Abstract
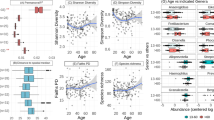
The prevalence of chronic, non-communicable diseases has risen sharply in recent decades, especially in industrialized countries. While several studies implicate the microbiome in this trend, few have examined the evolutionary history of industrialized microbiomes. Here we sampled 235 ancient dental calculus samples from individuals living in Great Britain (∼2200 bce to 1853 ce), including 127 well-contextualized London adults. We reconstructed their microbial history spanning the transition to industrialization. After controlling for oral geography and technical biases, we identified multiple oral microbial communities that coexisted in Britain for millennia, including a community associated with Methanobrevibacter, an anaerobic Archaea not commonly prevalent in the oral microbiome of modern industrialized societies. Calculus analysis suggests that oral hygiene contributed to oral microbiome composition, while microbial functions reflected past differences in diet, specifically in dairy and carbohydrate consumption. In London samples, Methanobrevibacter-associated microbial communities are linked with skeletal markers of systemic diseases (for example, periostitis and joint pathologies), and their disappearance is consistent with temporal shifts, including the arrival of the Second Plague Pandemic. This suggests pre-industrialized microbiomes were more diverse than previously recognized, enhancing our understanding of chronic, non-communicable disease origins in industrialized populations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All trimmed and merged DNA sequences (fastq) are available in the SRA database (BioProject PRJNA780005) of NCBI. The 2017 NCBI nr database and the 2017 NCBI RefSeq GCS database were used in this study. Unmerged reads can be made available upon request, as only merged sequences were assessed in full for this publication.
Code availability
The analysis pipelines are available in the microARCH GitHub page (@microARCHlab/BritishDentalCalculus_2021), as well as in https://github.com/michellepistner/ancientDNA.
References
Sonnenburg, E. D. & Sonnenburg, J. L. The ancestral and industrialized gut microbiota and implications for human health. Nat. Rev. Microbiol. 17, 383–390 (2019).
Broussard, J. L. & Devkota, S. The changing microbial landscape of Western society: diet, dwellings and discordance. Mol. Metab. 5, 737–742 (2016).
Abegunde, D. O., Mathers, C. D., Adam, T., Ortegon, M. & Strong, K. The burden and costs of chronic diseases in low-income and middle-income countries. Lancet 370, 1929–1938 (2007).
Mathers, C. D. & Loncar, D. Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med. 3, e442 (2006).
Schnorr, S. L. et al. Gut microbiome of the Hadza hunter-gatherers. Nat. Commun. 5, 3654 (2014).
Clemente, J. C. et al. The microbiome of uncontacted Amerindians. Sci. Adv. 1, e1500183 (2015).
Martínez, I. et al. The gut microbiota of rural Papua New Guineans: composition, diversity patterns, and ecological processes. Cell Rep. 11, 527–538 (2015).
Clayton, J. B. et al. Captivity humanizes the primate microbiome. Proc. Natl Acad. Sci. USA 113, 10376–10381 (2016).
Obregon-Tito, A. J. et al. Subsistence strategies in traditional societies distinguish gut microbiomes. Nat. Commun. 6, 6505 (2015).
Lokmer, A. et al. Response of the human gut and saliva microbiome to urbanization in Cameroon. Sci. Rep. 10, 2856 (2020).
Jha, A. R. et al. Gut microbiome transition across a lifestyle gradient in Himalaya. PLoS Biol. 16, e2005396 (2018).
Vangay, P. et al. U.S. immigration westernizes the human gut microbiome. Cell 175, 962–972.e10 (2018).
Bello, M. G. D., Knight, R., Gilbert, J. A. & Blaser, M. J. Preserving microbial diversity. Science 362, 33–34 (2018).
Skelly, E., Kapellas, K., Cooper, A. & Weyrich, L. S. Consequences of colonialism: a microbial perspective to contemporary Indigenous health. Am. J. Biol. Anthropol. 167, 423–437 (2018).
Schnorr, S. L. Meanings, measurements, and musings on the significance of patterns in human microbiome variation. Curr. Opin. Genet. Dev. 53, 43–52 (2018).
Carmody, R. N., Sarkar, A. & Reese, A. T. Gut microbiota through an evolutionary lens. Science 372, 462–463 (2021).
Tsosie, K. S., Fox, K. & Yracheta, J. M. Genomics data: the broken promise is to Indigenous people. Nature 591, 529–529 (2021).
Wibowo, M. C. et al. Reconstruction of ancient microbial genomes from the human gut. Nature 594, 234–239 (2021).
Weyrich, L. S., Dobney, K. & Cooper, A. Ancient DNA analysis of dental calculus. J. Hum. Evol. 79, 119–124 (2015).
Weyrich, L. S. et al. Neanderthal behaviour, diet, and disease inferred from ancient DNA in dental calculus. Nature 544, 357–361 (2017).
Warinner, C., Speller, C. & Collins, M. J. A new era in palaeomicrobiology: prospects for ancient dental calculus as a long-term record of the human oral microbiome. Philos. Trans. R. Soc. Lond. B 370, 20130376 (2015).
Hillmann, B. et al. Evaluating the information content of shallow shotgun metagenomics. mSystems 3, e00069-18 (2018).
Liu, Y. The Role of Epigenetic Modications and Microbiome Evolution in Bovid Adaptation to Environmental Changes. PhD thesis, University of Adelaide (2019).
Jónsson, H., Ginolhac, A., Schubert, M., Johnson, P. L. F. & Orlando, L. mapDamage2.0: fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics 29, 1682–1684 (2013).
Simón-Soro, A. et al. Microbial geography of the oral cavity. J. Dent. Res. 92, 616–621 (2013).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Eisenhofer, R., Anderson, A., Dobney, K., Cooper, A. & Weyrich, L. S. Ancient microbial DNA in dental calculus: a new method for studying rapid human migration events. J. Isl. Coast. Archaeol. 14, 149–162 (2019).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Velsko, I. M. et al. Ancient dental calculus preserves signatures of biofilm succession and interindividual variation independent of dental pathology. PNAS Nexus 1, pgac148 (2022).
Yates, J. A. F. et al. The evolution and changing ecology of the African hominid oral microbiome. Proc. Natl Acad. Sci. 118, e2021655118 (2021).
Velsko, I. M., Gallois, S., Stahl, R., Henry, A. G. & Warinner, C. High conservation of the dental plaque microbiome across populations with differing subsistence strategies and levels of market integration. Mol. Ecol. https://doi.org/10.1111/mec.16988 (2023).
Morton, J. T. et al. Uncovering the horseshoe effect in microbial analyses. mSystems 2, e00166-16 (2017).
Granehäll, L. et al. Metagenomic analysis of ancient dental calculus reveals unexplored diversity of oral archaeal Methanobrevibacter. Microbiome 9, 197 (2021).
Downes, J., Munson, M. A., Radford, D. R., Spratt, D. A. & Wade, W. G. Shuttleworthia satelles gen. nov., sp. nov., isolated from the human oral cavity. Int. J. Syst. Evol. Microbiol. 52, 1469–1475 (2002).
Siqueira, J. F. & Rôças, I. N. Pseudoramibacter alactolyticus in primary endodontic infections. J. Endod. 29, 735–738 (2003).
Huang, S. et al. Preliminary characterization of the oral microbiota of Chinese adults with and without gingivitis. BMC Oral Health 11, 33 (2011).
Gaci, N., Borrel, G., Tottey, W., O’Toole, P. W. & Brugère, J.-F. Archaea and the human gut: new beginning of an old story. World J. Gastroenterol. 20, 16062–16078 (2014).
Handsley-Davis, M. et al. Heritage-specific oral microbiota in Indigenous Australian dental calculus. Evol. Med. Public Health 10, 352–362 (2022).
Adler, C. J. et al. Sequencing ancient calcified dental plaque shows changes in oral microbiota with dietary shifts of the Neolithic and Industrial revolutions. Nat. Genet. 45, 450–455 (2013).
Lepp, P. W. et al. Methanogenic Archaea and human periodontal disease. Proc. Natl Acad. Sci. USA 101, 6176–6181 (2004).
Creeth, J. E. et al. The effect of brushing time and dentifrice on dental plaque removal in vivo. J. Dent. Hyg. 83, 111–116 (2009).
Zanatta, F. B., Bergoli, A. D., Werle, S. B. & Antoniazzi, R. P. Biofilm removal and gingival abrasion with medium and soft toothbrushes. Oral Health Prev. Dent. 9, 177–183 (2011).
Mann, A. E. et al. Do I have something in my teeth? The trouble with genetic analyses of diet from archaeological dental calculus. Quat. Int. 653–654, 33–46 (2023).
Ozga, A. T. & Ottoni, C. Dental calculus as a proxy for animal microbiomes. Quat. Int. 653–654, 47–52 (2023).
Hillson, S. Archaeology and the study of teeth. Endeavour 10, 145–149 (1986).
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011).
Ley, R. E. et al. Evolution of mammals and their gut microbes. Science 320, 1647–1651 (2008).
Warinner, C. et al. Direct evidence of milk consumption from ancient human dental calculus. Sci. Rep. 4, 7104 (2014).
Ottoni, C. et al. Tracking the transition to agriculture in southern Europe through ancient DNA analysis of dental calculus. Proc. Natl Acad. Sci. USA 118, e2102116118 (2021).
Lee, Y.-H. et al. Progress in oral microbiome related to oral and systemic diseases: an update. Diagnostics 11, 1283 (2021).
Peng, X. et al. Oral microbiota in human systematic diseases. Int. J. Oral. Sci. 14, 1–11 (2022).
Schuenemann, V. J. et al. Targeted enrichment of ancient pathogens yielding the pPCP1 plasmid of Yersinia pestis from victims of the Black Death. Proc. Natl Acad. Sci. USA 108, E746–E752 (2011).
Shrewsbury, J. F. D. A History of Bubonic Plague in the British Isles (Cambridge Univ. Press, 2005).
DeWitte, S. N. & Wood, J. W. Selectivity of Black Death mortality with respect to preexisting health. Proc. Natl Acad. Sci. USA 105, 1436–1441 (2008).
Yaussy, S. L. & DeWitte, S. N. Calculus and survivorship in medieval London: the association between dental disease and a demographic measure of general health. Am. J. Phys. Anthropol. 168, 552–565 (2019).
DeWitte, S. N. Health in post-Black Death London (1350–1538): age patterns of periosteal new bone formation in a post-epidemic population. Am. J. Phys. Anthropol. 155, 260–267 (2014).
Moore, N. E. & Weyrich, L. S. A. in The Oral Microbiome: Methods and Protocols (ed. Adami, G. R.) 93–118 (Springer, 2021); https://doi.org/10.1007/978-1-0716-1518-8_7
Brotherton, P. et al. Neolithic mitochondrial haplogroup H genomes and the genetic origins of Europeans. Nat. Commun. 4, 1764 (2013).
Eisenhofer, R. et al. Contamination in low microbial biomass microbiome studies: issues and recommendations. Trends Microbiol. 27, 105–117 (2019).
Lindgreen, S. AdapterRemoval: easy cleaning of next-generation sequencing reads. BMC Res. Notes 5, 337 (2012).
Knights, D. et al. Bayesian community-wide culture-independent microbial source tracking. Nat. Methods 8, 761–763 (2011).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Mitra, S. et al. Functional analysis of metagenomes and metatranscriptomes using SEED and KEGG. BMC Bioinform. 12, S21 (2011).
Archer, L., Dawson, E., DeWitt, J., Seakins, A. & Wong, B. “Science capital”: a conceptual, methodological, and empirical argument for extending bourdieusian notions of capital beyond the arts. J. Res. Sci. Teach. 52, 922–948 (2015).
Acknowledgements
We thank C. Stringer and R. Kruszynski of the Natural History Museum, London; S. Schiffels; D. Sayer; Oxford Archaeology East; M. Farrell of the Royal College of Surgeons of England; J. Pearson of the Inverness Museum; and all of the museums for access to samples. We also thank the Museum of London for allowing us to collect and destructively analyse archaeological dental calculus samples from their collections from London, particularly J. Bekvalac and R. Redfern. We would also like to acknowledge J. VanderBerg at EnDev Geographic for producing the map used in Fig. 1. A.C., C.A. and L.W. thank the Australian Research Council for research funding (DP110105038) and Laureate (FL140100260). The work was also supported by an Australian Research Council Future Fellowship Award to L.S.W. (FT180100407). This material is also based on work supported by the National Science Foundation Graduate Research Fellowship Program awarded to A.S.G. under Grant No. DGE1255832. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation.
Author information
Authors and Affiliations
Contributions
A.G.F., N.G., A.C., K.D. and L.S.W. conceived of the study and developed the experimental design. A.C., A.G.F., C.A., K.B., K.D. and L.S.W. worked on sample acquisition. A.G.F. completed the laboratory analysis. A.S.G., A.G.F., S.W. and L.A. completed the bioinformatics and computational analysis. A.S.G., M.P.N., E.R.D., J.D.S. and L.S.W. performed the statistical analysis. A.S.G., A.G.F. and L.S.W. wrote the paper, and all authors edited and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Marcus de Goffau, Aleksandar Kostic and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text Sections 1–9 and Supplementary Figs. 1–19.
Supplementary Tables
Supplementary Tables 1–23.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gancz, A.S., Farrer, A.G., Nixon, M.P. et al. Ancient dental calculus reveals oral microbiome shifts associated with lifestyle and disease in Great Britain. Nat Microbiol 8, 2315–2325 (2023). https://doi.org/10.1038/s41564-023-01527-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01527-3