Abstract
The coronavirus disease-19 (COVID-19) is caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The long incubation period of this new virus, which is mostly asymptomatic yet contagious, is a key reason for its rapid spread across the world. Currently, there is no worldwide-approved treatment for COVID-19. Therefore, the clinical and scientific communities have joint efforts to reduce the severe impact of the outbreak. Research on previous emerging infectious diseases have created valuable knowledge that is being exploited for drug repurposing and accelerated vaccine development. Nevertheless, it is important to generate knowledge on SARS-CoV-2 mechanisms of infection and its impact on host immunity, to guide the design of COVID-19 specific therapeutics and vaccines suitable for mass immunization. Nanoscale delivery systems are expected to play a paramount role in the success of these prophylactic and therapeutic approaches. This Review provides an overview of SARS-CoV-2 pathogenesis and examines immune-mediated approaches currently explored for COVID-19 treatments, with an emphasis on nanotechnological tools.
Similar content being viewed by others
Main
The coronavirus disease-19 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), was first reported in Wuhan, China in December 2019. Since then, it has spread globally, already infecting millions of people worldwide. As of 30 June 2020, 213 countries have reported COVID-19 cases, with a total number that reached above 10.3 million, the most being in the USA (2.6 million), Brazil (1.4 million), Russia (640 thousand), India (548 thousand) and UK (314 thousand). USA has the highest number of deaths (126 thousand) followed by Brazil (58 thousand), UK (44 thousand) and Italy (35 thousand). The worldwide case fatality rate across all communities is 4.9%.
Coronaviruses (CoVs) are enveloped viruses entrapping non-segmented, positive-sense and single-stranded ribonucleic acid (ssRNA). Their genome size ranges from 26 to 32 kb, being the largest known RNA virus. SARS-CoV-2 3’ terminus encodes structural proteins, including spike (S) glycoproteins1,2, membrane (M) glycoproteins3, as well as envelope (E)4 and nucleocapsid (N) proteins2,5 (Fig. 1). In addition to the genes encoding structural proteins, there are specific genomic regions encoding for viral proteins required for replication6, in addition to other non-structural proteins, such as the papain-like protease (PLpro)7 and coronavirus main protease (3CLpro)8.
According to the Center for Disease Control and Prevention (CDC), the incubation period following infection is 2–14 days, with an estimated median of 5.1 days9,10. However, cases with longer incubation of 24 days have been reported11. The long incubation period is the primary reason for the massive infection, as it is mostly asymptomatic yet contagious10. Although the estimated patients’ age average is ~70, all age groups are susceptible to this virus. However, the elder population (>60) and people with comorbidities are more likely to develop severe symptoms upon infection12.
Much like previous CoVs, severe acute respiratory syndrome (SARS) and Middle East respiratory ryndrome (MERS), SARS-CoV-2 is predominantly infecting the lower airways, ranging from mild respiratory illness to severe acute respiratory syndrome and septic shock in advanced stages6. The most commonly reported symptoms are fever, dry cough, dyspnea, fatigue and myalgia, which are early characteristics of the most frequent manifestation of SARS-CoV-2 infection, pneumonia13,14,15. Physicians and pathologists are also reporting devastating damage to the cardiovascular system, gut, kidneys and brain16,17. Of importance is the recently observed tendency of COVID-19 patients´ blood to clot, which leads to vessel constriction and ultimately can result in pulmonary embolism or large-vessel ischemic stroke, in addition to ischemia in fingers and toes16,18.
On 7 May, remdesivir was approved for COVID-19 in Japan19. In the rest of the world, there are no specific treatments or vaccines available for COVID-19. To the best of our knowledge, the only available options are the condition marketing authorization to remdesivir recommended by the European Medicines Agency (EMA) on 25 June, as well as the emergency use authorizations (EUA) provided by the US Food and Drug Administration (FDA) on 1 May for remdesivir20, on 30 March for the decades-old malaria drugs hydroxychloroquine sulfate and chloroquine phosphate21, and on 24 March for the infusion of plasma of COVID-19 convalescent patients22. The latter was supported by data obtained on only 5 critically ill COVID-19 patients23, which had high levels of neutralizing antibodies, raising the realistic hope of producing virus-specific enriched immunoglobulins, similarly to the ones used for tetanus, hepatitis A/B, cytomegalovirus, varicella and measles. On 2 June, a clinical trial designed to evaluate the safety and tolerability of the antibody LY-CoV555 was posted. This IgG1 antibody is the first potential preventive and therapeutic medicine developed to specifically target the spike protein on the SARS-CoV-224.
The rapid spread of SARS-CoV-2 requires accelerated development timelines for COVID-19 vaccines and therapeutic candidates to enter expeditiously into phase 1 clinical trials. Hence, the type and extent of preclinical and preliminary clinical data needed to inform these clinical development programs must be weighed against the overall risk–benefit assessment of this unmet medical need. In fact, so far, most published clinical trials on prophylactic and therapeutic strategies against COVID-19 have countless weaknesses, such as not being randomized nor controlled, studying small populations, and/or lacking a placebo group, making it difficult to draw conclusive outcomes.
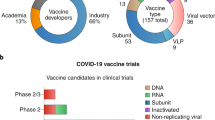
As of 30 June, 2,351 clinical trials are listed in ClinicalTrials.gov, using COVID-19, 2019-nCOV, SARS-CoV-2 or 2019 novel coronavirus as search terms (Fig. 2). Most of trial locations are reported in Europe (709), North America (441), Asia (200) and the Middle East (94).
Summary of COVID-19 trials first posted on ClinicalTrials.gov per month (30 June 2020). The search terms COVID-19, 2019-nCOV, SARS-CoV-2 or 2019 novel coronavirus resulted in 2,351 trials, including 185 with hydroxycholoroquine/chloroquine, 25 with remdesivir, 375 with other repurposed drugs, 48 with vaccines, 217 with immune-modulatory approaches, 118 with convalescent plasma, 202 with diagnostic tests and 41 with dietary supplements.
The strong commitment of the scientific community in addressing COVID-19 disease and global social and economic impact led already to the publication of 27,255 manuscripts (as of 30 June, using keywords COVID-19 or SARS-CoV-2) and submission of 5,891 preprints (4,701 medRxiv, 1,190 bioRxiv). Thus, the large amount of information generated and reported in the last months raises the need for a consensus analysis.
Potential repurposed treatments
The similarities found between SARS-CoV-2 and SARS-CoV, MERS-CoV, or human immunodeficiency viruses (HIV) can enhance the development of potential therapeutic approaches and advance the understanding of the virus mechanism of action25,26,27,28. Currently, major efforts are being invested around the world by commercial vaccine manufacturers, pharmaceutical companies, biotech entities and laboratories at academic institutions to accelerate the discovery and development of prophylactic and therapeutic solutions against SARS-CoV-2 infection using technologies and platforms mostly based on therapeutic modalities developed against these previous infectious diseases. These include repurposing of antivirals (for example, viral polymerase and protease inhibitors29,30), anti-inflammatory drugs31,32, monoclonal antibodies (mAb)27, antibiotics and immune modulators (for example, anti-interleukin-6 (IL-6) agents33,34, PEGylated interferon (IFN)-alpha and beta (PEG-IFN-α/β)35,36), activators of toll-like receptors (TLR)37 and vaccines (for example, based on whole virus38, messenger RNA (mRNA)39, DNA40,41, recombinant proteins42 and peptides43, including when packaged in diverse delivery systems, as well as viral-vectored vaccines44,45 and virus-like particles (VLP)46).
Hydroxychloroquine sulfate and chloroquine phosphate are anti-malaria drugs shown to change the acidic conditions of organelles in mammalian cell culture studies47, as well as to inhibit the terminal glycosylation of ACE2 in vitro against SARS-CoV48, indicating its possible role in preventing the fusion of the virus with the cell membrane and thus blocking SARS-CoV-2 infection. Hydroxychloroquine sulfate was shown to affect immune cell activation and phenotype against lupus erythematosus49, which indicates its possible impact on the modulation of host immune response during SARS-CoV-2 infection50,51,52. Hydroxychloroquine sulfate is also being tested in combination with an array of drugs, amongst them the antibiotic azithromycin, to treat COVID-19 patients53.
Viruses commonly use specific enzymes that act in mechanisms distinct from human cell biology, hence they may be used as specific targets for the treatment of viral infections. In vitro studies have reported that umifenovir inhibits SARS entry into target cells by disturbing the angiotensin-converting enzyme 2 (ACE2)/S protein interaction, further inhibiting fusion of viral envelope54, also being effective in vivo against H1N155 and influenza A viruses56. Umifenovir is now being considered as a COVID-19 treatment in combination with protease inhibitors14,57.
Lopinavir and ritonavir (LPV/RTV) are protease inhibitors used for the treatment of HIV infections by inhibiting the HIV protease activity, which is essential for virus maturation. Both are effective against MERS-CoV in mice58. Cobicistat is a cytochrome P450 (CYP) 3A inhibitor used as a pharmacokinetic enhancer in combination with ritonavir for HIV treatment29,30. Other protease inhibitors are under investigation, such as ASC09 and darunavir59. Ribavirin is a broad-spectrum antiviral drug currently used as a standard of care against hepatitis C virus, in combination with PEG-IFN-α60. It is a ribonucleic analogue that prevents viral RNA replication61.
One of the most investigated drugs at the moment is remdesivir, an adenosine nucleotide analogue that interferes with the viral RNA-polymerase activity, recently approved for COVID-19 treatment in Japan. Remdesivir was shown to be effective against the SARS-CoV virus in vitro and is under evaluation in clinical trials62. Preliminary data on a randomized, placebo-controlled trial of remdesivir enrolling 1063 patients have been released on 22 May, showing that the drug was overall safe and effective for COVID-19 patients, leading to a shorter time to recovery than that obtained in the placebo group63. Similarly, favipiravir is a guanine analogue that also acts as an RNA polymerase inhibitor, currently used for influenza treatment, and 38 clinical trials were recently initiated to assess its effect against COVID-1964,65. Other antivirals currently being considered in COVID-19 clinical trials include the neuraminidase inhibitor oseltamivir and the viral endonuclease inhibitor baloxavir marboxil, both used against influenza infection66. In addition, azvudine is a reverse transcriptase inhibitor developed for HIV treatment that may also interfere with the viral replication67.
Additional clinical trials are exploring the use of anti-inflammatory agents to prevent COVID-19 patients from developing severe lung inflammation. Those include corticosteroids and non-steroid immune suppressive agents, such as indomethacin and barcitinib68, in addition to mAb targeting inflammatory cytokines, such as IL-6 and complement protein 5 (C5). On 16 June, the RECOVERY study announced the first findings of this randomized and controlled clinical trial, which show that the drug dexamethasone reduced the death of COVID-19 ventilated patients by one-third, supporting its potential use as standard of care in these patients69.
Interestingly, angiotensin-receptor blockers (ARB) and angiotensin-converting enzyme (ACE) inhibitors were shown to increase the expression of ACE2, thus potentially enhancing cell infection by SARS-CoV-2. However, a large population-based study, carried out on 6272 COVID-19 patients, did not find a correlation between the use of ACE inhibitors or ARB and SARS-CoV-2 infection70. Therefore, besides the limited data available, currently, there is no clear evidence that supports the discontinuation of these drugs, considering the potential risk that this action could bring to patients with a history of hypertension, heart failure, post-myocardial infarction state and chronic kidney disease.
SARS-CoV-2 mechanisms of infection and immunity
Cell infection by pathogenic agents may trigger host humoral and cellular immunities essential to eliminate the viral infection. However, an uncontrolled or insufficient immune response may lead to immunopathology and cause severe damage to patients71. A deeper understanding of the immune response induced by SARS-CoV-2 infection may lead to new immunotherapies while reducing the potential risk of inflammation.
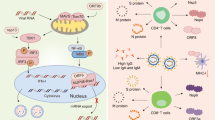
The best known cellular infection mechanism of SARS-CoV-2 is mediated by the cell surface receptor ACE2 (Fig. 3), similarly to SARS-CoV65,72. Several studies have shown that the SARS-CoV-2 was able to infect cells that were genetically modified to express solely the ACE2 receptor30,73. As this ACE2 receptor is predominantly found in the human epithelia of lung and small intestine, the SARS-CoV-2 is more likely to infect the respiratory and gastrointestinal tracts72,74. The preprint posted by Hikmet et al. indicates that the expression of ACE2 was also found in glandular cells of the seminal vesicle, renal proximal tubules, cardiomyocytes, testicular Sertoli cells, Leydig cells and gallbladder epithelium75. Moreover, it has been suggested that the brain may also be infected by this virus, as COVID-19 patients present neurological signs, such as the hyposmia at early stages of infection, but also nausea, vomiting, headache and cerebral damage14 at severe situations. SARS-CoV was indeed found in the brain of animals and patients76. It has been suggested that SARS-CoV-2 enters into the brain through the cerebral circulation, due to the presence of the ACE2 receptor on the endothelium, or from the cribriform plate close to the olfactory bulb, which can justify at least in part the altered sense of smell. In fact, glial cells and neurons are known to express ACE277, being, therefore, potential targets for SARS-CoV-2.
a–d, SARS-CoV-2 is internalized by the cell via (i) membrane fusion or (ii) endocytosis. The SARS-CoV-2 spike binds to the angiotensin-converting enzyme 2 (ACE2) via its receptor-binding domain (RBD) and further releases its RNA (b), which will be translated into viral proteins (c,d). e–h, These proteins will form a replication complex to create additional RNA (e) that will further assemble with the viral proteins into a new virus (f), which will be released (g,h). The transmembrane protease serine 2 (TMPRSS2) is a protease shown to affect virus entry, even though its knockout does not inhibit cell infection by SARS-CoV-2. TMPRSS2, transmembrane protease serine 2; ACE2, angiotensin-converting enzyme 2.
L-SIGN (CD209L) is a less explored potential SARS-CoV-2 receptor. It is a type II transmembrane glycoprotein of the C-type lectin family, mainly expressed in human lung alveolar epithelial type II cells and endothelial cells. This receptor was shown to mediate SARS-CoV entry into host cells, but at a lesser extent than the ACE2 receptor78,79. Therefore, CD209L is likely to mediate SARS-CoV-2 entry as well.
The surface-exposed S glycoprotein is the first viral component that interacts with host cells, being encoded by the S gene, the most variable region of the SARS-CoV genome30. The S protein is a major component accounting for viral binding, fusion and further entry into target cells. It comprises two main functional units. The SARS-CoV-2 S1 subunit contains the receptor-binding domain (RBD), which affinity for binding to host receptor varies among different CoVs, therefore affecting the infection and thereby dictating disease severity6,80. The SARS-CoV-2 S2 subunit is involved in the virus fusion with the host-cell membrane6.
The binding and/or entering mechanisms suggested for SARS-CoV-2 include several steps similar to those identified for SARS-CoV and MERS-CoV infection due to the RBD genomic similarity26. The virus entry is a complex process that starts with the recognition of the RBD 394 glutamine residue by the lysine 31 residue on the human ACE2, further adopting specific conformations that will allow a fast dissociation of the S1 and S2 subunits80,81 (Fig. 3). This proteolytic priming of the S protein by host proteases is also crucial to release the fusion peptide, enabling the viral and host cell membrane fusion, and subsequent release of SARS-CoV-2 RNA into the cytoplasm28,82,83.
It has been shown that the inhibition or depletion of host proteases blocked the entry of the virus in vitro, while protease treatment of target-cell-associated virus led to the opposite effect82,84,85,86. The transmembrane protease serine 2 (TMPRSS2) expressed in human lung cells is one of these proteases. The knockout of TMPRSS2 decreased the viral entry in cell cultures86, but it was not enough to prevent the SARS-CoV-2 infection. An additional study showed that the complete inhibition of viral entry was achieved when the camostat mesylate, a clinically approved serine protease inhibitor, was combined with cathepsin B/L inhibitors. Previous research on SARS-CoV infection exploited other proteases, such as cathepsin L and B, for cell entry87. This study showed that the inhibition of the pH-sensitive endosomal protease cathepsin L activated SARS-CoV membrane fusion in vitro, indicating that the proteolysis of this cathepsin within the endosomes may be required for viral fusion and entry. Therefore, combinations of host cell protease inhibitors constitute potential therapeutics against COVID-19, becoming an active area of research.
Upon virus entry into host cells, genome RNA serves as mRNA for the first open reading frame (ORF1), being thus translated into viral replicase polyproteins that are later cleaved into small products by viral proteinases. These assemble on double-membrane vesicles that become sites for RNA viral synthesis, which has two stages. The first one constitutes the genome replication, while the second one includes the subgenomic RNA transcription and further translation into structural and accessory proteins from additional ORF78,88. These structural proteins are crucial for RNA synthesis by RNA-dependent RNA polymerases (RdRP) in order to replicate the genomic RNA that will be subsequently released upon fusion with plasma membrane89. The double-membranes enable the viral evasion of host immune responses by lacking the pattern recognition receptors (PRR), the activation of which is crucial for triggering host innate immunity against these viral invading pathogens90.
The first line of defence mounted by the host at the entry site, encompassing the induced expression of type I IFN (IFN-I) and other pro-inflammatory cytokines, was reported to be suppressed by SARS-CoV and MERS-CoV36, being related to disease severity.
The modulation of host IFN-I response involves the interference with the ubiquitination and degradation of RNA sensor molecules and the nuclear translocation of the IFN regulatory factor 3 (IRF3), as well as with the reduction of the signal transducer and activator of transcription 1 (SATA1) phosphorylation91,92. It has been already reported that high levels of the cytokine IL-6 were correlated with SARS-CoV-2 viral load in the blood of critically ill COVID-19 patients80.
In addition, the inhibition of cellular components of host immunity has also been reported for SARS-CoV-2 and MERS-CoV infections. The latter led to the downregulation of gene expression related to antigen presentation93, while increased levels of exhausted CD8+ T cells and loss of CD4+ T cell function were found in the peripheral blood of patients infected with SARS-CoV-294.
Therefore, the induction of a balanced host immune response against pathogens in general, and SARS-CoV-2 in particular, is crucial to control and eliminate infection, employing adaptive and innate immune responses, as well as events mediated by the complement system (Supplementary Table 1). On the one hand, an uncontrolled immunity may result in pulmonary tissue damage, functional impairment and reduced lung capacity71. On the other hand, immune insufficiency or misdirection may increase viral replication and cause tissue damage90.
COVID-19 vaccine—a real unmet need
Considering the infection rate of SARS-CoV-2, the prolonged incubation time and the associated high fatality rates, especially in those vulnerable high-risk populations, currently the only possible option to control COVID-19 pandemic is by quarantine, physical distancing and face masks. These measures have been adopted worldwide, but it is becoming an immense economic burden. Historically, vaccines were shown to be effective tools to abolish pandemics, and protect the population, especially high-risk groups, against a variety of viral infections95. Individuals that recover from SARS-CoV-2 infection may develop a protective immunity, considering the immune response triggered by other CoVs and recent findings reported in a small study performed in rhesus macaques, posted as a preprint to bioRxiv96. Even though it is currently unknown if humans previously exposed to the virus are protected against SARS-CoV-2, and there is no sufficient information on the impact of this infection on host immunity, the kinetics of the immune response detected in a patient with mild-to-moderate COVID-19 was recently reported97. The blood of this patient contained increased levels of activated CD4+ and CD8+ T cells, follicular T-helper cells, antibody-secreting cells and IgM/IgG SARS CoV-2-binding antibodies. However, it is not known if those titres are high enough and will remain for a period required to confer protection against re-infection. The antibody response against SARS-CoV was detected at 10–20 days98 and lasts for 2–3 years80.
Importantly, the clinical condition of five patients infected with SARS-CoV-2 already presenting acute respiratory distress syndrome (ARDS), the most severe acute lung injury, was improved following the administration of plasma containing neutralizing antibodies collected from patients that recovered from this disease99. Caution should be taken while drawing conclusions from a clinical study covering such a small number of individuals, however, these observations support the enthusiasm associated with the development of COVID-19 vaccines.
Elucidation of SARS-CoV-2 potential epitopes
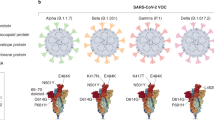
Comparison of a consensus SARS-CoV-2 protein sequence to sequences of SARS-CoV, bat-SARS-like CoV, and MERS-CoV revealed a high degree of similarity between SARS-CoV and bat-SARS-like CoV. This is in contrast to the low similarity to MERS-CoV (Supplementary Table 2)30,63,100.
Taken together, the limited information regarding SARS-CoV-2 and the similarity between the two viruses (SARS-CoV and SARS-CoV-2), led to the rational development of vaccines for SARS-CoV-2 based on the already available knowledge on the Betacoronavirus comparisons.
Among the SARS-CoV-2 proteins, multiple studies suggest that the S and N structural proteins are the most promising targets for vaccine design1,2,26,63,80,101. Since S protein mediates the virus entry into host cells, it is the main target of neutralizing antibodies upon infection1,2. The N protein is a highly immunogenic and abundantly expressed protein during infection of SARS-CoV1,2. In addition, T cell responses against the S, M and N proteins of the SARS-CoV-2 have been reported to be among the most dominant and long-lasting100.
Diverse SARS-CoV epitope sequences have been defined and compared to the SARS-CoV-2 sequence, most of them being derived from S and N proteins. In addition to the characterization of SARS-CoV-2 genome sequences, vaccine design requires the evaluation of their inherent antigenicity to identify T cell and B cell epitope candidates (Fig. 4). To this end, it is fundamental to understand if these sequences bind to major histocompatibility complex (MHC) class I or class II molecules, and further confirm their recognition and subsequent anti-SARS-CoV-2 immune response induction following the activation of human T cell and B cells.
Bioinformatics-assisted prediction of SARS-CoV-2 T cell and B cell epitope candidates given the similarity of the SARS-CoV and SARS-CoV-2 genomic structure. Potential antigen epitope sequences are represented for N protein (PBD: 6VYO) and for S protein (PBD: 6VXX). Potential antigen epitope sequences are not presented for membrane (M) and envelope (E) proteins, as the crystallographic structures of these proteins are not yet available at Protein Data Bank.
Across the world, many research groups addressed several B and T cell epitopes, mainly using bioinformatic tools (Virus Pathogen Database and Analysis Resource (ViPR) or The Immune Epitope Database (IEDB)). Although this is a topic under active investigation, the first genome sequences became available on 11 January 202026,100 and those constitute a valuable source to support the development of COVID-19 vaccine candidates.
The large amount of information generated and reported in the last months suggests the need for a consensus analysis. We summarized the most described B and T cell epitopes that may have the potential to elicit a cross-reactive and effective response against SARS-CoV-2 (Supplementary Table 3).
SARS-CoV-2 vaccine development
The urgent development of a COVID-19 vaccine has prompted industries and researchers worldwide to leverage vaccine platforms developed in the past against other viral infections, such as SARS-CoV and MERS-CoV. Different approaches have been devised to boost and/or reprogramme host immunity against viral diseases, in order to mostly induce Th1-type immune responses and improve the production of binding and also neutralizing antibodies against targeted proteins. The data generated following the clinical development of these platforms is crucial to identify the best COVID-19 vaccine candidates by providing evidence for safety collected during previous biodistribution and toxicity studies.
Most of vaccine candidates102 developed to control SARS-CoV and MERS-CoV infections are based on the whole virus (live-attenuated or inactivated), as well as subunits or full-length viral proteins, DNA and RNA, frequently encompassing the use of non-replicating or replicating viral vectors, VLP or synthetic delivery systems (Fig. 5). The classical inactivated or live-attenuated based vaccines raise safety concerns due to a possible induction of the disease, besides triggering a strong protective immune response, including that observed against SARS-CoV infection103. The efficacy of viral vectors may be compromised by a potentially existing pre-immunity, as the one triggered against the adenovirus. Accordingly, vaccine candidates, for example, based on chimpanzee adenovirus with low seroprevalence in human population104 have been explored against MERS, even if their safety profile remains to be fully addressed in humans.
Proteins, fragments of proteins and peptides, RNAi or DNA- and mRNA-coding antigens can be stabilized while entrapped in different delivery systems, including lipid and polymeric nanoparticles, or conjugated to lipids or polymers. Viral-vectored vaccines and virus-like particles also constitute an active area of research for vaccine development.
The use of peptides, proteins, mRNA- or DNA-based antigens, as well as oligonucleotide-based adjuvants for vaccination is limited by their inherent instability when systemically administered. Advanced formulation methods allied to a rational design of nanocarrier systems have been developed for the systemic delivery of these different bioactive compounds, thus overcoming their rapid degradation by nucleases, fast clearance, limited cellular uptake and off-target effects105,106.
Nanotechnology-based107,108,109,110,111,112,113 approaches hold several advantages that are expected to play a paramount role in the development of an effective vaccine (Box 1).
The FDA- and EMA- approved nanosized delivery systems (for example, liposomes, polymeric nanoparticles) for drug delivery in humans nicely support the impact of these advanced carriers on addressing especially life-threatening human diseases. The bioconvergence merge of nanotechnology, bioengineering, and drug delivery is opportune to address unmet needs, as each of these fields has matured on their own to the point that can now be used to complement others substantively and rationally, rather than modestly and empirically. The expectation is that these nano-delivery platforms along with the recently developed techniques to detect the most appropriate targets will enable potent antigen-specific humoral and cellular immune responses and will allow next-generation vaccines to be devised against a range of viral diseases, including SARS-CoV-2. The use of nano-delivery carriers to advance the development of vaccines have been based on synthetic recombinant proteins or peptide sequences of structural (for example, S or N SARS-CoV proteins114) or non-structural viral targets previously identified as antigens, often combined with adjuvants, such as TLR agonists (for example, CpG43, imiquimod115, Poly (I:C)116, Monophosphoryl Lipid A (MPLA)117 or glucopyranosyl lipid adjuvant (GLA)118, and other immune regulators, such as montanide119 and inflammasome inducers (for example, Chitosan120)) or retinoic acid-inducible protein 1 (RIG-I) ligands121. In fact, our own previous findings demonstrated that the combined delivery of antigens and TLR agonists were fundamental to achieve a strong humoral and cell‐mediated cytotoxic immune response for cancer prevention and therapy43. We now re-orient our efforts to exploit our mannosylated polymeric nanoplatform to develop a COVID-19 vaccine. Our nano-vaccine enable at once to programme the location, pharmacokinetics and co-deliver immunomodulatory compounds, promoting responses that cannot be attained upon the administration of such compounds in solution.
The use of RNA for vaccine development constitutes a revolutionizing technology that involves the use of an adequate delivery system to improve the stability and thereby the translatability of this oligonucleotide intracellularly39,122,123. The non-replicating mRNA vaccines encode the antigen of interest, while self-amplifying RNA will lead to the translation of both the antigen and the viral replication machinery that will allow the intracellular RNA amplification and protein expression. Vaccines based on mRNA, DNA, VLP or other delivery systems became popular candidates especially in light of the recent approvals of 2 small interfering RNA (siRNA) products of Alnylam (Onpattro (patisiran) and Givlaari (givosiran)). No mRNA and DNA-based vaccines have reached the market and therefore, their safety profile in large populations is currently unknown124,125. Nevertheless, the COVID-19 mRNA-loaded cationic lipid nanoparticle (mRNA-1273) that encodes for a stabilized SARS-CoV-2 S protein developed by ModernaTX, Inc126., entered phase I clinical trials127 on 16 March, and phase II clinical trials on 28 May.
While it is crucial to accelerate the development of these immune-mediated approaches, it is important to bear in mind that there are important open questions related to the SARS-CoV-2 infection development and impact on host immunity that must be addressed. This advanced knowledge is fundamental, on the one hand, to elucidate the different mechanisms of the host immune response involved in the neutralization of the virus and/or the eradication of infected cells. However, on the other hand, it will also predict the outcome obtained when massively applied in the targeted population. Accordingly, the gender and age effect on the ability of the virus to modulate host immune response must be fully addressed, as well as the impact of SARS-CoV-2 infection on the immunity of patients with chronic diseases, namely diabetes, hypertension, chronic obstructive pulmonary disease (COPD), among others.
At early stages of SARS-CoV infection, macrophages display a pro-inflammatory phenotype (M1) and the production of nitric oxide, IL-6, IL-8, IL-1, ROS, MCP-1, CXCL-10 and TNF mediate host fight against the virus, but also promotes lung injury128. In the meanwhile, anti-inflammatory macrophages (M2) become activated and these will regulate wound healing once the pathogenic agent is eliminated, restoring the lung tissue128. It has been observed that patients defeated by the SARS-CoV infection presented a faster increase in anti-S neutralizing antibody titres than those that recovered from this disease129. Furthermore, studies on SARS-CoV-infected mice and monkeys reported an association between the induction of a SARS-CoV-specific immune memory and enhanced lung inflammation46,130. However, T cells have also been related to murine protection against this infection131,132. A recent study128 showed that the vaccination of SARS-CoV infected macaques with a modified vaccinia Ankara (MVA) virus encoding full-length SARS-CoV S glycoprotein (ADS-MVA) was able to improve the production of anti-SARS-CoV S protein neutralizing antibodies, but these animals presented different degrees of alveolar damage. This S-IgG specific immunity was related to a decrease in viral load, in parallel to an improved infiltration of inflammatory monocytes/macrophages in the lungs and abrogation of wound-healing macrophage response at the early stages of infection128. Moreover, the medRxiv preprint posted by Wu et al. indicates that the anti-S specific antibody levels were correlated to lymphopenia and disease severity in COVID-19 patients133. Other studies have reported that the S-specific immunity successfully assisted in viral clearance and protected experimental animals against SARS-CoV infection134,135.
Therefore, systematic studies must address the SARS-CoV-2 immune-mediated impact at different stages of the disease, correlating those findings associated to cellular and humoral responses with their impact on pulmonary immunopathology, but also taking into consideration the type of vaccine and the viral strain used for infections.
Depending on the vaccine and related nature of the antigen used to trigger a specific immune response, it is important to demonstrate the extension of the response triggered under COVID-19 disease upon immunization (Fig. 6). These required preclinical studies are currently limited as it was previously shown that the CoV S protein does not bind to mouse counterpart due to structural differences already identified between the human and mouse ACE2136. However, even if the information available on preclinical studies performed to specifically address immune-mediated response against COVID-19 is limited, including in non-human primate models, several mouse models previously developed to address the development of therapeutic and prophylactic solutions against SARS-CoV infection can now be explored to advance drug discovery and development against COVID-19. Examples include the ACE2, TMPRSS2 and STAT1 knockout mouse models. ACE2137,138 and TMPRSS286,138 have been identified as being related to SARS-CoV-2 entry into cells, while STAT1139,140 favours progressive lung disease by increasing viral replication in the lungs. These experimental mouse models are particularly suitable for the study of SARS-CoV-2 pathogenesis and development of new therapeutic options. BALB/c and C57BL/6 mice may be used to characterize the immune response against potential vaccine candidates, even in the absence of disease141. Particularly relevant for characterizing the human immune response triggered by SARS-CoV-2 infection and therefore to guide the design of effective and safe vaccines may be the transgenic human leucocyte antigen (HLA) class I and class II mouse models142 harbouring HLA genes covering high percentages of human population. Additional tools to support preclinical development include the transgenic mouse model bearing human ACE2 for SARS-CoV-2 infection (SARS-CoV-2 hACE2), including the recently developed by Bao et al.51 and Jiang et al.143, as well as, by the Jackson Laboratories following a model previously established by the Pearlman research group144,145. Moreover, mice that support the development of immune cells following the engraftment of human peripheral blood mononuclear cells (PBMC)146,147 or human hematopoietic stem cells (HSC)148 should also be considered.
These assays include the evaluation of dendritic cell (DC) and T cell function upon immunization by flow cytometry and quantification of levels of antigen-specific binding and neutralizing antibodies at different time-points. These studies are still limited by the mouse models of SARS disease currently available, but different options are emerging as potentially useful for the study of SARS-CoV-2 infection mechanisms and COVID-19 vaccine development. DC, dendritic cell; FACS, fluorescence-activated cell sortin; ELISA, enzyme-linked immunosorbent assay; Ig, immunoglobulins (Ig); hACE2, human angiotensin-converting enzyme 2; HLA, human leukocyte antigen; PBMC, peripheral blood mononuclear cell.
Host immune-modulation beyond vaccines
The homology of MERS-CoV, SARS-CoV, and the newly emerging SARS-CoV-2, as well as the similar clinical development of these diseases that starts by a pulmonary inflammatory state followed by fibrosis and subsequent compromise of lung function, are driving the attention of the scientific and clinical communities to the use of immune-modulators, which has already achieved remarkable success149. The immune-mediated targets vary between the different types of viruses, as in some infections this pathogen has a major role in host tissue/organ damage, while in others it is the systemic and local immune activation mounted upon infection that impairs host repair function due to hyperstimulation149.
The scientific and clinical communities have been exploring mostly two approaches to achieve balanced immunity to reduce viral load, while preventing end-stage failure of the infected organ90. The first one encompasses the administration of immune suppressors150 to reduce the hyperinflammation induced by the viral infection, that in turn can provoke damage to the inflamed area90. Corticosteroids are widely used to suppress lung inflammation and reduce the risk for pulmonary damage in critically ill patients affected by respiratory viral infections, such as those induced by MERS-CoV32,151 and SARS-CoV152. However, the real risk-benefit underlying the use of corticosteroids is yet unclear. In fact, corticosteroids also limit immune responses and thereby might counteract viral clearance, as it was observed in some individuals affected by MERS-CoV infection31,153. Taking this into account, together with ARDS developed in patients affected by SARS-CoV-2 infection, the recommendation of corticosteroid treatment to these patients is still controversial, being currently addressed in ongoing clinical trials (NCT04244591)13,154. Nevertheless, the RECOVERY trial results point toward the potential use of dexamethasone to reduce the death rate of patients that require respiratory support69.
The second therapeutic approach is based on the use of immune potentiators, such as cytokines, immune checkpoint inhibitors, signalling proteins, antimicrobial peptides and PRR ligands. The main goal is to eradicate the virus following the stimulation of host innate and adaptive immune responses against the virus.
Patients with severe COVID-19 develop viral sepsis following the excessive inflammatory response syndrome, as well as inflammation-lung injury, which often leads to septic shock and ARDS, the leading cause of death of these individuals155. Viral sepsis induces the dysfunction of several physiological responses, including both innate and adaptive immune responses, that reduces host ability to eradicate the viral infection156. However, treatments addressing this excessive inflammation against other microbial infections did not improve patient prognosis157.
The immunosuppression, mainly lymphocytopenia, is recognized as a leading cause of increased morbidity and mortality during sepsis158, enhancing patient vulnerability to bacterial or viral infections159. One of the main hypotheses for this immunosuppression occurred during sepsis is the increased expression of the immune regulatory checkpoints156, such as programmed death-1 (PD-1) and programmed death ligand-1 (PD-L1), which have a crucial role in T-cell depletion in septic patients160. Preclinical and clinical studies have shown that the inhibition of those immune checkpoint molecules can reverse the sepsis-induced immunosuppression, overcoming lymphocytopenia and thereby improving host resistance to infection161. These effects suggest that PD-1/PDL-1 inhibition constitutes a potential treatment against SARS-CoV-2- induced sepsis, and anti-PD-1 blocking antibodies are currently being investigated, as single agents or in combination with thymosin, that has also been shown to reduce the mortality in these patients (NCT04268537). In addition, anti-PD-L1-treated DC induced T cell priming and proliferation162. Therefore, the blockage of PD-1 associated with a vaccine may also be considered in future endeavours.
Additional approaches focus on the reduction of cytokine secretion. Cytokines are protein mediators that provide signals crucial for key biological processes including immunity, cell proliferation and inflammation, wound healing and repair, cell migration, fibrosis and angiogenesis163. High levels of transforming growth factor beta-1 (TGF-β1)26,27 have been detected in SARS patients, being related to the development of pulmonary fibrosis and to the reduction in apoptosis of SARS-CoV-infected cells164. Similar consequences of high levels of TGF-β1 were found in SARS-CoV-2-infected patients. This suggests that targeting pro-inflammatory cytokines that induce pulmonary fibrosis may be useful. One such therapeutic option is pirfenidone, for which different mechanisms have been suggested for its activity. Specifically, it reduces the levels of fibrosis-associated proteins and cytokines, as well as the extracellular matrix accumulation in response to TGF-β1 and platelet-derived growth factor (PDGF)165. The effect of pirfenidone is currently being studied in a clinical trial (NCT04282902) taking into account encouraging results previously reported against idiopathic pulmonary fibrosis, in addition to its pronounced anti-inflammatory and anti-oxidant effects. Furthermore, the impact of the suppression of the TGF-β pathway on overall T-cell function should also be considered in these patients, as it may limit the regulatory T cell (Treg)-mediated immunosuppression166.
In contrast to what was observed regarding the overexpression of TGF-β1 in SARS-CoV and MERS-CoV infected patients, the analysis of the expression profiles of immune-related genes indicated the downregulation of IFN-α/β genes35,36. The SARS-CoV virus does not indeed trigger the activation of the IFN-α/β pathway97, which has a relevant role on reducing viral replication and dissemination at an early stage167, in addition to the overall stimulatory effect on host immune system168. Therefore, several ongoing clinical trials are exploring the efficacy and safety of different IFN subtypes for COVID-19 patients as therapeutic and prophylactic agents (NCT04254874, NCT04293887, NCT04315948).
In addition to this passive administration of IFN, alternative approaches for inducing the stimulation of TLR3 and TLR7 are particularly promising. These are transmembrane upstream regulatory factors that upon recognition of a specific pattern, such as a single and double stranded RNA viruses, will induce the transcription of pro-inflammatory cytokines, including IFN91, increasing host ability to eliminate the pathogen169.
TLR bridge innate and adaptive immune responses as they do not only activate the innate immune system but can also determine the nature of the adaptive immune response by upregulating MHC on DC, in addition to the secretion of costimulatory molecules and proinflammatory cytokine release. This process leads to the differentiation of CD4+ T cells into Th1 cells, which in turn produce IFN-γ, and lead to class switch from IgM to IgG2 antibodies by B cells170. Therefore, an ongoing clinical trial in China (ChiCTR2000029776) is evaluating the therapeutic effect of triggering TLR signalling pathways in COVID-19 patients by an analogue of double-stranded RNA [Poly(I:C)].
Another immunological approach for treating COVID-19 is based on the use of endogenous molecules of the innate immune system such as defensins. Defensins are a family of endogenous antibiotic peptides with a wide antimicrobial and antiviral spectrum activity, being important for the function of the innate defence system171. It was previously observed that the treatment of SARS-CoV-infected mice with defensin caused alterations in the cytokine profile at the lung parenchyma, but it did not seem to affect lung pathology, inhibit proliferation or eradicate the SARS‐CoV virus. However, the intranasal administration of defensin significantly protected mice against SARS-CoV infection172, being therefore potentially beneficial for high-risk groups.
COVID-19 preclinical prophylactic immune-based approaches
Since the end of 2019, multiple strategies are being adopted for the SARS-CoV-2 vaccine development74,173. The majority of these approaches target the surface-exposed S glycoprotein or the S protein (full-length or specific subunits) to induce a potent neutralizing effect by eliciting specific T-cell responses and neutralizing antibodies174,175. As of 8 April 2020, there were 115 vaccine candidates, of which 37 have not been confirmed as active against SARS-CoV-2 infection, and 73 out of the 78 reported as active, are in very early stage phases with no preclinical studies yet97.
Supplementary Tables 4a,b summarize diverse vaccine candidates developed to overcome SARS-CoV-2 infection, of which efficacy and safety are being evaluated at a preclinical stage. It is reasonable to anticipate that even those candidates that will successfully demonstrate a safety profile and considerable ability to modulate host immunity against this virus, it will still take about a year to start phase I clinical trials.
COVID-19 preclinical therapeutic immune-based approaches
Therapeutic options that could be used against SARS-CoV-2 include virus-binding molecules, inhibitors that target specific enzymes involved in viral replication and transcription, small-molecule inhibitors targeting helicase, proteases or other proteins crucial for the survival of the virus, host cell protease and endocytosis inhibitors, siRNA, anti-sense RNA and ribozyme, neutralizing antibodies, mAb against host receptor or S1 RBD, anti-viral peptide targeting S2 and natural products173,176,177. For now, remdesivir, lopinavir/ritonavir alone or in combination with IFN-β, mAb or convalescent plasma are among the most investigated therapeutic options against the SARS-CoV-2173,178. In addition, the characterization of the effects induced by mAb in COVID-19 patients may also advance the development of vaccines, and increasingly specific diagnostics179. However, every single one of these tools needs to be evaluated regarding clinical efficacy and safety, before being used to treat infected patients. Supplementary Tables 5a,d list the advanced therapeutic approaches under preclinical development for SARS-CoV-2.
COVID-19 prophylactic approaches in clinical development
Prevention of SARS-CoV-2 new infections may be the most effective approach, not only to prevent COVID-19 but also to block the spreading of the virus worldwide. Since SARS-CoV-2 virus was only recently identified, large efforts dedicated to vaccine development are being pursued. As of 30 June, 48 clinical trials are already evaluating the efficacy of COVID-19 vaccines (Supplementary Table 6a). One approach, developed by ModernaTX, Inc. uses lipid nanoparticles (LNP) encapsulating mRNA-1273 that encodes full-length, SARS-CoV-2 S protein (NCT04283461). Cells expressing this viral protein will be able to present T cell-recognized SARS-CoV-2 antigen and induce an immune response against the virus. This may be an effective and safe approach since it does not use viral particles, but rather delivers mRNA that can be expressed by immune cells and non-immune cells, leading to both MCH class I and MHC class II antigen presentation. Furthermore, encapsulating viral mRNA in LNP will protect the mRNA from degradation and enhance delivery efficiency180. ModernaTX’s press release181 published on 18 May announced the interim data of this ongoing phase I trial for mRNA-1273 vaccine, which indicate that the vaccine was generally well-tolerated and safe, and led to the production of neutralizing antibodies in eight initial patients enrolled in this study, two weeks after receiving the second dose of this COVID-19 vaccine candidate.
Recombinant novel CoV vaccine using adenovirus type 5 vector, also called Ad5-nCoV (NCT04313127) was proposed by Tianjin-based CanSino Biologics Inc. This product uses a genetically-modified adenovirus type 5 that is replication-defective and expresses the SARS-CoV-2 S protein. The ChAdOx1 nCoV-19 (NCT04324606) developed by the University of Oxford is also an adenovirus-vectored vaccine in clinical development, which has been explored as a vaccine against other infectious diseases, such as influenza, tuberculosis, Chikungunya virus (CHIKV), Zika, meningitis B and the plague.
There are other clinical trials exploring the use of genetically modified, activated immune cells, which recognize SARS-CoV-2 antigens (Supplementary Table 6a). Shenzhen Geno-Immune Medical Institute engineered minigenes (exon gene fragments) based on multiple viral genes, which are introduced into artificial antigen presenting cells (aAPC) using a lentiviral vector (NCT04299724). When injected into the patient, the aAPC will activate T cells and generate an immune response against the virus. In addition, LV-SMENP-DC, also from Shenzhen Geno-Immune Medical Institute, is a modified DC that expresses viral antigens (NCT04276896). These DC are used to activate cytotoxic T cells (CTL) ex vivo. The modified DC and activated CTL will be both injected into the patients. On 6 April 2020, INOVIO Pharmaceuticals announced the initiation of phase 1 clinical trial of its INO-4800 DNA plasmid vaccine encoding S protein delivered by electroporation (NCT04336410), based on their technology previously exploited for Lassa, Nipah, HIV, filovirus, human papillomavirus, cancer indications, Zika and hepatitis B. An additional prophylactic approach to prevent COVID-19 disease in adults exposed to the virus, encompasses the inhalation of the PUL-042 solution, which is a TLR 2/6/9 agonist182. By binding to these TLR, PUL-042 can activate immune cells, such as natural killer (NK) cells, macrophages and DC, and stimulate lung epithelial cells to produce diverse factors against photogenic infection183.
Although promising, these clinical trials are now recruiting for phase I or phase II, and efficacy and safety in human patients have not yet been confirmed.
COVID-19 therapeutic approaches in clinical development
Most of the clinical studies test current immunotherapies or re-purposing of existing drugs for the treatment of COVID-19 patients (Supplementary Tables 6a–d). As of 30 June, there are 217 registered clinical trials using immune modulators against COVID-19, while over 476 investigate the added-value of repurposing drugs.
The injection of immunoglobulins from convalescent patients has been tested to induce a specific immune response against SARS-COV-2 (NCT04264858). Other trials are trying to induce a broad immune response through the non-specific activation of the immune system. For instance, a combination of a standard treatment with anti-PD-1 antibody is under clinical investigation (NCT04268537). Another example is the recombinant IFN-α2β, an immune-modulator approved for the treatment of viral infections, such as hepatitis B and C, and for the treatment of different types of cancer, such as metastatic melanoma (NCT04293887)184,185.
Other clinical studies are testing different anti-inflammatory agents to reduce lung inflammation (pneumonia), the leading cause of death for COVID-19 patients. These include antibodies targeting inflammatory factors, such as IL-6 and complement protein C5, or the CD24Fc conjugate that blocks TLR activation. Two clinical studies are using the anti-angiogenic drug bevacizumab (anti-VEGF mAb) to reduce lung oedema. Another antibody in clinical development is the meplazumab, which blocks the binding of SARS-CoV-2 S protein to CD147 molecule on human cells, thus reducing the virus infection ability34. Other immunosuppressive agents are also being tested, such as the JAK1/JAK2 inhibitor baricitinib and the anti-malaria drug hydroxychloroquine sulfate. Although optimal treatment regimens are still under investigation, different dosing and schedules are being reported by clinicians.
Few clinical studies are exploring cell therapies, such as NK cells to support and strengthen the immune response (NCT04280224), while others use mesenchymal stem cells, which can facilitate tissue regeneration and immune suppression186.
Facing the tragic numbers that are arriving every day from all over the world, on 18 March 2020, the United Nations Director-General announced an unprecedented action where international cooperation made possible the design of the Solidarity Trial. The World Health Organization (WHO)187 announced on 27 March 2020 that 45 countries were already contributing, and many others had demonstrated interest in joining this historic trial, where the major aim is to rapidly understand the safety and effectiveness of remdesivir, lopinavir/ritonavir, lopinavir/ritonavir with IFN-β1a and chloroquine or hydroxychloroquine188. The contribution of data obtained by this multi-country trial in a concerted manner enables the direct comparison between those drugs, which will drastically cut the time needed to provide clear and robust data to demonstrate which drugs will be actually effective against COVID-19.
Conclusions and perspectives
The COVID-19 pandemic has spread to 213 countries and territories, and the number of cases reported as of 30 June 2020 surpasses 8.5 million worldwide. More than 5.6 million people recovered from SARS-CoV-2 infection, but 506 thousand people were defeated by this virus. The number of new cases continuously rises.
The severity of the situation has already re-shaped our society. Reports are arriving from most countries testifying the tremendous commitment of the governmental, scientific and clinical communities, in making concerted efforts to help local populations to deal with the pandemic.
Reorientation of research at many academic and industrial institutions is also currently in place, taking advantage of already existing knowledge and experience in fighting previous infectious diseases that may bring new solutions to the COVID-19 pipeline. History also shows that these crises create unique opportunities for the development of new technologies or the flexible use of existing ones.
The SARS-CoV-2 is indeed a new strain of coronavirus, and even if the scientific community is rapidly gaining tremendous knowledge about this virus, the world population has not yet acquired immunity. The development of an effective vaccine will be particularly important to protect high-risk patients, taking into account their decaying immune function. Therefore, the development of a COVID-19 vaccine suitable for mass immunization is urgent and companies and research institutes have already reported the development of more than 115 vaccine candidates. Researchers worldwide are exploiting platform technologies previously developed to control CoV infections, as well as other diseases, such as HIV, influenza, Zika, Ebola, plague, tuberculosis and meningitis. The development of these vaccine candidates is expected to be particularly fast as their safety and efficacy in modulating host immune response have already been attested, even if against other agents that may not exactly present the same pathogenesis but that modulate physiological functions in a similar manner. However, despite the diverse platform technologies available to produce these vaccines, all developers agree that it is highly unlikely that a vaccine will be available for worldwide use in less than 12 months, in particular taking into account that the development of an Ebola vaccine took 5 years and a mumps vaccine took 4 years in a record time.
The tremendous advances in molecular engineering and biotechnology for the last years have led to engineered biotech compounds, such as peptide and protein antigens, multiple copies of antigen-encoding mRNA, in addition to gene regulators of immune cell function, such as siRNA to suppress the expression of immunosuppression-related genes.
However, as candidates move towards clinical investigation, it becomes clear that their biological effects depend on the development of a tool able to attain their transport across biological barriers. In fact, low response rate of patients has been related to the limited intrinsic immunogenicity of antigens, and therefore, the added value of adjuvants, such as TLR ligands, are commonly considered to overcome sub-optimal efficacy. However, oligonucleotides, such as the TLR ligands CpG or Poly (I:C), or siRNA and mRNA candidates, are limited by low stability under physiological conditions, off-target effects and limited cellular uptake189. The systemic effect of oligonucleotides demands the design of safe and effective delivery systems to target and cross plasma membranes, but also to escape from endosomal compartments into the cytoplasm. The key to unlock their potential has pointed nanomedicines as an approach able to guarantee the target selectivity required for their efficacy, while ensuring patients’ safety. Accordingly, nanotechnology-based systems incorporating combinations of antigen epitopes and adjuvants are being developed to improve vaccine delivery. These nano-based vaccines orchestrate a broad immunity by modulating B cell-mediated responses towards an increased and fast production of high-affinity neutralizing antibodies upon re-encountering cognate antigens, but also by allowing an adequate activation and expansion of T cell-function directed, for example, to the destruction of virus-infected cells. Nanotechnology-based strategies also constitute a potentially useful tool to enhance the stability and improve the pharmacokinetics of therapeutic antibodies. The entrapment of antibody derivatives has been explored to improve their delivery and endosomal escape, which will result in improved intracellular antigen recognition190. It has also been shown that the entrapment of immune checkpoint monoclonal antibody modulators by polymeric nanoparticles improved the immune-mediated responses to these immunotherapies, such as to the agonist anti-OX40191 and to the anti-CTLA4 blocker192. Nanoscale delivery systems improve the delivery of active compounds to specific cells and tissues, and decrease adverse effects cause by systemic administration, for example of FDA-approved cytokines, such as TNF-α193 and IL-2194.
The similarity between the infection mechanisms of SARS-CoV-2 and other CoVs and HIV also indicates that drugs already clinically-approved to target different steps of the infection pathway constitute promising potential solutions against COVID-19. Accordingly, ritonavir and lopinavir are currently under consideration. However, caution should be taken due to distinct HIV and SARS proteases, which can limit the specificity of their therapeutic activity. Chloroquine phosphate and hydroxychloroquine sulfate, as well as remdesivir may lead to better clinical outcomes as these will probably have a broad-spectrum activity by eventually inhibiting SARS-CoV-2 cell entry and the RNA polymerase, respectively195. Time is also crucial; vaccines are intended for administration before the infection (or as intervention in case of a therapeutic vaccine), antivirals need to be administered as soon as possible after infection, whereas immunostimulants or -inhibitors could be given later.
Despite lack of medical evidence, the severe respiratory symptoms presented by an 80-year old critically ill COVID-19 patient were rapidly attenuated upon the administration of recombinant human erythropoietin (EPO)196. Previous preclinical studies reported the effect of EPO on septic cases197,198, and on the inhibition of pro-inflammatory cytokine expression and increase of B cells, as well as CD8+ and CD4+ T cells in circulation199. Even though this patient also received hydroxychloroquine, oseltamivir and lopinavir/ritonavir, these effects were observed following EPO and red blood cell transfusion, which led to an increase in the levels of haemoglobin. Clinical studies on the use of EPO in COVID-19 patients are required, but its use should be carefully considered, for example in patients with anaemia associated with renal chronic disease due to already known side effects such as thrombotic events.
It is important to bear in mind that unfortunately, we are still far from having a full picture of the pathophysiology of this dangerous disease, including its long-term implications on individuals that did not experience the severe form of this infection. What the clinical and scientific communities first thought of as being an infectious disease that, similarly to other CoV-related infections, would mostly affect the respiratory tract, soon became a multifaceted pathology, which major features include the massive accumulation of blood clots. It is now clear that the SARS-CoV-2 infection causes heart attacks, coronary-related kidney damage, severe stroke, including in young people with no previous history of cardiovascular disease, being all related to extensive clot formation. In addition, several countries, such as Italy200, UK201, Canada202 and USA202, reported the hospitalization of children and young adults presenting severe Kawasaki-like disease related to COVID-19, showing inflammation in medium-size blood vessels, rash, fever and, in some cases, shock. Children are accepted to be the least affected by this disease, but this emerging link between SARS-CoV-2 infection and this rare inflammatory condition associated with systemic blood vessel damage in COVID-19 positive young patients, is raising awareness towards the urgent need for collecting data covering clinical outcomes and presentations of this emerging phenomenon. Clinicians and scientist worldwide are driving concerted efforts to correlate the behaviour of the immune system components with SARS-CoV-2 infection, as well as to decipher genes related to high-risk of disease severity, fundamental to point out children who are at high-risk and help on clarifying the chaotic/uncontrolled immunity found in adults.
The search for life-saving solutions to treat COVID-19 patients or achieve the control of SARS-CoV-2 transmission is crucial to win the race against the rapid transmission of this virus, and regulatory bodies worldwide have joined efforts to respond to this outbreak by providing new mechanisms to support a faster development and approval of the most promising therapeutics and vaccine candidates. However, it is fundamental to simultaneously ensure that adequate measures are being taken to thoroughly demonstrate their safety and efficacy. For example, the European Medicines Agency (EMA) released on 27 March an updated version of the Guidance on the management of clinical trials during COVID-19203, namely addressing communication with the governmental authorities, informed consent and distribution channels of investigational medicines. The guidance further includes information regarding the distribution of diagnosis tests, safety reporting and auditing.
The COVID-19 pandemic reminds us of the devastating effect that previous emerging diseases had in our history until the development of effective vaccines that greatly reduced the burden caused by those infections. As recognized by the WHO204, only access to clean water has a greater impact on disease and death of populations. Anti-vaccine movements have raised their voices against vaccination programmes worldwide, which has already increased the incidence of diseases that were already rare in developed countries. Therefore, the measures now being taken by society to limit the severe economic, social and cultural consequences of the COVID-19 pandemic, has the potential to make the national vaccination programmes strong again. Sadly, however, physical distancing measures and lockdown have already heavily compromised the vaccination of children against other important infectious diseases, namely measles, polio, diphtheria, tetanus and pertussis205. WHO indicates that 100 million children are expected to miss measles vaccination due to COVID-19 lockdown. As a result, as the pandemic starts to reach a containment phase in several countries, it is important to analyse data gathered for the last 5 months and the expected impact of a prolonged lockdown, looking also at potential effect on social and economic levels. It is also important to investigate medical care procedures and the treatment of patients affected by other diseases that have been abruptly interrupted due to the COVID-19 pandemic. For example, cancelled procedures that limit disease diagnosis and cancer treatment schedules that have been affected, such as surgeries, chemotherapy or radiotherapy. Taking these facts into account, several countries are already taking careful measures to relax COVID-19 lockdown, to simultaneously control SARS-CoV-2 spread among the population.
The management of highly contagious and potentially fatal infectious diseases has shaped human history over the centuries, and the COVID-19 pandemic will not be an exception. However, we are now in an era in which highly advanced technological resources are available, which can be our weapon to change the course of this pandemic. Therefore, it is important to invest in health systems and in basic and innovative science to find diagnostic and therapeutic solutions. It is heart-warming to see the collaborative efforts amongst all sectors involved in the global healthtech ecosystem. We only hope that these insights will still resonate after this pandemic is behind us.
References
Jiang, S., He, Y. & Liu, S. SARS vaccine development. Emerg. Infect. Dis. 11, 1016–1020 (2005).
Leung, D. T. et al. Antibody response of patients with severe acute respiratory syndrome (SARS) targets the viral nucleocapsid. J. Infect. Dis. 190, 379–386 (2004).
He, Y., Zhou, Y., Siddiqui, P., Niu, J. & Jiang, S. Identification of immunodominant epitopes on the membrane protein of the severe acute respiratory syndrome-associated coronavirus. J. Clin. Microbiol. 43, 3718–3726 (2005).
Schoeman, D. & Fielding, B. C. Coronavirus envelope protein: current knowledge. Virol. J. 16, 69 (2019).
Yu, F., Du, L., Ojcius, D. M., Pan, C. & Jiang, S. Measures for diagnosing and treating infections by a novel coronavirus responsible for a pneumonia outbreak originating in Wuhan, China. Microbes Infect. 22, 74–79 (2020).
Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Baez-Santos, Y. M., St John, S. E. & Mesecar, A. D. The SARS-coronavirus papain-like protease: structure, function and inhibition by designed antiviral compounds. Antiviral Res. 115, 21–38 (2015).
Ziebuhr, J., Snijder, E. J. & Gorbalenya, A. E. Virus-encoded proteinases and proteolytic processing in the Nidovirales. J. Gen. Virol. 81, 853–879 (2000).
Lauer, S. A. et al. The incubation period of coronavirus disease 2019 (covid-19) from publicly reported confirmed cases: Estimation and application. Ann. Intern. Med. 172, 577–582 (2020).
Li, Q. et al. Early transmission dynamics in Wuhan, China, of bovel Coronavirus-infected pneumonia. N. Engl. J. Med. 382, 1199–1207 (2020).
Bai, Y. et al. Presumed asymptomatic carrier transmission of COVID-19. JAMA 323, 1406–1407 (2020).
Kluge, H. Older people are at highest risk from COVID-19, but all must act to prevent community spread. World Health Organization for Europe http://www.euro.who.int/en/health-topics/health-emergencies/coronavirus-covid-19/statements/statement-older-people-are-at-highest-risk-from-covid-19,-but-all-must-act-to-prevent-community-spread (2020).
Huang, C. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506 (2020).
Baig, A. M., Khaleeq, A., Ali, U. & Syeda, H. Evidence of the COVID-19 virus targeting the CNS: Tissue distribution, host-virus interaction, and proposed neurotropic mechanisms. ACS Chem. Neurosci. 11, 995–998 (2020).
Yang, X. et al. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir. Med. 8, 475–481 (2020).
Wadman, M. C.-F., J, Kaiser, J & Matacic, C. How does coronavirus kill? Clinicians trace a ferocious rampage through the body, from brain to toes. Science https://www.sciencemag.org/news/2020/04/how-does-coronavirus-kill-clinicians-trace-ferocious-rampage-through-body-brain-toes (2020).
Shi, S. et al. Association of cardiac injury with mortality in hospitalized patients with COVID-19 in Wuhan, China. JAMA Cardiol. 25, e200950 (2020).
Oxley, T. J. et al. Large-vessel stroke as a presenting feature of Covid-19 in the young. N. Engl. J. Med. 382, e60 (2020).
Gilead announces approval of Veklury® (remdesivir) in Japan for patients with severe COVID-19. GILEAD https://www.gilead.com/news-and-press/press-room/press-releases/2020/5/gilead-announces-approval-of-veklury-remdesivir-in-japan-for-patients-with-severe-covid19 (2020).
Coronavirus (COVID-19) update: FDA issues emergency use authorization for potential COVID-19tTreatment. FDA https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-issues-emergency-use-authorization-potential-covid-19-treatment (2020).
Drugs@FDA: FDA-approved drugs. FDA https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&varApplNo=009768 (2020).
Recommendations for investigational COVID-19 convalescent plasma. FDA https://www.fda.gov/vaccines-blood-biologics/investigational-new-drug-ind-or-device-exemption-ide-process-cber/investigational-covid-19-convalescent-plasma-emergency-inds (2020).
Shen, C. et al. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA 323, 1582–1589 (2020).
A study of LY3819253 (LY-CoV555) in participants hospitalized for COVID-19. NIH https://clinicaltrials.gov/ct2/show/NCT04411628?term=LY-CoV555&cond=COVID-19&draw=2&rank=1 (2020).
Lu, R. et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395, 565–574 (2020).
Ahmed, S. F., Quadeer, A. A. & McKay, M. R. Preliminary identification of potential vaccine targets for the COVID-19 coronavirus (SARS-CoV-2) based on SARS-CoV immunological studies. Viruses 12, 254 (2020).
Xu, J. et al. Systematic comparison of two animal-to-human transmitted human coronaviruses: SARS-CoV-2 and SARS-CoV. Viruses 12, 244 (2020).
Kliger, Y. & Levanon, E. Y. Cloaked similarity between HIV-1 and SARS-CoV suggests an anti-SARS strategy. BMC Microbiol. 3, 20 (2003).
Martinez, M. A. Compounds with therapeutic potential against novel respiratory 2019 coronavirus. Antimicrob. Agents Chemother. 64, e00399–00320 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Arabi, Y. M. et al. Corticosteroid therapy for critically ill patients with Middle East Respiratory Syndrome. Am. J. Respir. Crit. Care Med. 197, 757–767 (2018).
Hui, D. S. Systemic corticosteroid therapy may delay viral clearance in patients with Middle East Respiratory Syndrome coronavirus infection. Am. J. Respir. Crit. Care Med. 197, 700–701 (2018).
To, K. K. et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect. Dis. 20, 565–574 (2020).
Fleischmann, R. et al. Long-term safety of sarilumab in rheumatoid arthritis: an integrated analysis with up to 7 years’ follow-up. Rheumatology (Oxford) 59, 292–302 (2020).
Hu, W., Yen, Y. T., Singh, S., Kao, C. L. & Wu-Hsieh, B. A. SARS-CoV regulates immune function-related gene expression in human monocytic cells. Viral Immunol. 25, 277–288 (2012).
Prompetchara, E., Ketloy, C. & Palaga, T. Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic. Asian Pac. J. Allergy Immunol. 38, 1–9 (2020).
Leiva-Juarez, M. M. et al. Combined aerosolized Toll-like receptor ligands are an effective therapeutic agent against influenza pneumonia when co-administered with oseltamivir. Eur. J. Pharmacol. 818, 191–197 (2018).
Spruth, M. et al. A double-inactivated whole virus candidate SARS coronavirus vaccine stimulates neutralising and protective antibody responses. Vaccine 24, 652–661 (2006).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines - a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Yang, Z. Y. et al. A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 428, 561–564 (2004).
Shim, B. S. et al. Intranasal immunization with plasmid DNA encoding spike protein of SARS-coronavirus/polyethylenimine nanoparticles elicits antigen-specific humoral and cellular immune responses. BMC Immunol. 11, 65 (2010).
Pitcovski, J. et al. Development and large-scale use of recombinant VP2 vaccine for the prevention of infectious bursal disease of chickens. Vaccine 21, 4736–4743 (2003).
Conniot, J. et al. Immunization with mannosylated nanovaccines and inhibition of the immune-suppressing microenvironment sensitizes melanoma to immune checkpoint modulators. Nat. Nanotechnol. 14, 891–901 (2019).
Lauer, K. B., Borrow, R. & Blanchard, T. J. Multivalent and multipathogen viral vector vaccines. Clin. Vaccine Immunol. 24, e00298–16 (2017).
Draper, S. J. & Heeney, J. L. Viruses as vaccine vectors for infectious diseases and cancer. Nat. Rev. Microbiol. 8, 62–73 (2010).
Tseng, C. T. et al. Immunization with SARS coronavirus vaccines leads to pulmonary immunopathology on challenge with the SARS virus. PLoS One 7, e35421 (2012).
Savarino, A., Boelaert, J. R., Cassone, A., Majori, G. & Cauda, R. Effects of chloroquine on viral infections: an old drug against today’s diseases? Lancet Infect. Dis. 3, 722–727 (2003).
Vincent, M. J. et al. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol. J. 2, 69 (2005).
Alunno, A., Padjen, I., Fanouriakis, A. & Boumpas, D. T. Pathogenic and therapeutic relevance of JAK/STAT signaling in Systemic Lupus Erythematosus: Integration of distinct inflammatory pathways and the prospect of their inhibition with an oral agent. Cells 8, 898 (2019).
Hu, T., Frieman, M. & Wolfram, J. Insights from nanomedicine into chloroquine efficacy against COVID-19. Nat. Nanotechnol. 15, 247–249 (2020).
Bao, L. et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature https://doi.org/10.1038/s41586-020-2312-y (2020).
Devaux, C. A., Rolain, J. M., Colson, P. & Raoult, D. New insights on the antiviral effects of chloroquine against coronavirus: what to expect for COVID-19? Int. J. Antimicrob. Agents 55, 105938 (2020).
Molina, J. D. et al. No evidence of rapid antiviral clearance or clinical benefit with the combination of hydroxychloroquine and azithromycin in patients with severe COVID-19 infection. Médecine et Maladies Infectieuses 50, 384 (2020).
Boriskin, Y. S., Leneva, I. A., Pecheur, E. I. & Polyak, S. J. Arbidol: a broad-spectrum antiviral compound that blocks viral fusion. Curr. Med. Chem. 15, 997–1005 (2008).
Liu, Q. et al. Antiviral and anti-inflammatory activity of arbidol hydrochloride in influenza A (H1N1) virus infection. Acta Pharmacol. Sin. 34, 1075–1083 (2013).
Shi, L. et al. Antiviral activity of arbidol against influenza A virus, respiratory syncytial virus, rhinovirus, coxsackie virus and adenovirus in vitro and in vivo. Arch. Virol. 152, 1447–1455 (2007).
Kadam, R. U. & Wilson, I. A. Structural basis of influenza virus fusion inhibition by the antiviral drug Arbidol. Proc. Natl Acad. Sci. USA 114, 206–214 (2017).
Sheahan, T. P. et al. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat. Commun. 11, 222 (2020).
Harrison, C. Coronavirus puts drug repurposing on the fast track. Nat. Biotechnol. 38, 379–381 (2020).
Fried, M. W. et al. Peginterferon alfa-2a plus ribavirin for chronic hepatitis C virus infection. N. Engl. J. Med. 347, 975–982 (2002).
Te, H. S., Randall, G. & Jensen, D. M. Mechanism of action of ribavirin in the treatment of chronic hepatitis C. Gastroenterol. Hepatol. (N Y) 3, 218–225 (2007).
Agostini, M. L. et al. Coronavirus susceptibility to the antiviral remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. mBio 9, e00221–00218 (2018).
Beigel, J. et al. Remdesivir for the treatment of Covid-19 — preliminary report. N. Engl. J. Med. 22, NEJMoa2007764 (2020).
Maxmen, A. More than 80 clinical trials launch to test coronavirus treatments. Nature 578, 347–348 (2020).
Petitprez, F. et al. B cells are associated with survival and immunotherapy response in sarcoma. Nature 577, 556–560 (2020).
Wang, D. et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 323, 1061–1069 (2020).
Wang, R. R. et al. Azvudine, a novel nucleoside reverse transcriptase inhibitor showed good drug combination features and better inhibition on drug-resistant strains than lamivudine in vitro. PLoS One 9, e105617 (2014).
Richardson, P. et al. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet 395, e30–e31 (2020).
Low-cost dexamethasone reduces death by up to one third in hospitalised patients with severe respiratory complications of COVID-19. RECOVERY https://www.recoverytrial.net/news/low-cost-dexamethasone-reduces-death-by-up-to-one-third-in-hospitalised-patients-with-severe-respiratory-complications-of-covid-19 (2020).
Reynolds, H. R. et al. Renin-angiotensin-aldosterone system inhibitors and risk of Covid-19. N. Engl. J. Med. 383, 2441–2448 (2020).
Kruse, R. L. Therapeutic strategies in an outbreak scenario to treat the novel coronavirus originating in Wuhan, China. F1000Res 9, 72 (2020).
Malik, Y. S. et al. Emerging novel coronavirus (2019-nCoV)-current scenario, evolutionary perspective based on genome analysis and recent developments. Vet Q 40, 68–76 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020).
Chen, W.-H., Strych, U., Hotez, P. J. & Bottazzi, M. E. The SARS-CoV-2 vaccine pipeline: An overview. Current Tropical Medicine Reports 3, 1–4 (2020).
Hikmet, F., Méar, L, Uhlén, M & Lindskog, C. The protein expression profile of ACE2 in human tissues. Preprint at https://www.biorxiv.org/content/10.1101/2020.03.31.016048v1 (2020).
Li, Y. C., Bai, W. Z. & Hashikawa, T. The neuroinvasive potential of SARS-CoV2 may play a role in the respiratory failure of COVID-19 patients. J. Med. Virol. 92, 55–555 (2020).
Xia, H. & Lazartigues, E. Angiotensin-converting enzyme 2 in the brain: properties and future directions. J. Neurochem. 107, 1482–1494 (2008).
Kliger, Y., Levanon, E. Y. & Gerber, D. From genome to antivirals: SARS as a test tube. Drug Discov. Today 10, 345–352 (2005).
Jeffers, S. A. et al. CD209L (L-SIGN) is a receptor for severe acute respiratory syndrome coronavirus. Proc. Natl Acad. Sci. USA 101, 15748–15753 (2004).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292 (2020).
Tai, W. et al. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 17, 613–620 (2020).
Letko, M., Marzi, A. & Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol 5, 562–569 (2020).
Yuan, Y. et al. Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat. Commun. 8, 15092 (2017).
Kam, Y. W. et al. Cleavage of the SARS coronavirus spike glycoprotein by airway proteases enhances virus entry into human bronchial epithelial cells in vitro. PLoS One 4, e7870 (2009).
Matsuyama, S., Ujike, M., Morikawa, S., Tashiro, M. & Taguchi, F. Protease-mediated enhancement of severe acute respiratory syndrome coronavirus infection. Proc. Natl Acad. Sci. USA 102, 12543–12547 (2005).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280–e278 (2020).
Simmons, G. et al. Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc. Natl Acad. Sci. USA 102, 11876–11881 (2005).
Ahlquist, P. & RNA-dependent, R. N. A. polymerases, viruses, and RNA silencing. Science 296, 1270–1273 (2002).
Li, X., Geng, M., Peng, Y., Meng, L. & Lu, S. Molecular immune pathogenesis and diagnosis of COVID-19. J. Pharm. Anal. 10, 102–108 (2020).
Li, G. et al. Coronavirus infections and immune responses. J. Med. Virol. 92, 424–432 (2020).
de Wit, E., van Doremalen, N., Falzarano, D. & Munster, V. J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 14, 523–534 (2016).
Kindler, E., Thiel, V. & Weber, F. Interaction of SARS and MERS coronaviruses with the antiviral interferon response. Adv. Virus Res. 96, 219–243 (2016).
Menachery, V. D. et al. MERS-CoV and H5N1 influenza virus antagonize antigen presentation by altering the epigenetic landscape. Proc. Natl Acad. Sci. USA 115, E1012–E1021 (2018).
Zheng, H. Y. et al. Elevated exhaustion levels and reduced functional diversity of T cells in peripheral blood may predict severe progression in COVID-19 patients. Cell. Mol. Immunol. 17, 541–543 (2020).
Graham, B. S. Advances in antiviral vaccine development. Immunol. Rev. 255, 230–242 (2013).
Bao, L. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. Preprint at https://www.biorxiv.org/content/10.1101/2020.03.13.990226v1 (2020).
Thevarajan, I. et al. Breadth of concomitant immune responses prior to patient recovery: a case report of non-severe COVID-19. Nat. Med. 26, 453–455 (2020).
Zhao, J.et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin. Infect. Dis. ciaa344 (2020).
Shen, C. Treatment of 5 critically ill Patients with COVID-19 with convalescent plasma. JAMA 323, 1582–1589 (2020).
Grifoni, A. W. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 182, 1–9 (2020).
Letko, M., Marzi, A. & Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol. 5, 562–569 (2020).
Reed, S. G., Orr, M. T. & Fox, C. B. Key roles of adjuvants in modern vaccines. Nat. Med. 19, 1597–1608 (2013).
Luo, F. et al. Evaluation of antibody-dependent enhancement of SARS-CoV infection in rhesus macaques immunized with an inactivated SARS-CoV vaccine. Virol. Sin. 33, 201–204 (2018).
Jia, W. et al. Single intranasal immunization with chimpanzee adenovirus-based vaccine induces sustained and protective immunity against MERS-CoV infection. Emerg. Microbes Infect. 8, 760–772 (2019).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines — a new era in vaccinology. Nat. Rev. Drug. Discov. 17, 261–279 (2018).
Tiram, G., Scomparin, A., Ofek, P. & Satchi-Fainaro, R. Interfering cancer with polymeric siRNA nanomedicines. J. Biomed. Nanotechnol. 10, 50–66 (2014).
Panyam, J. & Labhasetwar, V. Biodegradable nanoparticles for drug and gene delivery to cells and tissue. Adv. Drug Deliv. Rev. 55, 329–347 (2003).
Veiseh, O., Tang, B. C., Whitehead, K. A., Anderson, D. G. & Langer, R. Managing diabetes with nanomedicine: challenges and opportunities. Nat. Rev. Drug Discov. 14, 45–57 (2015).
Mograo, J., da Costa, C. A., Gaspar, R. & Florindo, H. F. Modulation of dendritic cells by nanotechnology-based immunotherapeutic strategies. J. Biomed. Nanotechnol. 12, 405–434 (2016).
Sainz, V. et al. Regulatory aspects on nanomedicines. Biochem. Biophys. Res. Commun. 468, 504–510 (2015).
Shen, Y., Hao, T., Ou, S., Hu, C. & Chen, L. Applications and perspectives of nanomaterials in novel vaccine development. Medchemcomm 9, 226–238 (2018).
Gregory, A. E., Titball, R. & Williamson, D. Vaccine delivery using nanoparticles. Front. Cell. Infect. Microbiol. 3, 13 (2013).
Qi, F., Wu, J., Li, H. & Ma, G. Recent research and development of PLGA/PLA microspheres/nanoparticles: A review in scientific and industrial aspects. Front. Chem. Sci. Eng. 13, 14–27 (2019).
Chang, C. K., Lo, S. C., Wang, Y. S. & Hou, M. H. Recent insights into the development of therapeutics against coronavirus diseases by targeting N protein. Drug Discov. Today 21, 562–572 (2016).
Van Hoeven, N. et al. A formulated TLR7/8 agonist is a flexible, highly potent and effective adjuvant for pandemic influenza vaccines. Sci Rep 7, 46426 (2017).
Kumar, A., Zhang, J. & Yu, F. S. Toll-like receptor 3 agonist poly(I:C)-induced antiviral response in human corneal epithelial cells. Immunology 117, 11–21 (2006).
Reed, S. G., Bertholet, S., Coler, R. N. & Friede, M. New horizons in adjuvants for vaccine development. Trends Immunol. 30, 23–32 (2009).
Coler, R. N. et al. Development and characterization of synthetic glucopyranosyl lipid adjuvant system as a vaccine adjuvant. PLoS One 6, e16333 (2011).
Azizi, A. et al. A combined nucleocapsid vaccine induces vigorous SARS-CD8+ T-cell immune responses. Genet. Vaccines Ther. 3, 7 (2005).
Zupancic, E. et al. Rational design of nanoparticles towards targeting antigen-presenting cells and improved T cell priming. J. Control. Release 258, 182–195 (2017).
Linehan, M. M. et al. A minimal RNA ligand for potent RIG-I activation in living mice. Sci. Adv. 4, e1701854 (2018).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Zhang, C., Maruggi, G., Shan, H. & Li, J. Advances in mRNA vaccines for infectious diseases. Front. Immunol. 10, 594 (2019).
Iavarone, C., O’Hagan D, T., Yu, D., Delahaye, N. F. & Ulmer, J. B. Mechanism of action of mRNA-based vaccines. Expert. Rev. Vaccines 16, 871–881 (2017).
Alberer, M. et al. Safety and immunogenicity of a mRNA rabies vaccine in healthy adults: an open-label, non-randomised, prospective, first-in-human phase 1 clinical trial. Lancet 390, 1511–1520 (2017).
Corbett K. S. et al. SARS-CoV-2 mRNA vaccine development enabled by prototype pathogen 1 preparedness. bioRxiv https://doi.org/10.1101/2020.06.11.145920 (2020)
Moderna ships mRNA vaccine against novel coronavirus (mRNA-1273) for phase 1 study. Moderna https://investors.modernatx.com/news-releases/news-release-details/moderna-ships-mrna-vaccine-against-novel-coronavirus-mrna-1273 (2020).
Liu, L. et al. Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight 4, e123158 (2019).
Zhang, L. et al. Antibody responses against SARS coronavirus are correlated with disease outcome of infected individuals. J. Med. Virol. 78, 1–8 (2006).
Clay, C. et al. Primary severe acute respiratory syndrome coronavirus infection limits replication but not lung inflammation upon homologous rechallenge. J. Virol. 86, 4234–4244 (2012).
Zhao, J., Zhao, J. & Perlman, S. T cell responses are required for protection from clinical disease and for virus clearance in severe acute respiratory syndrome coronavirus-infected mice. J. Virol. 84, 9318–9325 (2010).
Zhao, J. et al. Airway memory CD4(+) T cells mediate protective immunity against emerging respiratory coronaviruses. Immunity 44, 1379–1391 (2016).
Wu, F. et al. Neutralizing antibody responses to SARS-CoV-2 in a COVID-19 recovered patient cohort and their implications. Preprint at https://www.medrxiv.org/content/10.1101/2020.03.30.20047365v2 (2020).
Du, L. et al. Priming with rAAV encoding RBD of SARS-CoV S protein and boosting with RBD-specific peptides for T cell epitopes elevated humoral and cellular immune responses against SARS-CoV infection. Vaccine 26, 1644–1651 (2008).
Fett, C., DeDiego, M. L., Regla-Nava, J. A., Enjuanes, L. & Perlman, S. Complete protection against severe acute respiratory syndrome coronavirus-mediated lethal respiratory disease in aged mice by immunization with a mouse-adapted virus lacking E protein. J. Virol. 87, 6551–6559 (2013).
Yang, X. H. et al. Mice transgenic for human angiotensin-converting enzyme 2 provide a model for SARS coronavirus infection. Comp Med 57, 450–459 (2007).
Kuba, K. et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat. Med. 11, 875–879 (2005).
Imai, Y. et al. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature 436, 112–116 (2005).
Graham, R. L. et al. A live, impaired-fidelity coronavirus vaccine protects in an aged, immunocompromised mouse model of lethal disease. Nat. Med. 18, 1820–1826 (2012).
Frieman, M. B. et al. SARS-CoV pathogenesis is regulated by a STAT1 dependent but a type I, II and III interferon receptor independent mechanism. PLoS Pathog. 6, e1000849 (2010).
Roberts, A. et al. Animal models and vaccines for SARS-CoV infection. Virus Res. 133, 20–32 (2008).
Shultz, L. D. et al. Generation of functional human T-cell subsets with HLA-restricted immune responses in HLA class I expressing NOD/SCID/IL2r gamma(null) humanized mice. Proc. Natl Acad. Sci. USA 107, 13022–13027 (2010).
Jiang, R. L. et al. Pathogenesis of SARS-CoV-2 in transgenic mice expressing human angiotensin-converting enzyme 2. Cell 182, 1–9 (2020).
Netland, J., Meyerholz, D. K., Moore, S., Cassell, M. & Perlman, S. Severe acute respiratory syndrome coronavirus infection causes neuronal death in the absence of encephalitis in mice transgenic for human ACE2. J. Virol. 82, 7264–7275 (2008).
McCray, P. B. Jr. et al. Lethal infection of K18-hACE2 mice infected with severe acute respiratory syndrome coronavirus. J. Virol. 81, 813–821 (2007).
Spengler, J. R. et al. Ebola virus replication and disease without immunopathology in mice expressing transgenes to support human myeloid and lymphoid cell engraftment. J. Infect. Dis. 214, S308–S318 (2016).
Shultz, L. D., Brehm, M. A., Garcia-Martinez, J. V. & Greiner, D. L. Humanized mice for immune system investigation: progress, promise and challenges. Nat. Rev. Immunol. 12, 786–798 (2012).
Brehm, M. A., Wiles, M. V., Greiner, D. L. & Shultz, L. D. Generation of improved humanized mouse models for human infectious diseases. J. Immunol. Methods 410, 3–17 (2014).
Braciale, T. J. & Hahn, Y. S. Immunity to viruses. Immunol. Rev. 255, 5–12 (2013).
Barnard, D. L. et al. Evaluation of immunomodulators, interferons and known in vitro SARS-coV inhibitors for inhibition of SARS-coV replication in BALB/c mice. Antivir. Chem. Chemother. 17, 275–284 (2006).
Arabi, Y. M. et al. Corticosteroid therapy for critically ill patients with Middle East Respiratory Syndrome. Am. J. Respir. Crit. Care Med 197, 757–767 (2018).
Auyeung, T. W. et al. The use of corticosteroid as treatment in SARS was associated with adverse outcomes: a retrospective cohort study. J. Infect. 51, 98–102 (2005).
Russell, C. D., Millar, J. E. & Baillie, J. K. Clinical evidence does not support corticosteroid treatment for 2019-nCoV lung injury. Lancet 395, 473–475 (2020).
Li, G. & De Clercq, E. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat. Rev. Drug Discov. 19, 149–150 (2020).
Zhou, F. et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395, 1054–1062 (2020).
Hotchkiss, R. S., Coopersmith, C. M., McDunn, J. E. & Ferguson, T. A. The sepsis seesaw: tilting toward immunosuppression. Nat. Med. 15, 496–497 (2009).
Chousterman, B. G., Swirski, F. K. & Weber, G. F. Cytokine storm and sepsis disease pathogenesis. Semi.n Immunopathol. 39, 517–528 (2017).
Cavaillon, J. M. & Adib-Conquy, M. Immune status in sepsis: the bug, the site of infection and the severity can make the difference. Crit. Care 14, 167 (2010).
Otto, G. P. et al. The late phase of sepsis is characterized by an increased microbiological burden and death rate. Crit. Care 15, R183 (2011).
Zhang, X. et al. Structural and functional analysis of the costimulatory receptor programmed death-1. Immunity 20, 337–347 (2004).
Patil, N. K., Guo, Y., Luan, L. & Sherwood, E. R. Targeting immune cell checkpoints during sepsis. Int. J. Mol. Sci. 18, 2413 (2017).
Mayoux, M. et al. Dendritic cells dictate responses to PD-L1 blockade cancer immunotherapy. Sci. Transl. Med. 12, eaav7431 (2020).
Naran, K., Nundalall, T., Chetty, S. & Barth, S. Principles of immunotherapy: implications for treatment strategies in cancer and infectious diseases. Front. Microbiol. 9, 3158 (2018).
Zhao, X., Nicholls, J. M. & Chen, Y. G. Severe acute respiratory syndrome-associated coronavirus nucleocapsid protein interacts with Smad3 and modulates transforming growth factor-beta signaling. J. Biol. Chem. 283, 3272–3280 (2008).
Stahnke, T. et al. Suppression of TGF-beta pathway by pirfenidone decreases extracellular matrix deposition in ocular fibroblasts in vitro. PLoS One 12, e0172592 (2017).
Wrzesinski, S. H., Wan, Y. Y. & Flavell, R. A. Transforming growth factor-beta and the immune response: implications for anticancer therapy. Clin. Cancer Res. 13, 5262–5270 (2007).
Sen, G. C. Viruses and interferons. Annu. Rev. Microbiol. 55, 255–281 (2001).
Zhou, J. H. et al. Type III interferons in viral infection and antiviral immunity. Cell. Physiol. Biochem. 51, 173–185 (2018).
Honda, K. & Taniguchi, T. IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat. Rev. Immunol. 6, 644–658 (2006).
Schnare, M. et al. Toll-like receptors control activation of adaptive immune responses. Nat. Immunol. 2, 947–950 (2001).
White, M. R., Tecle, T., Crouch, E. C. & Hartshorn, K. L. Impact of neutrophils on antiviral activity of human bronchoalveolar lavage fluid. Am. J. Physiol. Lung Cell. Mol. Physiol. 293, L1293–1299 (2007).
Wohlford-Lenane, C. L. et al. Rhesus theta-defensin prevents death in a mouse model of severe acute respiratory syndrome coronavirus pulmonary disease. J. Virol. 83, 11385–11390 (2009).
Dhama, K. et al. COVID-19, an emerging coronavirus infection: advances and prospects in designing and developing vaccines, immunotherapeutics, and therapeutics. Hum. Vaccin. Immunother. 18, 1–7 (2020).
Ji, W., Wang, W., Zhao, X., Zai, J. & Li, X. Cross-species transmission of the newly identified coronavirus 2019-nCoV. J. Med. Virol. 92, 433–440 (2020).
Widjaja, I. et al. Towards a solution to MERS: protective human monoclonal antibodies targeting different domains and functions of the MERS-coronavirus spike glycoprotein. Emerg. Microbes Infect. 8, 516–530 (2019).
Kumar, V., Jung, Y.-S. & Liang, P.-H. Anti-SARS coronavirus agents: a patent review (2008 – present). Expert Opin. Ther. Pat. 23, 1337–1348 (2013).
Lu, H. Drug treatment options for the 2019-new coronavirus (2019-nCoV). BioSci. Trends 14, 69–71 (2020).
Pang, J. et al. Potential rapid diagnostics, vaccine and therapeutics for 2019 novel coronavirus (2019-nCoV): A systematic review. J. Clin. Med. 9, 623 (2020).
Marston, H. D., Paules, C. I. & Fauci, A. S. Monoclonal Antibodies for Emerging Infectious Diseases - Borrowing from History. N. Engl J. Med. 378, 1469–1472 (2018).
Zhang, C., Maruggi, G., Shan, H. & Li, J. Advances in mRNA Vaccines for Infectious Diseases. Front. Immunol. 10, 594–594 (2019).
Moderna announces positive interim phase 1 data for its mRNA vaccine (mRNA-1273) against novel coronavirus. Moderna https://investors.modernatx.com/news-releases/news-release-details/moderna-announces-positive-interim-phase-1-data-its-mrna-vaccine (2020).
Leiva-Juarez, M. M. et al. Combined aerosolized Toll-like receptor ligands are an effective therapeutic agent against influenza pneumonia when co-administered with oseltamivir. Eur. J. Pharmacol. 818, 191–197 (2018).
Thompson, J. M. & Iwasaki, A. Toll-like receptors regulation of viral infection and disease. Adv. Drug Deliv. Rev. 60, 786–794 (2008).
Tarhini, A. A., Gogas, H. & Kirkwood, J. M. IFN-α in the treatment of melanoma. J. Immunol. 189, 3789–3793 (2012).
Pestka, S. The interferons: 50 years after their discovery, there is much more to learn. J. Biol. Chem. 282, 20047–20051 (2007).
Saeedi, P., Halabian, R. & Imani Fooladi, A. A. A revealing review of mesenchymal stem cells therapy, clinical perspectives and Modification strategies. Stem Cell Investig. 6, 34–34 (2019).
WHO Director-General’s opening remarks at the media briefing on COVID-19 - 27 March 2020. World Health Organization https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19-27-march-2020 (2020).
“Solidarity” clinical trial for COVID-19 treatments. World Health Organization (https://www.who.int/emergencies/diseases/novel-coronavirus-2019/global-research-on-novel-coronavirus-2019-ncov/solidarity-clinical-trial-for-covid-19-treatments (2020).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nat. Rev. Drug Discov. 8, 129–138 (2009).
Kim, A. et al. Intracellular delivery of charge-converted monoclonal antibodies by combinatorial design of block/homo polyion complex micelles. Biomacromolecules 17, 446–453 (2016).
Chen, M., Ouyang, H., Zhou, S., Li, J. & Ye, Y. PLGA-nanoparticle mediated delivery of anti-OX40 monoclonal antibody enhances anti-tumor cytotoxic T cell responses. Cell. Immunol. 287, 91–99 (2014).
Lei, C. et al. Local release of highly loaded antibodies from functionalized nanoporous support for cancer immunotherapy. J. Am. Chem. Soc. 132, 6906–6907 (2010).
ten Hagen, T. L., Seynhaeve, A. L., van Tiel, S. T., Ruiter, D. J. & Eggermont, A. M. Pegylated liposomal tumor necrosis factor-alpha results in reduced toxicity and synergistic antitumor activity after systemic administration in combination with liposomal doxorubicin (Doxil) in soft tissue sarcoma-bearing rats. Int. J. Cancer 97, 115–120 (2002).
Neelapu, S. S. et al. A novel proteoliposomal vaccine induces antitumor immunity against follicular lymphoma. Blood 109, 5160–5163 (2007).
Hu, T. Y., Frieman, M. & Wolfram, J. Insights from nanomedicine into chloroquine efficacy against COVID-19. Nat. Nanotechnol. 15, 247–249 (2020).
Hadadi, A., Mortezazadeh, M., Kolahdouzan, K. & Alavian, G. Does recombinant human Erythropoietin administration in critically ill COVID-19 patients have miraculous therapeutic effects? J. Med. Virol. 92, 915–918 (2020).
Zhang, X. & Dong, S. Protective effects of erythropoietin towards acute lung injuries in rats with sepsis and its related mechanisms. Ann. Clin. Lab. Sci. 49, 257–264 (2019).
Zhang, G. X. et al. Protective effect of erythropoietin against lipopolysaccharide induced inflammation and mitochondrial damage in liver. J. Biol. Regul. Homeos. Agents 32, 199–206 (2018).
Ito, T., Hamazaki, Y., Takaori-Kondo, A. & Minato, N. Bone Marrow Endothelial Cells Induce Immature and Mature B Cell Egress in Response to Erythropoietin. Cell. Struct. Funct. 42, 149–157 (2017).
Verdoni, L. et al. An outbreak of severe Kawasaki-like disease at the Italian epicentre of the SARS-CoV-2 epidemic: an observational cohort study. Lancet 395, 1771–1778 (2020).
Viner, R. M. & Whittaker, E. Kawasaki-like disease: emerging complication during the COVID-19 pandemic. Lancet 395, 1741–1743 (2020).
Couzin-Frankel, J. Doctors race to understand rare inflammatory condition associated with coronavirus in young people. Science https://www.sciencemag.org/news/2020/05/doctors-race-understand-rare-inflammatory-condition-associated-coronavirus-young-people (2020).
Guidance on the Management of Clinical Trials during the COVID-19 (Coronavirus) pandemic (European Medicines Agency, 2020); https://ec.europa.eu/health/sites/health/files/files/eudralex/vol-10/guidanceclinicaltrials_covid19_en.pdf
Andre, F. E. et al. Vaccination greatly reduces disease, disability, death and inequity worldwide. World Health Organization https://www.who.int/bulletin/volumes/86/2/07-040089/en/ (2008).
Communities vulnerable without immunization against infectious diseases. World Health Organization https://www.who.int/malaysia/news/detail/23-04-2020-communities-vulnerable-without-immunization-against-infectious-diseases (2020).
Acknowledgements
R.S.-F. and H.F.F. thank the following funding agencies for their generous support: The MultiNano@MBM project supported by The Israeli Ministry of Health and The Fundação para a Ciência e Tecnologia-Ministério da Ciência, Tecnologia e Ensino Superior (FCT-MCTES) under the framework of EuroNanoMed-II (ENMed/0051/2016); “La Caixa” Foundation under the framework of the Healthcare Research call 2019 (LCF/PR/HR19/52160021; NanoPanther), CaixaImpulse (CF01-00014; CoVax). In addition, R.S.-F. thanks the generous financial support from the European Research Council (ERC) Proof of Concept (PoC) grant (862580; 3DCanPredict) and ERC Advanced grant (835227; 3DBrainStrom), The Israel Science Foundation (1969/18), The Melanoma Research Alliance-Established Investigator Award (615808), the Israel Cancer Research Fund (ICRF) Professorship award (PROF-18-682), the Morris Kahn Foundation and The NOFAR incentive program on COVID-19 by the Israel Innovation Authority supported by Merck Group. D.V. is grateful to The Rothschild Foundation (IL) for funding her PhD scholarship. B.C. is supported by the FCT-MCTES (PhD Fellowship SFRH/BD/131969/2017).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
R.S.-F. is a Director at the Board of Teva Pharmaceutical Industries Ltd.
Additional information
Peer review information Nature Nanotechnology thanks Erik De Clercq and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and refs. 1–22.
Rights and permissions
About this article
Cite this article
Florindo, H.F., Kleiner, R., Vaskovich-Koubi, D. et al. Immune-mediated approaches against COVID-19. Nat. Nanotechnol. 15, 630–645 (2020). https://doi.org/10.1038/s41565-020-0732-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-020-0732-3
This article is cited by
-
Surveillance of SARS-CoV-2 RNA in wastewater matrix: a review
Environmental Monitoring and Assessment (2024)
-
Development of a Single-Chain Fragment Variable that Binds to the SARS-CoV-2 Spike Protein Produced by Genetically Modified Lactic Acid Bacteria
Molecular Biotechnology (2024)
-
Adipose-derived, autologous mesenchymal stem cell therapy for patients with post-COVID-19 syndrome: an intermediate-size expanded access program
Stem Cell Research & Therapy (2023)
-
Amyloidogenic proteins in the SARS-CoV and SARS-CoV-2 proteomes
Nature Communications (2023)
-
Immunoregulatory nanomedicine for respiratory infections
Nature Reviews Bioengineering (2023)