Abstract
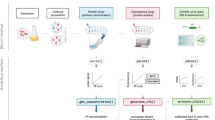
Quantitative models of Förster resonance energy transfer (FRET)—pioneered by Förster—define our understanding of FRET and underpin its widespread use. However, multicolour FRET (mFRET), which arises between multiple, stochastically distributed fluorophores, lacks a mechanistic model and remains intractable. mFRET notably arises in fluorescently barcoded microparticles, resulting in a complex, non-orthogonal fluorescence response that impedes their encoding and decoding. Here, we introduce an ensemble mFRET (emFRET) model, and apply it to guide barcoding into regimes with extreme FRET. We further introduce a facile, proportional multicolour labelling method using oligonucleotides as homogeneous linkers. A total of 580 barcodes were rapidly designed and validated using four dyes—with FRET efficiencies reaching 76%—and used for multiplexed immunoassays with cytometric readout and fully automated decoding. The emFRET model helps to expand the barcoding capacity of barcoded microparticles using common organic dyes and will benefit other applications subject to stochastic mFRET.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Nolan, J. P. & Sklar, La Suspension array technology: evolution of the flat-array paradigm. Trends Biotechnol. 20, 9–12 (2002).
Wilson, R., Cossins, A. R. & Spiller, D. G. Encoded microcarriers for high-throughput multiplexed detection. Angew. Chem. Int. Ed. 45, 6104–6117 (2006).
Fulton, R. J., McDade, R. L., Smith, P. L., Kienker, L. J. & Kettman, J. R. Advanced multiplex analysis with the FlowMetrix system. Clin. Chem. 43, 1749–1756 (1997).
Wu, W. et al. Antibody array analysis with label-based detection and resolution of protein size. Mol. Cell. Proteomics 8, 245–257 (2009).
Wang, L. & Tan, W. Multicolor FRET silica nanoparticles by single wavelength excitation. Nano. Lett. 6, 84–88 (2006).
Vaidya, S. V., Couzis, A. & Maldarelli, C. Reduction in aggregation and energy transfer of quantum dots incorporated in polystyrene beads by kinetic entrapment due to cross-linking during polymerization. Langmuir 31, 3167–3179 (2015).
Clapp, A. R., Medintz, I. L. & Mattoussi, H. Förster resonance energy transfer investigations using quantum-dot fluorophores. ChemPhysChem 7, 47–57 (2006).
Wagh, A. et al. Polymeric nanoparticles with sequential and multiple FRET cascade mechanisms for multicolor and multiplexed imaging. Small 9, 2129–2139 (2013).
Stuchlý, J. et al. An automated analysis of highly complex flow cytometry-based proteomic data. Cytom. A 81, 120–129 (2012).
Han, M., Gao, X., Su, J. Z. & Nie, S. Quantum-dot-tagged microbeads for multiplexed optical coding of biomolecules. Nat. Biotechnol. 19, 631–635 (2001).
Wang, G. et al. Highly efficient preparation of multiscaled quantum dot barcodes for multiplexed hepatitis B detection. ACS Nano 7, 471–481 (2013).
Ferguson, J. A., Steemers, F. J. & Walt, D. R. High-density fiber-optic DNA random microsphere array. Anal. Chem. 72, 5618–5624 (2000).
Förster, T. Experimentelle und theoretische Untersuchung des zwischengmolekularen übergangs von Elektronenanregungsenergie. Naturforsch. A Astrophys. Phys. Phys. Chem. 4a, 321–327 (1949).
Wolber, P. & Hudson, B. An analytic solution to the Förster energy transfer problem in two dimensions. Biophys. J. 28, 197–210 (1979).
Watrob, H. M., Pan, C. P. & Barkley, M. D. Two-step FRET as a structural tool. J. Am. Chem. Soc. 125, 7336–7343 (2003).
Fabian, A., Horvath, G., Vamosi, G., Vereb, G. & Szollosi, J. TripleFRET measurements in flow cytometry. Cytom. A 83, 375–385 (2013).
Stein, I. H., Steinhauer, C. & Tinnefeld, P. Single-molecule four-color FRET visualizes energy-transfer paths on DNA origami. J. Am. Chem. Soc. 133, 4193–4195 (2011).
Lee, J. et al. Single-molecule four-color FRET. Angew. Chem. Int. Ed. 49, 9922–9925 (2010).
Uphoff, S. et al. Monitoring multiple distances within a single molecule using switchable FRET. Nat. Methods 7, 831–836 (2010).
Bunt, G. & Wouters, F. S. FRET from single to multiplexed signaling events. Biophys. Rev. 9, 119–129 (2017).
Buckhout-White, S. et al. Assembling programmable FRET-based photonic networks using designer DNA scaffolds. Nat. Commun. 5, 5615–5630 (2014).
Spillmann, C. M. et al. Extending FRET cascades on linear DNA photonic wires. Chem. Commun. 50, 7246–7249 (2014).
Raicu, V. Efficiency of resonance energy transfer in homo-oligomeric complexes of proteins. J. Biol. Phys. 33, 109–127 (2007).
Liu, J. & Lu, Y. FRET study of a trifluorophore-labelled DNAzyme. J. Am. Chem. Soc. 124, 15208–15216 (2002).
Shapiro, H. M. Practical Flow Cytometry (Wiley, Hoboken, NJ, 2003).
Mátyus, L. Fluorescence resonance energy transfer measurements on cell surfaces. A spectroscopic tool for determining protein interactions. J. Photochem. Photobiol. 12, 323–337 (1992).
Berney, C. & Danuser, G. FRET or no FRET: a quantitative comparison. Biophys. J. 84, 3992–4010 (2003).
Corry, B., Jayatilaka, D. & Rigby, P. A flexible approach to the calculation of resonance energy transfer efficiency between multiple donors and acceptors in complex geometries. Biophys. J. 89, 3822–3836 (2005).
Koppel, D. E., Fleming, P. J. & Strittmatter, P. Intramembrane positions of membrane-bound chromophores determined by excitation energy transfer. Biochemistry 18, 5450–5457 (1979).
Nguyen, H. Q. et al. Programmable microfluidic synthesis of over one thousand uniquely identifiable spectral codes. Adv. Opt. Mater. 5, 1600548 (2016).
Lee, J. et al. Universal process-inert encoding architecture for polymer microparticles. Nat. Mater. 13, 524–529 (2014).
Fielding, A. H. Cluster and Classification Techniques for the Biosciences Vol. 53 (Cambridge Univ. Press, Cambridge, 2007).
King, L. E. et al. Ligand binding assay critical reagents and their stability: recommendations and best practices from the Global Bioanalysis Consortium Harmonization Team. AAPS J. 16, 504–515 (2014).
Uhlen, M. et al. A proposal for validation of antibodies. Nat. Methods 13, 823–827 (2016).
Pla-Roca, M. et al. Antibody colocalization microarray: a scalable technology for multiplex protein analysis in complex samples. Mol. Cell. Proteomics 11, https://doi.org/10.1074/mcp.M111.011460 (2012).
Juncker, D., Bergeron, S., Laforte, V. & Li, H. Cross-reactivity in antibody microarrays and multiplexed sandwich assays: shedding light on the dark side of multiplexing. Curr. Opin. Chem. Biol. 18, 29–37 (2014).
Schweitzer, B. et al. Multiplexed protein profiling on microarrays by rolling-circle amplification. Nat. Biotechnol. 20, 359–365 (2002).
Gonzalez, R. M. et al. Development and validation of sandwich ELISA microarrays with minimal assay interference. J. Proteome Res. 7, 2406–2414 (2008).
Hardin, B. E. et al. Increased light harvesting in dye-sensitized solar cells with energy relay dyes. Nat. Photon. 3, 406–411 (2009).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy (Springer, New York, NY, 2006).
Förster, T. in Modern Quantum Chemistry Part III: Action of Light and Organic Crystals (ed. Sinanglu, O.) 93–137 (Academic, New York, NY, 1965)..
Dempster, A., Laird, N. & Donald, R. Maximum likelihood from incomplete data via the EM algorithm. J. R. Statist. Soc. Ser. B 39, 1–38 (1977).
Acknowledgements
The authors thank T. Gervais for discussions, and J. Munzar for proofreading our manuscript. The authors thank NSERC and FQRNT for funding. M.D. acknowledges the NSERC-CREATE ISS programme for support. The flow cytometry work was performed at two McGill core flow facilities, namely the Microbiology and Immunology (MIMM) department and the Life Science Complex, which is supported by funding from the Canadian Foundation for Innovation.
Author information
Authors and Affiliations
Contributions
M.D. and D.J. developed the approach. M.D. and A.N. conceived the experiments. M.D. developed the models and performed the experiments. M.K. and M.D. developed the decoding algorithm. M.D and D.J. analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.J. and M.D. are inventors on a provisional US patent application 62/568,998 that covers some of the aspects reported here and was filed by McGill University on 6 October 2017. M.D. and D.J. are founders and shareholders of nplex biosciences inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–14, Supplementary Tables 1–2, Supplementary Notes and Supplementary References
Supplementary Table 3
Designed barcodes
Supplementary Table 4
35-plex sandwich assay: barcodes and reagents
Rights and permissions
About this article
Cite this article
Dagher, M., Kleinman, M., Ng, A. et al. Ensemble multicolour FRET model enables barcoding at extreme FRET levels. Nature Nanotech 13, 925–932 (2018). https://doi.org/10.1038/s41565-018-0205-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-018-0205-0
This article is cited by
-
Reconfigurable photoswitchable multistate fluorescent polymer-based information encryption toolbox
Science China Materials (2023)
-
Spatial transcriptomics using combinatorial fluorescence spectral and lifetime encoding, imaging and analysis
Nature Communications (2022)
-
Fibrillin-1-regulated miR-122 has a critical role in thoracic aortic aneurysm formation
Cellular and Molecular Life Sciences (2022)
-
MRBLES 2.0: High-throughput generation of chemically functionalized spectrally and magnetically encoded hydrogel beads using a simple single-layer microfluidic device
Microsystems & Nanoengineering (2020)
-
Highly multiplexed spatial mapping of microbial communities
Nature (2020)