Abstract
Human-associated bacteria secrete modified peptides to control host physiology and remodel the microbiota species composition. Here we scanned 2,229 Human Microbiome Project genomes of species colonizing skin, gastrointestinal tract, urogenital tract, mouth and trachea for gene clusters encoding RiPPs (ribosomally synthesized and post-translationally modified peptides). We found 218 lanthipeptides and 25 lasso peptides, 70 of which were synthesized and expressed in E. coli and 23 could be purified and functionally characterized. They were tested for activity against bacteria associated with healthy human flora and pathogens. New antibiotics were identified against strains implicated in skin, nasal and vaginal dysbiosis as well as from oral strains selectively targeting those in the gut. Extended- and narrow-spectrum antibiotics were found against methicillin-resistant Staphylococcus aureus and vancomycin-resistant Enterococci. Mining natural products produced by human-associated microbes will enable the elucidation of ecological relationships and may be a rich resource for antimicrobial discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The RiPPs biosynthetic pathways found in this study are available in NCBI (https://www.ncbi.nlm.nih.gov/), with identifiers listed in Supplementary Table 5 and Source data. All DNA sequences are listed in Supplementary Table 5. Data supporting the findings of this study are available within the paper and supplementary materials. Additional data, strains and plasmids are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
Python scripts used for LC–MS processing have been released as open source software under the MIT license (GitHub repository: https://github.com/dantheand/msms_structure_annot).
References
Sassone-Corsi, M. & Raffatellu, M. No vacancy: how beneficial microbes cooperate with immunity to provide colonization resistance to pathogens. J. Immunol. 194, 4081–4087 (2015).
Heilbronner, S., Krismer, B., Brötz-Oesterhelt, H. & Peschel, A. The microbiome-shaping roles of bacteriocins. Nat. Rev. Microbiol. 19, 726–739 (2021).
Russell, A. H. & Truman, A. W. Genome mining strategies for ribosomally synthesised and post-translationally modified peptides. Comput. Struct. Biotechnol. J. 18, 1838–1851 (2020).
Shepherd, E. S., DeLoache, W. C., Pruss, K. M., Whitaker, W. R. & Sonnenburg, J. L. An exclusive metabolic niche enables strain engraftment in the gut microbiota. Nature 557, 434–438 (2018).
Kearney, S. M., Gibbons, S. M., Erdman, S. E. & Alm, E. J. Orthogonal dietary niche enables reversible engraftment of a gut bacterial commensal. Cell Rep. 24, 1842–1851 (2018).
Isabella, V. M. et al. Development of a synthetic live bacterial therapeutic for the human metabolic disease phenylketonuria. Nat. Biotechnol. 36, 857–864 (2018).
Cotter, P. D., Paul Ross, R. & Hill, C. Bacteriocins—a viable alternative to antibiotics? Nat. Rev. Microbiol. 11, 95–105 (2012).
Montalbán-López, M. et al. New developments in RiPP discovery, enzymology and engineering. Nat. Prod. Rep. https://doi.org/10.1039/d0np00027b (2020).
Klaus, M. & Grininger, M. Engineering strategies for rational polyketide synthase design. Nat. Prod. Rep. 35, 1070–1081 (2018).
Cai, W. & Zhang, W. Engineering modular polyketide synthases for production of biofuels and industrial chemicals. Curr. Opin. Biotechnol. 50, 32–38 (2018).
Cubillos-Ruiz, A., Berta-Thompson, J. W., Becker, J. W., van der Donk, W. A. & Chisholm, S. W. Evolutionary radiation of lanthipeptides in marine cyanobacteria. Proc. Natl Acad. Sci. USA 114, E5424–E5433 (2017).
Sardar, D., Pierce, E., McIntosh, J. A. & Schmidt, E. W. Recognition sequences and substrate evolution in cyanobactin biosynthesis. ACS Synth. Biol. 4, 167–176 (2015).
Yang, X. et al. A lanthipeptide library used to identify a protein–protein interaction inhibitor. Nat. Chem. Biol. 14, 375–380 (2018).
Hetrick, K. J., Walker, M. C. & van der Donk, W. A. Development and application of yeast and phage display of diverse lanthipeptides. ACS Cent. Sci. 4, 458–467 (2018).
Urban, J. H. et al. Phage display and selection of lanthipeptides on the carboxy-terminus of the gene-3 minor coat protein. Nat. Commun. 8, 1500 (2017).
King, A. M. et al. Selection for constrained peptides that bind to a single target protein. Nat. Commun. 12, 6343 (2021).
Cheung-Lee, W. L. & Link, A. J. Genome mining for lasso peptides: past, present, and future. J. Ind. Microbiol. Biotechnol. 46, 1371–1379 (2019).
van Staden, A. D. P., van Zyl, W. F., Trindade, M., Dicks, L. M. T. & Smith, C. Therapeutic application of lantibiotics and other lanthipeptides: old and new findings. Appl. Environ. Microbiol. 87, e0018621 (2021).
Montalbán-López, M. et al. New developments in RiPP discovery, enzymology and engineering. Nat. Prod. Rep. 38, 130–239 (2021).
Ongpipattanakul, C. et al. Mechanism of action of ribosomally synthesized and post-translationally modified peptides. Chem. Rev. 122, 14722–14814 (2022).
Cao, L., Do, T. & Link, A. J. Mechanisms of action of ribosomally synthesized and posttranslationally modified peptides (RiPPs). J. Ind. Microbiol. Biotechnol. https://doi.org/10.1093/jimb/kuab005 (2021).
Hetrick, K. J. & van der Donk, W. A. Ribosomally synthesized and post-translationally modified peptide natural product discovery in the genomic era. Curr. Opin. Chem. Biol. 38, 36–44 (2017).
Barbour, A., Wescombe, P. & Smith, L. Evolution of lantibiotic salivaricins: new weapons to fight infectious diseases. Trends Microbiol. 28, 578–593 (2020).
Donia, M. S. et al. A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell 158, 1402–1414 (2014).
Metelev, M. et al. Acinetodin and klebsidin, RNA polymerase targeting lasso peptides produced by human isolates of Acinetobacter gyllenbergii and Klebsiella pneumoniae. ACS Chem. Biol. 12, 814–824 (2017).
Zhong, Z., He, B., Li, J. & Li, Y.-X. Challenges and advances in genome mining of ribosomally synthesized and post-translationally modified peptides (RiPPs). Synth. Syst. Biotechnol. 5, 155–172 (2020).
van Heel, A. J., de Jong, A., Montalbán-López, M., Kok, J. & Kuipers, O. P. BAGEL3: automated identification of genes encoding bacteriocins and (non-)bactericidal posttranslationally modified peptides. Nucleic Acids Res. 41, W448–453 (2013).
Blin, K., Kazempour, D., Wohlleben, W. & Weber, T. Improved lanthipeptide detection and prediction for antiSMASH. PLoS ONE 9, e89420 (2014).
Mohimani, H. et al. Automated genome mining of ribosomal peptide natural products. ACS Chem. Biol. 9, 1545–1551 (2014).
Merwin, N. J. et al. DeepRiPP integrates multiomics data to automate discovery of novel ribosomally synthesized natural products. Proc. Natl Acad. Sci. USA 117, 371–380 (2019).
Santos-Aberturas, J. et al. Uncovering the unexplored diversity of thioamidated ribosomal peptides in Actinobacteria using the RiPPER genome mining tool. Nucleic Acids Res. 47, 4624–4637 (2019).
Tietz, J. I. et al. A new genome-mining tool redefines the lasso peptide biosynthetic landscape. Nat. Chem. Biol. 13, 470–478 (2017).
Sugimoto, Y. et al. A metagenomic strategy for harnessing the chemical repertoire of the human microbiome. Science https://doi.org/10.1126/science.aax9176 (2019).
Walker, M. C. et al. Precursor peptide-targeted mining of more than one hundred thousand genomes expands the lanthipeptide natural product family. BMC Genomics 21, 387 (2020).
Ongey, E. L. & Neubauer, P. Lanthipeptides: chemical synthesis versus in vivo biosynthesis as tools for pharmaceutical production. Microb. Cell Fact. 15, 97 (2016).
Wilson, M. R., Zha, L. & Balskus, E. P. Natural product discovery from the human microbiome. J. Biol. Chem. 292, 8546–8552 (2017).
Bauman, K. D., Butler, K. S., Moore, B. S. & Chekan, J. R. Genome mining methods to discover bioactive natural products. Nat. Prod. Rep. https://doi.org/10.1039/d1np00032b (2021).
Ayikpoe, R. S. et al. A scalable platform to discover antimicrobials of ribosomal origin. Nat. Commun. 13, 6135 (2022).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded Human Microbiome Project. Nature 550, 61–66 (2017).
Temme, K., Zhao, D. & Voigt, C. A. Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca. Proc. Natl Acad. Sci. USA 109, 7085–7090 (2012).
Blin, K. et al. antiSMASH 4.0–improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 45, W36–W41 (2017).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Cimermancic, P. et al. Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158, 412–421 (2014).
Glassey, E., King, A. M., Anderson, D. A., Zhang, Z. & Voigt, C. A. Functional expression of diverse post-translational peptide-modifying enzymes in Escherichia coli. PLoS One 17, e0266488 (2022).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli ‘Marionette’ strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Lou, C., Stanton, B., Chen, Y.-J., Munsky, B. & Voigt, C. A. Ribozyme-based insulator parts buffer synthetic circuits from genetic context. Nat. Biotechnol. 30, 1137–1142 (2012).
Shin, J., Zhang, S., Der, B. S., Nielsen, A. A. & Voigt, C. A. Programming Escherichia coli to function as a digital display. Mol. Syst. Biol. 16, e9401 (2020).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Zhang, Q. et al. Structural investigation of ribosomally synthesized natural products by hypothetical structure enumeration and evaluation using tandem MS. Proc. Natl Acad. Sci. USA 111, 12031–12036 (2014).
Dufour, A., Hindré, T., Haras, D. & Le Pennec, J.-P. The biology of lantibiotics from the lacticin 481 group is coming of age. FEMS Microbiol. Rev. 31, 134–167 (2007).
Knerr, P. J. & van der Donk, W. A. Chemical synthesis of the lantibiotic lacticin 481 reveals the importance of lanthionine stereochemistry. J. Am. Chem. Soc. 135, 7094–7097 (2013).
Smith, M., Kassam, Z., Edelstein, C., Burgess, J. & Alm, E. OpenBiome remains open to serve the medical community. Nat. Biotechnol. 32, 867 (2014).
Xie, L. et al. Lacticin 481: in vitro reconstitution of lantibiotic synthetase activity. Science 303, 679–681 (2004).
Amabebe, E. & Anumba, D. O. C. The vaginal microenvironment: the physiologic role of lactobacilli. Front. Med. 5, 181 (2018).
Petrova, M. I., Reid, G., Vaneechoutte, M. & Lebeer, S. Lactobacillus iners: friend or foe? Trends Microbiol. 25, 182–191 (2017).
Severgnini, M. et al. Gardnerella vaginalis clades in pregnancy: new insights into the interactions with the vaginal microbiome. PLoS ONE 17, e0269590 (2022).
Vidor, C. J., Bulach, D., Awad, M. & Lyras, D. Paeniclostridium sordellii and Clostridioides difficile encode similar and clinically relevant tetracycline resistance loci in diverse genomic locations. BMC Microbiol. 19, 53 (2019).
Funke, G., von Graevenitz, A., Clarridge, J. E. 3rd & Bernard, K. A. Clinical microbiology of coryneform bacteria. Clin. Microbiol. Rev. 10, 125–159 (1997).
Martinez-Martinez, L., Suarez, A. I., Rodriguez-Bano, J., Bernard, K. & Muniain, M. A. Clinical significance of Corynebacterium striatum isolated from human samples. Clin. Microbiol. Infect. 3, 634–639 (1997).
Brugger, S. D. et al. Dolosigranulum pigrum cooperation and competition in human nasal microbiota. mSphere 5, e00852-20 (2020).
Rinninella, E. et al. What is the healthy gut microbiota composition? A changing ecosystem across age, environment, diet, and diseases. Microorganisms https://doi.org/10.3390/microorganisms7010014 (2019).
Antibiotic Resistance Threats in the United States, 2019 (US Centers for Disease Control and Prevention, 2019).
Pettigrew, M. M. et al. Upper respiratory tract microbial communities, acute otitis media pathogens, and antibiotic use in healthy and sick children. Appl. Environ. Microbiol. 78, 6262–6270 (2012).
Kafil, H. S. & Asgharzadeh, M. Vancomycin-resistant Enteroccus faecium and Enterococcus faecalis isolated from education hospital of Iran. Maedica 9, 323–327 (2014).
van den Hooven, H. W. et al. The structure of the lantibiotic lacticin 481 produced by Lactococcus lactis: location of the thioether bridges. FEBS Lett. 391, 317–322 (1996).
Walker, G. V., Heng, N. C. K., Carne, A., Tagg, J. R. & Wescombe, P. A. Salivaricin E and abundant dextranase activity may contribute to the anti-cariogenic potential of the probiotic candidate Streptococcus salivarius JH. Microbiology 162, 476–486 (2016).
Khan, S., Voordouw, M. J. & Hill, J. E. Competition among Gardnerella subgroups from the human vaginal microbiome. Front. Cell Infect. Microbiol. 9, 374 (2019).
O’Neill, A. M. et al. Identification of a human skin commensal bacterium that selectively kills Cutibacterium acnes. J. Invest. Dermatol. 140, 1619–1628.e2 (2020).
Janek, D., Zipperer, A., Kulik, A., Krismer, B. & Peschel, A. High frequency and diversity of antimicrobial activities produced by nasal Staphylococcus strains against bacterial competitors. PLoS Pathog. 12, e1005812 (2016).
Kaur, H. et al. The antibiotic darobactin mimics a β-strand to inhibit outer membrane insertase. Nature 593, 125–129 (2021).
Gavrish, E. et al. Lassomycin, a ribosomally synthesized cyclic peptide, kills Mycobacterium tuberculosis by targeting the ATP-dependent protease ClpC1P1P2. Chem. Biol. 21, 509–518 (2014).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature 517, 455–459 (2015).
Krismer, B., Weidenmaier, C., Zipperer, A. & Peschel, A. The commensal lifestyle of Staphylococcus aureus and its interactions with the nasal microbiota. Nat. Rev. Microbiol. 15, 675–687 (2017).
Ochi, K. & Hosaka, T. New strategies for drug discovery: activation of silent or weakly expressed microbial gene clusters. Appl. Microbiol. Biotechnol. 97, 87–98 (2013).
Zhang, X., Hindra & Elliot, M. A. Unlocking the trove of metabolic treasures: activating silent biosynthetic gene clusters in bacteria and fungi. Curr. Opin. Microbiol. 51, 9–15 (2019).
Frank, D. N. et al. The human nasal microbiota and Staphylococcus aureus carriage. PLoS ONE 5, e10598 (2010).
Otto, M. Staphylococci in the human microbiome: the role of host and interbacterial interactions. Curr. Opin. Microbiol. 53, 71–77 (2020).
Edlund, A. et al. Uncovering complex microbiome activities via metatranscriptomics during 24 hours of oral biofilm assembly and maturation. Microbiome 6, 217 (2018).
van Hal, S. J., Lodise, T. P. & Paterson, D. L. The clinical significance of vancomycin minimum inhibitory concentration in Staphylococcus aureus infections: a systematic review and meta-analysis. Clin. Infect. Dis. 54, 755–771 (2012).
King, E. A., McCoy, D., Desai, S., Nyirenda, T. & Bicking, K. Vancomycin-resistant enterococcal bacteraemia and daptomycin: are higher doses necessary? J. Antimicrob. Chemother. 66, 2112–2118 (2011).
Hegemann, J. D., Zimmermann, M., Xie, X. & Marahiel, M. A. Lasso peptides: an intriguing class of bacterial natural products. Acc. Chem. Res. 48, 1909–1919 (2015).
Piewngam, P. et al. Pathogen elimination by probiotic Bacillus via signalling interference. Nature 562, 532–537 (2018).
Pinho-Ribeiro, F. A. et al. Blocking neuronal signaling to immune cells treats streptococcal invasive infection. Cell 173, 1083–1097.e22 (2018).
Duan, Y. et al. Bacteriophage targeting of gut bacterium attenuates alcoholic liver disease. Nature 575, 505–511 (2019).
Meijerink, M. et al. Identification of genetic loci in Lactobacillus plantarum that modulate the immune response of dendritic cells using comparative genome hybridization. PLoS ONE 5, e10632 (2010).
Raffatellu, M. Learning from bacterial competition in the host to develop antimicrobials. Nat. Med. 24, 1097–1103 (2018).
Shi, B., Leung, D. Y. M., Taylor, P. A. & Li, H. Methicillin-resistant Staphylococcus aureus colonization is associated with decreased skin commensal bacteria in atopic dermatitis. J. Invest. Dermatol. 138, 1668–1671 (2018).
Strauss, J. et al. Invasive potential of gut mucosa-derived Fusobacterium nucleatum positively correlates with IBD status of the host. Inflamm. Bowel Dis. 17, 1971–1978 (2011).
Ismail, Y. et al. Investigation of the enteric pathogenic potential of oral Campylobacter concisus strains isolated from patients with inflammatory bowel disease. PLoS ONE 7, e38217 (2012).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–298 (2012).
Farrell, J. J. et al. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 61, 582–588 (2012).
Fan, X. et al. Human oral microbiome and prospective risk for pancreatic cancer: a population-based nested case-control study. Gut 67, 120–127 (2018).
Lu, H. et al. Deep sequencing reveals microbiota dysbiosis of tongue coat in patients with liver carcinoma. Sci. Rep. 6, 33142 (2016).
Bajaj, J. S. et al. Salivary microbiota reflects changes in gut microbiota in cirrhosis with hepatic encephalopathy. Hepatology 62, 1260–1271 (2015).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64 (2014).
Li, B. et al. Oral bacteria colonize and compete with gut microbiota in gnotobiotic mice. Int. J. Oral. Sci. 11, 10 (2019).
Bajer, L. et al. Distinct gut microbiota profiles in patients with primary sclerosing cholangitis and ulcerative colitis. World J. Gastroenterol. 23, 4548–4558 (2017).
Brugger, S. D., Bomar, L. & Lemon, K. P. Commensal–pathogen interactions along the human nasal passages. PLoS Pathog. 12, e1005633 (2016).
Zheng, J., Ganzle, M. G., Lin, X. B., Ruan, L. & Sun, M. Diversity and dynamics of bacteriocins from human microbiome. Environ. Microbiol. 17, 2133–2143 (2015).
Guzzetta, M., Williamson, A. & Duong, S. Clostridium sordellii as an uncommon cause of fatal toxic shock syndrome in a postpartum 33-year-old asian woman, and the need for antepartum screening for this clostridia species in the general female population. Lab. Med. 47, 251–254 (2016).
Laux, C., Peschel, A. & Krismer, B. Staphylococcus aureus colonization of the human nose and interaction with other microbiome members. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.GPP3-0029-2018 (2019).
Nakatsuji, T. et al. Development of a human skin commensal microbe for bacteriotherapy of atopic dermatitis and use in a phase 1 randomized clinical trial. Nat. Med. 27, 700–709 (2021).
Zipperer, A. et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature 535, 511–516 (2016).
Schilling, N. A. et al. Synthetic lugdunin analogues reveal essential structural motifs for antimicrobial action and proton translocation capability. Angew. Chem. Int. Ed. Engl. 58, 9234–9238 (2019).
Dhanda, G., Sarkar, P., Samaddar, S. & Haldar, J. Battle against vancomycin-resistant bacteria: recent developments in chemical strategies. J. Med. Chem. 62, 3184–3205 (2019).
Claesen, J. et al. A Cutibacterium acnes antibiotic modulates human skin microbiota composition in hair follicles. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aay5445 (2020).
Nakatsuji, T. et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aah4680 (2017).
Bessesen, M. T. et al. MRSA colonization and the nasal microbiome in adults at high risk of colonization and infection. J. Infect. 71, 649–657 (2015).
Delanghe, L. et al. The role of lactobacilli in inhibiting skin pathogens. Biochem. Soc. Trans. 49, 617–627 (2021).
Lebeer, S. et al. Selective targeting of skin pathobionts and inflammation with topically applied lactobacilli. Cell Rep Med. 15, 100521 (2022).
Yim, S. S. & Wang, H. H. Exploiting interbacterial antagonism for microbiome engineering. Curr. Opin. Biomed. Eng. 19, 100307 (2021).
Albright, M. B. N. et al. Solutions in microbiome engineering: prioritizing barriers to organism establishment. ISME J. 16, 331–338 (2022).
Corr, S. C. et al. Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc. Natl Acad. Sci. USA 104, 7617–7621 (2007).
Leventhal, D. S. et al. Immunotherapy with engineered bacteria by targeting the STING pathway for anti-tumor immunity. Nat. Commun. 11, 2739 (2020).
Bloom, S. M. et al. Cysteine dependence of Lactobacillus iners is a potential therapeutic target for vaginal microbiota modulation. Nat. Microbiol. 7, 434–450 (2022).
Espah Borujeni, A., Channarasappa, A. S. & Salis, H. M. Translation rate is controlled by coupled trade-offs between site accessibility, selective RNA unfolding and sliding at upstream standby sites. Nucleic Acids Res. 42, 2646–2659 (2014).
Patiny, L. & Borel, A. ChemCalc: a building block for tomorrow’s chemical infrastructure. J. Chem. Inf. Model. 53, 1223–1228 (2013).
Piard, J. C., Muriana, P. M., Desmazeaud, M. J. & Klaenhammer, T. R. Purification and partial characterization of Lacticin 481, a lanthionine-containing bacteriocin produced by Lactococcus lactis subsp. lactis CNRZ 481. Appl. Environ. Microbiol. 58, 279–284 (1992).
Acknowledgements
We thank C. A. Sheahan (Harvard Medical School) for help with nuclear magnetic resonance. This research was funded by a research award from Novartis Institute for BioMedical Research (Cambridge, USA), US Defense Advanced Research Projects Agency’s Living Foundries programme award HR0011-15-C-0084, and the Banting Fellowships Program (A.M.K).
Author information
Authors and Affiliations
Contributions
Z.Z., A.M.K., P.S., J.C. and C.A.V. conceived the study and designed the experiments. Z.Z. and A.M.K. performed experiments. E.G. performed the bioinformatics. Z.Z., A.M.K. and C.A.V. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Marnix Medema and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Network map of sequence similarities between RiPP gene clusters from human microbiome genomes.
antiSMASH 4.0 was used to identify BGCs from 2,229 HMP genome sequences. 2,248 RiPP BGCs were then clustered using BiG-SCAPE with default cutoff parameters (0.30) and visualized with Cytoscape. Nodes represent individual clusters colored by biosynthetic class. BGC nodes with similar cluster architecture are attached by edges using cutoff values as described in Methods.
Extended Data Fig. 2 Network map of sequence similarities between RiPP gene clusters selected for expression.
The networks for (a) lasso peptides and (b) lanthipeptides are shown. BGC nodes from the same network generated in Extended Data Fig. 1 were extracted to create the simplified visualization shown here. BGCs selected for gene synthesis and heterologous expression are colored with green circles.
Extended Data Fig. 3 Genome mining and reconstruction of lasso peptide gene clusters.
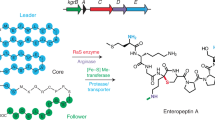
(a) An example of a native gene cluster. The precursor peptide is expressed, the leader cleaved (LasB), and the resulting free amine cyclized to the side chain of an Asp/Glu residue (LasC). The grey gene encodes a transporter. To build the synthetic gene cluster, each gene is codon optimized (dashed lines) and placed under the control of a synthetic ribozyme insulator, RBS, and terminator. The precursor peptide is fused to RSTC. (b) An example synthetic gene cluster is shown along with synthetic genetic parts (Supplementary Table 4). The components of the RSTN tag are shown (black, His6-tag; dark blue, SUMO; orange, thrombin protease site; white, linkers). A thrombin protease site (orange) is added before SUMO (dark blue) in the precursor peptide, and this leaves a C-terminal RVLP on the core (black circle). Dashed lines in the genome indicate the presence of other inducible systems in the E. coli Marionette X strain.
Extended Data Fig. 4 Mass spectrometry data for LANII-286.
(a) Peptide and biosynthetic gene cluster details. The genes synthesized are shown in blue and sequences are provided in Supplementary Table 3. (b) Proposed annotated structure. (c) High-res mass spectrometry traces of peptides selected for tandem MS/MS. An arrow indicates the monoisotopic mass chosen for fragmentation (m/z 1793.7391). (d) N-ethylmaleimide (NEM) labeling of peptides. Top trace, TCEP-treated peptide without the addition of NEM (black). Bottom trace, TCEP-treated peptides with the addition of NEM (purple). Each NEM adduct would lead to an increase in mass of 125.05 Da. For each peptide, the predicted masses for up to the maximum number of NEM adducts were used to generate extracted ion chromatograms. An extraction window of the mass +/- 0.25 Da was used. The grey bars correspond to the predicted masses for 1, 2,… n non-cyclized residues (no grey bars means that no adduct is detected, meaning the product is completely cyclized). (e) MS/MS spectrum of the modified peptide. The amino acid sequence of the peptide is shown with observed b and y ions mapped. The blue lines capped with a blue dot mark the labeled experimental peaks.
Extended Data Fig. 5 Mass spectrometry data for LANII-287.
(a) Peptide and biosynthetic gene cluster details. The genes synthesized are shown in blue and sequences are provided in Supplementary Table 3. (b) Proposed annotated structure. (c) High-res mass spectrometry traces of peptides selected for tandem MS/MS. An arrow indicates the monoisotopic mass chosen for fragmentation (m/z 1776.2837). (d) N-ethylmaleimide (NEM) labeling of peptides. Top trace, TCEP-treated peptide without the addition of NEM (black). Bottom trace, TCEP-treated peptides with the addition of NEM (purple). Each NEM adduct would lead to an increase in mass of 125.05 Da. For each peptide, the predicted masses for up to the maximum number of NEM adducts were used to generate extracted ion chromatograms. An extraction window of the mass +/- 0.25 Da was used. (e) MS/MS spectrum of the modified peptide. The amino acid sequence of the peptide is shown with observed b and y ions mapped. The blue lines capped with a blue dot mark the labeled experimental peaks.
Extended Data Fig. 6 Mass spectrometry data for LANII-916.
(a) Peptide and biosynthetic gene cluster details. The genes synthesized are shown in blue and sequences are provided in Supplementary Table 3. (b) Proposed annotated structure. (c) High-res mass spectrometry traces of peptides selected for tandem MS/MS. An arrow indicates the monoisotopic mass chosen for fragmentation (m/z 1163.8450). (d) N-ethylmaleimide (NEM) labeling of peptides. Top trace, TCEP-treated peptide without the addition of NEM (black). Bottom trace, TCEP-treated peptides with the addition of NEM (purple). Each NEM adduct would lead to an increase in mass of 125.05 Da. For each peptide, the predicted masses for up to the maximum number of NEM adducts were used to generate extracted ion chromatograms. An extraction window of the mass +/- 0.25 Da was used. The grey bars correspond to the predicted masses for 1, 2,… n non-cyclized residues (no grey bars means that no adduct is detected, meaning the product is completely cyclized). (e) MS/MS spectrum of the modified peptide. The amino acid sequence of the peptide is shown with observed b and y ions mapped. The blue lines capped with a blue dot mark the labeled experimental peaks.
Extended Data Fig. 7 Mass spectrometry data for LANII-417.
(a) Peptide and biosynthetic gene cluster details. The genes synthesized are shown in blue and sequences are provided in Supplementary Table 3. (b) Proposed annotated structure. (c) High-res mass spectrometry traces of peptides selected for tandem MS/MS. An arrow indicates the monoisotopic mass chosen for fragmentation (m/z 1381.5389). (d) N-ethylmaleimide (NEM) labeling of peptides. Top trace, TCEP-treated peptide without the addition of NEM (black). Bottom trace, TCEP-treated peptides with the addition of NEM (purple). Each NEM adduct would lead to an increase in mass of 125.05 Da. For each peptide, the predicted masses for up to the maximum number of NEM adducts were used to generate extracted ion chromatograms. An extraction window of the mass +/- 0.25 Da was used. (e) MS/MS spectrum of the modified peptide. The amino acid sequence of the peptide is shown with observed b and y ions mapped. The blue lines capped with a blue dot mark the labeled experimental peaks.
Extended Data Fig. 8 Mass spectrometry data for LANII-687.
(a) Peptide and biosynthetic gene cluster details. The genes synthesized are shown in blue and sequences are provided in Supplementary Table 3. (b) Proposed annotated structure. (c) High-res mass spectrometry traces of peptides selected for tandem MS/MS. An arrow indicates the monoisotopic mass chosen for fragmentation (m/z 1619.2318). (d) N-ethylmaleimide (NEM) labeling of peptides. Top trace, TCEP-treated peptide without the addition of NEM (black). Bottom trace, TCEP-treated peptides with the addition of NEM (purple). Each NEM adduct would lead to an increase in mass of 125.05 Da. For each peptide, the predicted masses for up to the maximum number of NEM adducts were used to generate extracted ion chromatograms. An extraction window of the mass +/- 0.25 Da was used. The grey bars correspond to the predicted masses for 1, 2,… n non-cyclized residues (no grey bars means that no adduct is detected, meaning the product is completely cyclized). (e) MS/MS spectrum of the modified peptide. The amino acid sequence of the peptide is shown with observed b and y ions mapped. The blue lines capped with a blue dot mark the labeled experimental peaks.
Extended Data Fig. 9 Mass spectrometry data for LANII-691.
(a) Peptide and biosynthetic gene cluster details. The genes synthesized are shown in blue and sequences are provided in Supplementary Table 3. (b) Proposed annotated structure. (c) High-res mass spectrometry traces of peptides selected for tandem MS/MS. An arrow indicates the monoisotopic mass chosen for fragmentation (m/z 1426.6661). (d) N-ethylmaleimide (NEM) labeling of peptides. Top trace, TCEP-treated peptide without the addition of NEM (black). Bottom trace, TCEP-treated peptides with the addition of NEM (purple). Each NEM adduct would lead to an increase in mass of 125.05 Da. (e) MS/MS spectrum of the modified peptide. The amino acid sequence of the peptide is shown with observed b and y ions mapped. The blue lines capped with a blue dot mark the labeled experimental peaks.
Extended Data Fig. 10 MIC assay for RiPPs against human pathogens and commensals.
The antimicrobial activity of (a) LANII-286, (b) LANII-287, (c) LANII-417, (d) LANII-687, (e) LANII-691 and (f) LANII-916 were tested against human pathogens (Methods). The data points show three replicates performed on different days and the lines show the means of these data. The % Growth was calculated for each day and then these values were averaged.
Supplementary information
Supplementary Information
Supplementary Text, Figs. 1–20, Tables 1–5 and references.
Source data
Source Data Fig. 2
Antismash files for RiPP gene clusters.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
King, A.M., Zhang, Z., Glassey, E. et al. Systematic mining of the human microbiome identifies antimicrobial peptides with diverse activity spectra. Nat Microbiol 8, 2420–2434 (2023). https://doi.org/10.1038/s41564-023-01524-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01524-6