Abstract
Staphylococcus aureus invades cells and persists intracellularly, causing persistent inflammation that is notoriously difficult to treat. Here we investigated host–pathogen interactions underlying intracellular S. aureus infection in macrophages and discovered that the endoplasmic reticulum (ER) is an important cellular compartment for intracellular S. aureus infection. Using CRISPR–Cas9 guide RNA library screening, we determined that the autocrine motility factor receptor (AMFR), an ER-resident E3 ubiquitin ligase, played an essential role in mediating intracellular S. aureus-induced inflammation. AMFR directly interacted with TAK1-binding protein 3 (TAB3) in the ER, inducing K27-linked polyubiquitination of TAB3 on lysine 649 and promoting TAK1 activation. Moreover, the virulence factor γ-haemolysin B (HIgB) of S. aureus bound to the AMFR and regulated TAB3. Our findings highlight an unknown role of AMFR in intracellular S. aureus infection-induced pneumonia and suggest that pharmacological interruption of AMFR-mediated TAB3 signalling cascades and HIgB targeting may prevent invasive staphylococci-mediated pneumonia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the article. MRSA strain sequences used in this work are available at https://www.ncbi.nlm.nih.gov/. Source data are provided with this paper.
References
Tong, S. Y., Davis, J. S., Eichenberger, E., Holland, T. L. & Fowler, V. G. Jr. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol Rev. 28, 603–661 (2015).
Lee, A. S. et al. Methicillin-resistant Staphylococcus aureus. Nat. Rev. Dis. Prim. 4, 18033 (2018).
Peyrusson, F. et al. Intracellular Staphylococcus aureus persisters upon antibiotic exposure. Nat. Commun. 11, 2200 (2020).
Sendi, P. & Proctor, R. A. Staphylococcus aureus as an intracellular pathogen: the role of small colony variants. Trends Microbiol. 17, 54–58 (2009).
Strobel, M. et al. Post-invasion events after infection with Staphylococcus aureus are strongly dependent on both the host cell type and the infecting S. aureus strain. Clin. Microbiol Infect. 22, 799–809 (2016).
Ziegler, C. et al. The dynamics of T cells during persistent Staphylococcus aureus infection: from antigen-reactivity to in vivo anergy. EMBO Mol. Med. 3, 652–666 (2011).
Stelzner, K. et al. Intracellular Staphylococcus aureus employs the cysteine protease staphopain A to induce host cell death in epithelial cells. PLoS Pathog. 17, e1009874 (2021).
Peschel, A. & Otto, M. Phenol-soluble modulins and staphylococcal infection. Nat. Rev. Microbiol. 11, 667–673 (2013).
Schwarz, D. S. & Blower, M. D. The endoplasmic reticulum: structure, function and response to cellular signaling. Cell. Mol. Life Sci. 73, 79–94 (2016).
Pillich, H., Loose, M., Zimmer, K. P. & Chakraborty, T. Diverse roles of endoplasmic reticulum stress sensors in bacterial infection. Mol. Cell Pediatr. 3, 9 (2016).
Moretti, J. et al. STING senses microbial viability to orchestrate stress-mediated autophagy of the endoplasmic reticulum. Cell 171, 809–823 e813 (2017).
Magadan, J. G. et al. Multilayered mechanism of CD4 downregulation by HIV-1 Vpu involving distinct ER retention and ERAD targeting steps. PLoS Pathog. 6, e1000869 (2010).
Celli, J. & Tsolis, R. M. Bacteria, the endoplasmic reticulum and the unfolded protein response: friends or foes? Nat. Rev. Microbiol. 13, 71–82 (2015).
Tilney, L. G., Harb, O. S., Connelly, P. S., Robinson, C. G. & Roy, C. R. How the parasitic bacterium Legionella pneumophila modifies its phagosome and transforms it into rough ER: implications for conversion of plasma membrane to the ER membrane. J. Cell Sci. 114, 4637–4650 (2001).
Robinson, C. G. & Roy, C. R. Attachment and fusion of endoplasmic reticulum with vacuoles containing Legionella pneumophila. Cell Microbiol. 8, 793–805 (2006).
Menzies, S. A. et al. The sterol-responsive RNF145 E3 ubiquitin ligase mediates the degradation of HMG-CoA reductase together with gp78 and Hrd1. eLife 7, e40009 (2018).
Joshi, V. et al. Gp78 involvement in cellular proliferation: can act as a promising modulator for cell cycle regulatory proteins? J. Cell. Physiol. 233, 6352–6368 (2018).
Li, L. et al. p38 MAP kinase-dependent phosphorylation of the Gp78 E3 ubiquitin ligase controls ER–mitochondria association and mitochondria motility. Mol. Biol. Cell 26, 3828–3840 (2015).
Joshi, V., Upadhyay, A., Kumar, A. & Mishra, A. Gp78 E3 ubiquitin ligase: essential functions and contributions in proteostasis. Front. Cell Neurosci. 11, 259 (2017).
Zhang, H. et al. AMFR drives allergic asthma development by promoting alveolar macrophage-derived GM-CSF production. J. Exp. Med. 219, e20211828 (2022).
Kanayama, A. et al. TAB2 and TAB3 activate the NF-κB pathway through binding to polyubiquitin chains. Mol. Cell 15, 535–548 (2004).
Chen, Z., Zhou, Y., Zhang, Z. & Song, J. Towards more accurate prediction of ubiquitination sites: a comprehensive review of current methods, tools and features. Brief. Bioinformatics 16, 640–657 (2015).
Kaufmann, S. H. Immunity to intracellular bacteria. Annu. Rev. Immunol. 11, 129–163 (1993).
Fraunholz, M. & Sinha, B. Intracellular Staphylococcus aureus: live-in and let die. Front Cell Infect. Microbiol. 2, 43 (2012).
Soe, Y. M., Bedoui, S., Stinear, T. P. & Hachani, A. Intracellular Staphylococcus aureus and host cell death pathways. Cell Microbiol. 23, e13317 (2021).
Bravo-Santano, N. et al. Intracellular Staphylococcus aureus modulates host central carbon metabolism to activate autophagy. mSphere 3, e00374–18 (2018).
Stelzner, K. et al. Intracellular Staphylococcus aureus perturbs the host cell Ca2+ homeostasis to promote cell death. mBio 11, e02250–20 (2020).
Bravo-Santano, N. et al. Identification of novel targets for host-directed therapeutics against intracellular Staphylococcus aureus. Sci. Rep. 9, 15435 (2019).
Abuaita, B. H., Burkholder, K. M., Boles, B. R. & O’Riordan, M. X. The endoplasmic reticulum stress sensor inositol-requiring enzyme 1α augments bacterial killing through sustained oxidant production. mBio 6, e00705 (2015).
Lizak, M. & Yarovinsky, T. O. Phospholipid scramblase 1 mediates type i interferon-induced protection against staphylococcal alpha-toxin. Cell Host Microbe 11, 70–80 (2012).
Wang, Q. et al. The E3 ubiquitin ligase AMFR and INSIG1 bridge the activation of TBK1 kinase by modifying the adaptor STING. Immunity 41, 919–933 (2014).
Jacobs, J. L., Zhu, J., Sarkar, S. N. & Coyne, C. B. Regulation of mitochondrial antiviral signaling (MAVS) expression and signaling by the mitochondria-associated endoplasmic reticulum membrane (MAM) protein Gp78. J. Biol. Chem. 289, 1604–1616 (2014).
Xu, Y. R. & Lei, C. Q. TAK1–TABs complex: a central signalosome in inflammatory responses. Front. Immunol. 11, 608976 (2020).
Ishitani, T. et al. Role of the TAB2-related protein TAB3 in IL-1 and TNF signaling. EMBO J. 22, 6277–6288 (2003).
Cheung, P. C., Nebreda, A. R. & Cohen, P. TAB3, a new binding partner of the protein kinase TAK1. Biochem. J. 378, 27–34 (2004).
Wang, J. et al. Mycobacterium tuberculosis suppresses innate immunity by coopting the host ubiquitin system. Nat. Immunol. 16, 237–245 (2015).
Qian, Y., Commane, M., Ninomiya-Tsuji, J., Matsumoto, K. & Li, X. IRAK-mediated translocation of TRAF6 and TAB2 in the interleukin-1-induced activation of NFκB. J. Biol. Chem. 276, 41661–41667 (2001).
Zhou, Q. & Zhang, J. K27-linked noncanonic ubiquitination in immune regulation. J. Leukoc. Biol. 111, 223–235 (2022).
Cheung, G. Y. C., Bae, J. S. & Otto, M. Pathogenicity and virulence of Staphylococcus aureus. Virulence 12, 547–569 (2021).
McGilligan, V. E. et al. Staphylococcus aureus activates the NLRP3 inflammasome in human and rat conjunctival goblet cells. PLoS ONE 8, e74010 (2013).
Hodille, E. et al. Delta hemolysin and phenol-soluble modulins, but not alpha hemolysin or Panton–Valentine leukocidin, induce mast cell activation. Front. Cell Infect. Microbiol 6, 180 (2016).
Parcina, M. et al. Pathogen-triggered activation of plasmacytoid dendritic cells induces IL-10-producing B cells in response to Staphylococcus aureus. J. Immunol. 190, 1591–1602 (2013).
Ishii, K. et al. Induction of virulence gene expression in Staphylococcus aureus by pulmonary surfactant. Infect. Immun. 82, 1500–1510 (2014).
Gronnemose, R. B. et al. Bacteria-host transcriptional response during endothelial invasion by Staphylococcus aureus. Sci. Rep. 11, 6037 (2021).
Zhu, Z. et al. Nedd8 modification of Cullin-5 regulates lipopolysaccharide-induced acute lung injury. Am. J. Physiol. Lung Cell. Mol. Physiol. 313, L104–L114 (2017).
Zhu, Z. et al. Cutting edge: a cullin-5–TRAF6 interaction promotes TRAF6 polyubiquitination and lipopolysaccharide signaling. J. Immunol. 197, 21–26 (2016).
Huang, L.N. et al. p38α MAP kinase promotes asthmatic inflammation through modulation of alternatively activated macrophages. J. Mol. Cell. Biol. 11, 1095–1097 (2019).
Charan, J. & Kantharia, N. D. How to calculate sample size in animal studies? J. Pharm. Pharmacother. 4, 303–306 (2013).
Qian, F. et al. A non-redundant role for MKP5 in limiting ROS production and preventing LPS-induced vascular injury. EMBO J. 28, 2896–2907 (2009).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein–protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Williamson, C. D., Wong, D. S., Bozidis, P., Zhang, A. & Colberg-Poley, A. M. Isolation of endoplasmic reticulum, mitochondria, and mitochondria-associated membrane and detergent resistant membrane fractions from transfected cells and from human cytomegalovirus-infected primary fibroblasts. Curr. Protoc. Cell Biol. 68, 3 27 21–23 27 33 (2015).
Chen, W., Zhang, Y., Yeo, W. S., Bae, T. & Ji, Q. Rapid and efficient genome editing in Staphylococcus aureus by using an engineered CRISPR/Cas9 system. J. Am. Chem. Soc. 139, 3790–3795 (2017).
Wang, R. et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat. Med. 13, 1510–1514 (2007).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (grant numbers 81973329 and 82173821 to F.Q. and grant numbers 82073858 and 82273934 to L.S.) and Natural Science Foundation of Shanghai (21ZR1432700, to L.S.).
Author information
Authors and Affiliations
Contributions
L.S. and F.Q. conceived the study. Ha.Z., Hu.Z., X.L., Z.W., Y.W., X.Y. and B.G. designed, performed and interpreted experimental data. L.S. D.C., A.Z. and F.Q. wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Michael Otto, Soumen Basak and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 AMFR deficiency in macrophages ameliorates intracellular MRSA-induced cytokine production.
(a) BMDMs were infected with MRSA43300 (dot) and clinical MRSA strain S1-2-4 (circle), S1-2-12 (triangle), and S1-2-22 (square) with an MOI of 8 for 1 h, extracellular MRSA was removed and killed with 300 μg/mL gentamicin and incubated in medium containing gentamicin (50 µg/mL). The number of gentamicin-protected bacteria was determined at 1, 3, and 5 day post-infection by plating for colony forming units. (b) BMDMs were treated with MRSA43300 (MOI 8:1) for 1 h. The gentamicin-protected intracellular MRSA were examined by TEM at 2 h post-infection. Triangles indicate ER membranes, Arrowheads indicate living cells. (c) THP-1 cells were transfected with lentivirus-mediated AMFR shRNA (AMFR shRNA) for 48 h. The knockdown efficiency was detected by western blot. Scrambled shRNA (Scr shRNA) was used as control. (d) qRT-PCR analysis of TNF, IL1B, and IL6 mRNA expression in THP-1 cells transfected with lentivirus-mediated AMFR shRNA or Scrambled shRNA for 48 h and infected with clinical MRSA strains (S1-2-4, S1-2-12, S1-2-14, and S1-2-22) (MOI = 8). The results are shown as the relative levels of gene transcripts, with that of uninfected cells set as 1. (e) Human CD14+ monocytes were transfected with lentivirus-mediated AMFR shRNA. The knockdown efficiency was detected by western blot. Scrambled shRNA (Scr shRNA) was used as control. Data are representative of three independent experiments (b, c, e) and means ± SEM (a-d), n = 3 biological replicates, unpaired two-tailed Student’s t test.
Extended Data Fig. 2 AMFR does not regulate the engulf and killing activity of macrophages against MRSA.
(a) BMDMs were isolated from Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice. Then, cells were incubated with mCherry-MRSA43300 (MOI = 8) for 45 min for phagocytosis assay. (b) Amfr+/+ and Amfr−/− macrophages were infected with mCherry-MRSA43300 (MOI = 8) for 1 h. The cells were then incubated with gentamicin for 1 hours, and intracellular bacterial CFUs were determined at the end of the gentamicin incubation period. Data shown are representative of three independent experiments (a), or presented as means ± SEM (b), n = 3 biological replicates, unpaired two-tailed Student’s t test.
Extended Data Fig. 3 AMFR deficiency in macrophages ameliorates intracellular MRSA-induced pneumonia.
(a) Gating strategy for flow cytometry analysis of the neutrophils (Neu., Ly-6G+) and alveolar macrophages (AMs, CD11c+ F4/80+) in the BALF of MRSA-infected mice. (b) BALF cells were sorted by Flow cytometry following Cytospin preparations and Giemsa staining. Representative neutrophils (Neu.) and alveolar macrophages (AMs) are shown. (c) BALF neutrophils and AMs’ ratios were estimated. Data shown are representative of three independent experiments, or presented as means ± SD from three independent experiments (n = 5 mice per group per experiment), two-way ANOVA (Bonferroni test).
Extended Data Fig. 4 AMFR deficiency dose not alter the antimicrobial functions of neutrophils.
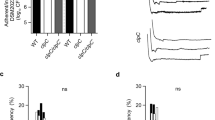
(a) Bone marrow neutrophils were isolated from Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice. Then, cells were infected with mCherry-MRSA43300 (MOI = 8) for 15 min for phagocytosis assay. (b) Amfr+/+ and Amfr−/− neutrophils were infected with mCherry-MRSA43300 (MOI = 8) for 1 h. The cells were then incubated with gentamicin for 1 hours, and intracellular bacterial CFUs were determined at the end of the gentamicin incubation period. (c) The Amfr+/+ and Amfr−/− neutrophils were stimulated with MRSA43300 (MOI = 8), and the superoxide production were measured. Representative tracing showing the production of superoxide. (d) Cumulative superoxide production was quantified based the area under tracing line in c. Data shown are representative of three independent experiments (a, c), or presented as means ± SEM (b, d), n = 3 biological replicates, unpaired two-tailed Student’s t test.
Extended Data Fig. 5 AMFR deficiency have no effect on UPR and ER stress.
(a) qRT-PCR analysis of Chop, Bip, and Atf4 mRNA expression in BMDMs isolated Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice and infected with MRSA43300 (MOI = 8) for 1 h following gentamicin incubation for 11 h or ER stress activator thapsigargin (Thap., 1 mM) for 12 h. (b) Immunoblot analysis of the phosphorylation level of IRE1 in BMDMs isolated from Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice and infected with MRSA (MOI = 8) for 1 h following gentamicin incubation for 11 h or ER stress activator thapsigargin (Thap., 1 mM) for 12 h. Data shown are representative of three independent experiments. (c) qRT-PCR analysis of Edem1 expression in BMDMs isolated Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice and infected with MRSA as above. The results are shown as the relative levels of gene transcripts, with that of unstimulated cells set as 1. Data are means ± SEM (a and c), n = 3 biological replicates, two-way ANOVA (Sidak’s test).
Extended Data Fig. 6 AMFR deficience in macrophages attenuates NF-kB and MAPK signaling activation under MRSA stimulation.
(a) Densitometric analysis of immunoblot analysis of phosphorylated (p-) p65, p38, JNK, ERK and IκBa in Fig. 3a. (b) Immunoblot analysis of indicated protein in cell lysates of THP-1 cells transfected with lentivirus-mediated AMFR shRNA (AMFR shRNA) or Scrambled shRNA (Scr shRNA) and infected with MRSA43300 for the indicated time. (c) Densitometric analysis of immunoblot analysis in b. The samples derive from the same experiment and that gels/blots were processed in parallel. (a, c). Data shown are representative of three independent experiments (b), or presented as means ± SD (a, c), n = 3 biological replicates, two-way ANOVA (Sidak’s test).
Extended Data Fig. 7 AMFR interacts with TAB3 and recruits TAB3 to the ER.
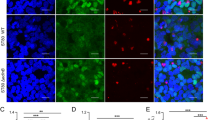
(a) Coimmunoprecipitation and immunoblot analysis of HEK293T cells co-transfected with MYC-TAB3, plus FLAG-AMFR, and FLAG-AMFR mutant vectors. (b) Coimmunoprecipitation and immunoblot analysis of HEK293T cells co-transfected with FLAG-TAB3, plus MYC-TAB3, and MYC-TAB3 mutant vectors. (c) Confocal microscopic imaging of HeLa cells expressing MYC-TAB3 (green) and FLAG-AMFR or FLAG-AMFR mutant vectors (red). Blue indicates ER-Tracker stain of ER. Yellow in merge 1 indicates the co-localization of TAB3 and AMFR. White in merge 2 indicates the co-localization of TAB3, AMFR, and ER. Scale bar, 10 mm. (d) Immunoblot analysis of TAB3 expression in cytoplasm and light membranes (LM, containing ER) of Amfr−/− BMDMs transfected with AMFR, or AMFR mutants infected with MRSA43300. (e) Immunoblot analysis of TAB3 expression in cytoplasm and light membranes (LM, containing ER) of Tab3−/− BMDMs transfected with TAB3, or TAB3 mutants infected with MRSA43300. Data shown are representative of three independent experiments.
Extended Data Fig. 8 AMFR mediates K27-linked polyubiquitination of TAB3 on Lys649, but not regulates the degradation of TAB3.
(a) Immunoblot analysis of ubiquitination of TAB3 in HEK293T cells co-transfected with MYC-TAB3 mutant (K649R) and FLAG-AMFR, along with the mutant ubiquitin HA-Ub (K63). (b) Luciferase assay of NF-κB activation in HEK293T cells transfected with indicated plasmids. (c) Immunoblot analysis of the expression of TAB3 in cell lysates of HEK293T cells co-transfected with MYC-TAB3 along with increasing amount of FLAG-AMFR for 48 h. Before harvest, cells were treated with or without proteasome inhibitor MG-132 (25 mM) or lysosomal inhibitor (NH4CL 10 mM or leupeptin 20 mM) for 6 h. Densitometric analysis of immunoblot analysis was shown below. (d) Coimmunoprecipitation and immunoblot analysis of HEK293T cells transfected with indicated plasmids. Data shown are representative of three independent experiments, or presented as means ± SD, n = 3 biological replicates, two-way ANOVA (Sidak’s test).
Extended Data Fig. 9 MRSA g-hemolysin B (HlgB) binds and promotes AMFR-mediated NF-kB activation.
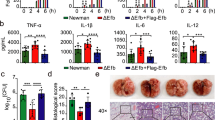
(a) qRT-PCR analysis of Tnf, Il1b, and Il6 mRNA expression in BMDMs isolated from Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice and treated with heat-killed MRSA43300 (HKSA) (MOI = 8). (b) qRT-PCR analysis of Tnf, Il1b, and Il6 mRNA expression in BMDMs isolated from Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice and treated with LPS, or E.coli. (c) qRT-PCR analysis of Tnf, Il1b, and Il6 mRNA expression in BMDMs isolated from Amfrfl/fl (Amfr+/+) and LysM-Cre Amfrfl/fl (Amfr−/−) mice and treated with Poly(I:C). The results are shown as the relative levels of gene transcripts, with that of unstimulated cells set as 1. (d) Luciferase assay of NF-kB activation in HEK293T cells transfected with TAB3 and together with different MRSA leucotoxins expression vector (FLAG-tagged). The expression of each protein was shown below. (e) Densitometric analysis of immunoblot analysis of phosphorylated (p-) p65, p38, JNK, and ERK in Fig. 5c. The samples derive from the same experiment and that gels/blots were processed in parallel. (f) Coimmunoprecipitation and immunoblot analysis of HEK293T cells co-transfected with MYC-AMFR plus FLAG-Hld, or FLAG-HlgC, followed by IP with an anti-MYC Ab. (g) qRT-PCR analysis of Tnf, Il1b, and Il6 mRNA expression in Amfr+/+ and Amfr−/− BMDMs and infected with 43300∆hld. Data are representative of three independent experiments (f), means ± SEM in a-d and g, or means ± SD in e, one-way ANOVA (Tukey’s test), two-way ANOVA (Sidak’s test), or unpaired two-tailed Student’s t test.
Extended Data Fig. 10 Staphylococcus aureus g-hemolysin B (HlgB) plays a critical role in AMFR-mediated pneumonia.
(a) Hematoxylin and eosin staining of lung tissues from indicated mice after MRSA USA300 or USA300∆hlgB exposure (n = 5 per group). (b) Lung injury was assessed by histological scores in different groups. (c) Neutrophils (Neu., Ly-6G+) and alveolar macrophages (AMs, CD11c+ F4/80+) in BALF were analyzed using flow cytometry. (d) Total cell, neutrophils, and alveolar macrophages counts in BALF were estimated. (e) BALF protein were measured. (f) Bacterial burden in lung tissues from Amfrfl/fl and LysM-Cre Amfrfl/fl mice intratracheally injected with USA300 or USA300∆hlgB for the indicated times. Data are representative of three independent experiments (means ± SD) (n = 5 mice per group per experiment), two-way ANOVA (Sidak’s test).
Supplementary information
Supplementary Information
Supplementary Table 1.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, L., Zhang, H., Zhang, H. et al. Staphylococcal virulence factor HlgB targets the endoplasmic-reticulum-resident E3 ubiquitin ligase AMFR to promote pneumonia. Nat Microbiol 8, 107–120 (2023). https://doi.org/10.1038/s41564-022-01278-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01278-7
This article is cited by
-
Cullin5 drives experimental asthma exacerbations by modulating alveolar macrophage antiviral immunity
Nature Communications (2024)
-
Gp78 deficiency in hepatocytes alleviates hepatic ischemia-reperfusion injury via suppressing ACSL4-mediated ferroptosis
Cell Death & Disease (2023)
-
Geometric constraint-triggered collagen expression mediates bacterial-host adhesion
Nature Communications (2023)