophage VLPs has also been utilized to achieve multivalent antigen display for enhanced immunogenicity52,53,54.
And finally, subunit vaccines can constitute viral proteins incorporated in synthetic nanomaterials, protein cages and VLPs, which serve as adjuvants and/or delivery vehicles, in addition to conferring other benefits inherent to each nanocarrier platform55,56,57. For example, the influenza virus vaccine Crucell (Janssen, Johnson & Johnson) is a liposomal formulation that incorporates influenza protein haemagglutinin58. Figure 3a, b highlights some nanotechnologies used in subunit vaccine design. Besides the aforementioned antigen multivalency, nanocarriers enable efficient co-delivery of antigen/adjuvant to secondary lymphoid organs59, exhibit size-dependent lymphatic trafficking and preferential uptake by antigen presenting cells (APCs), create depot effects for sustained immune stimulus, and facilitate antigen cross presentation—enabling extracellular antigens to be presented via the MHC-I pathway for CD8+ T cell engagement60 (Fig. 3c).
Peptide-based vaccines
An important consideration for vaccine design is safety. Many vaccines rely on immunological presentation of whole structural motifs, for example, full-length S protein, which will present a large repertoire of potent epitopes leading to a broad spectrum of antibody and cellular responses. However, earlier studies on SARS and MERS vaccine candidates have pointed to risks of antibody-dependent enhancement (ADE) of infection61,62,63. In the former, presence of non-neutralizing antibodies contributes to increased infections whereas the latter can lead to life-threatening allergic inflammations64,65. While there is no clear evidence yet, immunological data from patients may point toward possible ADE for SARS-CoV-2, suggesting that high IgG titers correlate with worse outcomes66,67. Therefore, developing peptide epitope vaccine strategies targeting the SARS-CoV-2 S protein may yield a safer vaccine. Various B- and T-cell epitopes of the SARS-CoV-2 S protein have already been identified and predicted in silico15,16,17,18. Importantly, when serum from convalescent COVID-19 patients is screened for neutralizing antibodies, experimentally-derived peptide epitopes will confirm useful epitope regions and inform more optimal antigens in second-generation SARS-CoV-2 peptide-vaccines; the National Institutes of Health (NIH) recently funded La Jolla Institute for Immunology (LJI) in this endeavour68.
Peptide-based vaccines represent the simplest form of vaccines that are easily designed, readily validated and rapidly manufactured69. Peptide-based vaccines can be formulated as peptides plus adjuvant mixtures or peptides can be delivered by an appropriate nanocarrier or be encoded by nucleic acid vaccine formulations. Several peptide-based vaccines as well as peptide–nanoparticle conjugates targeting chronic diseases and cancer are in clinical testing and development70,71. In addition to the development of peptide-based COVID-19 vaccines, industry and academic efforts leverage predicted B- and T-cell epitopes in their subunit vaccines against SARS-CoV-2; for example OncoGen, and University of Cambridge/DIOSynVax are using immunoinformatics-derived peptide sequences of S protein in their vaccine formulations (https://oncogen.ro/oncogen-vaccine-design-for-coronavirus/; https://www.cam.ac.uk/research/news/cambridge-research-team-working-towards-vaccine-against-covid-19). Within the DNA vaccine domain, Immunonomic Therapeutics/EpiVax/PharmaJet are leveraging in silico T-cell epitope prediction72. The use of B-cells epitopes appears practical because of its universality compared to the HLA-restricted T-cell specific vaccines16,18.
Peptide based vaccines are dependent on adjuvants and delivery systems for efficacy, and nanoparticles can serve both these roles. By incorporating emerging strategies for targeting lymph nodes (LNs) or cellular subsets and subcellular locations, nanoparticle vaccine efficacies can be improved and their immune profiles tailored to address specific diseases. For example, the innovative strategy of ‘albumin hitchhiking’ exploits the natural trafficking ability of albumin to LNs73. Recently, the intrinsic ability of nanoparticles to target specific subsets of LN-resident dendritic cells (DCs) and macrophages was utilized to design a dual targeting Hepatitis B virus (HBV) vaccine. The complementing immune responses generated by these cellular subsets resulted in an enhanced efficacy of viral clearance in a chronic HBV mouse model74. Subcellular localization of the antigen is also a critical determinant of the ensuing immune response. Vaccine design parameters such as encapsulated antigens versus surface displayed antigens govern the processing and presentation of the antigen. While the former requires degradation or disassembly of the nanocarrier and therefore mimic viral infection leading to cellular immune response, the latter leads primarily to humoral immune response generated by the externally displayed viral proteins56. However, nanocarriers such as polymeric micelles (PEG-PE) that transform antigenic peptides conformation to facilitate cytosolic delivery could be reliably used for LN targeting, APC uptake and antigen cross presentation75.
VLPs from mammalian viruses, insect viruses, plant viruses and bacteriophages have been developed as peptide display nanotechnologies for various vaccine and immunotherapy applications76 (Fig. 3a,b). While non-infectious toward mammals, VLPs mimic the molecular patterns associated with pathogens, making them highly visible to the immune system. Therefore, VLP display and delivery platforms also serve as adjuvants, making them efficient activators and amplifiers of antigen-specific immune response (Box 1). Several VLP platforms are in the development pipeline and some have entered clinical trials70,77. The beauty of the VLP display platform technology is the scalability and modularity; the peptides can be rapidly adapted as new information about SARS-CoV-2 and its immunogenicity is made available; it is also possible to rapidly adapt the technology should novel or mutated strains emerge. Over the past years, our laboratory has developed plant virus-based nanotechnologies for cancer vaccines and immunotherapy78,79,80. We are now putting this technology toward development of COVID-19 vaccines. An established delivery platform would ensure that the next wave of SARS-CoV-2 infections and other emerging viruses, potentially including novel betacoronaviruses, are met with a more efficient and rapid response.
Vaccine scalability and manufacturing
The rapid emergence of the COVID-19 pandemic has also raised concerns regarding critical deficiencies in manufacturing and distribution of vaccines. Even when an effective vaccine is developed, considerations of cost, formulation and scale-up manufacturing must be taken into account. Large-scale production and worldwide distribution of a potent COVID-19 vaccine(s) will be governed by economic disparities between nations. When demand exceeds supply, developing countries are in a disadvantaged position for the bidding contest to procure the highly sought after vaccine, a situation already seen with personal protective equipment and other critical goods even between industrialized nations. Therefore, it is critical to also consider technologies and platforms suited for developing countries.
Recombinant protein production can be carried out in a variety of platforms, each coming with its own advantages and disadvantages regarding yields, regulatory compliance, cost, scalability, flexibility, speed and safety (Table 2)81,82. While traditional manufacturing processes using bioreactors and mammalian, bacterial or yeast cell cultures are well-established in the pharmaceutical sector, these platforms are expensive, and production can be hampered by human pathogen contamination. Innovative manufacturing technologies that can meet the required global demand and distribution in response to outbreak have recently been deployed with success.
Plant-based expression systems have emerged in the past decade and already made an appearance during the 2014 Ebola epidemic when patients were treated with ZMapp, an antibody cocktail manufactured through molecular farming83,84. Plant molecular farming approaches offer scalability: while in fermentation-based platform, every scale-up step needs to be carefully verified—in molecular farming, each plant is a bioreactor. The more plants are grown, the more product is made; scale-up does not change the upstream production processes82,85. Other positive attributes of the molecular farming platform are the low manufacturing costs, the inability of human pathogens to replicate in plant cells (hence safety), and relatively non-sophisticated infrastructure that could be implemented worldwide also in low-resource countries. While the Bill and Melinda Gates Foundation is funding new factories for potential coronavirus vaccines86, efforts should also be made to prepare for large-scale manufacture of plant molecular farming solutions. Many entities have already announced COVID-19 responses, including for example, Medicago, iBio and Kentucky BioProcessing, all of who set out to develop subunit vaccines87 (https://www.medicago.com/en/newsroom/; https://ir.ibioinc.com/press-releases) as well as Plantform and South Africa-based CapeBiopharms, who set out to produce antibodies and test reagents. Adding to that, further developments and innovations are sprouting from academic labs; including a team at the Spanish Research Council (CSIC) at the Centre for Research in Agricultural Genomics and our laboratories at the University of California, San Diego88,89.
The practical vaccine delivery, distribution and administration
Challenges of vaccine administration for a global pandemic must inform the selection of vaccine platforms. An ideal vaccine platform would have facile integration into devices designed to be supplied far and wide, manufactured at low cost and administered with minimal supervision. (Table 3 and Fig. 4). Several vaccine formulations require constant refrigeration. The need for a cold chain makes their global distribution and application logistically difficult and for under-developed and developing nations with tropical climates, nearly impossible. In fact, one of the largest challenges has been the reliance on refrigerated transport of solution-based vaccines. The WHO also reported that 2.8 million vaccines were lost in five countries due to cold chain failures, and less than 10% of countries met WHO recommendations for effective vaccine management practices. While some lyophilized vaccines are available that may be stored at room temperature, such solutions are difficult to produce and present challenges for the healthcare professionals who must reconstitute them on site90.
Vaccines are encapsulated in polymeric components in either an implant or microneedle patch. Over time the polymer will hydrolyse in the aqueous environment of the body and release the active vaccine. The degradation rate of the device and subsequent release rate of the vaccine can be tuned based on the material in which the vaccine is embedded. The primary difference between the two devices is the means of administration. Implants are administered subcutaneously by a qualified healthcare provider, while microneedle patches can be painlessly self-applied.
A highly suitable nanotechnology platform is derived from plant viruses and bacteriophages that evolved as stable nanocontainers protecting their genome cargo under various environmental conditions. Cowpea mosaic virus, for example, is stable at temperatures above 60°C in buffered solution for at least one hour and pH values from 3.5–9.0 indefinitely at room temperature91. Furthermore, plant virus nanoparticles are stable under gastrointestinal conditions92 and orally bioavailable93, therefore opening the door for global distribution and oral vaccination. Vaccines could be produced in edible leaf tissue to enable vaccination of the human population but also livestock, since SARS-CoV-2 is a zoonotic virus that can infect humans and animals. This would be a step forward to meet the goals of the One Health Initiative to unite human and veterinary medicine, which will likely be important to prevent future outbreaks.
Effective vaccination campaigns also require access to health care professionals (HCP), which is challenging in resource-poor or densely populated developing countries under normal circumstances but presents a greater challenge during a global pandemic where the health care system is already strained or breaking. Recently, modern alternatives to such distribution and access challenges have come to light, such as single-dose slow-release implants, film-based vaccines94, and microneedle-based patches that could reduce reliance on the cold chain and ensure vaccination even in situations where qualified HCP are rare or in high demand. Microneedle-based patches could even be self-administered, which would dramatically hasten roll-out and dissemination of such vaccines as well as reduce the burden on the healthcare system. Such modern vaccine delivery devices can be made by solution methods or fabricated via traditional polymer melt-processing (for example, injection moulding). The advantage of melt-processed devices lies in their potential for rapid manufacturing at large scale and their long-term stability independent of the cold chain95,96,97,98. The potential to break the cold chain and ease the burden on the medical system by offering a safe and effective self-administered prophylactic vaccine has been capitalized on by a number of companies such as Veleritas Inc., Zosano, Corium International and Debiotech, and has led to the filing of over 10,000 patents worldwide99.
Concluding remarks
Advances in bio/nanotechnology and advanced nano/manufacturing coupled with open reporting and data sharing lay the foundation for rapid development of innovative vaccine technologies to make an impact during the COVID-19 pandemic. Within 40 days of initial structural and genomic reports of SARS-CoV-2, the first vaccine candidate entered into the clinical development pipeline and as of 1 June 2020, there are already 16 vaccine candidates in clinical trials, many in Phase II and even one in Phase III. While any vaccine is still months-to-years away from clinical reality, the parallel and rapid efforts from academic laboratories and industry provide hope for success. A plethora of nanotechnology platforms are being pivoted against SARS-CoV-2; while highly promising, many of these may be several years away from deployment and therefore may not have an impact on the SARS-CoV-2 pandemic. Nevertheless, as devastating as COVID-19 is, it may serve as an impetus for the scientific community, funding bodies, and stakeholders to commit more focused efforts toward development of platform technologies that bolster the preparedness for future pandemics. Several nanomaterials afford platform technologies that are amenable to scalability, stability, portability, distribution and device incorporation for self-administration. Moreover, several platform technologies described herein may serve as plug-and-play technologies that can be tailored to seasonal or new strains of coronaviruses. Indeed, COVID-19 harbours the potential to become a seasonal disease; underscoring the need for continued investment in coronavirus vaccines. SARS and MERS vaccine candidates did not make it to market due to lack of financial incentive given the low infection numbers, and because the risk of a global pandemic from a newly emerged virus were largely ignored. Yet, because there is some conservation between the coronaviruses, continued research and product development is critical to tackle any new version of coronavirus that emerges in the future.
References
Wang, C., Horby, P. W., Hayden, F. G. & Gao, G. F. A novel coronavirus outbreak of global health concern. Lancet 395, 470–473 (2020).
Gorbalenya, A. E. et al. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 5, 536–544 (2020).
Kim, J. M. et al. Identification of coronavirus isolated from a patient in Korea with COVID-19. Osong Public Health Res. Perspect. 11, 3–7 (2020).
Institute of Medicine (US) Forum on Microbial Threats. The Threat of Pandemic Influenza: Are We Ready? Workshop Summary (National Academies Press, 2005).
Kissler, S. M., Tedijanto, C., Goldstein, E., Grad, Y. H. & Lipsitch, M. Projecting the transmission dynamics of SARS-CoV-2 through the postpandemic period. Science 368, 860–868 (2020).
Day, M. Covid-19: four fifths of cases are asymptomatic, China figures indicate. BMJ 369, 1375 (2020).
Sutton, D., Fuchs, K., D’Alton, M. & Goffman, D. Universal screening for SARS-CoV-2 in women admitted for delivery. N. Engl. J. Med. 382, 2163–2164 (2020).
Mizumoto, K., Kagaya, K., Zarebski, A. & Chowell, G. Estimating the asymptomatic proportion of coronavirus disease 2019 (COVID-19) cases on board the Diamond Princess cruise ship, Yokohama, Japan, 2020. Eur. Surveill. 25, 2000180 (2020).
Cheng, V. C. C., Lau, S. K. P., Woo, P. C. Y. & Yuen, K. Y. Severe acute respiratory syndrome coronavirus as an agent of emerging and reemerging infection. Clin. Microbiol. Rev. 20, 660–694 (2007).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Andersen, K. G., Rambaut, A., Lipkin, W. I., Holmes, E. C. & Garry, R. F. The proximal origin of SARS-CoV-2. Nat. Med. 26, 450–452 (2020).
Benvenuto, D. et al. The 2019-new coronavirus epidemic: Evidence for virus evolution. J. Med. Virol. 92, 455–459 (2020).
Yuan, M. et al. A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV. Science 368, 630–633 (2020).
Yan, R. et al. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448 (2020).
Lucchese, G. Epitopes for a 2019-nCoV vaccine. Cell. Mol. Immunol. 17, 539–540 (2020).
Grifoni, A. et al. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host Microbe 27, 671–680e2 (2020).
Baruah, V. & Bose, S. Immunoinformatics-aided identification of T cell and B cell epitopes in the surface glycoprotein of 2019-nCoV. J. Med. Virol. 92, 495–500 (2020).
Ahmed, S. F., Quadeer, A. A. & McKay, M. R. Preliminary identification of potential vaccine targets for the COVID-19 coronavirus (SARS-CoV-2) based on SARS-CoV immunological studies. Viruses 12, 254 (2020).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292 (2020).
Hoffmann, M. et al. SARS-CoV-2 Cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Lei, C. et al. Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat. Commun. 11, 2070 (2020).
Wang, C. et al. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat. Commun. 11, 2251 (2020).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Enjuanes, L. et al. Molecular basis of coronavirus virulence and vaccine development. Adv. Virus Res. 96, 245–286 (2016).
Song, Z. et al. From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses 11, 59 (2019).
Schoeman, D. & Fielding, B. C. Coronavirus envelope protein: current knowledge. Virol. J. 16, 69 (2019).
Xia, S. et al. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell. Mol. Immunol. https://doi.org/10.1038/s41423-020-0374-2 (2020).
Wang, N., Shang, J., Jiang, S. & Du, L. Subunit vaccines against emerging pathogenic human coronaviruses. Front. Microbiol. 11, 298 (2020).
Wu, F. et al. Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. Preprint at https://www.medrxiv.org/content/10.1101/2020.03.30.20047365v2 (2020).
Watanabe, Y., Allen, J. D., Wrapp, D., McLellan, J. S. & Crispin, M. Site-specific glycan analysis of the SARS-CoV-2 spike. Science https://doi.org/10.1126/science.abb9983 (2020).
Banerjee, A., Santra, D. & Maiti, S. Energetics based epitope screening in SARS CoV-2 (COVID 19) spike glycoprotein by Immuno-informatic analysis aiming to a suitable vaccine development. Preprint at https://www.biorxiv.org/content/10.1101/2020.04.02.021725v1 (2020).
Zhou, D., Qi, R., Zhang, W., Tian, X. & Peng, C. Identification of 22 N-glycosites on spike glycoprotein of SARS-CoV-2 and accessible surface glycopeptide motifs: implications on vaccination and antibody therapeutics. Preprint at https://www.preprints.org/manuscript/202002.0381/v2 (2020).
Bull, J. J. Evolutionary reversion of live viral vaccines: Can genetic engineering subdue it? Virus Evol. 1, 1 (2015).
Si, L. et al. Generation of influenza A viruses as live but replication-incompetent virus vaccines. Science 354, 1170–1173 (2016).
Thao, T. T. N. et al. Rapid reconstruction of SARS-CoV-2 using a synthetic genomics platform. Nature 582, 561–565 (2020).
Xie, X. et al. An infectious cDNA clone of SARS-CoV-2. Cell Host Microbe 27, 841–848e3 (2020).
Ciabattini, A. et al. Vaccination in the elderly: The challenge of immune changes with aging. Semin. Immunol. 40, 83–94 (2018).
Dicks, M. D. J. et al. A novel chimpanzee adenovirus vector with low human seroprevalence: improved systems for vector derivation and comparative immunogenicity. PloS One 7, e40385 (2012).
Fausther-Bovendo, H. & Kobinger, G. P. Pre-existing immunity against Ad vectors. Hum. Vaccines Immunother. 10, 2875–2884 (2014).
Alberer, M. et al. Safety and immunogenicity of a mRNA rabies vaccine in healthy adults: an open-label, non-randomised, prospective, first-in-human phase 1 clinical trial. Lancet 390, 1511–1520 (2017).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Smith, T. R. F. et al. Immunogenicity of a DNA vaccine candidate for COVID-19. Nat. Commun. 11, 2601 (2020).
Iavarone, C., O’hagan, D. T., Yu, D., Delahaye, N. F. & Ulmer, J. B. Mechanism of action of mRNA-based vaccines. Expert Rev. Vaccines 16, 871–881 (2017).
BioNTech and Pfizer announce regulatory approval from German authority Paul-Ehrlich-Institut to commence first clinical trial of COVID-19 vaccine candidates. BIONTECH https://investors.biontech.de/news-releases/news-release-details/biontech-and-pfizer-announce-regulatory-approval-german (2020).
Zeng, C. et al. Leveraging mRNAs sequences to express SARS-CoV-2 antigens in vivo. Preprint at https://www.biorxiv.org/content/10.1101/2020.04.01.019877v1 (2020).
Arcturus Therapeutics and Duke-NUS Medical School partner to develop a coronavirus (COVID-19) vaccine using STARR™ Technology. ARCTURUS therapeutics https://ir.arcturusrx.com/news-releases/news-release-details/arcturus-therapeutics-and-duke-nus-medical-school-partner (2020).
Lim, M. et al. Engineered nanodelivery systems to improve DNA vaccine technologies. Pharmaceutics 12, 30 (2020).
Takashima, Y., Osaki, M., Ishimaru, Y., Yamaguchi, H. & Harada, A. Artificial molecular clamp: A novel device for synthetic polymerases. Angew. Chem. Int. Ed. 50, 7524–7528 (2011).
Liu, H. et al. Improvement of pharmacokinetic profile of TRAIL via trimer-tag enhances its antitumor activity in vivo. Sci. Rep. 7, 8953 (2017).
Hotez, P. J. & Bottazzi, M. E. Developing a low-cost and accessible COVID-19 vaccine for global health. Preprint at https://www.preprints.org/manuscript/202003.0464/v1 (2020).
Kanekiyo, M. et al. Rational design of an Epstein-Barr virus vaccine targeting the receptor-binding site. Cell 162, 1090–1100 (2015).
Kanekiyo, M. et al. Mosaic nanoparticle display of diverse influenza virus hemagglutinins elicits broad B cell responses. Nat. Immunol. 20, 362–372 (2019).
Sharma, J. et al. A Self-adjuvanted, modular, antigenic VLP for rapid response to influenza virus variability. ACS Appl. Mater. Interfaces 12, 18211–18224 (2020).
Brune, K. D. & Howarth, M. New routes and opportunities for modular construction of particulate vaccines: stick, click, and glue. Front. Immunol. 9, 1–15 (2018).
Ross, K. et al. Single dose combination nanovaccine provides protection against influenza A virus in young and aged mice. Biomater. Sci. 7, 809–821 (2019).
Patterson, D. P., Rynda-Apple, A., Harmsen, A. L., Harmsen, A. G. & Douglas, T. Biomimetic antigenic nanoparticles elicit controlled protective immune response to influenza. ACS Nano 7, 3036–3044 (2013).
Bachmann, M. F. & Jennings, G. T. Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat. Rev. Immunol. 10, 787–796 (2010).
Herzog, C. et al. Eleven years of Inflexal® V—a virosomal adjuvanted influenza vaccine. Vaccine 27, 4381–4387 (2009).
Wang, S. et al. Rational vaccinology with spherical nucleic acids. Proc. Natl Acad. Sci. 116, 10473–10481 (2019).
Smith, D. M., Simon, J. K. & Baker, J. R. Jr Applications of nanotechnology for immunology. Nat. Rev. Immunol. 13, 592–605 (2013).
Wang, Q. et al. Immunodominant SARS coronavirus epitopes in humans elicited both enhancing and neutralizing effects on infection in non-human primates. ACS Infect. Dis. 2, 361–376 (2016).
Quinlan, B. D. et al. The SARS-CoV-2 receptor-binding domain elicits a potent neutralizing response without antibody-dependent enhancement. Preprint at https://www.biorxiv.org/content/10.1101/2020.04.10.036418v1 (2020).
Chen, W. H. et al. Optimization of the production process and characterization of the yeast-expressed SARS-CoV recombinant receptor-binding domain (RBD219-N1), a SARS vaccine candidate. J. Pharm. Sci. 106, 1961–1970 (2017).
Iwasaki, A. & Yang, Y. The potential danger of suboptimal antibody responses in COVID-19. Nat. Rev. Immunol. 20, 339–341 (2020).
Peeples, L. News feature: Avoiding pitfalls in the pursuit of a COVID-19. Vaccin. Proc. Natl Acad. Sci. 117, 8218–8221 (2020).
Zhao, J. et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin. Infect. Dis. (2020).
Zhang, B. et al. Immune phenotyping based on neutrophil-to-lymphocyte ratio and IgG predicts disease severity and outcome for patients with COVID-19. Preprint at https://www.medrxiv.org/content/10.1101/2020.03.12.20035048v1 (2020).
LJI scientists awarded new funding to COMBAT COVID-19. La Jolla Institute for Immunology https://www.lji.org/news-events/news/post/lji-scientists-awarded-new-funding-to-combat-covid-19 (2020).
Li, W., Joshi, M. D., Singhania, S., Ramsey, K. H. & Murthy, A. K. PeptidevVaccine: Progress and challenges. Vaccines 2, 515–536 (2014).
Mohsen, M. O., Zha, L., Cabral-Miranda, G. & Bachmann, M. F. Major findings and recent advances in virus–like particle (VLP)-based vaccines. Semin. Immunol. 34, 123–132 (2017).
Bezu, L. et al. Trial watch: Peptide-based vaccines in anticancer therapy. Oncoimmunology 7, e1511506 (2018).
ITI forms collaboration with EpiVax & PharmaJet to develop novel vaccine candidate against COVID-19 using its investigational UNITE platform. IMMUNOMIC https://www.immunomix.com/immunomic-therapeutics-forms-collaboration-with-epivax-and-pharmajet-to-develop-novel-vaccine-candidate-against-covid-19-using-its-investigational-unite-platform (2020).
Liu, H. et al. Structure-based programming of lymph-node targeting in molecular vaccines. Nature 507, 519–522 (2014).
Wang, W. et al. Dual-targeting nanoparticle vaccine elicits a therapeutic antibody response against chronic hepatitis B. Nat. Nanotechnol. 15, 406–416 (2020).
Liu, Z. et al. Coordinating antigen cytosolic delivery and danger signaling to program potent cross-priming by micelle-based nanovaccine. Cell Discov. 3, 17007 (2017).
Plummer, E. M. & Manchester, M. Viral nanoparticles and virus-like particles: platforms for contemporary vaccine design. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 3, 174–196 (2011).
Shoeb, E. & Hefferon, K. Future of cancer immunotherapy using plant virus-based nanoparticles. Future Sci. OA 5, FSO401 (2019).
Shukla, S. et al. A viral nanoparticle cancer vaccine delays tumor progression and prolongs survival in a HER2+ tumor mouse model. Adv. Ther. 2, 1800139 (2019).
Cai, H., Shukla, S., Wang, C., Masarapu, H. & Steinmetz, N. F. Heterologous prime-boost enhances the antitumor immune response elicited by plant-virus-based cancer vaccine. J. Am. Chem. Soc. 141, 6509–6518 (2019).
Murray, A. A., Wang, C., Fiering, S. & Steinmetz, N. F. In situ vaccination with cowpea vs tobacco mosaic virus against melanoma. Mol. Pharm. 15, 3700–3716 (2018).
Clemente, M. & Corigliano, M. G. Overview of plant-made vaccine antigens against malaria. J. Biomed. Biotechnol. 2012, 206918 (2012).
Nandi, S. et al. Techno-economic analysis of a transient plant-based platform for monoclonal antibody production. mAbs 8, 1456–1466 (2016).
Qiu, X. et al. Reversion of advanced Ebola virus disease in nonhuman primates with ZMapp. Nature 514, 47–53 (2014).
PREVAIL, I. I. writing group. a randomized, controlled trial of ZMapp for Ebola virus. Infect. N. Engl. J. Med. 375, 1448–1456 (2016).
Rosales-Mendoza, S. Will plant-made biopharmaceuticals play a role in the fight against COVID-19? Expert Opin. Biol. Ther. 20, 545–548 (2020).
Bill Gates is able—and willing—to lose big money funding factories for COVID-19 vaccines. FIERCE Pharma https://www.fiercepharma.com/vaccines/bill-gates-plans-to-help-fund-factories-for-7-covid-19-vaccines-but-expects-only-2-will (2020).
BAT biotech using tobacco to make COVID-19 vaccine candidate. BioProcess International https://bioprocessintl.com/bioprocess-insider/therapeutic-class/bat-biotech-using-tobacco-to-make-covid-19-vaccine-candidate (2020).
Researchers in Spain use biotech to produce SARS-CoV-2 vaccine in plants. International Service for the Acquisition of Agri-biotech Applications http://www.isaaa.org/kc/cropbiotechupdate/article/default.asp?ID=18065.
Marrying molecular farming and advanced manufacturing to develop a COVID-19 vaccine. UC San Diego https://ucsdnews.ucsd.edu/pressrelease/marrying-molecular-farming-and-advanced-manufacturing-to-develop-a-covid-19-vaccine (2020).
Kristensen, D. D., Lorenson, T., Bartholomew, K. & Villadiego, S. Can thermostable vaccines help address cold-chain challenges? Results from stakeholder interviews in six low- and middle-income countries. Vaccine 34, 899–904 (2016).
Wang, Q., Lin, T., Tang, L., Johnson, J. E. & Finn, M. G. Icosahedral virus particles as addressable nanoscale building blocks. Angew. Chem. Int. Ed. 41, 459–462 (2002).
Berardi, A., Evans, D. J., Bombelli, F. B. & Lomonossoff, G. P. Stability of plant virus-based nanocarriers in gastrointestinal fluids. Nanoscale 10, 1667–1679 (2018).
Rae, C. S. et al. Systemic trafficking of plant virus nanoparticles in mice via the oral route. Virology 343, 224–235 (2005).
Bajrovic, I., Schafer, S. C., Romanovicz, D. K. & Croyle, M. A. Novel technology for storage and distribution of live vaccines and other biological medicines at ambient temperature. Sci. Adv. 6, eaau4819 (2020).
Lee, P. W. et al. Biodegradable viral nanoparticle/polymer implants prepared via melt-processing. ACS Nano 11, 8777–8789 (2017).
Wirth, D. M. & Pokorski, J. K. Design and fabrication of a low-cost pilot-scale melt-processing system. Polymer 181, 121802 (2019).
Juster, H., Aar, Bvander & Brouwer, Hde A review on microfabrication of thermoplastic polymer-based microneedle arrays. Polym. Eng. Sci. 59, 877–890 (2019).
Bediz, B. et al. Dissolvable microneedle arrays for intradermal delivery of biologics: Fabrication and application. Pharm. Res. 31, 117–135 (2014).
Parhi, R. Review of microneedle based transdermal drug delivery systems. Int. J. Pharm. Sci. Nano. 12, 4511–24 (2019).
Donini, M. & Marusic, C. Current state-of-the-art in plant-based antibody production systems. Biotechnol. Lett. 41, 335–346 (2019).
Vieira Gomes, A. M., Souza Carmo, T., Silva Carvalho, L., Mendonça Bahia, F. & Parachin, N. S. Comparison of yeasts as hosts for recombinant protein production. Microorganisms 6, 38 (2018).
Schneemann, A. & Young, M. J. Viral assembly using heterologous expression systems and cell extracts. Adv. Protein Chem. 64, 1–36 (2003).
Buyel, J. F., Twyman, R. M. & Fischer, R. Very-large-scale production of antibodies in plants: The biologization of manufacturing. Biotechnol. Adv. 35, 458–465 (2017).
Craven, J. COVID-19 vaccine tracker. Regulatory Affairs Professionals Society https://www.raps.org/news-and-articles/news-articles/2020/3/covid-19-vaccine-tracker (2020).
Acknowledgements
This work is supported by a grant from the National Science Foundation NSF CMMI-2027668 (to J.K.P. and N.F.S.). M.S. was supported in part by CBI training program NIH T32GM135142. O.A.O.R. is supported by a UC MEXUS-CONACYT Postdoctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Nanotechnology thanks Yizhou Dong, Mingzhao Zhu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shin, M.D., Shukla, S., Chung, Y.H. et al. COVID-19 vaccine development and a potential nanomaterial path forward. Nat. Nanotechnol. 15, 646–655 (2020). https://doi.org/10.1038/s41565-020-0737-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-020-0737-y
This article is cited by
-
Double-layered N-S1 protein nanoparticle immunization elicits robust cellular immune and broad antibody responses against SARS-CoV-2
Journal of Nanobiotechnology (2024)
-
Continuous and size-control synthesis of lipopolyplex nanoparticles enabled by controlled micromixing performance for mRNA delivery
Journal of Flow Chemistry (2024)
-
The use of RNA-based treatments in the field of cancer immunotherapy
Molecular Cancer (2023)
-
Liver in infections: a single-cell and spatial transcriptomics perspective
Journal of Biomedical Science (2023)
-
mRNA-based cancer therapeutics
Nature Reviews Cancer (2023)