Abstract
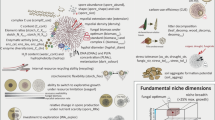
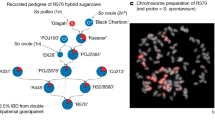
Hyaloperonospora arabidopsidis (Hpa) is an obligately biotrophic downy mildew that is routinely cultured on Arabidopsis thaliana hosts that harbour complex microbiomes. We hypothesized that the culturing procedure proliferates Hpa-associated microbiota (HAM) in addition to the pathogen and exploited this model system to investigate which microorganisms consistently associate with Hpa. Using amplicon sequencing, we found nine bacterial sequence variants that are shared between at least three out of four Hpa cultures in the Netherlands and Germany and comprise 34% of the phyllosphere community of the infected plants. Whole-genome sequencing showed that representative HAM bacterial isolates from these distinct Hpa cultures are isogenic and that an additional seven published Hpa metagenomes contain numerous sequences of the HAM. Although we showed that HAM benefit from Hpa infection, HAM negatively affect Hpa spore formation. Moreover, we show that pathogen-infected plants can selectively recruit HAM to both their roots and shoots and form a soil-borne infection-associated microbiome that helps resist the pathogen. Understanding the mechanisms by which infection-associated microbiomes are formed might enable breeding of crop varieties that select for protective microbiomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study and isolates are available from the corresponding author upon reasonable request. Moreover, the raw amplicon sequence data generated by this study are available at https://www.ncbi.nlm.nih.gov/sra/PRJNA944652; raw WGS data are available at http://www.ncbi.nlm.nih.gov/bioproject/1011197; genome assemblies generated in this study are available at http://www.ncbi.nlm.nih.gov/bioproject/1011284. Whenever possible, post-processing amplicon sequencing (ASV) count tables are also included together with the processing code at the Zenodo-archived GitHub page (https://doi.org/10.5281/zenodo.8307753). Source data are provided with this paper.
Code availability
The code used to analyse data and generate figures can be found at https://doi.org/10.5281/zenodo.8307753. No unpublished algorithms or methods were used.
References
Kamoun, S. et al. The top 10 oomycete pathogens in molecular plant pathology. Mol. Plant Pathol. 16, 413–434 (2015).
Thines, M. & Choi, Y.-J. Evolution, diversity, and taxonomy of the Peronosporaceae, with focus on the genus Peronospora. Phytopathology 106, 6–18 (2016).
Ngou, B. P. M., Ding, P. & Jones, J. D. G. Thirty years of resistance: zig-zag through the plant immune system. Plant Cell 34, 1447–1478 (2022).
Parker, J. E. et al. Characterization of eds1, a mutation in Arabidopsis suppressing resistance to Peronospora parasitica specified by several different RPP genes. Plant Cell 8, 2033–2046 (1996).
Holub, E. B. in The Downy Mildews—Genetics, Molecular Biology and Control (eds Lebeda, A., Spencer-Phillips, P. T. N. & Cooke, B. N.) 91–109 (Springer, 2008).
Vorholt, J. A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 10, 828–840 (2012).
Bakker, P. A. H. M., Berendsen, R. L., Doornbos, R. F., Wintermans, P. C. A. & Pieterse, C. M. J. The rhizosphere revisited: root microbiomics. Front. Plant Sci. 4, 165 (2013).
Rolfe, S. A., Griffiths, J. & Ton, J. Crying out for help with root exudates: adaptive mechanisms by which stressed plants assemble health-promoting soil microbiomes. Curr. Opin. Microbiol. 49, 73–82 (2019).
Liu, H., Brettell, L. E. & Singh, B. Linking the phyllosphere microbiome to plant health. Trends Plant Sci. 25, 841–844 (2020).
Pieterse, C. M. J. et al. Induced systemic resistance by beneficial microbes. Annu. Rev. Phytopathol. 52, 347–375 (2014).
Raaijmakers, J. M. & Mazzola, M. Diversity and natural functions of antibiotics produced by beneficial and plant pathogenic bacteria. Annu. Rev. Phytopathol. 50, 403–424 (2012).
Teixeira, P. J. P. L., Colaianni, N. R., Fitzpatrick, C. R. & Dangl, J. L. Beyond pathogens: microbiota interactions with the plant immune system. Curr. Opin. Microbiol. 49, 7–17 (2019).
Berendsen, R. L. et al. Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J. 12, 1496–1507 (2018).
Yuan, J. et al. Root exudates drive the soil-borne legacy of aboveground pathogen infection. Microbiome 6, 156 (2018).
Wen, T., Zhao, M., Yuan, J., Kowalchuk, G. A. & Shen, Q. Root exudates mediate plant defense against foliar pathogens by recruiting beneficial microbes. Soil Ecol. Lett. 3, 42–51 (2021).
Wang, Q. et al. Tea plants with gray blight have altered root exudates that recruit a beneficial rhizosphere microbiome to prime immunity against aboveground pathogen infection. Front. Microbiol. 12, 3775 (2021).
Friman, J. et al. Shoot and root insect herbivory change the plant rhizosphere microbiome and affects cabbage–insect interactions through plant–soil feedback. New Phytol. 232, 2475–2490 (2021).
Bakker, P. A. H. M., Pieterse, C. M. J., de Jonge, R. & Berendsen, R. L. The soil-borne legacy. Cell 172, 1178–1180 (2018).
Raaijmakers, J. M. & Mazzola, M. Soil immune responses. Science 352, 1392–1393 (2016).
Vismans, G. et al. Coumarin biosynthesis genes are required after foliar pathogen infection for the creation of a microbial soil-borne legacy that primes plants for SA-dependent defenses. Sci. Rep. 12, 22473 (2022).
Carrión, V. J. et al. Pathogen-induced activation of disease-suppressive functions in the endophytic root microbiome. Science 366, 606–612 (2019).
Schlatter, D., Kinkel, L., Thomashow, L., Weller, D. & Paulitz, T. Disease suppressive soils: new insights from the soil microbiome. Phytopathology 107, 1284–1297 (2017).
McDowell, J. M., Hoff, T., Anderson, R. G. & Deegan, D. in Plant Immunity: Methods and Protocols (ed McDowell, J. M.) 137–151 (Springer, 2011).
Holub, E. B., Beynon, J. L. & Crute, I. R. Phenotypic and genotypic characterization of interactions between isolates of Peronospora parasitica and accessions of Arabidopsis thaliana. Mol. Plant Microbe Interact. 7, 223–239 (1994).
Parker, J. E. et al. Phenotypic characterization and molecular mapping of the Arabidopsis thaliana locus RPP5, determining disease resistance to Peronospora parasitica. Plant J. 4, 821–831 (1993).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Lapin, D., Meyer, R. C., Takahashi, H., Bechtold, U. & Van den Ackerveken, G. Broad-spectrum resistance of Arabidopsis C24 to downy mildew is mediated by different combinations of isolate-specific loci. New Phytol. 196, 1171–1181 (2012).
Nemri, A. et al. Genome-wide survey of Arabidopsis natural variation in downy mildew resistance using combined association and linkage mapping. Proc. Natl Acad. Sci. USA 107, 10302–10307 (2010).
Olm, M. R. et al. InStrain enables population genomic analysis from metagenomic data and rigorous detection of identical microbial strains. Nat. Biotechnol. 39, 727–736 (2021).
Baxter, L. et al. Signatures of adaptation to obligate biotrophy in the Hyaloperonospora arabidopsidis genome. Science 330, 1549–1551 (2010).
Asai, S. et al. A downy mildew effector evades recognition by polymorphism of expression and subcellular localization. Nat. Commun. 9, 5192 (2018).
Schaeffer, L., Pimentel, H., Bray, N., Melsted, P. & Pachter, L. Pseudoalignment for metagenomic read assignment. Bioinformatics 33, 2082–2088 (2017).
Parker, J. E. et al. The Arabidopsis downy mildew resistance gene RPP5 shares similarity to the toll and interleukin-1 receptors with N and L6. Plant Cell 9, 879–894 (1997).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Becking, L. G. M. B. Geobiologie of inleiding tot de milieukunde (W.P. Van Stockum & Zoon, 1934).
De Zutter, N. et al. Shifts in the rhizobiome during consecutive in planta enrichment for phosphate‐solubilizing bacteria differentially affect maize P status. Microb. Biotechnol. 14, 1594–1612 (2021).
Harbort, C. J. et al. Root-secreted coumarins and the microbiota interact to improve iron nutrition in Arabidopsis. Cell Host Microbe 28, 825–837 (2020).
Santos-Medellín, C. et al. Prolonged drought imparts lasting compositional changes to the rice root microbiome. Nat. Plants 7, 1065–1077 (2021).
Xu, L. et al. Genome-resolved metagenomics reveals role of iron metabolism in drought-induced rhizosphere microbiome dynamics. Nat. Commun. 12, 3209 (2021).
Monohon, S. J., Manter, D. K. & Vivanco, J. M. Conditioned soils reveal plant-selected microbial communities that impact plant drought response. Sci. Rep. 11, 21153 (2021).
Yuan, Y., Brunel, C., van Kleunen, M., Li, J. & Jin, Z. Salinity-induced changes in the rhizosphere microbiome improve salt tolerance of Hibiscus hamabo. Plant Soil 443, 525–537 (2019).
Gao, M. et al. Disease-induced changes in plant microbiome assembly and functional adaptation. Microbiome 9, 187 (2021).
Mendes, R. et al. Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332, 1097–1100 (2011).
Weller, D. M., Raaijmakers, J. M., Gardener, B. B. M. & Thomashow, L. S. Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu. Rev. Phytopathol. 40, 309–348 (2002).
Liu, X. et al. Phyllosphere microbiome induces host metabolic defence against rice false-smut disease. Nat. Microbiol. 8, 1419–1433 (2023).
Bass, D., Stentiford, G. D., Wang, H.-C., Koskella, B. & Tyler, C. R. The pathobiome in animal and plant diseases. Trends Ecol. Evol. 34, 996–1008 (2019).
Vayssier-Taussat, M. et al. Shifting the paradigm from pathogens to pathobiome: new concepts in the light of meta-omics. Front. Cell. Infect. Microbiol. 4, 29 (2014).
Xu, S. et al. Fusarium fruiting body microbiome member Pantoea agglomerans inhibits fungal pathogenesis by targeting lipid rafts. Nat. Microbiol. 7, 831–843 (2022).
Aarts, N. et al. Different requirements for EDS1 and NDR1 by disease resistance genes define at least two R gene-mediated signaling pathways in Arabidopsis. Proc. Natl Acad. Sci. USA 95, 10306–10311 (1998).
Pieterse, C. M. J., Van Wees, S. C. M., Hoffland, E., Van Pelt, J. A. & Van Loon, L. C. Systemic resistance in Arabidopsis induced by biocontrol bacteria is independent of salicylic acid accumulation and pathogenesis-related gene expression. Plant Cell 8, 1225–1237 (1996).
Lindsey III, B. E., Rivero, L., Calhoun, C. S., Grotewold, E. & Brkljacic, J. Standardized method for high-throughput sterilization of Arabidopsis seeds. J. Vis. Exp. 17, 56587 (2017).
Vismans, G., Spooren, J., Pieterse, C. M. J., Bakker, P. A. H. M. & Berendsen, R. L. in The Plant Microbiome: Methods and Protocols Vol. 2232 (eds Carvalhais, L. C. & Dennis, P. G.) 209–218 (Springer, 2021).
Anderson, R. G. & McDowell, J. M. A PCR assay for the quantification of growth of the oomycete pathogen Hyaloperonospora arabidopsidis in Arabidopsis thaliana. Mol. Plant Pathol. 16, 893–898 (2015).
de Muinck, E. J., Trosvik, P., Gilfillan, G. D., Hov, J. R. & Sundaram, A. Y. M. A novel ultra high-throughput 16S rRNA gene amplicon sequencing library preparation method for the Illumina HiSeq platform. Microbiome 5, 68 (2017).
Lundberg, D. S., Yourstone, S., Mieczkowski, P., Jones, C. D. & Dangl, J. L. Practical innovations for high-throughput amplicon sequencing. Nat. Methods 10, 999–1002 (2013).
Ihrmark, K. et al. New primers to amplify the fungal ITS2 region—evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 82, 666–677 (2012).
Agler, M. T., Mari, A., Dombrowski, N., Haquard, S. & Kemen, E. M. New insights in host-associated microbial diversity with broad and accurate taxonomic resolution. Preprint at bioRxiv https://doi.org/10.1101/050005 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet j. 17, 10–12 (2011).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
King, E. O., Ward, M. K. & Raney, D. E. Two simple media for the demonstration of pyocyanin and fluorescin. J. Lab. Clin. Med. 44, 301–307 (1954).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Acknowledgements
This study was sponsored by the TopSector Horticulture and Starting Materials (TKI grant number 1605-106), the Dutch Research Council (NWO) through the Gravitation programme MiCRop (grant number 024.004.014) and the XL programme ‘Unwiring beneficial functions and regulatory networks in the plant endosphere’ (grant number OCENW.GROOT.2019.063). The TKI project was carried out in collaboration with four industrial partners; DSM, Enza Zaden, Pop Vriend Seeds and RijkZwaan Breeding B.V. We thank J. Parker for providing Col-0 RPP5 and Ler rpp5 Arabidopsis seeds, P. Bakker and R. de Jonge for valuable input on experimental designs and bioinformatic approaches, and J. Elberse, D. Duijker, L. Pronk, C. Molina Ruiz, M. Alderkamp, X. Pan, L. Wagenaar and T. Tarrant for excellent technical assistance.
Author information
Authors and Affiliations
Contributions
P.G., J.S., G.A., C.M.J.P. and R.L.B. designed the experiments and wrote the paper. R.L.B., G.A. and C.M.J.P. supervised the project and edited the paper. P.G. performed the experiments and data analysis shown in Figs. 1–4. D.L. performed the experiments in Cologne, Germany (Fig. 2). P.G. and N.E. performed the experiments shown in Fig. 5a,b. J.S. performed the experiments shown in Fig. 5c–e. P.G. and J.S. performed the experiments and data analysis shown in Fig. 6. K.C.M.B. and A.A. provided technical support during the conduct of the experiments and in the maintenance of Hpa and gnoHpa cultures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Yang Bai, Mengcen Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effect of water- and wind-transmitted Hpa infections on bacterial phyllosphere community structure.
(a) Quantification of Hpa DNA relative to Arabidopsis DNA by qPCR for samples taken 7 days post-inoculation. Bars represent average Hpa abundance. Error bars show standard error. N = 7 biologically independent samples. PCoA ordination plot based on Bray-Curtis dissimilarities of the (b) bacterial phyllosphere communities and (c) fungal phyllosphere communities of Arabidopsis thaliana Col-0 untreated control plants (blue symbols), or plants inoculated with Hpa spores via wind (orange stroke) or water (orange fill).
Extended Data Fig. 2 Compatibility of Noco2 and Cala2 with susceptible and resistant Arabidopsis accessions.
qPCR quantification of Hpa abundance in the susceptible and resistant Hpa-Arabidopsis interactions indicated, confirming that C24 is resistant to both Noco2 and Cala2, that Col-0 is susceptible to Noco2, that Ler is susceptible to Cala2, and that Pro-0 is susceptible to both Noco2 and Cala2. qPCR quantification was performed on total genomic DNA from inoculated leaves that were also used for 16 S rDNA amplicon sequencing (Fig. 1). Hpa abundance was calculated as a ratio of the levels of ACTIN in Hpa and Arabidopsis. Bars represent average ratios, error bars represent standard error. N = 4 biologically independent samples, except for C24 mock-, Noco2-, and Cala2- treated (N = 3 biologically independent samples).
Extended Data Fig. 3 Schematic overview of the ‘9-passages experiment’, which tests the effect of the removal of Hpa on its associated microbiome.
A uniform HAM (uHAM) containing a mix of of Noco2 and Cala2 spore suspensions was spray-inoculated on different Arabidopsis genotypes to selectively remove Noco2 and/or Cala2 from the microbiome (HAM) that travels together with these Hpa isolates upon passaging to new host plants. One week post-inoculation, Col-0 (Lineage 1) and Ler plants (Lineage 2) sporulated with Noco2 and Cala2, respectively, and Col-0/RPP5 transgenic plants (Lineage 3) did not sporulate. From each Arabidopsis genotype, a leaf wash-off was obtained, containing Noco2, Cala2, or no Hpa, and sprayed on eight pots containing small fields of Ler/rpp5 mutant plants, which are susceptible to both Noco2 and Cala2. All pots were then placed in individual plastic containers, to prevent cross-contamination between pots. One week post-inoculation, the Ler/rpp5 plants that were inoculated with Noco2 (Lineage 1) or Cala2 (Lineage 2), sporulated, while the Ler/rpp5 plants that were inoculated with the leaf wash-off without Hpa spores (Lineage 3) did not display disease symptoms. From each individual pot, the leaf wash-off was sprayed on a new pot containing Ler/rpp5 plants, thereby propagating eight separate phyllosphere microbiomes or Hpa cultures per lineage. This process was maintained for nine consecutive weeks, allowing eight separate lineages of Noco2, Cala2, or the uHAM without Hpa to develop independently. Eight untreated control pots with Ler/rpp5 plants were included for all planting cycles.
Extended Data Fig. 4 The Hpa-culture bacterial community is largely unaffected by removal of Hpa, but nonetheless there are community shifts in the absence of Hpa.
PcoA plots based on Bray-Curtis dissimilarities of (a) all samples from Lineages 1–3 and untreated plants of passages 1 (circles), passage 5 (triangles) and passage 9 (diamonds); and of all inoculated samples of Lineage 1–3 from (b) passage 1, (c) passage 5, and (d) passage 9. Plants were left untreated (black symbols) or were inoculated with leaf wash-offs from Lineage 1 containing Noco2 (orange symbols), from Lineage 2 containing Cala2 (green symbols), or from Lineage 3 which remained Hpa free (blue symbols).
Extended Data Fig. 5 Setup of soil-borne legacy experiments.
A conditioning population of two-week-old Arabidopsis thaliana Col- seedlings was inoculated with mock, Hpa Noco2 or gnoHpa Noco2. After one week of infection, shoots were cut-off and a response population of plants was directly sown on the conditioned soil and again mock- Hpa- or gnoHpa-inoculated. Figure created with BioRender.com.
Supplementary information
Supplementary Information
Supplementary Figs. 1–18 and Tables 1–11.
Source data
Source Data Fig. 1
(Grouped) ASV abundance data.
Source Data Fig. 2
(Grouped) ASV abundance data and ASV-of-interest abundance data.
Source Data Fig. 4
ASV abundance data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Goossens, P., Spooren, J., Baremans, K.C.M. et al. Obligate biotroph downy mildew consistently induces near-identical protective microbiomes in Arabidopsis thaliana. Nat Microbiol 8, 2349–2364 (2023). https://doi.org/10.1038/s41564-023-01502-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01502-y
This article is cited by
-
Leaf microbiome dysbiosis triggered by T2SS-dependent enzyme secretion from opportunistic Xanthomonas pathogens
Nature Microbiology (2024)
-
Disease resistance through M genes
Nature Plants (2024)