Abstract
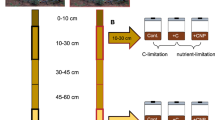
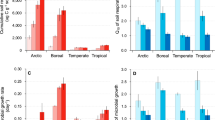
Perturbation of soil microbial communities by rising temperatures could have important consequences for biodiversity and future climate, particularly in tropical forests where high biological diversity coincides with a vast store of soil carbon. We carried out a 2-year in situ soil warming experiment in a tropical forest in Panama and found large changes in the soil microbial community and its growth sensitivity, which did not fully explain observed large increases in CO2 emission. Microbial diversity, especially of bacteria, declined markedly with 3 to 8 °C warming, demonstrating a breakdown in the positive temperature-diversity relationship observed elsewhere. The microbial community composition shifted with warming, with many taxa no longer detected and others enriched, including thermophilic taxa. This community shift resulted in community adaptation of growth to warmer temperatures, which we used to predict changes in soil CO2 emissions. However, the in situ CO2 emissions exceeded our model predictions threefold, potentially driven by abiotic acceleration of enzymatic activity. Our results suggest that warming of tropical forests will have rapid, detrimental consequences both for soil microbial biodiversity and future climate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Trimmed (primers removed) sequence data generated in this study are deposited in the European Nucleotide Archive (ENA) under Project Accession number PRJEB45074 (ERP129199), sample accession numbers ERS6485270–ERS6485284 (16S rRNA) and sample accession numbers ERS6485285–ERS6485299 (ITS). Raw fastq files can be accessed through the Smithsonian figshare at https://doi.org/10.25573/data.14686665 (16S rRNA) and https://doi.org/10.25573/data.14686755 (ITS). Related data and data products for individual analysis workflows are available through the Smithsonian figshare under the collection https://doi.org/10.25573/data.c.5667571.
Code availability
All code, reproducible workflows and further information on data availability can be found on the project website at https://sweltr.github.io/high-temp/. The code embedded in the website is available on GitHub (https://github.com/sweltr/high-temp/) in R Markdown format. The version of code used in this study is archived under SWELTR Workflows v1.0 (https://github.com/sweltr/high-temp), DOI identifier https://zenodo.org/badge/latestdoi/368915237.
References
Cavicchioli, R. et al. Scientists’ warning to humanity: microorganisms and climate change. Nat. Rev. Microbiol. 17, 569–586 (2019).
Jackson, R. B. et al. The ecology of soil carbon: pools, vulnerabilities, and biotic and abiotic controls. Annu. Rev. Ecol. Evol. Syst. 48, 419–445 (2017).
Pan, Y. et al. A large and persistent carbon sink in the world’s forests. Science 333, 988–993 (2011).
Myers, N., Mittermeier, R. A., Mittermeier, C. G., da Fonseca, G. A. B. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853–858 (2000).
IPCC. Climate Change 2021: The Physical Science Basis. (eds Masson-Delmotte, V. et al.) (Cambridge Univ. Press, in press).
Mora, C. et al. The projected timing of climate departure from recent variability. Nature 502, 183–187 (2013).
Wood, T. E. et al. in Ecosystem Consequences of Soil Warming: Microbes, Vegetation, Fauna and Soil Biogeochemistry (ed. Mohan, J.) Ch. 14 (Academic Press, 2019).
Davidson, E. A. & Janssens, I. A. Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature 440, 165–173 (2006).
van Gestel, N. et al. Predicting soil carbon loss with warming. Nature 554, E4–E5 (2018).
Melillo, J. M. et al. Long-term pattern and magnitude of soil carbon feedback to the climate system in a warming world. Science 358, 101–104 (2017).
Romero-Olivares, A. L., Allison, S. D. & Treseder, K. K. Soil microbes and their response to experimental warming over time: a meta-analysis of field studies. Soil Biol. Biochem. 107, 32–40 (2017).
Anderson-Teixeira, K. J., Wang, M. M. H., McGarvey, J. C. & LeBauer, D. S. Carbon dynamics of mature and regrowth tropical forests derived from a pantropical database (TropForC-db). Glob. Change Biol. 22, 1690–1709 (2016).
Nottingham, A. T., Meir, P., Velasquez, E. & Turner, B. L. Soil carbon loss by experimental warming in a tropical forest. Nature 584, 234–237 (2020).
Kimball, B. A. et al. Infrared heater system for warming tropical forest understory plants and soils. Ecol. Evol. 8, 1932–1944 (2018).
DeAngelis, K. M. et al. Long-term forest soil warming alters microbial communities in temperate forest soils. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00104 (2015)
Bååth, E. Temperature sensitivity of soil microbial activity modeled by the square root equation as a unifying model to differentiate between direct temperature effects and microbial community adaptation. Glob. Change Biol. 24, 2850–2861 (2018).
Wieder, W. R., Bonan, G. B. & Allison, S. D. Global soil carbon projections are improved by modelling microbial processes. Nat. Clim. Change 3, 909–912 (2013).
Ratkowsky, D. A., Olley, J., Mcmeekin, T. A. & Ball, A. Relationship between temperature and growth-rate of bacterial cultures. J. Bacteriol. 149, 1–5 (1982).
Rinnan, R., Rousk, J., Yergeau, E., Kowalchuk, G. A. & Bååth, E. Temperature adaptation of soil bacterial communities along an Antarctic climate gradient: predicting responses to climate warming. Glob. Change Biol. 15, 2615–2625 (2009).
Nottingham, A. T., Bååth, E., Reischke, S., Salinas, N. & Meir, P. Adaptation of soil microbial growth to temperature: using a tropical elevation gradient to predict future changes. Glob. Change Biol. https://doi.org/10.1111/gcb.14502 (2019).
Li, J. Q., Bååth, E., Pei, J. M., Fang, C. M. & Nie, M. Temperature adaptation of soil microbial respiration in alpine, boreal and tropical soils: an application of the square root (Ratkowsky) model. Glob. Change Biol. 27, 1281–1292 (2021).
Rousk, J., Frey, S. D. & Bååth, E. Temperature adaptation of bacterial communities in experimentally warmed forest soils. Glob. Change Biol. 18, 3252–3258 (2012).
Nottingham, A. T. et al. Annual to decadal temperature adaptation of the soil bacterial community after translocation across an elevation gradient in the Andes. Soil Biol. Biochem. 158, 108217 (2021).
Nottingham, A. T. et al. Microbial responses to warming enhance soil carbon loss following translocation across a tropical forest elevation gradient. Ecol. Lett. 22, 1889–1899 (2019).
Donhauser, J., Niklaus, P. A., Rousk, J., Larose, C. & Frey, B. Temperatures beyond the community optimum promote the dominance of heat-adapted, fast growing and stress resistant bacteria in alpine soils. Soil Biol. Biochem. 148, 107873 (2020).
Mangan, S. A. et al. Negative plant–soil feedback predicts tree-species relative abundance in a tropical forest. Nature 466, 752–755 (2010).
Pold, G., Melillo, J. M. & DeAngelis, K. M. Two decades of warming increases diversity of a potentially lignolytic bacterial community. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00480 (2015).
Zhou, J. Z. et al. Temperature mediates continental-scale diversity of microbes in forest soils. Nat. Commun. 7, 12083 (2016).
Tedersoo, L. et al. Global diversity and geography of soil fungi. Science 346, 1256688 (2014).
Wu, L. et al. Reduction of microbial diversity in grassland soil is driven by long-term climate warming. Nat. Microbiol. 7, 1054–1062 (2022).
Oliverio, A. M., Bradford, M. A. & Fierer, N. Identifying the microbial taxa that consistently respond to soil warming across time and space. Glob. Change Biol. 23, 2117–2129 (2017).
Bahram, M. et al. Structure and function of the global topsoil microbiome. Nature 560, 233–237 (2018).
Spracklen, D. V., Baker, J. C. A., Garcia-Carreras, L. & Marsham, J. H. The effects of tropical vegetation on rainfall. Annu. Rev. Env. Resour. 43, 193–218 (2018).
Bradford, M. A. Thermal adaptation of decomposer communities in warming soils. Front. Microbiol. https://doi.org/10.3389/Fmicb.2013.00333 (2013).
Pietikäinen, J., Pettersson, M. & Bååth, E. Comparison of temperature effects on soil respiration and bacterial and fungal growth rates. FEMS Microbiol. Ecol. 52, 49–58 (2005).
Mori, A. S. et al. Biodiversity–productivity relationships are key to nature-based climate solutions. Nat. Clim. Change 11, 543–550 (2021).
Delgado-Baquerizo, M. et al. Multiple elements of soil biodiversity drive ecosystem functions across biomes. Nat. Ecol. Evol. 4, 210–220 (2020).
Wagg, C., Bender, S. F., Widmer, F. & van der Heijden, M. G. A. Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc. Natl Acad. Sci. USA 111, 5266–5270 (2014).
Nottingham, A. T. et al. Microbes follow Humboldt: temperature drives plant and soil microbial diversity patterns from the Amazon to the Andes. Ecology 99, 2455–2466 (2018).
Brown, J. H., Gillooly, J. F., Allen, A. P., Savage, V. M. & West, G. B. Toward a metabolic theory of ecology. Ecology 85, 1771–1789 (2004).
Brown, J. H. Why are there so many species in the tropics? J. Biogeogr. 41, 8–22 (2014).
LaManna, J. A. et al. Plant diversity increases with the strength of negative density dependence at the global scale. Science 356, 1389–1392 (2017).
Bagchi, R. et al. Pathogens and insect herbivores drive rainforest plant diversity and composition. Nature 506, 85–88 (2014).
Lapebie, P., Lombard, V., Drula, E., Terrapon, N. & Henrissat, B. Bacteroidetes use thousands of enzyme combinations to break down glycans. Nat. Commun. https://doi.org/10.1038/s41467-019-10068-5 (2019).
Makhalanyane, T. P. et al. Microbial ecology of hot desert edaphic systems. FEMS Microbiol. Rev. 39, 203–221 (2015).
Aydogan, E. L., Moser, G., Muller, C., Kampfer, P. & Glaeser, S. P. Long-term warming shifts the composition of bacterial communities in the phyllosphere of Galium album in a permanent grassland field-experiment. Front. Microbiol. https://doi.org/10.3389/fmicb.2018.00144 (2018).
Hu, D. Y., Zang, Y., Mao, Y. J. & Gao, B. L. Identification of molecular markers that are specific to the class thermoleophilia. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.01185 (2019).
Mohan, J. E. et al. Mycorrhizal fungi mediation of terrestrial ecosystem responses to global change: mini-review. Fungal Ecol. 10, 3–19 (2014).
Manzoni, S., Taylor, P., Richter, A., Porporato, A. & Agren, G. I. Environmental and stoichiometric controls on microbial carbon-use efficiency in soils. New Phytol. 196, 79–91 (2012).
Allison, S. D., Wallenstein, M. D. & Bradford, M. A. Soil-carbon response to warming dependent on microbial physiology. Nat. Geosci. 3, 336–340 (2010).
Reed, S. C. et al. Soil biogeochemical responses of a tropical forest to warming and hurricane disturbance. Adv. Ecol. Res. 62, 225–252 (2020).
Nottingham, A. T., Turner, B. L., Stott, A. W. & Tanner, E. V. J. Nitrogen and phosphorus constrain labile and stable carbon turnover in lowland tropical forest soils. Soil Biol. Biochem. 80, 26–33 (2015).
Walker, T. W. N. et al. Microbial temperature sensitivity and biomass change explain soil carbon loss with warming. Nat. Clim. Change 8, 885–889 (2018).
Kemmitt, S. J. et al. Mineralization of native soil organic matter is not regulated by the size, activity or composition of the soil microbial biomass—a new perspective. Soil Biol. Biochem. 40, 61–73 (2008).
Nannipieri, P., Trasar-Cepeda, C. & Dick, R. P. Soil enzyme activity: a brief history and biochemistry as a basis for appropriate interpretations and meta-analysis. Biol. Fert. Soils 54, 11–19 (2018).
Wallenstein, M., Allison, S., Ernakovich, J., Steinweg, J. M. & Sinsabaugh, R. in Soil Enzymology. Soil Biology Vol. 22 (eds Shukla, G. & Varma, A.) Ch. 13 (Springer, 2011).
Zhou, X. Y., Chen, L., Xu, J. M. & Brookes, P. C. Soil biochemical properties and bacteria community in a repeatedly fumigated-incubated soil. Biol. Fert. Soils 56, 619–631 (2020).
Sanchez-Julia, M. & Turner, B. L. Abiotic contribution to phenol oxidase activity across a manganese gradient in tropical forest soils. Biogeochemistry https://doi.org/10.1007/s10533-021-00764-0 (2021).
Razavi, B. S., Liu, S. B. & Kuzyakov, Y. Hot experience for cold-adapted microorganisms: temperature sensitivity of soil enzymes. Soil Biol. Biochem. 105, 236–243 (2017).
Pinney, M. M. et al. Parallel molecular mechanisms for enzyme temperature adaptation. Science 371, eaay2784 (2021).
Fanin, N. et al. Soil enzymes in response to climate warming: mechanisms and feedbacks. Funct. Ecol. https://doi.org/10.1111/1365-2435.14027 (2022).
Hall, S. J. & Silver, W. L. Iron oxidation stimulates organic matter decomposition in humid tropical forest soils. Glob. Change Biol. 19, 2804–2813 (2013).
Freeman, C., Ostle, N. & Kang, H. An enzymic ‘latch’ on a global carbon store. Nature 409, 149 (2001).
Sarmiento, C. et al. Soilborne fungi have host affinity and host-specific effects on seed germination and survival in a lowland tropical forest. Proc. Natl Acad. Sci. USA 114, 11458–11463 (2017).
Condit, R., Perez, R., Lao, S., Aguilar, S. & Hubbell, S. P. Demographic trends and climate over 35 years in the Barro Colorado 50 ha plot. For. Ecosyst. https://doi.org/10.1186/s40663-017-0103-1 (2017).
Woodring, W. P. Geology of Barro Colorado Island. Smithson. Misc. Collect. 135, 1–39 (1958).
Sanchez, P. A. & Logan, T. J. Myths and science about the chemistry and fertility of soils in the tropics. SSSA Spec. Publ. 29, 35–46 (1992).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Brookes, P. C., Landman, A., Pruden, G. & Jenkinson, D. S. Chloroform fumigation and the release of soil nitrogen: a rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol. Biochem. 17, 837–842 (1985).
Vance, E. D., Brookes, P. C. & Jenkinson, D. S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 19, 703–707 (1987).
Jenkinson, D. S., Brookes, P. C. & Powlson, D. S. Measuring soil microbial biomass. Soil Biol. Biochem. 36, 5–7 (2004).
Kouno, K., Tuchiya, Y. & Ando, T. Measurement of soil microbial biomass phosphorus by an anion-exchange membrane method. Soil Biol. Biochem. 27, 1353–1357 (1995).
Tabatabai, M. A. in Methods of Soil Analysis. Part 2. Microbiological and Biochemical Properties (ed. Page, A.L.) 778–833 (SSSA, 1994).
Marx, M. C., Wood, M. & Jarvis, S. C. A microplate fluorimetric assay for the study of enzyme diversity in soils. Soil Biol. Biochem. 33, 1633–1640 (2001).
Price, N. & Stevens, L. Fundamentals of Enzymology: Cell and Molecular Biology of Catalytic Proteins (Oxford Univ. Press, 1999).
Hagerty, S. B., Allison, S. D. & Schimel, J. P. Evaluating soil microbial carbon use efficiency explicitly as a function of cellular processes: implications for measurements and models. Biogeochemistry 140, 269–283 (2018).
Frey, S. D., Lee, J., Melillo, J. M. & Six, J. The temperature response of soil microbial efficiency and its feedback to climate. Nat. Clim. Change 3, 395–398 (2013).
Spohn, M. et al. Soil microbial carbon use efficiency and biomass turnover in a long-term fertilization experiment in a temperate grassland. Soil Biol. Biochem. 97, 168–175 (2016).
Sinsabaugh, R. L. et al. Stoichiometry of microbial carbon use efficiency in soils. Ecol. Monogr. 86, 172–189 (2016).
Geyer, K. M., Dijkstra, P., Sinsabaugh, R. & Frey, S. D. Clarifying the interpretation of carbon use efficiency in soil through methods comparison. Soil Biol. Biochem. 128, 79–88 (2019).
Bååth, E., Pettersson, M. & Söderberg, K. H. Adaptation of a rapid and economical microcentrifugation method to measure thymidine and leucine incorporation by soil bacteria. Soil Biol. Biochem. 33, 1571–1574 (2001).
Bárcenas-Moreno, G., Gomez-Brandon, M., Rousk, J. & Bååth, E. Adaptation of soil microbial communities to temperature: comparison of fungi and bacteria in a laboratory experiment. Glob. Change Biol. 15, 2950–2957 (2009).
Smirnova, E., Huzurbazar, S. & Jafari, F. PERFect: PERmutation Filtering test for microbiome data. Biostatistics 20, 615–631 (2019).
Alberdi, A. & Gilbert, M. T. P. hilldiv: an R package for the integral analysis of diversity based on Hill numbers. Preprint at bioRxiv https://doi.org/10.1101/545665 (2019).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2011).
Oksanen, J. et al. vegan: Community ecology package, R Package version 2 https://cran.r-project.org/web/packages/vegan/ (2018).
Dufrene, M. & Legendre, P. Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol. Monogr. 67, 345–366 (1997).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Roesch, L. F. W. et al. PIME: a package for discovery of novel differences among microbial communities. Mol. Ecol. Resour. 20, 415–428 (2020).
Roberts, D.W. labdsv: Ordination and multivariate analysis for ecology. R package version 2.0-1 https://cran.r-project.org/web/packages/labdsv/ (2019).
Cao, Y. et al. microbiomeMarker: an R/Bioconductor package for microbiome marker identification and visualization. Bioinformatics 38, 4027–4029 (2022).
Eren, A. M. et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. Peerj 3, e1319 (2015).
Peterson, R. A. & Cavanaugh, J. E. Ordered quantile normalization: a semiparametric transformation built for the cross-validation era. J. Appl. Stat. 47, 2312–2327 (2020).
Acknowledgements
This study was supported by three fellowships to A.T.N.: UK NERC grant NE/T012226, European Union Marie-Curie Fellowship FP7-2012-329360 and an STRI Tupper Fellowship. Further support came from UK NERC grant NE/K01627X/1 to P.M., an ANU Biology Innovation grant to P.M. and Simons Foundation grant No. 429440 to W. Wcislo, STRI, and support from the US Department of Agriculture (USDA), Agricultural Research Service to K.B. We thank O. Acevado, D. Agudo, A. Bielnicka, G. Broders, M. Cano, D. Dominguez, M. Garcia, M. Larsen, J. Rodriguez, H. Szczygiel, I. Torres, E. Velasquez, W. Wcislo, K. Winter and J. Wright for support. We thank B. Turner for his contribution to SWELTR, especially during its initial phase of operation. Sequencing analyses were conducted on the Smithsonian High-Performance Cluster (SI/HPC), Smithsonian Institution (https://doi.org/10.25572/SIHPC). For the purpose of open access, the authors have applied a Creative Commons Attribution (CC BY) licence to any author-accepted manuscript version arising from this submission. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the USDA. The USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Contributions
A.T.N. conceived the study. A.T.N., J.J.S., M.M.-S., J.P., E.B., K.B. and K.S. performed the study. A.T.N. and J.J.S. analysed the data. A.T.N. wrote the paper with input from J.J.S., E.B., K.S., K.B. and P.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Nadia Maaroufi, Ashish Malik and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 One of five warmed plots at SWELTR.
The images show the soil surface temperature shortly after the warming structure was switched on (a and c) and after a period of thermal equilibration‘ (b and d). The circular heating structure was 3.5 m in diameter and extended to 1.2 m depth, which resulted in an effective heated plot of approximately 5 m diameter x > 1.5 m depth (that is to the bedrock, situated at around 1.5–2.0 m across the study site). The experiment consisted of five warmed and control plot-pairs in total. For this study we had three treatment levels, +3 °C warming (within the warmed plots), +8 °C (within a high-temperature buffer zone close [~10 cm] to the heating source for each warmed plot) and ambient temperature controls (within the control plots). Therefore, all analyses are for n = 5 independent sampling locations for each treatment level. Image credit: J. Bujan and E. Velasquez.
Extended Data Fig. 2 Diversity response of soil bacteria (a–c) and fungi (d–f) to two years of warming by +3 °C and +8 °C.
Shapiro-Wilk Normality and Bartlett tests indicated all alpha diversity estimates (following PERfect filtering) were normally distributed and differences were assessed for (a) bacteria and (d) fungi using analysis of variance (ANOVA) followed by Tukey HSD post-hoc tests. Compositional similarity of microbial communities (beta-diversity) represented as PCoA ordination plots of PERfect filtered data for (b) bacteria—estimated using Unweighted (left) and Weighted Unifrac (right) distance matrices; and (e) fungi estimated—using Jensen–Shannon divergence (left) and Bray-Curtis (right) distance matrices. Within group distances for the (c) bacteria and (f) fungi datasets. The centre line of each box plot represents the median, the lower and upper hinges represent the first and third quartiles and whiskers represent + 1.5 the interquartile range. For panels (a), (c), (d), and (f), only significant differences between treatments are shown.
Extended Data Fig. 3 The response of select soil bacteria taxa to two years of warming by +3 °C and +8 °C.
Differences assessed for multiple-group pair-wise comparisons using ANOVA followed by Tukey HSD post hoc tests. PERfect filtered read count data was log10 transformed and normalized using total sum scaling (TSS). The centre line of each box plot represents the median, the lower and upper hinges represent the first and third quartiles and whiskers represent + 1.5 the interquartile range. Only significant differences between treatments are shown.
Extended Data Fig. 4 The response of select soil fungal taxa to two years of warming by +3 °C and +8 °C.
Differences assessed for multiple-group pair-wise comparisons using ANOVA followed by Tukey HSD post hoc tests. PERfect filtered read count data was log10 transformed and normalized using total sum scaling (TSS). The centre line of each box plot represents the median, the lower and upper hinges represent the first and third quartiles and whiskers represent + 1.5 the interquartile range. Only significant differences between treatments are shown.
Extended Data Fig. 5 Distance-based Redundancy Analysis (db-RDA).
Distance-based Redundancy Analysis (db-RDA) of PIME filtered data based on Bray-Curtis dissimilarity showing the relationships between community composition change for (a) bacteria and (b) fungi versus edaphic properties (left) and microbial functional response (right). All analyses are for soil collected from n = 5 independent sampling locations for each treatment level.
Extended Data Fig. 6 Soil, enzyme, and microbial responses to +3 °C and +8 °C in situ soil warming.
Data are grouped by (a) soil properties, (b) microbial functional responses, and (c) microbial temperature adaptive responses; we used the same grouping to test three hypotheses on how each of these responses were correlated to changes in microbial diversity and community composition (Fig. 2; Extended Data Table 2, Fig. 5). All properties were determined for soil samples collected during the 2018 wet season (June and November); see methods. Units for enzyme Vmax are nmol MU g−1 min−1, except Phenol oxidase in μmol g−1 h−1 and Leucine aminopeptidase in nmol AMC g−1 min−1. The centre line of each box plot represents the median, the lower and upper hinges represent the first and third quartiles and whiskers represent + 1.5 the interquartile range. Significant differences between treatments and controls are highlighted by asterisks (ANOVA; * p ≤ 0.05, ** p ≤ 0.01, *** p ≤ 0.001). All analyses are for soil collected from n = 5 independent sampling locations for each treatment level.
Extended Data Fig. 7 Soil enzyme activities in response to incubation temperature (that is instantaneous temperature response determined in laboratory assays).
Data are maximum potential enzyme activity (Vmax), determined by activity under saturating substrate conditions. Enzymes are: α-glucosidase (AGase), β-glucosidase (BGase), phospho-diesterase (BPase), cellolbiohydrolase (CEase), leucine aminopeptidase (LPase), phosphomonoesterase (Pase), N-acetyl β-glucosaminidase (Nase), phenol oxidase (PXase), sulfatase (Sase) and β-xylanase (XYase). Units for enzyme Vmax are nmol MU g−1 min−1, except Phenol oxidase in μmol g−1 h−1 and Leucine aminopeptidase in nmol AMC g−1 min−1. The error bars represent mean ± one standard error, for n = 5 plots. Fitted lines depict quadratic functions with 95% confidence intervals. All analyses are for soil collected from n = 5 independent sampling locations for each treatment level. Activity was determined during the wet season 2018 for the following sampling periods: controls include 4 sampling periods (June, Sept, Oct, Dec 2018); +3 °C include 3 sampling periods (June, Sept, Dec 2018); +8 °C include 1 sampling period (Sept 2018).
Supplementary information
Supplementary Information
Supplementary Methods, Results, Figs. 1–14 and Tables 1–15.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nottingham, A.T., Scott, J.J., Saltonstall, K. et al. Microbial diversity declines in warmed tropical soil and respiration rise exceed predictions as communities adapt. Nat Microbiol 7, 1650–1660 (2022). https://doi.org/10.1038/s41564-022-01200-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01200-1
This article is cited by
-
Gut microbiota modulation enhances the immune capacity of lizards under climate warming
Microbiome (2024)
-
Elevated methane flux in a tropical peatland post-fire is linked to depth-dependent changes in peat microbiome assembly
npj Biofilms and Microbiomes (2024)
-
Quantifying thermal adaptation of soil microbial respiration
Nature Communications (2023)
-
Viral lysing can alleviate microbial nutrient limitations and accumulate recalcitrant dissolved organic matter components in soil
The ISME Journal (2023)
-
The Biological Properties of Rice Paddy Fields in Different Depths Affected by Pretilachlor Herbicide
Journal of Soil Science and Plant Nutrition (2023)