Abstract
Emerging severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants with antigenic changes in the spike protein are neutralized less efficiently by serum antibodies elicited by legacy vaccines against the ancestral Wuhan-1 virus. Nonetheless, these vaccines, including mRNA-1273 and BNT162b2, retained their ability to protect against severe disease and death, suggesting that other aspects of immunity control infection in the lung. Vaccine-elicited antibodies can bind Fc gamma receptors (FcγRs) and mediate effector functions against SARS-CoV-2 variants, and this property correlates with improved clinical coronavirus disease 2019 outcome. However, a causal relationship between Fc effector functions and vaccine-mediated protection against infection has not been established. Here, using passive and active immunization approaches in wild-type and FcγR-knockout mice, we determined the requirement for Fc effector functions to control SARS-CoV-2 infection. The antiviral activity of passively transferred immune serum was lost against multiple SARS-CoV-2 strains in mice lacking expression of activating FcγRs, especially murine FcγR III (CD16), or depleted of alveolar macrophages. After immunization with the pre-clinical mRNA-1273 vaccine, control of Omicron BA.5 infection in the respiratory tract also was lost in mice lacking FcγR III. Our passive and active immunization studies in mice suggest that Fc–FcγR engagement and alveolar macrophages are required for vaccine-induced antibody-mediated protection against infection by antigenically changed SARS-CoV-2 variants, including Omicron strains.
Similar content being viewed by others
Main
Since the emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in late 2019, 757 million infections and 6.8 million deaths have been reported (https://covid19.who.int/). Vaccines were rapidly generated and deployed resulting in reductions in symptomatic infections, hospitalizations and deaths. Initial SARS-CoV-2 vaccines all targeted the viral spike protein derived from strains that circulated in early 2020. However, the continuing evolution of SARS-CoV-2 has jeopardized the immunity generated by these vaccines and the control of virus infection and transmission.
SARS-CoV-2 vaccination can induce neutralizing antibodies that inhibit infection1,2. Correlates of vaccine protection initially focused on neutralizing activity of elicited anti-spike antibodies3. Variants of concern, which have amino acid substitutions in regions targeted by neutralizing antibodies including the receptor binding domain (RBD) and N-terminal domain (NTD)4,5, have prompted the development of boosters with components that match more recent strains6. Indeed, a substantial decrease in the neutralizing activity of serum antibodies elicited by vaccines against the ancestral Wuhan-1 virus has been observed against emerging variants, which has correlated with symptomatic breakthrough infections, especially with Omicron lineage viruses7,8. The large number of substitutions in the spike protein in Omicron strains has been termed by some as an antigenic shift9,10. Despite the loss in serum neutralizing activity against variants in the Omicron lineage, most immunized individuals remain protected against severe disease. The basis for this protection has not been fully determined but could be due to beneficial effects of non-neutralizing antibodies, cross-reactive T-cell responses or anamnestic memory B-cell responses4,11,12,13.
Fc effector function activity of non-neutralizing, cross-reactive, anti-spike antibodies is one hypothesized mechanism for protection against antigenically shifted SARS-CoV-2 variants5. In patients with moderate to severe coronavirus disease 2019 (COVID-19), the ability of antibodies to bind Fc gamma receptors (FcγR) and mediate effector functions correlated with increased survival14. Interactions of the conserved Fc region of immunoglobulin G (IgG) antibodies with FcγR or complement can promote clearance of virally infected cells through antibody-dependent cellular cytotoxicity, antibody-dependent cellular phagocytosis (ADCP) or complement-dependent deposition and phagocytosis or lysis. Indeed, monoclonal antibodies (mAbs) that lose their ability to neutralize SARS-CoV-2 variants yet still bind the spike protein avidly enough to trigger Fc effector functions retain protective activity11,15. Analogously, non-neutralizing antibodies induced by SARS-CoV-2 vaccines have been linked to protection against variant Omicron strains by virtue of their ability to engage FcγRs and promote clearance16,17. Furthermore, the depletion of RBD-specific antibodies from serum of mRNA-1273 or BNT162b2 vaccinated individuals did not appreciably impact Fc-mediated effector function activity in cell culture4, suggesting that antibodies recognizing conserved, non-neutralizing epitopes may contribute to protection against variant strains.
Although serum-derived anti-SARS-CoV-2 antibody-mediated Fc effector functions can activate complement deposition, immune cell phagocytosis and target cell killing in vitro, their contribution to protection in vivo remains uncertain. In this Article, to address this gap, we evaluated the impact of Fc effector functions in the context of passive transfer of vaccine-elicited antibodies or active immunization with mRNA-1273 vaccine using wild-type, C1q knockout (KO) and FcγR KO C57BL/6 mice and challenge with SARS-CoV-2 viruses. In passive serum transfer experiments, we found that activating FcγRs but not C1q were required to control SARS-CoV-2 infection in the lower respiratory tract, and protection was lost in mice depleted of alveolar macrophages but not neutrophils and monocytes. Experiments with mice lacking individual FcγRs showed the protective effect of passively transferred serum antibody on viral load reduction required expression of FcγR III. To determine the impact of FcγRs in the context of active immunization, wild-type, FcγR I KO, FcγR III KO and FcγR I/III/IV KO mice were administered a two-dose primary vaccination series with mRNA-1273, evaluated for immunogenicity and then challenged with the antigenically shifted Omicron BA.5 strain. Although the levels of anti-RBD antibody, neutralizing antibody and spike-specific T cells were similar in all tested strains of mice, protection against infection was lost in FcγR III KO and FcγR I/III/IV KO mice. Overall, our results in mice suggest that Fc–FcγR interactions contribute to antibody-mediated control in vivo in the context of both passive and active immunization with legacy vaccines, particularly when neutralizing antibody levels are low against antigenically distant SARS-CoV-2 variant strains.
Results
Immunoglobulin subclass and FcγR binding of immune serum
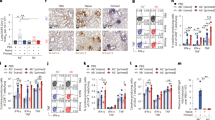
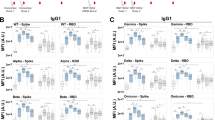
To begin to evaluate the contribution of Fc effector functions to antibody protection against SARS-CoV-2 infection, we profiled vaccine-induced antibodies from sera pooled from immunized C57BL/6 mice using a systems serology assay18. We measured the binding of antibodies to several spike proteins (Wuhan-1, B.1.617.2, BA.1 and BA.4/5) and determined their IgG subclass specificity (IgG1, IgG2b or IgG2c) and ability to interact with specific FcγRs (FcγR IIb, FcγR III or FcγR IV). We used naïve sera and binding to influenza haemagglutinin (HA) protein as negative controls (Fig. 1a–f). Immune sera contained higher levels of IgG1, IgG2b and IgG2c antibodies against Wuhan-1, B.1.617.2, BA.1 and BA.4/.5 but not HA compared with naïve sera (Fig. 1a–c). Anti-spike antibody binding to inhibitory (FcγR IIb) and activating FcγRs (FcγR III and FcγR IV) was higher in immune sera compared with naïve sera for all SARS-CoV-2 spike variants (Fig. 1d–f). We also assessed the effector function activity of immune serum using assays that measure spike-specific antibody-dependent cellular phagocytosis in murine monocytes (ADCP) and neutrophils (ADNP) (Fig. 1g,h and Extended Data Fig. 1). Compared with naïve sera, vaccine-elicited immune sera promoted greater ADCP and ADNP activity against the Wuhan-1 and BA.4/.5 spike proteins. In comparison, immune sera did not enhance antibody-dependent natural killer cell activation (CD107a expression, Fig. 1i). Immune sera also promoted antibody-dependent complement deposition (ADCD) on beads coated with spike proteins compared with HA protein (Fig. 1j). Overall, these studies indicated that our pooled vaccine sera have a diversity of antibodies against spike proteins that enables binding to FcγRs, and most Fc-mediated effector functions in cell culture.
a–c, Levels of IgG1 (a), IgG2b (b) and IgG2c (c) that bind to SARS-CoV-2 spike (Wuhan-1, B.1.617.2, BA.1 and BA.4/5), or influenza HA in naïve and vaccine-induced immune sera. d–f, Levels of spike- or HA-binding IgG antibodies that engage FcγR IIb (d), FcγR III (e) or FcγR IV (f) in naïve and vaccine-induced immune sera. g–j, Antibody effector functions. Antibody-mediated cellular phagocytosis with monocytes (ADCP, g) or neutrophils (ADNP, h) using vaccine-induced immune or naïve sera and beads coated with SARS-CoV-2 Wuhan-1 and BA.4/5 spike proteins. CD107a surface expression on human natural killer cells (ADNKA) after incubation with beads coated with Wuhan-1 or BA4/5 spike proteins and immune sera (bars indicate median values; n = 2 donors per group, one experiment; one-way ANOVA with Tukey’s post-test; NS, not significant) (i). Deposition of complement (ADCD) on beads coated with indicated SARS-CoV-2 spike or influenza HA proteins after treatment with naïve or immune sera (j).
Protection conferred by sera requires Fc–FcγR engagement
To assess the impact of Fc effector functions in vivo in the context of immune anti-SARS-CoV-2 antibodies, pooled naïve or vaccine-elicited immune sera were transferred passively to 12-week-old male wild-type, FcγR I/III/IV KO or C1q KO C57BL/6 mice; FcγR I/III/IV KO mice lack the common γ chain present in all activating murine FcγRs, whereas C1q KO mice lack C1q, a protein required for initiation of antibody-dependent complement activation. One day after transfer, mice were inoculated with SARS-CoV-2 MA-10 (ref. 19), and 4 days post-infection (dpi), nasal wash, nasal turbinates and lungs were collected (Fig. 2a). We used the mouse-adapted MA-10 strain for these initial studies because it spreads to the lungs of conventional C57BL/6 mice without a need for ectopic expression of human ACE2 receptor. Pooled vaccine-induced immune sera neutralized MA-10 infection by 50% at a dilution of 1/3,300, and 1 day after transfer, serum from recipient mice neutralized MA-10 infection by 50% at a dilution of 1/36 (Fig. 2b). In the nasal washes and nasal turbinates of the upper airway of wild-type, FcγR I/III/IV KO and C1q KO mice, we observed no significant differences in levels of viral RNA or infectious virus among the three groups receiving naive or immune serum. Although there was a trend towards less viral infection in the nasal turbinates of animals receiving immune compared with naïve sera, the comparisons did not reach statistical significance (Fig. 2c–e); these results are consistent with the lower accumulation of IgG in the upper respiratory tract after passive transfer by a systemic route20. Results from lung tissues, however, showed a different pattern, with a loss of control of infection (viral RNA levels and infectious virus) by immune sera in FcγR I/III/IV KO but not in wild-type or C1q KO mice (Fig. 2f,g). Thus, FcγR expression in the lower respiratory tract appears important for restricting infection after passive antibody transfer.
a, Scheme of passive transfer, virus challenge and tissue collection. b, Neutralizing antibody responses against SARS-CoV-2 MA-10 using sera from naïve (circles) or Wuhan-1 spike protein vaccinated mice (pooled from animals immunized and boosted with mRNA-1273 or ChAd-SARS-CoV-2-S) (squares). Also shown is serum neutralizing antibody activity from recipient wild-type and FcγR I/III/IV KO mice 1 day after transfer of immune sera. c–g, Twelve-week-old male wild-type, C1q KO and FcγR I/III/IV KO C57BL/6 mice were passively transferred by intraperitoneal injection 60 μl of naïve or vaccine-induced immune sera 1 day before intranasal challenge with 103 FFU of SARS-CoV-2 MA-10. At 4 dpi, viral RNA in the nasal wash (c), nasal turbinates (d) and lungs (f) was quantified, and infectious virus in the nasal turbinates (e) and lungs (g) was determined (bars indicate mean ± standard error of the mean; in order left to right n = 5 (b); n = 6, 6, 7, 7, 6 and 7 (c); n = 6, 6, 7, 7, 6 and 7 (d); n = 6, 6, 7, 7, 6 and 7 (e); n = 6, 6, 7, 7, 6 and 7 (f); n = 6, 6, 7, 7, 6 and 7 (g) mice per group, one experiment (b), two experiments (c–g), dotted lines show limit of detection (LOD)). One-way ANOVA with Tukey’s post-test; NS, not significant; *P = 0.0188 (f); ****P < 0.0001 (g)); additional statistical comparisons are presented in Supplementary Table 3. In c–g, the LOD are weight and volume based, and vary on the basis of the amount of material collected for RNA extraction.
Antibody protection requires FcγR III engagement
As MA-10 is not matched to the vaccine antigen, in the context of passive transfer, we might underestimate the protection against infection afforded by serum neutralizing antibody. To address this issue and also identify which FcγR contributed to the protective phenotype, we passively transferred naïve or immune sera to 12-week-old male wild-type, FcγR I KO, FcγR II KO, FcγR III KO and FcγR I/III/IV KO congenic C57BL/6 male mice before inoculation with SARS-CoV-2 WA1/2020 N501Y/D614G (Fig. 3a), a more closely matched virus; because this suite of FcγR-deficient C57BL/6 mice lacks human ACE2 expression, we used a virus with a mouse-adapting N501Y mutation21,22. Pooled vaccine-elicited immune sera neutralized WA1/2020 N501Y/D614G more efficiently than MA-10 with half-maximal inhibition at a serum dilution of 1/16,750; moreover, 1 day after transfer, serum from recipient mice neutralized WA1/2020 N501Y/D614G with half-maximal inhibition at a serum dilution of 1/750 (Fig. 3b), which exceeds a ~1/50 presumptive correlate of protection in humans3. As seen with MA-10 infection, in the nasal washes and nasal turbinates of the upper respiratory tract, we did not observe reductions in WA1/2020 N501Y/D614G viral load by serum antibodies in wild-type C57BL/6 mice (Fig. 3c–e); thus, we focused analysis on the lung. Indeed, passive transfer of immune sera protected against SARS-CoV-2 WA1/2020 N501Y/D614G infection in the lungs of wild-type C57BL/6 mice as measured by viral RNA and infectious virus levels (Fig. 3f,g). Similar reductions in viral load were observed in FcγR I KO and FcγR II KO. However, reductions in SARS-CoV-2 lung infection were diminished or lost in FcγR III and FcγR I/III/IV KO mice. These data suggest that, even in the context of passive transfer of immune sera with neutralizing activity, protection against lower respiratory tract infection by SARS-CoV-2 is mediated at least in part by Fc interactions with activating FcγRs, particularly FcγR III.
a, Scheme of passive transfer, virus challenge and tissue collection. b, Neutralizing antibody response against SARS-CoV-2 WA1/2020 N501Y/D614G using sera from naïve (circles) or Wuhan-1 spike protein vaccinated (squares) mice. Also shown is serum neutralizing antibody activity from recipient wild-type and FcγR I/III/IV KO mice 1 day after transfer of immune sera. c–g, Twelve-week-old male wild-type, FcγR I KO, FcγR II KO, FcγR III KO and FcγR I/III/IV KO mice were passively transferred by intraperitoneal injection 60 μl of naïve or vaccine-immune sera 1 day before intranasal challenge with 104 FFU of WA1/2020 N501Y/D614G. At 4 dpi, viral RNA and infectious virus were measured in the upper respiratory tract (nasal wash (c); nasal turbinates (d and e); or lungs (f and g). In c–e, wild-type mice only; in f and g, wild-type, FcγR I KO, FcγR II KO, FcγR III KO and FcγR I/III/IV KO mice (bars indicate mean ± standard error of the mean; in order left to right n = 5 (b); n = 18 and 11 (c); n = 18 and 11 (d); n = 18 and 11 (e); n = 18, 9, 9, 12, 10, 11, 8, 10, 11 and 11 (f); n = 18, 9, 9, 12, 10, 11, 8, 10, 11 and 11 (g) mice per group, one experiment (b), three experiments (c–g), dotted lines show limit of detection (LOD)). One-way ANOVA with Tukey’s post-test (NS, not significant; **P = 0.0068, ****P < 0.0001 (f); ***P = 0.0002, ****P < 0.0001 (g)); additional statistical comparisons are presented in Supplementary Table 3.
Vaccine-elicited immunity requires Fc-FcγR engagement
We next evaluated the dependence on Fc-FcγR engagement on control of SARS-CoV-2 infection in the context of vaccine-elicited immunity, which induces both cellular and humoral responses. We immunized groups of 9-week-old male wild-type, FcγR I KO, FcγR III KO and FcγR I/III/IV KO C57BL/6 mice twice over 4 weeks with 0.25 μg of control or pre-clinical mRNA-1273 vaccine (Fig. 4a); we did not immunize FcγR II KO mice, since the virological phenotypes in the context of passive antibody transfer were present in mice lacking activating FcγRs but not FcγR II (Figs. 2 and 3). The 0.25 μg dose of mRNA vaccine was used because the B- and T-cell responses generated in C57BL/6 mice with this dose is approximate to those observed in humans receiving 100 μg doses23. One potential limitation of this experiment is that a loss of activating FcγRs could affect vaccine-induced immune responses, which might confound interpretation of challenge studies. To evaluate this first, 24 days following boosting, serum was obtained to measure binding and neutralizing antibody against WA1/2020 N501Y/D614G and BA.5. As expected, higher levels of serum IgG were detected against Wuhan-1 than BA.5 RBD protein (Fig. 4b,c), consistent with the antigenic shift of Omicron lineage strains9,10. However, no statistical differences in binding titres were observed between the groups of vaccinated wild-type and FcγR KO mice (Fig. 4b,c). Similarly, lower neutralization titres were detected against BA.5 than WA1/2020 N501Y/D614G, with no substantive differences observed between groups of vaccinated wild-type and FcγR KO (Fig. 4d,e). Thus, mRNA-1273 vaccination induced relatively similar humoral immune responses in mice that were sufficient or deficient in FcγR expression. We also measured spike-specific CD4+ and CD8+ T-cell responses in vaccinated mice using immunodominant peptides24,25. As expected, antigen-specific T-cell responses were greater in animals give the mRNA-1273 vaccine compared with the control mRNA vaccine. However, IFN-γ and TNF-α responses in CD4+ and CD8+ T cells after peptide restimulation were equivalent in FcγR III KO, FcγR I/III/IV KO and wild-type mice after mRNA-1273 vaccination (Extended Data Fig. 2). These experiments establish that FcγR KO and wild-type mice have similar serum antibody and T-cell responses after mRNA vaccination.
a, Scheme of immunization, serum sampling, virus challenge and tissue collection. b,c, Anti-Wuhan-1 (b) and BA.4/5 (c) RBD IgG responses from serum of mice immunized with control or mRNA-1273 vaccines. d,e, Neutralizing antibody responses against WA1/2020 N501Y/D614G (d) and BA.5 (e) from serum collected from wild-type, FcγR I KO, FcγR III KO and FcγR I/III/IV KO mice 25 days after completion of a two-dose primary vaccination series with control (closed circles) or mRNA-1273 (open circles). f,g, Nine-week-old male wild-type, FcγR I KO, FcγR III KO and FcγR I/III/IV KO mice were immunized twice at 4 week intervals with control or mRNA-1273 vaccine via intramuscular route. Four weeks after the primary vaccination series, mice were challenged via intranasal route with 104 FFU of BA.5. At 3 dpi, infectious virus in the nasal turbinates (f) and lungs (g) was determined (boxes illustrate geometric mean titres (GMT), dotted lines show limit of detection (LOD), bars indicate mean ± standard error of the mean; in order left to right n = 10, 10, 10, 10, 25, 10, 10 and 14 (b); n = 12, 10, 10, 10, 22, 10, 10 and 15 (c); n = 22, 9, 9, 12, 30, 15, 15 and 15 (d); n = 22, 9, 9, 12, 25, 10, 10 and 15 (e); n = 19, 10, 8, 8, 19, 9, 10 and 10 (f); n = 19, 10, 8, 8, 19, 9, 10 and 10 (g) mice per group, one experiment (b–e), two experiments (f and g), dotted lines show LOD, one-way ANOVA with Dunnett’s test (NS, not significant, ***P = 0.002, ****P < 0.0001 (f); **P = 0.0044, ****P < 0.0001 (g)); additional statistical comparisons are presented in Supplementary Table 3.
To assess whether vaccinated mice expressing FcγRs were differentially protected against SARS-CoV-2 infection, animals were challenged by intranasal route with 103 focus-forming units (FFU) of BA.5, and infectious virus in the nasal turbinates and lungs was measured at 3 dpi (Fig. 4a). For these experiments, we used BA.5 as the challenge virus because: (1) it encodes a mouse-adapting mutation (N501Y) that facilitates replication in mice lacking human ACE2 expression; (2) it allowed us to assess protection against infection under conditions when high levels of neutralizing antibody are absent (Fig. 4e); and (3) BA.5 and other Omicron variants are circulating, so use of this strain could provide insight as to how legacy vaccines directed against ancestral spikes protect against Omicron strain infections in humans. Notably, we observed protection against BA.5 infection in the upper and lower respiratory tracts of wild-type and FcγR I KO mice but not in FcγR III KO or FcγR I/III/IV KO mice (Fig. 4f,g). Thus, protection elicited by mRNA-1273 vaccine-induced immunity against infection by antigenically shifted BA.5 SARS-CoV-2 requires Fc–FcγR engagement, and FcγR III interactions in particular contribute to this phenotype in mice.
Alveolar macrophages are required for control of BA.5 infection
We next addressed which FcγR III-expressing immune cells in the lung were important for mediating antibody protection in the context of passive transfer of immune sera and BA.5 challenge (Fig. 5). Mice that received vaccine-elicited immune sera had detectable amounts of anti-BA.5 spike IgG but low levels of neutralizing activity (~1/10 titre), as expected (Extended Data Fig. 3). Flow cytometric analysis of CD45+ immune cells in the lungs of wild-type C57BL/6 mice showed that monocytes, neutrophils, interstitial macrophages and alveolar macrophages express FcγR III (Extended Data Fig. 4 and Supplementary Table 1). We first assessed the role of neutrophils and monocytes by depleting these cells in wild-type mice with an anti-Ly6C/Ly6G (Gr-1) antibody (Fig. 5a). Depletion of these cells in circulation (Fig. 5b–d and Extended Data Fig. 5a), which corresponds to depletion in the lung26, did not impact BA.5 infection in the nasal turbinates or lungs; decreased levels of infectious BA.5 virus in the lungs were seen after immune sera transfer regardless of whether neutrophils and monocytes were present (Fig. 5e,f). We next depleted alveolar macrophages (Fig. 5g–i and Extended Data Fig. 5b) in the lung using intranasal administration of clodronate liposomes27. Clodronate, but not control, liposomes specifically depleted alveolar macrophages in the lung, and was associated with a loss of control of BA.5 infection after passive transfer of immune but not non-immune (naïve) sera (Fig. 5j,k). Together, these experiments establish an important role for FcγR III-expressing alveolar macrophages in antibody-mediated control of BA.5 infection in mice.
a, Scheme for depletion of neutrophils and monocytes, passive transfer of immune sera and BA.5 challenge. b–d, Analysis of depletion of immune cell subsets in blood of mice receiving anti-Gr-1 (Ly6C/Ly6G) or isotype antibody at 3 dpi. Summary of different cell types (b) based on the gating strategy (Extended Data Fig. 5a). Representative flow cytometry dot plots showing depletion of neutrophils (c) and monocytes (d) with numbers indicating the cell population as a percentage of CD45+ cells. e,f, Analysis of infectious viral burden at 3 dpi in nasal turbinates (e) and lungs (f) after BA.5 challenge. g, Scheme for depletion of alveolar macrophages, passive transfer of immune sera and BA.5 challenge. h, Analysis of depletion of immune cell subsets in lungs of mice receiving liposomes at 3 dpi. Summary of different cell types (h) based on gating strategy (Extended Data Fig. 5b); DCs, dendritic cells. i, Representative flow cytometry dot plots showing depletion of alveolar macrophages after liposome administration with numbers indicating the cell population as a percentage of CD45+ cells. j,k, Analysis of infectious viral burden at 3 dpi in nasal turbinates (j) and lungs (k) after BA.5 challenge (boxes illustrate GMT, bars indicate mean; in order left to right n = 10, 10, 10 and 6 (b); n = 10, 9, 9 and 6 (e); n = 10, 9, 9 and 6 (f); n = 9, 10, 7 and 12 (h); n = 9, 10, 7 and 12 (j); n = 9, 10, 7 and 12 (k) mice per group, two experiments (b, e, f, h, j and k), dotted lines show limit of detection (LOD), Mann–Whitney test with Bonferroni post-test correction (b and h), Mann–Whitney test (e, f, j and k), NS, not significant, **P = 0.0077, ****P < 0.0001, ***P = 0.0003 (b); **P = 0.003, ***P = 0.0007, **P = 0.0034, **P = 0.0011 (h); ****P < 0.0001 (k).
Discussion
Despite the diminished neutralizing ability of vaccine-elicited antibodies against SARS-CoV-2 variants with amino acid substitutions in the RBD and NTD9,16, protection against severe disease is maintained in the majority of vaccine recipients28,29. Although neutralizing activity of antibodies is a correlate of vaccine-mediated protection3, the ability of monovalent COVID-19 vaccines to limit Omicron disease in the setting of waning serum antibody neutralization suggests additional protective immune mechanisms. These include anamnestic B-cell responses that rapidly generate cross-reactive neutralizing antibodies, cross-reactive T-cell responses and/or non-neutralizing, cross-reactive antibodies that promote Fc-mediated effector activities12,30,31,32. We focused on evaluating Fc mediated effector functions as a possible mechanism of vaccine-mediated protection against antigenic variants. In vitro studies with human convalescent sera have demonstrated that Fc effector functions are retained against antigenically variant strains, and that sera of patients with COVID-19 with more severe disease have compromised FcγR binding abilities and effector functions4,11,14. Studies in mice with passively transferred mAbs show that Fc effector functions contribute to protection33, and this activity is maintained against antigenically distant strains even when neutralizing capacity is compromised15. Our experiments in mice show that Fc–FcγR interactions contribute to control of SARS-CoV-2 infection in vivo in the context of active or passive immunization, and that alveolar macrophages are a key cell type required for this activity.
The increased binding of immune antibodies to spike proteins was associated with several Fc effector functions including ADCP, ADNP and ADCD. When immune serum was passively transferred to mice, control of infection by SARS-CoV-2 strains MA-10 or WA1/2020 N501Y/D614G in the lungs required FcγR expression, particularly FcγR III, even though serum antibody neutralizing activity was present after transfer. These data showing a requirement for Fc effector functions for optimal antibody-mediated control of virus infection in the lung are consistent with studies in mice and hamsters with neutralizing mAbs that bind epitopes in the RBD15,33. Moreover, Fc-engineered anti-SARS-CoV-2 non-neutralizing and neutralizing mAbs binding the NTD and RBD regions confer greater protection in mice and hamsters34,35. In the context of passive antibody transfer, we observed less impact in upper respiratory tract tissues, which could reflect the diminished ability of serum IgG antibodies to accumulate in airway spaces20. Nonetheless, we observed FcγR-dependent reductions in viral load in the nasal turbinates after active mRNA vaccination, which could be due to higher levels of systemic anti-spike IgG/IgA or production of antibody by tissue-resident B cells. Persistent SARS-CoV-2 IgG antibodies in oral mucosal fluid and upper respiratory tract specimens have been reported following mRNA vaccination36.
In wild-type and FcγR KO mice immunized with mRNA-1273, we observed similar levels of neutralizing and RBD or spike-specific antibodies, and vaccine-induced CD4+ and CD8+ T-cell responses. Although these results contrast with the idea that FcγRs can regulate adaptive immunity37,38, they are consistent with studies showing a lack of impairment of adaptive immune responses in FcγR KO mice to bacterial infection or IgG complexes39. Indeed, in control mRNA vaccinated animals, SARS-CoV-2 viral loads were similar in wild-type and FcγR KO mice. Thus, we attribute the diminished control of infection of the antigenically shifted BA.5 strain in the turbinates and lungs of mRNA-1273 vaccinated FcγR KO mice to the loss of Fc–FcγR interactions that mediate antibody effector functions. Notwithstanding these results, antigen-matched bivalent mRNA vaccines targeting BA.1 and BA.5 spike proteins can induce higher levels of neutralizing antibodies against Omicron strains40,41, which might result in less reliance on Fc effector functions for protection against infection. We also performed cellular depletions to investigate the cell type responsible for the protection conferred by passively transferred antibody. Although several myeloid cells in the lung express multiple FcγRs including FcγR III, depletion of alveolar macrophages compromised protection against infection. This effect was antibody-dependent since, in the absence of immune sera, depletion of alveolar macrophages did not affect the viral burden in the lungs. These results are consistent with studies of influenza virus in mice, which showed that protective immunity conferred by non-neutralizing antibodies required alveolar macrophages and other lung phagocytes27.
Limitations of study
We note several limitations in our study. (1) Our conclusions are based on experiments in mice. We used mice because of the availability of animals deficient in specific FcγRs, the reagents to achieve immune cell depletions, and the ability to perform both passive and active immunization and BA.5 challenge. However, our results may not directly correlate with those in humans because of species-dependent differences in FcγR subtypes, functions and expression on specific immune cells in the lung (Supplementary Tables 1 and 2). Future studies with human FcγR transgenic mice lacking individual human FcγRs may bridge this gap. (2) While we evaluated anti-spike antibody function in several FcγR KO mice, we did not directly assess a role for FcγR IV KO mice. Passive and active immunization studies in FcγR IV KO mice42 are warranted. (3) Although our studies indicate an importance of alveolar macrophages and FcγR III interactions with antibody, we did not identify a specific cellular mechanism of action. While our serum profiling analysis is consistent with an antibody- and Fc-dependent phagocytic mechanism, it remains unclear if this occurs with opsonized virus or infected cells. (4) Although challenge of wild-type and FcγR KO C57BL/6 mice with BA.5 allowed us to evaluate the effects on viral burden in the setting of low levels of transferred or induced serum antibody neutralization, Omicron strains are less virulent in most strains of laboratory mice43. Thus, we were not able to evaluate effects of FcγRs on antibody-mediated protection against clinical disease and lung pathology. (5) It will be important to determine whether vaccine-elicited antibodies engage FcγRs (for example, FcγR I and FcγR III) on specific myeloid cells and promote inflammation, as infection-induced antibodies from patients enhanced SARS-CoV-2 uptake by monocytes and macrophages, and triggered inflammasome activation, pyroptotic cell death and COVID-19 pathogenesis44,45. However, these studies also showed that immune plasma from mRNA vaccine recipients did not promote antibody-dependent monocyte infection and inflammation.
In summary, our experiments in mice provide insight and help to explain human studies that correlate Fc–FcγR interactions with clinical outcome against SARS-CoV-2 and emerging variants of concern. We provide an explanation as to how Fc–FcγR interactions might contribute to monovalent vaccine-mediated protection against severe infection by SARS-CoV-2 variants even in the setting when serum neutralizing antibody activity is lost5. Our work also suggests that targeting Fc effector functions in the context of vaccine design could be a strategy to generate more broadly protective immune responses. Future studies are warranted to define the epitopes targeted by antibodies with strong Fc effector functions and develop improved in vitro Fc effector function assays that correlate better with protection in vivo against infection by SARS-CoV-2 and variants.
Methods
Ethics statement
Animal studies were carried out in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The protocols were approved by the Institutional Animal Care and Use Committee at the Washington University School of Medicine (assurance number A3381–01).
Cells
Vero-TMPRSS2 (ref. 46) and Vero-hACE2-TMPRSS2 (ref. 47) cells were cultured at 37 °C in Dulbecco’s modified Eagle medium supplemented with 10% foetal bovine serum (FBS), 10 mM HEPES pH 7.3, 1 mM sodium pyruvate, 1× non-essential amino acids and 100 U ml−1 of penicillin–streptomycin. Vero-TMPRSS2 and Vero-hACE2-TMPRSS2 cells were supplemented with 5 μg ml−1 of blasticidin and 10 μg ml−1 of puromycin, respectively. All cells routinely tested negative for mycoplasma using a PCR-based assay.
Viruses
All SARS-CoV-2 strains used (WA1/2020 N501Y/D614G, mouse-adapted MA-10, B.1.351 and BA.5) have been previously described19,40,47,48. All viruses were subjected to next-generation deep sequencing to confirm the presence and stability of substitutions (Supplementary Table 4). All experiments with virus were performed in approved biosafety level 3 facilities with appropriate positive-pressure respirators, personal protective gear and containment.
Mice
Virus inoculations were performed under anaesthesia that was induced and maintained with ketamine hydrochloride and xylazine, and all efforts were made to minimize animal suffering. No sample size calculations were performed to power each study. Sample size for animal experiments was determined on the basis of criteria set by the institutional Animal Care and Use Committee and prior in vivo virus challenge experiments. Data distribution was assumed to be normal, although this was not formally tested. Experiments were neither randomized nor blinded, and mice were randomly assigned to treatment groups. C57BL/6 male mice (cat. no. 000664) were obtained from The Jackson Laboratory. FcγR I KO49, FcγR II KO (Taconic Biosciences; cat. no. 580), FcγR III KO (Jackson Laboratory; cat. no. 009637), FcγR I/III/IV (common γ-chain) KO (Taconic Biosciences; cat. no. 583) and C1q KO50 mice were obtained commercially or from collaborators and then backcrossed onto a C57BL/6 background (>99%) using Speed Congenics (Charles River Laboratories) and single nucleotide polymorphism analysis. Male mice were housed in groups of four to five; photoperiod, 12 h on:12 h off dark/light cycle. Ambient animal room temperature was at 70 °F, controlled within ±2 °F, and room humidity was 50%, controlled within ±5%. Animals were fed standard chow diets.
Preclinical mRNA and ChAd-SARS-CoV-2 vaccines
A sequence-optimized mRNA encoding prefusion-stabilized Wuhan-Hu-1 (mRNA-1273) was designed and synthesized in vitro using an optimized T7 RNA polymerase-mediated transcription reaction with complete replacement of uridine by N1m-pseudouridine51. A non-translating control mRNA was synthesized and formulated into lipid nanoparticles as previously described52. The reaction included a DNA template containing the immunogen open-reading frame flanked by 5′ untranslated region (UTR) and 3′ UTR sequences and was terminated by an encoded polyA tail. After RNA transcription, the cap-1 structure was added using the vaccinia virus capping enzyme and 2′-O-methyltransferase (New England Biolabs). The mRNA was purified by oligo-dT affinity purification, buffer exchanged by tangential flow filtration into sodium acetate, pH 5.0, sterile filtered and kept frozen at −20 °C until further use. The mRNA was encapsulated in a lipid nanoparticle through a modified ethanol-drop nanoprecipitation process described previously53. Ionizable, structural, helper and polyethylene glycol lipids were briefly mixed with mRNA in an acetate buffer, pH 5.0, at a ratio of 2.5:1 (lipid:mRNA). The mixture was neutralized with Tris–HCl, pH 7.5, sucrose was added as a cryoprotectan, and the final solution was sterile filtered. Vials were filled with formulated lipid nanoparticle and stored frozen at −20 °C until further use.
The ChAd-SARS-CoV-2 and ChAd-Control vaccine vectors were derived from simian Ad36 backbones, and the construction and validation has been described in detail previously54. The rescued replication-incompetent ChAd-SARS-CoV-2-S were scaled up in HEK293 cells (ATCC CRL-1573) and purified by CsCl density-gradient ultracentrifugation. For passive transfer studies, a large batch (8 ml) of immune sera (obtained ≥30 days post immunization) was pooled from C57BL/6 mice vaccinated with ChAd-SARS-CoV-2 or mRNA-1273.
Viral antigens
Recombinant RBD proteins from Wuhan-1 and BA.5 SARS-CoV-2 strains were expressed as described55. Recombinant proteins were produced in Expi293F cells (Thermo Fisher) by transfection of DNA using the ExpiFectamine 293 Transfection Kit (Thermo Fisher). Supernatants were collected 3 days post-transfection, recombinant proteins were purified using Ni-NTA agarose (Thermo Fisher) and buffer exchanged into PBS and concentrated using Amicon Ultracel centrifugal filters (EMD Millipore).
ELISA
Recombinant Wuhan-1, BA.4/5 RBD or BA.4/5 spike protein (4 μg ml−1) was immobilized on 96-well Maxisorp ELISA plates (Thermo Fisher) overnight at 4 °C in coating buffer (1X PBS supplemented with 0.05% Tween-20, 2% BSA and 0.02% NaN3). Plates were washed with PBS and blocked with 4% BSA for 1 h at 25 °C. Serum was serially diluted in 2% BSA and incubated on plates for 1 h at 25 °C. After washing with PBS, RBD or spike-bound serum antibodies were detected with horseradish peroxidase-conjugated goat anti-mouse IgG (1:500 dilution, Milipore Sigma) incubating for 2 h at 25 °C. Plates were washed and developed with 3,3′-5,5′ tetramethylbenzidine substrate (Thermo Fisher), halted with 2 N H2SO4 and read at 450 nm using a microplate reader (BioTek). Mean serum endpoint titres were calculated with curve fit analysis of optical density values set as the reciprocal value of the serum dilution equal to the mean plus six times the standard deviation of background signal.
Focus reduction neutralization test
Serial dilutions of sera were incubated with 102 FFU of MA-10, WA1/2020 N501Y/D614G or BA.5 for 1 h at 37 °C. Antibody–virus complexes were added to Vero-TMPRSS2 cell monolayers in 96-well plates and incubated at 37 °C for 1 h. Subsequently, cells were overlaid with 1% (w/v) methylcellulose in MEM. Plates were collected 30 h (MA-10 and WA1/2020 N501Y/D614G) or 72 h (BA.5) later by removing overlays and fixed with 4% paraformaldehyde (PFA) in PBS for 20 min at room temperature. Plates were washed and sequentially incubated with an oligoclonal pool (SARS2-02, −08, −09, −10, −11, −13, −14, −17, −20, −26, −27, −28, −31, −38, −41, −42, −44, −49, −57, −62, −64, −65, −67 and −71 (ref. 56) of anti-S murine antibodies at a dilution of 1:5,000 (including cross-reactive mAbs to SARS-CoV) and HRP-conjugated goat anti-mouse IgG at a dilution of 1:500 (Sigma cat. no. A8924, RRID: AB_258426) in PBS supplemented with 0.1% saponin and 0.1% bovine serum albumin. SARS-CoV-2-infected cell foci were visualized using TrueBlue peroxidase substrate (KPL) and quantitated on an ImmunoSpot microanalyzer (Cellular Technologies).
Viral plaque assay
Titration of infectious SARS-CoV-2 was performed as previously described57. Briefly, lung and nasal turbinate homogenates were serially diluted and added to Vero-TMPRSS2-hACE2 cell monolayers in 24-well tissue culture plates for 1 h at 37 °C. Cells were then overlaid with 1% (w/v) methylcellulose in MEM and incubated for 72 h (MA-10 and WA1/2020 N501Y/D614G) or 96 h (BA.5). Subsequently, cells were fixed with 4% PFA in PBS for 20 min at room temperature before staining with 0.05% (w/v) crystal violet in 20% methanol. Viral plaques were counted manually.
Mouse experiments
(1) Passive transfer studies. Twelve-week-old male C57BL/6, FcγR I KO, FcγR II KO, FcγR III KO, FcγR I/III/IV KO and C1q KO mice were administered 60 μl of sera (naïve or pooled from mice immunized and boosted with mRNA-1273 or ChAd-SARS-CoV-2-S) 1 day before challenge with 50 μl of 103 FFU of MA-10 or 104 FFU of WA1/2020 N501Y/D614G or BA.5 by intranasal administration. Lungs, nasal turbinates and nasal washes (collected in 500 μl of 0.5% bovine serum albumin in PBS) were collected 4 days after inoculation for virological analysis. (2) Immunization studies. Nine-week-old male C57BL/6, FcγR I KO, FcγR III KO and FcγR I/III/IV KO mice were immunized and boosted with 0.25 μg of control or mRNA-1273 vaccine by intramuscular injection 4 weeks apart. Animals were bled 24 days after boosting for immunogenicity analysis of sera and then challenged 4 days later with 50 μl of 104 FFU of BA.5 by intranasal administration. Lungs, nasal turbinates and nasal washes (collected in 500 μl of 0.5% bovine serum albumin in PBS) were collected 3 days after inoculation for virological analysis.
Immune cell depletions
For monocyte and neutrophil depletions, anti-Ly6G/Ly6C (BioXCell; clone RB8-8C5; 500 μg) or an IgG2b isotype control (BioXCell; clone LTF-2; 500 μg) were administered to mice by intraperitoneal injection at days −3 and −1 relative to SARS-CoV-2 inoculation. For alveolar macrophage depletion, clodronate liposomes (Liposoma; 250 μg) or control liposomes (Liposoma; 250 μg) were administered via intranasal route at day −3 relative to SARS-CoV-2 inoculation.
For analysis of neutrophil and monocyte depletion, peripheral blood was collected on the day of collection. Erythrocytes were lysed with ACK lysis buffer (GIBCO) at room temperature for 3 min and resuspended in RPMI 1640. Single-cell suspensions were pre-incubated with Fc block antibody (1:100; clone S17011E; BioLegend) in PBS + 2% heat-inactivated FBS + 1 mM EDTA for 20 min at 4 °C, stained with antibodies against CD45 (AF488; clone 30-F11; BioLegend), CD11b (APC/Fire 810; clone M1/70; BioLegend), Ly6G (Spark Blue 550; clone 1A8; BioLegend), Ly6C (APC-Fire 750; clone HK1.4; BioLegend), NK1.1 (BV570; clone PK136; BioLegend), B220 (Pacific Blue; clone RA3-6B2; BioLegend), fixable viability dye (eFluor 510; BD Biosciences), True-Stain Monocyte Blocker (BioLegend; 5 μl per sample) and Brilliant Stain Buffer Plus (BD Biosciences; 10 μl per sample), and then fixed with 4% PFA in PBS for 20 min at room temperature. All antibodies were used at a dilution of 1:200. The viability dye was used at 1:100. Absolute cell counts were determined using Precision Count Beads (BioLegend). Flow cytometry data were acquired on a 3 laser Cytek Aurora cytometer (Cytekbio) and analysed using FlowJo software v10.8 (Treestar).
For analysis of lung tissues, mice were killed by ketamine overdose, perfused with sterile PBS, and the right inferior lung lobes were digested in 50 μl of 5 mg ml−1 of Liberase Tm (Roche) and 12.5 μl of 10 mg ml−1 of DNase I (Sigma-Aldrich) in 5 ml of HBSS at 37 °C for 35 min. Single-cell suspensions of lung digest were pre-incubated with Fc block antibody (BioLegend) in PBS + 2% heat-inactivated FBS + 1 mM EDTA for 20 min at 4 °C. Cells were then stained with antibodies against CD45 (AF488), CD11b (APC/Fire 810), MHC II (BV711; clone M5/114.15.2; BioLegend), CD11c (PE-Cy7; clone N418; BioLegend), CD64 (BV421; clone X54-5/7.1; BioLegend), CD88 (Viogreen; clone REA1206; Miltenyi Biotec), Siglec-F (PE Dazzle 594; clone S17007L; BioLegend), Ly6G (Spark Blue 550; clone HK1.4; BioLegend), Ly6C (APC-Fire 750; clone HK1.4; BioLegend), NK1.1 (BV570; clone PK136; BioLegend), CD3 (BV650; clone 145-2C11; BD Biosciences), B220 (Pacific Blue; clone RA3-6B2; BioLegend), CD16.2 (APC; clone 9E9; BioLegend), CD16 (PE; clone S17014E; BioLegend), fixable viability dye (eFluor 510), True-Stain Monocyte Blocker (5 μl per sample), and Brilliant Stain Buffer Plus (10 μl per sample) and then fixed with 4% PFA in PBS for 20 min at room temperature. All antibodies were used at a dilution of 1:200 with the exceptions of MHC II BV711, which was used at 1:300 and the viability dye, used at 1:100. In experiments staining for mouse FcγR III, Fc block was not used. Absolute cell counts were determined using Precision Count Beads (Biolegend). Flow cytometry data were acquired on a 3 laser Cytek Aurora cytometer (Cytekbio) and analysed using FlowJo software (version 10.4.2).
Measurement of viral burden
Tissues were weighed and homogenized with zirconia beads in a MagNA Lyser instrument (Roche Life Science) in 1 ml of Dulbecco’s modified Eagle medium supplemented with 2% heat-inactivated FBS. Tissue homogenates were clarified by centrifugation at 10,000g for 5 min and stored at −80 °C. RNA was extracted using the MagMax mirVana Total RNA isolation kit (Thermo Fisher Scientific) on the Kingfisher Flex extraction robot (Thermo Fisher Scientific). RNA was reverse transcribed and amplified using the TaqMan RNA-to-CT 1-Step Kit (Thermo Fisher Scientific). Reverse transcription was carried out at 48 °C for 15 min followed by 2 min at 95 °C. Amplification was accomplished over 50 cycles as follows: 95 °C for 15 s and 60 °C for 1 min. Copies of SARS-CoV-2 N gene RNA in samples were determined using a published assay57.
Antibody isotype and Fc-receptor binding profiling
Serum samples were analysed by a customized Luminex assay to quantify the levels of antigen-specific antibody subclasses and FcγR binding profiles, as previously described58. Briefly, antigens were coupled to magnetic Luminex beads (Luminex Corp) by carbodiimide-NHS ester coupling (Thermo Fisher). The antigen-coupled microspheres were washed and incubated with heat-inactivated serum samples at an appropriate sample dilution (1:100–1:400 for antibody isotyping and 1:1,000 for all low-affinity FcγRs) overnight in 384-well plates with continuous shaking (Greiner Bio-One). Unbound antibodies were washed away using the magnetic 384-well HydroSpeed Plate Washer (Tecan) using 1× Luminex assay buffer (1× PBS, 0.1% BSA and 0.05% Tween-20). Secondary antibodies (all from Southern Biotech; PE-coupled anti-IgG1, IgG2b, IgG2c and IgG3) were added at a 1:500 dilution and incubated for 1 h at room temperature with continuous shaking. Unbound complexes were washed away using the magnetic 384-well Hydrospeed Plate Washer. Beads were resuspended in 40 µl of QSOL buffer (Sartorius) and then run on the iQue Screener PLUS (Intellicyt) using a customized gating strategy for each bead region (Extended Data Fig. 1). Median fluorescence intensity was calculated for all samples, which were run in technical replicates.
For FcγR binding, sera were incubated with antigen-coated beads and washed as described above. Custom synthesized FcγR (FcγR IIb, FcγR III and FcγR IV; Duke Protein Production facility) were biotinylated and then bound to PE–streptavidin. The labelled FcγRs were then incubated with the sera for 1 h at room temperature with continuous shaking. Unbound complexes were washed as indicated above. Beads were resuspended in 40 µl of QSOL buffer and then run on the iQue Screen PLUS (Intellicyt). All flow cytometry files were analysed using Intellicyt ForeCyt (v8.1).
All antigens and FcγRs were equilibrated in 1× PBS using Zeba-Spin desalting and size exclusion chromatography columns (Thermo Fisher) before bead coupling. Dilution curves for each antibody isotype and subclass and FcγRs were performed for each antigen to ensure reported values were within the linear range of detection. Binding for antigens was calculated as the fold increase relative to naïve levels, which arbitrarily were set to 1.
Evaluation of antibody-mediated effector functions
ADCP and ADNP were evaluated using a flow-cytometry-based phagocytic assay using fluorescently labelled microspheres. In brief, fluorescent neutravidin microspheres were coupled with biotinylated antigens and then incubated with serum (diluted 1:100 in PBS) to form immune complexes. Bead-bound immune complexes were incubated with monocytes (THP-1 cells) or neutrophils overnight at 37 °C in 96-well, round-bottom plates (Corning). After overnight incubation, plates were centrifuged and non-bound beads/immune complexes were removed. Cells were then fixed in 4% PFA and stained for indicated markers. Neutrophils were stained using anti-CD66b-Pacific Blue at a 1:20 dilution in PBS. Microsphere-positive cells were quantified through flow cytometry and phagocytic scores were calculated as (the percentage of microsphere-positive cells × mean fluorescence intensity of positive cells)/100,000.
ADCD was quantified through the coupling of biotinylated antigens to neutravidin microspheres with serum (diluted 1:10) and then incubated with guinea pig complement at 37 °C for 50 min. Reactions were quenched with 15 mM EDTA. Fluorescein-conjugated anti-C3b (diluted 1:500) was added to the mixture for 1 h with constant shaking. Plates were washed with 1× PBS, and complexes were fixed with 4% PFA and washed again with 1× PBS. Beads were resuspended in 35 µl of 1× PBS and then analysed by flow cytometry. Complement deposition was calculated as the fold increase relative to naïve levels, which arbitrarily were set to 1.
ADNKA was quantified through the surface expression of CD107a (as a marker for degranulation). In brief, buffy coats were obtained from healthy donors at Massachusetts General Hospital and enriched using the RosetteSep NK enrichment kit (Stemcell) in the presence of IL-15. Antigen-coated, 96-well ELISA plates were then incubated with serum (1:50 dilution in PBS) and the NK cell preparation. Plates were placed in a 37 °C incubator for 2 h. Reactions were stopped and cells were fixed and stained extracellularly with anti-CD107a-PeCy5 (diluted 1:15 in PBS), anti-CD3-PB (1:30 dilution), anti-CD56-PE-Cy7 (1:30 dilution) and anti-CD16-APC-Cy7 (1:30 dilution). Cells were washed three times with 1× PBS and resuspended in 30 µl of 1× PBS and analysed by flow cytometry.
T-cell analysis
Spleens were collected from control or mRNA-1273 vaccinated wild-type or FcγR KO mice at day 10 after boosting, and single-cell suspensions were generated after tissue disruption and passage through a 70 μm cell strainer. Splenocytes were pelleted by centrifugation (300g, 5 min), and erythrocytes were lysed using ACK lysis buffer (Thermo Fisher). Cells then were resuspended in RPMI 1640 medium supplemented with 10% FBS, 1% HEPES, 1% l-glutamine and 0.1% β-mercaptoethanol. For peptide stimulation, splenocytes were incubated separately with class I MHC (VL8, peptide sequence S539-546: VNFNFNGL25) or class II MHC ((#62, peptide sequence S62-76: VTWFHAIHVSGTNGT24) immunodominant spike peptides (1 μg ml−1) overnight at 37 °C in the presence of Brefeldin A (1:500, Invitrogen). The following day, cells were washed and stained with Fc block (1:200, Clone 93; cat. no. 101320; BioLegend), CD45 (1:200, BUV395; Clone 30-F11; cat. no. 564279; BD Biosciences), CD8β (1:200, PerCP/Cy5.5; Clone YTS156.7.7; cat. no. 126610; BioLegend), CD4 (1:200, FITC; Clone GK1.5; cat. no. 100406; BioLegend) and CD44 (1:100, APC/Cy7; Clone IM7; cat. no. 103028; BioLegend) for 30 min at 4 °C in FACS buffer (1× PBS with 2% FCS and 2 mM EDTA). Dead cells were excluded using Live/Dead (Thermo Fisher) that was added concurrently with staining. Following this, cells were washed and fixed with and stained for intracellular IFN-γ (1:200, APC; Clone XMG1.2; cat. no. 505810; BioLegend) and TNF-α (1:200, PE/Cy7; Clone MP6-XT22; cat. no. 25-7321-82; Invitrogen) using BD fixation/permeabilization kit (BD Biosciences) according to the manufacturer’s instructions. Cells were processed by flow cytometry on a Cytek Aurora and analysed using FlowJo software version 10.4.2.
Statistical analysis
Statistical significance was assigned when P values were <0.05 using GraphPad Prism version 9.3. Tests, number of animals, median values and statistical comparison groups are indicated in the figure legends. Changes in infectious virus titre, viral RNA levels or serum antibody responses were analysed by one-way analysis of variance (ANOVA) with a post-test correction when comparing three or more groups. When comparing two groups, a Mann–Whitney test was performed and a Bonferrroni correction was used to account multiple independent comparisons. Best-fit lines were calculated using non-linear regression analyses. Two mice from the passive transfer experiments were excluded from analysis after the testing of their sera and confirming an absence of immune antibodies, which established unsuccessful serum transfers.
Materials availability
All requests for resources and reagents should be directed to the Lead Contact author. This includes viruses, proteins, vaccines and primer-probe sets. All reagents will be made available on request after completion of a Materials Transfer Agreement. The pre-clinical mRNA vaccines (control and mRNA-1273) can be obtained under a Materials Transfer Agreement with Moderna (contact: Darin Edwards, Darin.Edwards@modernatx.com).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the paper, its Extended data or Source data files. Any additional information related to the study also is available from the corresponding author upon request. Source data are provided with this paper.
Code availability
No code was used in the course of the data acquisition or analysis.
References
Amanat, F. & Krammer, F. SARS-CoV-2 vaccines: status report. Immunity 52, 583–589 (2020).
Bates, T. A. et al. Neutralization of SARS-CoV-2 variants by convalescent and BNT162b2 vaccinated serum. Nat. Commun. 12, 5135 (2021).
Khoury, D. S. et al. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat. Med. 27, 1205–1211 (2021).
Kaplonek, P. et al. mRNA-1273 vaccine-induced antibodies maintain Fc effector functions across SARS-CoV-2 variants of concern. Immunity 55, 355–365.e354 (2022).
Hederman, A. P., et al. SARS-CoV-2 mRNA vaccination elicits broad and potent Fc effector functions to VOCs in vulnerable populations. Preprint at medRxiv https://doi.org/10.1101/2022.09.15.22280000 (2022).
Chalkias, S. et al. A bivalent Omicron-containing booster vaccine against Covid-19. N. Engl. J. Med. 387, 1279–1291 (2022).
Cele, S. et al. Omicron extensively but incompletely escapes Pfizer BNT162b2 neutralization. Nature 602, 654–656 (2022).
Andrews, N. et al. Covid-19 vaccine effectiveness against the Omicron (B.1.1.529) variant. N. Engl. J. Med. 386, 1532–1546 (2022).
Wang, Q. et al. Antibody evasion by SARS-CoV-2 Omicron subvariants BA.2.12.1, BA.4 and BA.5. Nature 608, 603–608 (2022).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature 602, 664–667 (2022).
Grunst, M. W. & Uchil, P. D. Fc effector cross-reactivity: a hidden arsenal against SARS-CoV-2’s evasive maneuvering. Cell Rep. Med. 3, 100540 (2022).
Zhu, D. Y. et al. Defining the determinants of protection against SARS-CoV-2 infection and viral control in a dose-down Ad26.CoV2.S vaccine study in nonhuman primates. PLoS Biol. 20, e3001609 (2022).
Tarke, A. et al. SARS-CoV-2 vaccination induces immunological T cell memory able to cross-recognize variants from Alpha to Omicron. Cell 185, 847–859.e811 (2022).
Zohar, T. et al. Compromised humoral functional evolution tracks with SARS-CoV-2 mortality. Cell 183, 1508–1519.e1512 (2020).
Case, J. B. et al. Resilience of S309 and AZD7442 monoclonal antibody treatments against infection by SARS-CoV-2 Omicron lineage strains. Nat. Commun. 13, 3824 (2022).
Bates, T. A. et al. BNT162b2-induced neutralizing and non-neutralizing antibody functions against SARS-CoV-2 diminish with age. Cell Rep. 41, 111544 (2022).
Richardson, S. I. et al. SARS-CoV-2 Omicron triggers cross-reactive neutralization and Fc effector functions in previously vaccinated, but not unvaccinated, individuals. Cell Host Microbe 30, 880–886.e884 (2022).
Ackerman, M. E., Barouch, D. H. & Alter, G. Systems serology for evaluation of HIV vaccine trials. Immunol. Rev. 275, 262–270 (2017).
Dinnon, K. H. 3rd et al. A mouse-adapted model of SARS-CoV-2 to test COVID-19 countermeasures. Nature 586, 560–566 (2020).
Cobb, R. R. et al. A combination of two human neutralizing antibodies prevents SARS-CoV-2 infection in cynomolgus macaques. Med 3, 188–203.e184 (2022).
Winkler, E. S. et al. SARS-CoV-2 causes lung infection without severe disease in human ACE2 knock-in mice. J. Virol. 96, Jvi0151121 (2021).
Gu, H. et al. Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy. Science 369, 1603–1607 (2020).
Ying, B. et al. Boosting with variant-matched or historical mRNA vaccines protects against Omicron infection in mice. Cell 185, 1572–1587.e1511 (2022).
Zhuang, Z. et al. Mapping and role of T cell response in SARS-CoV-2-infected mice. J. Exp. Med. 218, e20202187 (2021).
Li, C. et al. Mechanisms of innate and adaptive immunity to the Pfizer-BioNTech BNT162b2 vaccine. Nat. Immunol. 23, 543–555 (2022).
Chong, Z. et al. Nasally delivered interferon-λ protects mice against infection by SARS-CoV-2 variants including Omicron. Cell Rep. 39, 110799 (2022).
Laidlaw, B. J. et al. Cooperativity between CD8+ T cells, non-neutralizing antibodies, and alveolar macrophages is important for heterosubtypic influenza virus immunity. PLoS Pathog. 9, e1003207 (2013).
Accorsi, E. K. et al. Association between 3 doses of mRNA COVID-19 vaccine and symptomatic infection caused by the SARS-CoV-2 Omicron and Delta variants. JAMA 327, 639–651 (2022).
Chemaitelly, H. et al. mRNA-1273 COVID-19 vaccine effectiveness against the B.1.1.7 and B.1.351 variants and severe COVID-19 disease in Qatar. Nat. Med. 27, 1614–1621 (2021).
Wang, Z. et al. Memory B cell responses to Omicron subvariants after SARS-CoV-2 mRNA breakthrough infection in humans. J. Exp. Med. 219, e202210006 (2022).
Brasu, N. et al. Memory CD8+ T cell diversity and B cell responses correlate with protection against SARS-CoV-2 following mRNA vaccination. Nat. Immunol. 23, 1445–1456 (2022).
Kaplonek, P. et al. mRNA-1273 and BNT162b2 COVID-19 vaccines elicit antibodies with differences in Fc-mediated effector functions. Sci. Transl. Med. 14, eabm2311 (2022).
Winkler, E. S. et al. Human neutralizing antibodies against SARS-CoV-2 require intact Fc effector functions for optimal therapeutic protection. Cell 184, 1804–1820.e1816 (2021).
Beaudoin-Bussières, G. et al. A Fc-enhanced NTD-binding non-neutralizing antibody delays virus spread and synergizes with a nAb to protect mice from lethal SARS-CoV-2 infection. Cell Rep. 38, 110368 (2022).
Yamin, R. et al. Fc-engineered antibody therapeutics with improved anti-SARS-CoV-2 efficacy. Nature 599, 465–470 (2021).
Mades, A. et al. Detection of persistent SARS-CoV-2 IgG antibodies in oral mucosal fluid and upper respiratory tract specimens following COVID-19 mRNA vaccination. Sci. Rep. 11, 24448 (2021).
Heyman, B. Antibodies as natural adjuvants. Curr. Top. Microbiol. Immunol. 382, 201–219 (2014).
Hamano, Y., Arase, H., Saisho, H. & Saito, T. Immune complex and Fc receptor-mediated augmentation of antigen presentation for in vivo Th cell responses. J. Immunol. 164, 6113–6119 (2000).
Fransen, M. F. et al. A restricted role for FcγR in the regulation of adaptive immunity. J. Immunol. 200, 2615–2626 (2018).
Scheaffer, S. M. et al. Bivalent SARS-CoV-2 mRNA vaccines increase breadth of neutralization and protect against the BA.5 Omicron variant in mice. Nat. Med. 29, 247–257 (2022).
Davis-Gardner, M. E., et al. Neutralization against BA.2.75.2, BQ.1.1, and XBB from mRNA Bivalent Booster. N. Engl. J. Med. 388, 183–185 (2023).
Nimmerjahn, F. et al. FcγRIV deletion reveals its central role for IgG2a and IgG2b activity in vivo. Proc. Natl Acad. Sci. USA 107, 19396–19401 (2010).
Halfmann, P. J. et al. SARS-CoV-2 Omicron virus causes attenuated disease in mice and hamsters. Nature 603, 687–692 (2022).
Junqueira, C. et al. FcγR-mediated SARS-CoV-2 infection of monocytes activates inflammation. Nature 606, 576–584 (2022).
Sefik, E. et al. Inflammasome activation in infected macrophages drives COVID-19 pathology. Nature 606, 585–593 (2022).
Zang, R. et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 5, eabc3582 (2020).
Chen, R. E. et al. Resistance of SARS-CoV-2 variants to neutralization by monoclonal and serum-derived polyclonal antibodies. Nat. Med. 27, 717–726 (2021).
Chen, R. E. et al. In vivo monoclonal antibody efficacy against SARS-CoV-2 variant strains. Nature 596, 103–108 (2021).
Ioan-Facsinay, A. et al. FcgammaRI (CD64) contributes substantially to severity of arthritis, hypersensitivity responses, and protection from bacterial infection. Immunity 16, 391–402 (2002).
Botto, M. et al. Homozygous C1q deficiency causes glomerulonephritis associated with multiple apoptotic bodies. Nat. Genet. 19, 56–59 (1998).
Nelson, J. et al. Impact of mRNA chemistry and manufacturing process on innate immune activation. Sci. Adv. 6, eaaz6893 (2020).
Corbett, K. S. et al. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness. Nature 586, 567–571 (2020).
Hassett, K. J. et al. Optimization of lipid nanoparticles for intramuscular administration of mRNA vaccines. Mol. Ther. Nucleic Acids 15, 1–11 (2019).
Hassan, A. O. et al. A single intranasal dose of chimpanzee adenovirus-vectored vaccine protects against SARS-CoV-2 infection in rhesus macaques. Cell Rep. Med. 2, 100230 (2021).
Amanat, F. et al. SARS-CoV-2 mRNA vaccination induces functionally diverse antibodies to NTD, RBD, and S2. Cell 184, 3936–3948.e3910 (2021).
VanBlargan, L. A. et al. A potently neutralizing SARS-CoV-2 antibody inhibits variants of concern by utilizing unique binding residues in a highly conserved epitope. Immunity 54, 2399–2416.e6 (2021).
Case, J. B., Bailey, A. L., Kim, A. S., Chen, R. E. & Diamond, M. S. Growth, detection, quantification, and inactivation of SARS-CoV-2. Virology 548, 39–48 (2020).
Brown, E. P. et al. Multiplexed Fc array for evaluation of antigen-specific antibody effector profiles. J. Immunol. Methods 443, 33–44 (2017).
Acknowledgements
This study was supported by the NIH (R01 AI157155, NIAID Centers of Excellence for Influenza Research and Response (CEIRR) contracts 75N93021C00014 and 75N93019C00051, to M.S.D.; R01 AI110700 to R.S.B., T32 AI007172 to S.R.M. and P01 AI1650721 to G.A. and R.P.M.). Work at the Ragon Institute is also supported by M. and L. Schwartz and T. and S. Ragon. We thank M. Suthar (Emory University) for the BA.5 isolate used in this study and T. Wang (Stanford University) for critical comments on the manuscript. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.R.M. performed binding and neutralization assays, immunization, passive transfer, depletion studies, challenge experiments and flow cytometry. P.D. performed and analysed T-cell responses. B.M.W. performed mouse experiments. C.E.K. performed some of the flow cytometry experiments. M.L. performed immune cell processing and staining. R.S.B. provided the mouse-adapted MA-10 strain. R.P.M., T.M.C. and G.A. designed, performed and analysed the serological profiling experiments. D.K.E. provided mRNA vaccines and helped to design vaccination experiments. S.R.M. and M.S.D. designed studies and wrote the initial draft, with the other authors providing editorial comments.
Corresponding author
Ethics declarations
Competing interests
M.S.D. is a consultant for Inbios, Vir Biotechnology, Senda Biosciences, Moderna, Ocugen and Immunome. The Diamond laboratory has received unrelated funding support in sponsored research agreements from Vir Biotechnology, Emergent BioSolutions, Generate Biomedicines and Moderna. R.S.B. is a member of the Scientific Advisory Board of VaxArt and Adagio, has consulted for Takeda and received unrelated funding from J&J and Pfizer. G.A. is a founder/equity holder in Seroymx Systems and Leyden Labs and has served as a scientific advisor for Sanofi Vaccines. G.A. has collaborative agreements with GlaxoSmithKline, Merck, Abbvie, Sanofi, Medicago, BioNtech, Moderna, BMS, Novavax, SK Biosciences, Gilead and Sanaria. D.K.E. and G.A. are employees and shareholders in Moderna, Inc. All other authors declare no conflicts of interest.
Peer review
Peer review information
Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gating strategy for Luminex-based and Fc effector function assays.
(a) Gating for Luminex-bead based antibody binding to spike-coated beads. (b) Gating for ADNP assay showing CD66+ neutrophils with opsinophagocytosed beads. (c) Gating for ADCP assay showing THP-1 monocytes and opsinophagocytosed beads. (d) Gating for ADCD assay showing complement deposition on spike and antibody coated beads. (e) Gating for NK cell activation assay showing CD107a expression.
Extended Data Fig. 2 T cell responses in mRNA-1273 vaccinated wild-type and FcγR KO mice.
(a) Representative flow cytometry plots show gating scheme for quantification of spike-specific CD4+ and CD8+ T cell responses in the spleen of wild-type, FcγR III KO and FcγR I/III/IV KO mice at day 10 after boosting with control or mRNA-1273 vaccines. (b-c) At day 10 after boosting, the spleen of wild-type and FcγR KO mice were harvested and T cell responses were measured after spike peptide re-stimulation. Splenocytes were incubated overnight with class I MHC (b) or class II MHC (c) immunodominant spike peptides, and the percentages and numbers of IFNγ and TNFα positive CD8+ (b) or CD4+ (c) T cells were quantified by intracellular staining and flow cytometry. Data are pooled from two experiments (in order left to right n = 10, 10, 8, 10, 10, 9 (b-c)). Comparisons were made between groups that received the mRNA 1273 vaccine (one-way ANOVA with Tukey’s post-test; all comparisons were not significant; column height indicates mean values).
Extended Data Fig. 3 Levels of anti-BA.5 antibody in mice passively transferred vaccine-elicited serum antibody.
(a) Levels of antiBA.5 spike IgG in serum of mice that were passively transferred naïve or immune sera. Amounts are compared to those in vaccine-elicited immune serum before transfer (n = 5 mice per group, columns indicate mean values). (b) Neutralizing antibody response against SARS-CoV-2 BA.5 using sera from naïve (circles) or spike protein vaccinated (grey squares) mice. Also shown is serum neutralizing antibody activity from recipient wild-type mice one day after transfer of immune sera (black squares). Data are representative of results with n = 5 mice per group.
Extended Data Fig. 4 FcγR expression on myeloid cells in the lung.
Lung cells from wild-type, FcγR I KO, FcγR III KO, and FcγR I/III/IV KO mice were stained with antibodies for FcγR I, FcγR III, or FcγR IV. After gating on live cells, alveolar macrophages, neutrophils and monocytes were defined (see Extended Data Fig. 5). The data are representative of results with n = 3 mice per group, and histograms are shown.
Extended Data Fig. 5 Gating scheme for analysis of cell populations in the blood and lung.
(a) Immune cell populations in the blood of C57BL/6 mice were analyzed using the indicated gating scheme and conjugated antibodies. After gating on live single cells, monocytes (P5) were defined as CD45+ CD11bhi Ly6Chi; neutrophils (P6) were defined as CD45+ CD11bhi Ly6Ghi; natural killer (NK) cells (P7) were defined as CD45+ NK1.1+; and B cells (P8) were defined as CD45+ B220+ cells. (b) Immune cell populations in the lungs of C57BL/6 mice were analyzed using the indicated gating scheme and conjugated antibodies. After gating on live single cells, alveolar macrophages (P1) were defined as CD45+ CD11c+ Siglec-F+; interstitial macrophages (P2) were defined as CD45+ CD64+, eosinophils (P3) were defined as CD45+ CD11b+ Siglec-F+; CD11b dendritic cells (P4) were defined CD45+ CD11b+ CD11c+, Siglec-F-, MHC II+; monocytes (P5) were defined as CD45+ Ly6Chi; neutrophils (P6) were identified as CD45+ Ly6G+; natural killer (NK) cells (P7) were defined as CD45+ NK1.1+; and B cells (P8) were defined as CD45+ B220+.
Supplementary information
Supplementary Tables 1–4
Supplementary Table 1. Mouse FcγR expression in immune cells of the lung. Supplementary Table 2. Human FcγR expression in immune cells of the lung. Supplementary Table 3. Additional statistical comparisons. Supplementary Table 4. Deep sequencing results.
Source data
Source Data Fig. 1
Excel file with source data for graphs.
Source Data Fig. 2
Excel file with source data for graphs.
Source Data Fig. 3
Excel file with source data for graphs.
Source Data Fig. 4
Excel file with source data for graphs.
Source Data Fig. 5
Excel file with source data for graphs.
Source Data Extended Data Fig. 2
Excel file with source data for graphs.
Source Data Extended Data Fig. 3
Excel file with source data for graphs.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mackin, S.R., Desai, P., Whitener, B.M. et al. Fc-γR-dependent antibody effector functions are required for vaccine-mediated protection against antigen-shifted variants of SARS-CoV-2. Nat Microbiol 8, 569–580 (2023). https://doi.org/10.1038/s41564-023-01359-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01359-1
This article is cited by
-
Mucosal vaccine-induced cross-reactive CD8+ T cells protect against SARS-CoV-2 XBB.1.5 respiratory tract infection
Nature Immunology (2024)
-
B-cell and antibody responses to SARS-CoV-2: infection, vaccination, and hybrid immunity
Cellular & Molecular Immunology (2023)
-
An FcRn-targeted mucosal vaccine against SARS-CoV-2 infection and transmission
Nature Communications (2023)
-
Antibody Fc-binding profiles and ACE2 affinity to SARS-CoV-2 RBD variants
Medical Microbiology and Immunology (2023)