Abstract
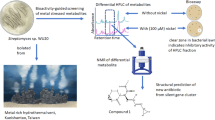
Industry screens of large chemical libraries have traditionally relied on rich media to ensure rapid bacterial growth in high-throughput testing. We used eukaryotic, nutrient-limited growth media in a compound screen that unmasked a previously unknown hyperactivity of the old antibiotic, rifabutin (RBT), against highly resistant Acinetobacter baumannii. In nutrient-limited, but not rich, media, RBT was 200-fold more potent than rifampin. RBT was also substantially more effective in vivo. The mechanism of enhanced efficacy was a Trojan horse-like import of RBT, but not rifampin, through fhuE, only in nutrient-limited conditions. These results are of fundamental importance to efforts to discover antibacterial agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Screening data are available on ReFRAMEdb.org. Genome sequencing data are available at NCBI accession no. PRJNA629056. Source data for the figures are provided with the paper.
Change history
17 June 2020
In the version of this Article originally published, the authors’ first names were formatted as initials. This has now been corrected and the authors’ full names are displayed.
References
European Committee for Antimicrobial Susceptibility Testing (EUCAST) of the European Society of Clinical Microbiology and Infectious Diseases (ESCMID). Determination of minimum inhibitory concentrations (MICs) of antibacterial agents by broth dilution. Clin. Microbiol. Infect. 9, ix–xv (2003).
Matuschek, E., Brown, D. F. J. & Kahlmeter, G. Development of the EUCAST disk diffusion antimicrobial susceptibility testing method and its implementation in routine microbiology laboratories. Clin. Microbiol. Infect. 20, O255–O266 (2014).
Brown, D. et al. Widespread implementation of EUCAST breakpoints for antibacterial susceptibility testing in Europe. Euro Surveill. 20, 21008 (2015).
Hombach, M., Courvalin, P. & Böttger, E. C. Validation of antibiotic susceptibility testing guidelines in a routine clinical microbiology laboratory exemplifies general key challenges in setting clinical breakpoints. Antimicrob. Agents Chemother. 58, 3921–3926 (2014).
Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically, Approved Standard, 9th edn. CLSI document M07-A9 (Clinical and Laboratory Standards Institute, 2018).
Fahnoe, K. C. et al. Non-traditional antibacterial screening approaches for the identification of novel inhibitors of the glyoxylate shunt in Gram-negative pathogens. PLoS ONE 7, e51732 (2012).
World Health Organization. Global Priority List of Antibiotic-resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics (WHO, 2017); www.who.int/medicines/publications/WHO-PPL-Short_Summary_25Feb-ET_NM_WHO.pdf?ua=1
Shlaes, D. M., Sahm, D., Opiela, C. & Spellberg, B. The FDA reboot of antibiotic development. Antimicrob. Agents Chemother. 57, 4605–4607 (2013).
Schäberle, T. F. & Hack, I. M. Overcoming the current deadlock in antibiotic research. Trends Microbiol. 22, 165–167 (2014).
Centers for Disease Control (CDC). Antibiotic Resistance Threats in the United States (US Department of Health and Human Services, CDC, 2013); www.cdc.gov/drugresistance/threat-report-2013.
Butler, M. S., Blaskovich, M. A. & Cooper, M. A. Antibiotics in the clinical pipeline at the end of 2015. J. Antibiot. 70, 3–24 (2017).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Spellberg, B. et al. The epidemic of antibiotic-resistant infections: a call to action for the medical community from the Infectious Diseases Society of America. Clin. Infect. Dis. 46, 155–164 (2008).
Janes, J. et al. The ReFRAME library as a comprehensive drug repurposing library and its application to the treatment of cryptosporidiosis. Proc. Natl Acad. Sci. USA 115, 10750–10755 (2018).
Nielsen, T. B. et al. Monoclonal antibody protects against Acinetobacter baumannii infection by enhancing bacterial clearance and evading sepsis. J. Infect. Dis. 216, 489–501 (2017).
Nielsen, T. B., Bruhn, K. W., Pantapalangkoor, P., Junus, J. L. & Spellberg, B. Cryopreservation of virulent Acinetobacter baumannii to reduce variability of in vivo studies. BMC Microbiol. 15, 252 (2015).
Bruhn, K. W. et al. Host fate is rapidly determined by innate effector-microbial interactions during Acinetobacter baumannii bacteremia. J. Infect. Dis. 211, 1296–1305 (2015).
Luo, G. et al. Active and passive immunization protects against lethal, extreme drug resistant-Acinetobacter baumannii infection. PLoS ONE 7, e29446 (2012).
Lin, L. et al. Inhibition of LpxC protects mice from resistant Acinetobacter baumannii by modulating inflammation and enhancing phagocytosis. mBio 3, e00312 (2012).
Jacobs, A. C. et al. Characterization of the Acinetobacter baumannii growth phase-dependent and serum responsive transcriptomes. FEMS Immunol. Med. Microbiol. 64, 403–412 (2012).
Blanchard, C., Barnett, P., Perlmutter, J. & Dunman, P. M. Identification of Acinetobacter baumannii serum-associated antibiotic efflux pump inhibitors. Antimicrob. Agents Chemother. 58, 6360–6370 (2014).
Asensio, C. & Pérez-Díaz, J. C. A new family of low molecular weight antibiotics from enterobacteria. Biochem. Biophys. Res. Commun. 69, 7–14 (1976).
Thulin, E., Thulin, M. & Andersson, D. I. Reversion of high-level mecillinam resistance to susceptibility in Escherichia coli during growth in urine. EBioMedicine 23, 111–118 (2017).
Parquet, M. D. C., Savage, K. A., Allan, D. S., Davidson, R. J. & Holbein, B. E. Novel iron-chelator DIBI inhibits Staphylococcus aureus growth, suppresses experimental MRSA infection in mice and enhances the activities of diverse antibiotics in vitro. Front. Microbiol. 9, 1811 (2018).
Parquet, M. D. C. et al. Antibiotic-resistant Acinetobacter baumannii is susceptible to the novel iron-sequestering anti-infective DIBI in vitro and in experimental pneumonia in mice. Antimicrob. Agents Chemother. 63, e00855-19 (2019).
Gallagher, L. A. et al. Resources for genetic and genomic analysis of emerging pathogen Acinetobacter baumannii. J. Bacteriol. 197, 2027–2035 (2015).
Funahashi, T. et al. Identification and characterization of an outer membrane receptor gene in Acinetobacter baumannii required for utilization of desferricoprogen, rhodotorulic acid, and desferrioxamine B as xenosiderophores. Biol. Pharm. Bull. 35, 753–760 (2012).
Durante-Mangoni, E. et al. Colistin and rifampicin compared with colistin alone for the treatment of serious infections due to extensively drug-resistant Acinetobacter baumannii: a multicenter, randomized clinical trial. Clin. Infect. Dis. 57, 349–358 (2013).
Payne, D. J., Gwynn, M. N., Holmes, D. J. & Pompliano, D. L. Drugs for bad bugs: confronting the challenges of antibacterial discovery. Nat. Rev. Drug Discov. 6, 29–40 (2007).
El Zahed, S. S., Kumar, G., Tong, M. & Brown, E. D. in Phenotypic Screening. Methods in Molecular Biology Vol. 1787 (ed. Wagner, B) 1–18 (Humana Press, 2018).
Zlitni, S., Ferruccio, L. F. & Brown, E. D. Metabolic suppression identifies new antibacterial inhibitors under nutrient limitation. Nat. Chem. Biol. 9, 796–804 (2013).
Hermanson, G. T. in Bioconjugate Techniques, 3rd edn (ed. Hermanson, G. T.) 127–228 (Academic Press, 2013).
Eoh, H. & Rhee, K. Y. Multifunctional essentiality of succinate metabolism in adaptation to hypoxia in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 110, 6554–6559 (2013).
Lee, J. J. et al. Transient drug-tolerance and permanent drug-resistance rely on the trehalose-catalytic shift in Mycobacterium tuberculosis. Nat. Commun. 10, 2928 (2019).
Lee, J. J. et al. Glutamate mediated metabolic neutralization mitigates propionate toxicity in intracellular Mycobacterium tuberculosis. Sci. Rep. 8, 8506 (2018).
Trebosc, V. et al. A novel genome-editing platform for drug-resistant Acinetobacter baumannii reveals an AdeR-unrelated tigecycline resistance mechanism. Antimicrob. Agents Chemother. 60, 7263–7271 (2016).
Luna, B. M. et al. Natural history of Acinetobacter baumannii infection in mice. PLoS ONE 14, e0219824 (2019).
Nielsen, T. B., Yan, J., Luna, B. & Spellberg, B. Murine oropharyngeal aspiration model of ventilator-associated and hospital-acquired bacterial pneumonia. J. Vis. Exp. 136, e57672 (2018).
Acknowledgements
We thank Calibr’s High Throughput Screening and Compound Management teams for assisting with the ReFRAME screen. We also thank B. Weiss at the University of North Texas for supporting the neutropenic lung infection model. This work was supported by the National Institute of Allergy and Infectious Diseases (grant nos. R01AI139052 to B.L., R.S. and B.S., and R01AI130060 and R01AI117211 to B.S.), and the FDA (BAA contract HHSF223201710199C to B.L. and B.S.). Calibr at Scripps Research Institute was supported by the Bill & Melinda Gates Foundation (OPP1107194).
Author information
Authors and Affiliations
Contributions
B.L., M.B., V.T., C.M. and B.S. designed the experiments and wrote the manuscript. M.B. and C.M. helped design the HTS assay and performed the compound screening and analysis. B.L., A.U., J.Y., T.N., P.L., J.L., J.C., W.K., H.E., N.S., R.S., C.K., S.L. and G.D. participated in performing experiments, contributed intellectually and interpreted results. B.L., A.U., P.L. and R.S. conducted the MIC testing. J.L., W.K. and H.E. conducted the LC–MS/MS experiments. J.Y., T.B.N., P.L. and B.L. conducted the in vivo experiments.
Corresponding authors
Ethics declarations
Competing interests
B.L., B.S. and T.N. own equity in ExBaq. The University of Southern California has a financial interest in ExBaq. G.E.D., V.T., C.K. and S.L. own equity in BioVersys.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
MIC of rifabutin in RPMI+10% FCS on non-A. baumannii Gram-negative ESKAPE species.
Extended Data Fig. 2
MIC of rifabutin in RPMI + 10% FCS supplemented with increasing amount of ammonium iron(III) citrate.
Extended Data Fig. 3 Rifabutin MICs against A. baumannii AB5075 transposon disruption mutants.
Mutants were deficient in amino acid transport genes. MIC assay was done in both MHII and RPMI media. AB5075-UW is the parent strain for the transposon mutants.
Extended Data Fig. 4 Rifabutin MICs against A. baumannii AB5075 transposon disruption mutants.
MIC assay was done by culturing the bacteria in either MHII or RPMI media.
Extended Data Fig. 5
MIC of rifabutin and rifampicin on HUMC1 and the serial passaged mutants, and WGS of mutants with increased rifabutin MIC.
Extended Data Fig. 6
MIC of rifabutin and rifampicin on the fhuE deleted mutants and their parental strains.
Extended Data Fig. 7 Clustering of fhuE by amino acid sequence.
MICs were done for 43 A. baumannii clinical isolates to determine if the isolates were hypersensitive to RBT in RPMI medium. 28 of the 33 isolates in the outer rings exhibit the hypersensitive phenotype (open circles) in RPMI (RBT MIC = 0.05 µg/mL). 9 of 10 isolates in the inner ring are not hypersensitive (filled-in shapes). Of the 41 carbapenem-resistant isolates tested, 26 of those are hypersensitive to RBT.
Extended Data Fig. 8
MIC of rifabutin on the plasmid mediated fhuE expressing ATCC-17978 strains.
Extended Data Fig. 9 RPMI MIC predicts in vivo response to treatment.
Galleria mellonella larvae (10 per group) were infected with A. baumannii. a, Larvae were infected with strain HUMC1 at 1.6 ×104 cfu/larvae and treated with rifabutin (plain lines) or rifampicin (dashed lines) at 0.1 mg/kg (blue lines), 1 mg/kg (red lines) and 10 mg/kg (green lines) and survival was measured over 72 hours. b, G. mellonella larvae were infected with A. baumannii LAC-4 and treated with RBT or c, RIF. Consistent with the RPMI MIC data, there was no difference in outcomes based on treatment.
Extended Data Fig. 10 Drug -drug interaction between rifabutin or rifampin and colistin in MHII The drug-drug interaction were evaluated by calculating the fractional inhibitory concentration index (FICI).
FICI = FICA + FICB = (CA/MICA) + (CB/MICB), in which CA and CB are drug concentration of drug A and drugB in combination and MICA and MICB are the MIC of drug A and drug B alone. Synergy was defined as FICI ≤ 0.5, no interaction was defined as FICI > 0.5–4.0 and antagonism was defined as FICI > 4.0.
Supplementary information
Supplementary Table 1
ReFRAME screen results.
Supplementary Table 2
WGS sequencing data summary.
Supplementary Table 3
List of bacteria strains used.
Source data
Source Data Fig. 1
Numerical data.
Source Data Fig. 2
Numerical data.
Source Data Fig. 3
Numerical data..
Source Data Fig. 4
Numerical data
Source Data Extended Data Fig. 7
Numerical data.
Source Data Extended Data Fig. 9
Numerical data.
Rights and permissions
About this article
Cite this article
Luna, B., Trebosc, V., Lee, B. et al. A nutrient-limited screen unmasks rifabutin hyperactivity for extensively drug-resistant Acinetobacter baumannii. Nat Microbiol 5, 1134–1143 (2020). https://doi.org/10.1038/s41564-020-0737-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0737-6
This article is cited by
-
Evybactin is a DNA gyrase inhibitor that selectively kills Mycobacterium tuberculosis
Nature Chemical Biology (2022)
-
In search for a synergistic combination against pandrug-resistant A. baumannii; methodological considerations
Infection (2022)