Abstract
Cell-intrinsic defences constitute the first line of defence against intracellular pathogens. The guanosine triphosphatase RAB32 orchestrates one such defence response against the bacterial pathogen Salmonella, through delivery of antimicrobial itaconate. Here we show that the Parkinson’s disease-associated leucine-rich repeat kinase 2 (LRRK2) orchestrates this defence response by scaffolding a complex between RAB32 and aconitate decarboxylase 1, which synthesizes itaconate from mitochondrial precursors. Itaconate delivery to Salmonella-containing vacuoles was impaired and Salmonella replication increased in LRRK2-deficient cells. Loss of LRRK2 also restored virulence of a Salmonella mutant defective in neutralizing this RAB32-dependent host defence pathway in mice. Cryo-electron tomography revealed tether formation between Salmonella-containing vacuoles and host mitochondria upon Salmonella infection, which was significantly impaired in LRRK2-deficient cells. This positions LRRK2 centrally within a host defence mechanism, which may have favoured selection of a common familial Parkinson’s disease mutant allele in the human population.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Subtomogram-average density maps and the raw cryo-ET tilt series have been deposited in EMDB (deposition ID numbers: EMD-41046, EMD-41047 and EMPIAR-11577). The rest of the data are available in the main text, supplementary materials and auxiliary files. Source data are provided with this paper.
References
Randow, F., MacMicking, J. & James, L. Cellular self-defense: how cell-autonomous immunity protects against pathogens. Science 340, 701–706 (2013).
Spanò, S. & Galán, J. A Rab32-dependent pathway contributes to Salmonella Typhi host restriction. Science 338, 960–963 (2012).
Tang, B. Rab32/38 and the xenophagic restriction of intracellular bacteria replication. Microbes Infect. 18, 595–603 (2016).
Li, Y. et al. Analysis of the Rab GTPase interactome in dendritic cells reveals anti-microbial functions of the Rab32 complex in bacterial containment. Immunity 44, 422–437 (2016).
Baldassarre, M. et al. The Rab32/BLOC-3-dependent pathway mediates host defense against different pathogens in human macrophages. Sci. Adv. 7, eabb1795 (2021).
Chen, M. et al. Itaconate is an effector of a Rab GTPase cell-autonomous host defense pathway against Salmonella. Science 369, 450–455 (2020).
Schuster, E. M. et al. TFEB induces mitochondrial itaconate synthesis to suppress bacterial growth in macrophages. Nat. Metab. 4, 856–866 (2022).
Michelucci, A. et al. Immune-responsive gene 1 protein links metabolism to immunity by catalyzing itaconic acid production. Proc. Natl Acad. Sci. USA 110, 7820–7825 (2013).
Ruetz, M. et al. Itaconyl-CoA forms a stable biradical in methylmalonyl-CoA mutase and derails its activity and repair. Science 366, 589–593 (2019).
Patel, T. & McFadden, B. Caenorhabditis elegans and Ascaris suum: inhibition of isocitrate lyase by itaconate. Exp. Parasitol. 44, 262–268 (1978).
McFadden, B. & Purohit, S. Itaconate, an isocitrate lyase-directed inhibitor in Pseudomonas indigofera. J. Bacteriol. 131, 136–144 (1977).
Cordes, T. et al. Immunoresponsive Gene 1 and itaconate inhibit succinate dehydrogenase to modulate intracellular succinate levels. J. Biol. Chem. 291, 14274–14284 (2016).
Wilson, R. & Maloy, S. Isolation and characterization of Salmonella Typhimurium glyoxylate shunt mutants. J. Bacteriol. 169, 3029–3034 (1987).
Fang, F., Libby, S., Castor, M. & Fung, A. Isocitrate lyase (AceA) is required for Salmonella persistence but not for acute lethal infection in mice. Infect. Immun. 73, 2547–2549 (2005).
McKinney, J. et al. Persistence of Mycobacterium tuberculosis in macrophages and mice requires the glyoxylate shunt enzyme isocitrate lyase. Nature 406, 735–738 (2000).
Mercado-Lubo, R., Gauger, E., Leatham, M., Conway, T. & Cohen, P. A Salmonella enterica serovar Typhimurium succinate dehydrogenase/fumarate reductase double mutant is avirulent and immunogenic in BALB/c mice. Infect. Immun. 76, 1128–1134 (2008).
Yimga, M. et al. Role of gluconeogenesis and the tricarboxylic acid cycle in the virulence of Salmonella enterica serovar Typhimurium in BALB/c mice. Infect. Immun. 74, 1130–1140 (2006).
Pecsi, I. et al. Essentiality of succinate dehydrogenase in Mycobacterium smegmatis and its role in the generation of the membrane potential under hypoxia. MBio 5, pii: e01093–01014 (2014).
Hartman, T. et al. Succinate dehydrogenase is the regulator of respiration in Mycobacterium tuberculosis. PLoS Pathog. 10, e1004510 (2014).
Wheeler, P. Oxidation of carbon sources through the tricarboxylic acid cycle in Mycobacterium leprae grown in armadillo liver. J. Gen. Microbiol. 130, 381–389 (1984).
Reddick, L. & Alto, N. Bacteria fighting back: how pathogens target and subvert the host innate immune system. Mol. Cell. 54, 321–328 (2014).
Finlay, B. & McFadden, G. Anti-immunology: evasion of the host immune system by bacterial and viral pathogens. Cell 124, 767–782 (2006).
Spanò, S., Gao, X., Hannemann, S., Lara-Tejero, M. & Galán, J. A bacterial pathogen targets a host Rab-family GTPase defense pathway with a GAP. Cell Host Microbe 19, 216–226 (2016).
Spano, S., Liu, X. & Galan, J. E. Proteolytic targeting of Rab29 by an effector protein distinguishes the intracellular compartments of human-adapted and broad-host Salmonella. Proc. Natl Acad. Sci. USA 108, 18418–18423 (2011).
Sasikaran, J., Ziemski, M., Zadora, P., Fleig, A. & Berg, I. Bacterial itaconate degradation promotes pathogenicity. Nat. Chem. Biol. 10, 371–377 (2014).
Martin, W. R., Frigan, F. & Bergman, E. H. Noninductive metabolism of itaconic acid by Pseudomonas and Salmonella species. J. Bacteriol. 82, 905–908 (1961).
Parkhill, J. et al. Complete genome sequence of a multiple drug resistant Salmonella enterica serovar Typhi CT18. Nature 413, 848–852 (2001).
Waschbüsch, D. et al. LRRK2 transport is regulated by its novel interacting partner Rab32. PLoS ONE 9, e111632 (2014).
McGrath, E., Waschbüsch, D., Baker, B. & Khan, A. LRRK2 binds to the Rab32 subfamily in a GTP-dependent manner via its armadillo domain. Small GTPases 12, 133–146 (2021).
Bui, M. et al. Rab32 modulates apoptosis onset and mitochondria-associated membrane (MAM) properties. J. Biol. Chem. 285, 31590–31602 (2010).
Zhang, F. et al. Identification of two new loci at IL23R and RAB32 that influence susceptibility to leprosy. Nat. Genet. 43, 1247–1251 (2011).
Singh, A., Zhi, L. & Zhang, H. LRRK2 and mitochondria: recent advances and current views. Brain Res. 1702, 96–104 (2019).
Gardet, A. et al. LRRK2 is involved in the IFN-gamma response and host response to pathogens. J. Immunol. 185, 5577–5585 (2010).
Shutinoski, B. et al. Lrrk2 alleles modulate inflammation during microbial infection of mice in a sex-dependent manner. Sci. Transl. Med. 11, eaas9292 (2019).
Liu, W. et al. LRRK2 promotes the activation of NLRC4 inflammasome during Salmonella Typhimurium infection. J. Exp. Med. 214, 3051–3066 (2017).
Weindel, C. et al. LRRK2 maintains mitochondrial homeostasis and regulates innate immune responses to Mycobacterium tuberculosis. eLife 9, e51071 (2020).
Fava, V. et al. A missense LRRK2 variant is a risk factor for excessive inflammatory responses in leprosy. PLoS Negl. Trop. Dis. 10, e0004412 (2016).
Gao, Y. et al. The emerging role of Rab GTPases in the pathogenesis of Parkinson’s disease. Mov. Disord. 33, 196–207 (2018).
Manzanillo, P. et al. The ubiquitin ligase parkin mediates resistance to intracellular pathogens. Nature 7468, 512–516 (2013).
Ali, S. et al. PARK2/PACRG polymorphisms and susceptibility to typhoid and paratyphoid fever. Clin. Exp. Immunol. 144, 425–431 (2006).
Pickrell, A. & Youle, R. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85, 257–273 (2015).
Nichols, R. LRRK2 phosphorylation. Adv. Neurobiol. 14, 51–70 (2017).
Galán, J. E. & Curtiss, R. III Cloning and molecular characterization of genes whose products allow Salmonella Typhimurium to penetrate tissue culture cells. Proc. Natl Acad. Sci. USA 86, 6383–6387 (1989).
Galan, J. E., Ginocchio, C. & Costeas, P. Molecular and functional characterization of the Salmonella invasion gene invA: homology of InvA to members of a new protein family. J. Bacteriol. 174, 4338–4349 (1992).
Reith, A. et al. GSK2578215A, a potent and highly selective 2-arylmethyloxy-5-substitutent-N-arylbenzamide LRRK2 kinase inhibitor. Bioorg. Med. Chem. Lett. 22, 5625–5629 (2012).
Leschziner, A. & Reck-Peterson, S. Structural biology of LRRK2 and its interaction with microtubules. Mov. Disord. 36, 2494–2504 (2021).
Harvey, K. & Outeiro, T. The role of LRRK2 in cell signalling. Biochem. Soc. Trans. 47, 197–207 (2019).
Degrandi, D., Hoffmann, R., Beuter-Gunia, C. & Pfeffer, K. The proinflammatory cytokine-induced IRG1 protein associates with mitochondria. J. Interferon Cytokine Res. 29, 55–67 (2009).
Szeligowski, R. et al. Molecular evolution of IRG1 shapes itaconate production in metazoans and alleviates the “double-edged dilemma” of innate immune defense. Preprint at bioRxiv https://doi.org/10.1101/2022.06.17.496652 (2022).
Martin, I. et al. Ribosomal protein s15 phosphorylation mediates LRRK2 neurodegeneration in Parkinson’s disease. Cell 157, 472–485 (2014).
Lee, J. et al. Parkinson’s disease-associated LRRK2-G2019S mutant acts through regulation of SERCA activity to control ER stress in astrocytes. Acta Neuropathol. Commun. 7, 68 (2019).
Gloeckner, C. & Porras, P. Guilt-by-association—functional insights gained from studying the LRRK2 interactome. Front. Neurosci. 14, 485 (2020).
Mosaoa, R., Kasprzyk-Pawelec, A., Fernandez, H. & Avantaggiati, M. The mitochondrial citrate carrier SLC25A1/CIC and the fundamental role of citrate in cancer, inflammation and beyond. Biomolecules 11, https://doi.org/10.3390/biom11020141 (2021).
Aluvila, S., Sun, J., Harrison, D. H., Walters, D. E. & Kaplan, R. S. Inhibitors of the mitochondrial citrate transport protein: validation of the role of substrate binding residues and discovery of the first purely competitive inhibitor. Mol. Pharmacol. 77, 26–34 (2010).
Soubannier, V. et al. A vesicular transport pathway shuttles cargo from mitochondria to lysosomes. Curr. Biol. 22, 135–141 (2012).
Soto-Heredero, G., Baixauli, F. & Mittelbrunn, M. Interorganelle communication between mitochondria and the endolysosomal system. Front. Cell Dev. Biol. 5, 95 (2017).
Abuaita, B., Schultz, T. & O’Riordan, M. Mitochondria-derived vesicles deliver antimicrobial reactive oxygen species to control phagosome-localized Staphylococcus aureus. Cell Host Microbe 24, 625–636 (2018).
Klecker, T., Böckler, S. & Westermann, B. Making connections: interorganelle contacts orchestrate mitochondrial behavior. Trends Cell Biol. 24, 537–545 (2014).
Wong, Y., Kim, S., Peng, W. & Krainc, D. Regulation and function of mitochondria–lysosome membrane contact sites in cellular homeostasis. Trends Cell Biol. 29, 500–513 (2019).
Wozny, M. R. et al. Supramolecular architecture of the ER–mitochondria encounter structure in its native environment. Preprint at bioRxiv https://doi.org/10.1101/2022.04.12.488000 (2022).
Collado, J. & Fernandez-Busnadiego, R. Deciphering the molecular architecture of membrane contact sites by cryo-electron tomography. Biochim. Biophys. Acta Mol. Cell. Res. 1864, 1507–1512 (2017).
Daniele, T. et al. Mitochondria and melanosomes establish physical contacts modulated by Mfn2 and involved in organelle biogenesis. Curr. Biol. 24, 393–403 (2014).
Li, M., Tripathi-Giesgen, I., Schulman, B., Baumeister, W. & Wilfling, F. In situ snapshots along a mammalian selective autophagy pathway. Proc. Natl Acad. Sci. USA 120, e2221712120 (2023).
Dhekne, H. et al. LRRK2-phosphorylated Rab10 sequesters Myosin Va with RILPL2 during ciliogenesis blockade. Life Sci. Alliance 16, e202101050 (2021).
Esteves, A. & Cardoso, S. LRRK2 at the crossroad between autophagy and microtubule trafficking: insights into Parkinson’s disease. Neuroscientist 23, 16–26 (2017).
Caesar, M. et al. Leucine-rich repeat kinase 2 functionally interacts with microtubules and kinase-dependently modulates cell migration. Neurobiol. Dis. 54, 280–288 (2013).
Toyofuku, T., Okamoto, Y., Ishikawa, T., Sasawatari, S. & Kumanogoh, A. LRRK2 regulates endoplasmic reticulum–mitochondrial tethering through the PERK-mediated ubiquitination pathway. EMBO J. 39, e100875 (2020).
Rocha, E. M., Keeney, M. T., Di Maio, R., De Miranda, B. R. & Greenamyre, J. T. LRRK2 and idiopathic Parkinson’s disease. Trends Neurosci. 45, 224–236 (2022).
Benson, D. L., Matikainen-Ankney, B. A., Hussein, A. & Huntley, G. W. Functional and behavioral consequences of Parkinson’s disease-associated LRRK2-G2019S mutation. Biochem. Soc. Trans. 46, 1697–1705 (2018).
Galan, J. E. & Curtiss, R. 3rd Distribution of the invA, -B, -C, and -D genes of Salmonella Typhimurium among other Salmonella serovars: invA mutants of Salmonella Typhi are deficient for entry into mammalian cells. Infect. Immun. 59, 2901–2908 (1991).
Hoiseth, S. K. & Stocker, B. A. Aromatic-dependent Salmonella Typhimurium are non-virulent and effective as live vaccines. Nature 291, 238–239 (1981).
Kaniga, K., Bossio, J. C. & Galan, J. E. The Salmonella Typhimurium invasion genes invF and invG encode homologues of the AraC and PulD family of proteins. Mol. Microbiol. 13, 555–568 (1994).
Demarre, G. et al. A new family of mobilizable suicide plasmids based on broad host range R388 plasmid (IncW) and RP4 plasmid (IncPα) conjugative machineries and their cognate Escherichia coli host strains. Res. Microbiol. 156, 245–255 (2005).
Galán, J. E. & Curtiss, R. III Expression of Salmonella Typhimurium genes required for invasion is regulated by changes in DNA supercoiling. Infect. Immun. 58, 1879–1885 (1990).
Gibson, D. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Chang, S., Song, J. & Galán, J. Receptor-mediated sorting of typhoid toxin during its export from Salmonella Typhi-infected cells. Cell Host Microbe 20, 682–689 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Obert, S., O’Connor, R. J., Schmid, S. & Hearing, P. The adenovirus E4-6/7 protein transactivates the E2 promoter by inducing dimerization of a heteromeric E2F complex. Mol. Cell. Biol. 14, 1333–1346 (1994).
Schnitzbauer, J., Strauss, M., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Lampropoulou, V. et al. Itaconate links inhibition of succinate dehydrogenase with macrophage metabolic remodeling and regulation of inflammation. Cell Metab. 24, 158–166 (2016).
Mastronarde, D. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Morado, D., Hu, B. & Liu, J. Using Tomoauto: a protocol for high-throughput automated cryo-electron tomography. J. Vis. Exp. 107, e53608 (2016).
Kremer, J., Mastronarde, D. & McIntosh, J. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Winkler, H. 3D reconstruction and processing of volumetric data in cryo-electron tomography. J. Struct. Biol. 157, 126–137 (2007).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Chen, M. et al. Convolutional neural networks for automated annotation of cellular cryo-electron tomograms. Nat. Methods 14, 983–985 (2017).
Liu, Y.-T. et al. Isotropic reconstruction for electron tomography with deep learning. Nat. Commun. 13, 6482 (2022).
Acknowledgements
We thank T. Lam and W. Wang from the WM Keck Foundation Biotechnology Resource Laboratory at the Yale University School of Medicine for assistance with the itaconate measurements. We also thank M. Shao (Yale University) for assistance with the cryo-ET experiments. F.S. was partially supported by a fellowship from the Human Frontiers Science Program (LT000056/2020-C). This work was supported by National Institutes of Health grants R01AI152421 and R01AI087946 to J.L. and R01AI114618 and R01AI055472 to J.E.G. and a pilot grant from the Parkinson’s Foundation (PF-RCE-1946). The Proteomics Resource of the WM Keck Foundation Biotechnology Resource Laboratory was partially supported by CTSA grant number UL1TR001863 from the National Center for Advancing Translational Sciences (of the National Institutes of Health).
Author information
Authors and Affiliations
Contributions
H.L. performed the functional and biochemical experiments. D.P. conducted all the cryo-ET experiments with the assistance of H.L. and M.C. and under the direction of J.L. F.S. performed the DNA-PAINT imaging experiments; M.L.-T. performed the liquid chromatography–tandem mass spectrometry experiments and coordinated the animal experiments. J.E.G. conceived and directed the project and wrote the manuscript with comments from all the authors.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Clare Bryant, Elizabeth Villa, Siyu Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LRRK2 is required for efficient itaconate delivery to the Salmonella-containing vacuole.
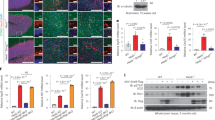
(a and b) LRRK2 is required for efficient itaconate delivery to the Salmonella-containing vacuole. Parental (control) and Lrrk2-/- Raw264.7 cells were infected with S. Typhi (MOI = 6) encoding an eGFP-based itaconate biosensor and the number of cells expressing eGFP was determined 20 hours after infection. Each square and circle represents the mean of an individual experiment experiments in which at least 200 infected cells were examined (b). The p value (unpaired two-tailed Student’s t test) of the indicated comparison is shown. Infected cells were fixed, stained with DAPI (blue) to visualize nuclei, and stained with an anti-Salmonella LPS antibody along with Alexa 594-conjugated anti-rabbit antibody (red) to visualize all bacteria. Representative fields of infected cells are shown (a) (scale bar = 5 µm). (c-g) Absence of LRRK2 does not influence the uptake of Salmonella into phagocytic cells. Raw264.7 or DC2.4 parental (control) and Lrrk2-/- cells, as well as bone marrow-derived macrophages (BMDM) derived from C57BL/6 and Lrrk2-/- mice were infected with either wild-type S. Typhi (MOI = 6) or a S. Typhimurium ∆gtgE ∆sopD2 mutant strain (MOI = 3) (as indicated) and the number of CFU was determined 1 hr after infection. Each square or circle represents the CFU in an independent measurement. The mean ± SD and p values (unpaired two-tailed Student’s t test) of the indicated comparisons are shown (n = 6 for each category).
Extended Data Fig. 2 Salmonella infection results in LRRK2 activation.
DC2.4 cells were treated with LPS or infected with the indicated bacterial strains for the indicated times. The activation of LRRK2, assessed by its phosphorylation at S935, was then analyzed by immunoblotting with the indicated antibodies.
Extended Data Fig. 3 LRRK2 scaffolds the formation of RAB32 and IRG1 complex.
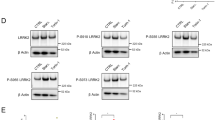
(a and b) LRRK2 interacts with RAB32 and IRG1. HEK293T cells were transiently co-transfected with a plasmid expressing GFP-LRRK2 and a plasmid expressing either FLAG-RAB32 (a) or FLAG-IRG1 (b). Twenty hours after transfection cells were infected with S. Typhi (MOI = 6) and 4 hs after infection, cell lysates were analyzed by immunoprecipitation and immunoblotting with antibodies against the FLAG epitope and GFP. (c-e) The kinase activity of LRRK2 is not required to form a complex with RAB32 and IRG1. (c and d) Raw264.7 (c) or DC2.4 (d) cells stably expressing FLAG-RAB32 or FLAG-IRG1 were pre−treated with the LRRK2 kinase inhibitor GSK2578215A for 90 min, infected with the S. Typhimurium ∆gtgE ∆sopD2 mutant strain (MOI = 3) (c) or treated with LPS (d). Eighteen hours after infection or 5 or 20 hs after LPS treatment, cell lysates were analyzed by immunoprecipitation and immunoblotting with the indicated antibodies. (e) HEK293T cells were transiently co-transfected with plasmids expressing GFP-RAB32, FLAG-Irg1, and the indicated forms of LRRK2: wild type (WT), kinase defective (3XKD = LRRK2K1906A/D1994A/D2017A), and constitutively active (LRRK2G2019S). Twenty hours after transfection, cell lysates were analyzed by immunoprecipitation and immunoblotting with the indicated antibodies. The quantification of the intensity of the RAB32 band relative to the intensity of the IRG1 band is shown in. Each circle, square, or triangle represents a measurement in an independent experiment. The mean ± SD and p values (unpaired two-tailed Student’s t test) of the indicated comparisons are shown (n = 3 for each category). (f and g) HEK293T parental or LRRK2-/- cells were transfected with GFP-RAB32 and FLAG-IRG1 for 20 hs. Cell lysates were then analyzed by immunoprecipitation with anti-FLAG and immunoblotting with anti-GFP antibody. The quantification of the intensity of the RAB32 band relative to the intensity of the IRG1 band is shown (f). Each circle or square represents a measurement in an independent experiment. The mean ± SD and p values (unpaired two-tailed Student’s t test) of the indicated comparisons are shown (n = 3 for each category). (g) Raw264.7 parental or Lrrk2-/- cells stably expressing FLAG-RAB32 were left untreated, treated with LPS, or infected with S. Typhimurium ∆gtgE ∆sopD2 mutant strain (MOI = 3) for 18 hs. Cell lysates were then analyzed by immunoprecipitation with anti-FLAG and immunoblotting with the indicated antibodies.
Extended Data Fig. 4 Localization of LRRK2, RAB32, and IRG1.
(a and b) LRRK2, RAB32, and IRG1are associated with the mitochondria accessible to protease digestion. DC2.4 cells stably expressing RAB32 (a), or DC2.4 parental (control) and Lrrk2-/- cells (b) were treated with LPS for 18 hs, mitochondria were purified and treated with proteinase K or left untreated, and subsequently analyzed by immunoblotting with the indicated antibodies. (c) Two color DNA-PAINT super-resolution image demonstrating that IRG1 does not co-localize with the mitochondrial matrix protein Cox IV. The top panel presents a HeLa cell expressing GFP-tagged IRG1 (green). Cells were fixed and stained with nanobodies to the GFP epitope, and primary and secondary antibodies to Cox IV (magenta). Nanobodies and secondary antibodies were labeled with a single stranded DNA oligomer acting as a docking site for DNA-PAINT super-resolution microscopy. First and second zoom levels show that Cox IV and IRG1 are spatially excluded from each other. The yellow arrows in zoom level two highlight examples of the spatial exclusion of Cox IV and IRG1. Scale bars 2 µm (top panel), 400 nm (zoom level 1) and 100 nm (zoom level 2). (d and e) Three-plex DNA-PAINT super-resolution image showing proximity of RAB32, LRRK2, and IRG1. (d) Hek293T cells expressing GFP-tagged LRRK2 (purple – DNA-PAINT), FLAG-tagged Rab32 (green – DNA-PAINT), and M45-tagged IRG1 (yellow – DNA-PAINT) were infected with S. Typhi carrying plasmid encoding an mCherry-based itaconate reporter (red – diffraction limited image). Cells were fixed and stained with nanobodies to the GFP epitope, and M45 and FLAG tags were labeled with primary antibodies and secondary antibodies conjugated to a single stranded DNA oligomer acting as a docking site for DNA-PAINT super-resolution microscopy. (e) The zoom in shows the spatial proximity of the three proteins in the proximity of S. Typhi expressing the itaconate reporter. The white arrows in the zoom-in highlight examples of the proximity cluster of the three proteins. Scale bars 5 µm (d), 1 µm (e).
Extended Data Fig. 5 Inhibition of the mitochondrial tricarboxylate transporter SLC25A1 impairs itaconate delivery to the Salmonella-containing vacuole.
(a and b) HeLa cells stably expressing EGFP-tagged IRG1 were pre-treated with the SLC25A1 transporter inhibitor CTPI-2 for 3, 6, or 18 hs (as indicated), and then infected with wild-type S. Typhi (MOI = 6) encoding a luciferase-based itaconate biosensor. The levels of luciferase activity in the cell lysates were then measured 3 hs after infection. Each circle or square represents a single luciferase measurement. The mean ± SD and p values (unpaired two-tailed Student’s t test) of the indicated comparisons are shown (n = 6 for each category). (a) and (b) show results of two independent experiments. (c and d) Inhibition of the mitochondrial tricarboxylate transporter SLC25A1 does not impair IRG1 expression or overall itaconate biosynthesis. (c) HeLa cells stably expressing EGFP-tagged IRG1 were pre-treated with the SLC25A1 transporter inhibitor CTPI-2 for 3, 6, or 18 hs (as indicated), and then infected with wild-type S. Typhi (MOI = 6) encoding a luciferase-based itaconate biosensor as indicated in Extended Data Fig. 10. The levels of IRG1 3, 6 or 18 hours after CTPI-2 treatment were evaluated by western immunoblot with the indicated antibodies. (d) HeLa cells stably expressing EGFP-tagged IRG1 were pre-treated with the SLC25A1 transporter inhibitor CTPI-2 for 18 hs, and the levels of itaconate were measured as indicated in Materials and Methods. Each square represents a single measurement and the mean and SD are shown (n = 3 for each category).
Extended Data Fig. 6 Tomographic slices of S. Typhi infected cells at different times after infection.
HeLa cells expression IRG1 (a and b) or BMDMs obtained from C57BL/6 mice (c and d) were infected with S. Typhi and 1 (a and c) and 3 (b and d) hs after infection were processed for cryo-ET imaging. Shown are representative tomographic slices showing that the appearance of S. Typhi within cells over time. Bacteria within HeLa-IRG1 cells 1 hr after infection appear normal, with many ribosomes and an intact bacterial envelope. However, bacteria within HeLa-IRG1 cells 3 hs post-infection or within BMDMs at 1 and 3 hs post infection exhibit altered morphology. Mi: mitochondria.
Extended Data Fig. 7 Visualization of tethers at the SCV-mitochondria interface.
(a-e) 3D renderings of the SCV-mitochondria interfaces shown in Figs. 4f, j, k, o, and p, respectively. Magenta, yellow, and green represent bacterial, vacuolar, and mitochondrial membranes, respectively. Intermembrane tethers are depicted in white. Please refer to the main Fig. 4 figure legend for experimental details. (f-j) Top-down views of the corresponding interfaces in Panels (a-e), revealing vacuolar membrane surfaces decorated with intermembrane tethers.
Extended Data Fig. 8 Itaconate delivery and bacterial growth in cells used for cryo-ET analysis.
(a) HeLa cells stably expressing IRG1 or BMDMs from C57BL/6 mice treated with LPS (200 ng/ml) for 3 hours were infected with S. Typhi (MOI = 10), and the number of CFU was determined 1 and 3 hs after infection. Each circle represents the CFU in an independent measurement; the mean ± SEM of all the measurements and p values of the indicated comparisons (two-sided Student’s t test) are shown. ns, not significant. ****p < 0.0001 (n = 6 for each category). (b and c) HeLa cells stably expressing IRG1 (b) or BMDMs from C57BL/6 mice treated with LPS (200 ng/ml) for 3 hs (c) were infected with S. Typhi (MOI = 10) carrying a plasmid encoding the itaconate nanoluciferase biosensor. One and three hours after infection, the levels of nanoluciferase were measured in lysates of the infected cells. Each circle represents a single luciferase measurement. The mean ± SD and p-values of the indicated comparisons (two-sided Student’s t-test) are shown. ****p < 0.0001 (n = 6 for each category). (d-g) BMDMs obtained from C57BL/6 (WT) or Hps4−/− were infected with S. Typhi (MOI = 10) carrying a plasmid encoding the itaconate nanoluciferase biosensor, and the number of CFU was determined 1 (a) or 3 (c) hs after infection. Alternatively, the levels of nanoluciferase were measured in lysates of the infected cells (b and d). Each circle represents the CFU in independent measurements (a and c) or a single luciferase measurement (b and d). Shown are the mean ± SEM of all the measurements (n = 6 for each category); p values of the indicated comparisons (two-sided Student’s t test) are shown. **p < 0.01 and ***p < 0.001, ****p < 0.0001. (h and i) Itaconate delivery and intracellular growth of S. Typhi expressing gtgE in cells used for cryo-ET analysis. (h and i) HeLa cells stably expressing IRG1 were infected with S. Typhi or S. Typhi- expressing gtgE (MOI = 10) carrying a plasmid encoding the itaconate nanoluciferase biosensor. The number of CFU (a) or the level of luciferase activity (b) was determined 1 hour or 3 hours after infection. Each circle represents a single measurement. Values are the mean ± SEM of all the measurements and p values of the indicated comparisons (two-sided Student’s t test) are shown. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 (n = 6 for each category).
Extended Data Fig. 9 Expression of the S. Typhimurium effector protein GtgE in S. Typhi does not prevent SCV-mitochondria association and tethering.
(a) Tomographic slice showing S. Typhi strain expressing GtgE within its replication vacuole and surrounding mitochondria (Mi) intimately interacting with the vacuolar membrane (VM). (b) 3D-rendering of the tomogram shown in panel (a)(z = 86 slices). Mitochondria is depicted in green, the SCV membrane in yellow, bacterial envelope in blue, inter membrane tethers in white, type III secretion machines in light blue, and bacterial ribosomes in grey (see close ups of the SCV-mitochondria interface in Fig. S14).
Extended Data Fig. 10 Model for the role of LRRK2 in itaconate delivery to the Salmonella containing vacuole.
LRRK2 may coordinate the close apposition between the Salmonella-containing vacuole (SCV) and the mitochondria (not depicted in this model) as it has been proposed to do with other intracellular organelles (64). In addition, as depicted in this model, through its ability to scaffold a complex between RAB32, IRG1, and SLC25A1, LRRK2 may coordinate the localized synthesis of itaconate at the mitochondria/SCV interface.(generated with the help of Biorender (www.biorender.com).
Supplementary information
Supplementary Information
Supplementary Tables 1–4.
Supplementary Video 1
Tethering of the SCV with the mitochondria observed by cryo-ET.
Source data
Source Data Figs. 1 and 3–5 and Extended Data Figs. 1, 3, 5 and 8
Numerical data for figures and extended data figures.
Source Data Figs. 1–3 and Extended Data Figs. 2–5
Uncropped western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lian, H., Park, D., Chen, M. et al. Parkinson’s disease kinase LRRK2 coordinates a cell-intrinsic itaconate-dependent defence pathway against intracellular Salmonella. Nat Microbiol 8, 1880–1895 (2023). https://doi.org/10.1038/s41564-023-01459-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01459-y