Abstract
Bacteria have evolved diverse mechanisms to fend off predation by bacteriophages. We previously identified the Dnd system, which uses DndABCDE to insert sulfur into the DNA backbone as a double-stranded phosphorothioate (PT) modification, and DndFGH, a restriction component. Here, we describe an unusual SspABCD–SspE PT system in Vibrio cyclitrophicus, Escherichia coli and Streptomyces yokosukanensis, which has distinct genetic organization, biochemical functions and phenotypic behaviour. SspABCD confers single-stranded and high-frequency PTs with SspB acting as a nickase and possibly introducing nicks to facilitate sulfur incorporation. Strikingly, SspABCD coupled with SspE provides protection against phages in unusual ways: (1) SspE senses sequence-specific PTs by virtue of its PT-stimulated NTPase activity to exert its anti-phage activity, and (2) SspE inhibits phage propagation by introducing nicking damage to impair phage DNA replication. These results not only expand our knowledge about the diversity and functions of DNA PT modification but also enhance our understanding of the known arsenal of defence systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available from the corresponding author on request. The atomic coordinates and structure factors of SspB and the magnesium ion-bound SspB from S. clavuligerus ATCC 27064 have been deposited in the Protein Data Bank with the accession numbers 6JUF and 6LB9, respectively, and those of SspE from S. yokosukanensis DSM 40224 have the accession number 6JIV. The genome sequence of phage JXY1 has been deposited in GenBank under accession number MN994275. Source data for Figs. 1b, 2a,b,e,f, 3a–d and 4a–d and Extended Data Figs. 2c, 3, 4a, 6a,b and 7a–e are included in this article and its Supplementary information.
Code availability
Custom codes or software used in this study are available from the authors on request.
References
Yamaguchi, Y., Park, J. H. & Inouye, M. Toxin-antitoxin systems in bacteria and archaea. Annu. Rev. Genet. 45, 61–79 (2011).
Tock, M. R. & Dryden, D. T. The biology of restriction and anti-restriction. Curr. Opin. Microbiol. 8, 466–472 (2005).
Dy, R. L., Przybilski, R., Semeijn, K., Salmond, G. P. & Fineran, P. C. A widespread bacteriophage abortive infection system functions through a Type IV toxin-antitoxin mechanism. Nucleic Acids Res. 42, 4590–4605 (2014).
Marraffini, L. A. CRISPR–Cas immunity in prokaryotes. Nature 526, 55–61 (2015).
Swarts, D. C. et al. DNA-guided DNA interference by a prokaryotic Argonaute. Nature 507, 258–261 (2014).
Goldfarb, T. et al. BREX is a novel phage resistance system widespread in microbial genomes. EMBO J. 34, 169–183 (2015).
Ofir, G. et al. DISARM is a widespread bacterial defence system with broad anti-phage activities. Nat. Microbiol. 3, 90–98 (2018).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Willbanks, A. et al. The evolution of epigenetics: from prokaryotes to humans and its biological consequences. Genet. Epigenet. 8, 25–36 (2016).
Blow, M. J. et al. The epigenomic landscape of prokaryotes. PLoS Genet. 12, e1005854 (2016).
Wang, L. et al. Phosphorothioation of DNA in bacteria by dnd genes. Nat. Chem. Biol. 3, 709–710 (2007).
Wang, L., Jiang, S., Deng, Z., Dedon, P. C. & Chen, S. DNA phosphorothioate modification—a new multi-functional epigenetic system in bacteria. FEMS Microbiol. Rev. 43, 109–122 (2018).
Thiaville, J. J. et al. Novel genomic island modifies DNA with 7-deazaguanine derivatives. Proc. Natl Acad. Sci. USA 113, E1452–E1459 (2016).
Zhou, X. et al. A novel DNA modification by sulphur. Mol. Microbiol. 57, 1428–1438 (2005).
Eckstein, F. & Gish, G. Phosphorothioates in molecular biology. Trends Biochem. Sci. 14, 97–100 (1989).
Gan, R. et al. DNA phosphorothioate modifications influence the global transcriptional response and protect DNA from double-stranded breaks. Sci. Rep. 4, 6642 (2014).
Cao, B. et al. Pathological phenotypes and in vivo DNA cleavage by unrestrained activity of a phosphorothioate-based restriction system in Salmonella. Mol. Microbiol. 93, 776–785 (2014).
Xu, T., Yao, F., Zhou, X., Deng, Z. & You, D. A novel host-specific restriction system associated with DNA backbone S-modification in Salmonella. Nucleic Acids Res. 38, 7133–7141 (2010).
Chen, C. et al. Convergence of DNA methylation and phosphorothioation epigenetics in bacterial genomes. Proc. Natl Acad. Sci. USA 114, 4501–4506 (2017).
Cao, B. et al. Genomic mapping of phosphorothioates reveals partial modification of short consensus sequences. Nat. Commun. 5, 3951 (2014).
Kellner, S. et al. Oxidation of phosphorothioate DNA modifications leads to lethal genomic instability. Nat. Chem. Biol. 13, 888–894 (2017).
Wang, L. et al. DNA phosphorothioation is widespread and quantized in bacterial genomes. Proc. Natl Acad. Sci. USA 108, 2963–2968 (2011).
Tong, T. et al. Occurrence, evolution, and functions of DNA phosphorothioate epigenetics in bacteria. Proc. Natl Acad. Sci. USA 115, E2988–E2996 (2018).
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Anantharaman, V., Makarova, K. S., Burroughs, A. M., Koonin, E. V. & Aravind, L. Comprehensive analysis of the HEPN superfamily: identification of novel roles in intra-genomic conflicts, defense, pathogenesis and RNA processing. Biol. Direct 8, 15 (2013).
You, D., Wang, L., Yao, F., Zhou, X. & Deng, Z. A novel DNA modification by sulfur: DndA is a NifS-like cysteine desulfurase capable of assembling DndC as an iron-sulfur cluster protein in Streptomyces lividans. Biochemistry 46, 6126–6133 (2007).
Yao, F., Xu, T., Zhou, X., Deng, Z. & You, D. Functional analysis of spfD gene involved in DNA phosphorothioation in Pseudomonas fluorescens Pf0-1. FEBS Lett. 583, 729–733 (2009).
Xia, S. et al. Tight control of genomic phosphorothioate modification by the ATP-modulated autoregulation and reusability of DndB. Mol. Microbiol. 111, 938–950 (2018).
Maindola, P. et al. Multiple enzymatic activities of ParB/Srx superfamily mediate sexual conflict among conjugative plasmids. Nat. Commun. 5, 5322 (2014).
Schumacher, M. A. & Funnell, B. E. Structures of ParB bound to DNA reveal mechanism of partition complex formation. Nature 438, 516–519 (2005).
Basu, M. K. & Koonin, E. V. Evolution of eukaryotic cysteine sulfinic acid reductase, sulfiredoxin (Srx), from bacterial chromosome partitioning protein ParB. Cell Cycle 4, 947–952 (2005).
He, X. et al. Expression and purification of a single-chain Type IV restriction enzyme Eco94GmrSD and determination of its substrate preference. Sci. Rep. 5, 9747 (2015).
Jablonska, J., Matelska, D., Steczkiewicz, K. & Ginalski, K. Systematic classification of the His-Me finger superfamily. Nucleic Acids Res. 45, 11479–11494 (2017).
Chen, F. et al. Crystal structure of the cysteine desulfurase DndA from Streptomyces lividans which is involved in DNA phosphorothioation. PLoS ONE 7, e36635 (2012).
Machnicka, M. A., Kaminska, K. H., Dunin-Horkawicz, S. & Bujnicki, J. M. Phylogenomics and sequence-structure-function relationships in the GmrSD family of Type IV restriction enzymes. BMC Bioinform. 16, 336 (2015).
Liu, Q. et al. Engineering an iterative polyketide pathway in Escherichia coli results in single-form alkene and alkane overproduction. Metab. Eng. 28, 82–90 (2015).
Colombo, C. V., Menin, L. & Clerici, M. Alkaline denaturing Southern blot analysis to monitor double-strand break processing. Methods Mol. Biol. 1672, 131–145 (2018).
Shi, H., Wang, X., Tan, D. X., Reiter, R. J. & Chan, Z. Comparative physiological and proteomic analyses reveal the actions of melatonin in the reduction of oxidative stress in Bermuda grass (Cynodon dactylon (L). Pers). J. Pineal Res. 59, 120–131 (2015).
Satoh, K., Maniwa, T., Oda, T. & Matsumoto, K. Proteomic profiling for the identification of serum diagnostic biomarkers for abdominal and thoracic aortic aneurysms. Proteome Sci. 11, 27 (2013).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Winn, M. D., Murshudov, G. N. & Papiz, M. Z. Macromolecular TLS refinement in REFMAC at moderate resolutions. Methods Enzymol. 374, 300–321 (2003).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McCoy, A. J., Storoni, L. C. & Read, R. J. Simple algorithm for a maximum-likelihood SAD function. Acta Crystallogr. D 60, 1220–1228 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Vriend, G. WHAT IF: a molecular modeling and drug design program. J. Mol. Graph. 8, 52–56 (1990).
Hunt, M. et al. Circlator: automated circularization of genome assemblies using long sequencing reads. Genome Biol. 16, 294 (2015).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
The PyMOL Molecular Graphics System, Version 1.8 (Schrödinger, LLC, 2015).
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005).
Acknowledgements
We thank C. Chen for his assistance in the preparation of the manuscript. We thank M. Polz at the Massachusetts Institute of Technology for his gift of the Vibrio strains. We thank the staff at the BL17U1 beamline at the SSRF. The T4gt phage was provided by S. Xu (New England Biolabs). This work was supported by grants from the National Natural Science Foundation of China (grant nos. 31720103906, 31925002, 31520103902 and 31872627), China National Key Research and Development Program (grant no. 2019YFA0904300), Open Funding Project of State Key Laboratory of Microbial Metabolism, US National Science Foundation (grant no. CHE-1019990), US National Institute of Allergy and Infectious Disease (grant no. AI112711) and the Singapore–MIT Alliance for Research and Technology, sponsored by the National Research Foundation of Singapore.
Author information
Authors and Affiliations
Contributions
L.W. supervised the study. X.X. and Y.Wei performed most of the biochemical and genetic experiments and analyses. L.L., Yubing Zhang and H.G. performed the crystallization experiments and analyses. X.J., M.L. and R.H. performed the phage-related assays. Si Chen conducted the bioinformatics analysis. X.T., Yizhou Zhang, L.H. and S.J. performed the mutation experiments. Y.T. performed the proteomics experiment. G.W., R.S., Z.L., Y.Wang, Z.D., J.W., P.D., Shi Chen and L.W. analysed the study data. G.W. designed the crystallization experiments. X.X., G.W., Y.Wei, P.C.D., Shi Chen and L.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Structural characterization of SspB (locus_tag SSCG_01550).
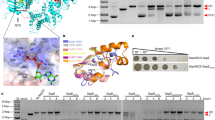
a, Sequence alignment of SspB with residue numbers and secondary structure elements indicated. Several putative dimerization-mediating residues in SspB, for example, H28, Y103, and E105, are replaced by similar amino acid residues in SspB homologues from other species; therefore, they are considered conserved. The imidazole ring of H28 resembles the benzene ring of F/Y in the SspB homologues of other species, which could also be involved in the hydrophobic interactions and thus serve a similar function. At the Y103 position of SspB homologues, there exists either a Y or an F residue, both of which possess benzene rings and can form hydrophobic interactions. The homologues of SspB have either E or D at E105. Both E and D have negatively charged carboxyl groups in their side chains and would play similar roles in the formation of salt bridges at the dimeric interface. b, Overall structure of SspB from S. clavuligerus ATCC 27064. α-helices, β-sheets, and loops are coloured red, yellow, and green, respectively. c, Conserved motifs 1, 2 and 3 are coloured magenta, green, and blue, respectively. d, The surface of the SspB structure is coloured according to the conservation score calculated by the ConSurf server. The “front” side of the SspB surface is highly conserved, while the “back” side is much less conserved. Motifs 1, 2, and 3 are mapped to the most conserved surface on the SspB structure.
Extended Data Fig. 2 Interaction of magnesium ions with SspB.
a, Ribbon diagram of magnesium ion-bound SspB from S. clavuligerus ATCC 27064. Magnesium ions are shown as small blue spheres. b, A magnesium ion is bound to E18 in each monomer. The SspB-Mg2+ complex was prepared by soaking the apo SspB crystals in reservoir solution containing 25% polyethylene glycol (PEG) 3350 and 1.0 M magnesium chloride for 6 h. c, The single-point mutation E18R remarkably impaired the DNA nicking activity of SspB. Data shown are representative images of two independent experiments.
Extended Data Fig. 3 Phylogenetic distribution of 234 homologues of the V. cyclitrophicus FF75 SspBCD–SspE in bacteria.
Phylogenetic groups are coloured by group (see the legend). Representative strains are labelled. For clarity, other strain names are not shown but are provided in Supplementary Data 2.
Extended Data Fig. 4 Sequence non-specific nicking activity of SspE.
a, Time-course treatment of pUC19 DNA by SspE. In this assay, 0.3 µg of pUC19 DNA was untreated or treated with 2 µM SspE at 28 °C in CutSmart (New England Biolabs, 50 mM potassium acetate, 20 mM Tris-acetate, pH 7.9, 10 mM magnesium acetate, 1.5 μM BSA). At the indicated time points, aliquots of the reaction mixture were withdrawn and analysed on a 1% agarose gel. L, linear pUC19 DNA. Nt.BspQI-nicked pUC19 was used as a control. Data shown are representative images of three independent experiments. b, Run-off sequencing of gel-purified SspE- or Nt.BspQI-nicked pUC19 DNA. The peaks in the sequencing chromatogram diminish sharply at the sequence-specific nicking site of Nt.BspQI. The extra adenine (A) base at the end of the run-off sequence was incorporated by the Taq DNA polymerase. In contrast, no such sequencing peak drops were observed when SspE-nicked pUC19 DNA was used as template despite the presence of the 5′-CCA-3′ motif.
Extended Data Fig. 5
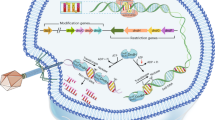
A schematic model of the Ssp system and defence against phage infection.
Extended Data Fig. 6 Interaction of SspB and SspC.
a, Pulldown of SspC with purified glutathione S-transferase (GST)-tagged SspB. BL21(DE3) cell lysate overexpressing 6xHis-SspC protein of S. clavuligerus ATCC 27064 was used as an input. A pulldown assay was performed by incubating the input with 56 pmol of purified GST-SspB of ATCC 27064 or GST, which was immobilized on glutathione (GSH) magnetic beads in PBS buffer at 4 °C overnight. Input and pulldown samples were examined via immunoblotting analysis. *, nonspecific bands. Data shown are representative images of three independent experiments. b, Nicking activity assays of SspB in the presence of SspC. Supercoiled pUC19 (0.3 µg) was incubated with 2 µM SspB, 2 µM SspB and 2 µM SspC or 2 µM SspBE204R and 2 µM SspC at 28 °C in CutSmart buffer. At the indicated time points, aliquots of the reaction mixture were withdrawn and analysed on a 1% agarose gel. Data shown are representative images of three independent experiments. c, Run-off sequencing results of gel-purified nicked pUC19 DNA. Nicked pUC19 DNA resulting from the treatment with SspB-SspC was gel-purified and subjected to run-off sequencing. The peaks in the sequencing chromatogram diminish sharply at the sequence-specific nicking site of Nt.BspQI. In contrast, no such sequencing peak drops were observed when SspB-SspC-nicked pUC19 DNA was used as template despite the presence of the 5′-CCA-3′ motif.
Extended Data Fig. 7 Effect of temperature on growth and ssp gene expression in Vibrio strains.
a, Growth of the wild-type and ssp mutants was assessed at 28 °C, 15 °C and 4 °C, respectively, on TSB agar plates supplemented with 2% NaCl. M565_ctg1P1910 = SspE. Results are representative of two independent experiments. b, The transcription of sspBCD and sspE at 15 °C and 28 °C was compared by quantitative real-time PCR with the housekeeping gene gapA, encoding glyceraldehyde 3-phosphate dehydrogenase (GAPDH), as a reference. Data are shown as the mean ± s.d. and based on three independent experiments (n = 3). Error bars are s.d. of the means. Two-sided Student t test was used for statistical analysis. **, p < 0.01. ns, not significant. In contrast to the enhanced transcription of sspE at 15 °C, sspBCD is constitutively expressed at all tested temperatures. c, E. coli Trans1-T1 (pWHU734) cells expressing His-tagged SspE under its native promoter were grown at 28 °C to an OD600 of 0.5 and were subsequently shifted to 15 °C for 1.5 h. The expression level of SspE was determined by Western blot. RpoB (RNA polymerase beta subunit) was used as an internal control. Results are representative of two independent experiments. d, Cell survival following a temperature downshift from 28 °C to 15 °C was assessed by determining the number of cells (colony-forming units, cfu) in the cultures at different time points. CFU numbers represent the mean values of three independent experiments (n = 3). Error bars are s.d. of the means. e, Cell survival after 6 h of antibiotic treatment (as a percentage on a log scale). Exponentially growing Vibrio cells were pretreated at 15 °C for 1.5 h, followed by exposure to 100 μg ml-1 ampicillin or 200 μg ml-1 streptomycin for 6 h. The concentrations of antibiotics used here are more than five times the minimum inhibitory concentration for V. cyclitrophicus FF75. Data are shown as mean ± s.d. and are representative of three independent experiments. Two-sided Student t test was used to measure significance. *, p < 0.05, ***, p < 0.001.
Supplementary information
Supplementary Information
Supplementary Results, Discussion, Tables 1–4, Figs. 1–12 and references.
Supplementary Data 1
Proteomics analysis of ΔsspC and wild-type FF75 at different temperatures.
Supplementary Data 2
SspBCDE homologues in 234 bacterial strains.
Supplementary Data 3
Unprocessed gels.
Supplementary Data 4
LC–MS/MS data.
Supplementary Data 5
Statistical source data.
Supplementary Data 6
LC–MS/MS data.
Supplementary Data 7
Unprocessed gels.
Supplementary Data 8
Unprocessed plate images.
Supplementary Data 9
Unprocessed plate images.
Supplementary Data 10
Unprocessed microscopy images.
Supplementary Data 11
Statistical source data.
Supplementary Data 12
Unprocessed gels.
Supplementary Data 13
Unprocessed gels.
Supplementary Data 14
Unprocessed gels.
Source data
Source Data Fig. 1
LC–MS/MS data.
Source Data Fig. 2
LC–MS/MS data.
Source Data Fig. 2
Unprocessed gels.
Source Data Fig. 3
Unprocessed plate images and Southern blots.
Source Data Fig. 4
Unprocessed plate images and gels.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed gels.
Source Data Extended Data Fig. 3
Strains for phylogenetic tree.
Source Data Extended Data Fig. 4
Unprocessed gels.
Source Data Extended Data Fig. 6
Unprocessed western blot and gel.
Source Data Extended Data Fig. 7
Unprocessed plate images and western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Xiong, X., Wu, G., Wei, Y. et al. SspABCD–SspE is a phosphorothioation-sensing bacterial defence system with broad anti-phage activities. Nat Microbiol 5, 917–928 (2020). https://doi.org/10.1038/s41564-020-0700-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-0700-6
This article is cited by
-
The highly diverse antiphage defence systems of bacteria
Nature Reviews Microbiology (2023)
-
Strain and process engineering toward continuous industrial fermentation
Frontiers of Chemical Science and Engineering (2023)
-
Nicking mechanism underlying the DNA phosphorothioate-sensing antiphage defense by SspE
Nature Communications (2022)
-
Systematic and quantitative view of the antiviral arsenal of prokaryotes
Nature Communications (2022)
-
The functional coupling between restriction and DNA phosphorothioate modification systems underlying the DndFGH restriction complex
Nature Catalysis (2022)