Abstract
Bacteria from the same species can differ widely in their gene content. In Escherichia coli, the set of genes shared by all strains, known as the core genome, represents about half the number of genes present in any strain. Although recent advances in bacterial genomics have unravelled genes required for fitness in various experimental conditions, most studies have focused on single model strains. As a result, the impact of the species’ genetic diversity on core processes of the bacterial cell remains largely under-investigated. Here, we have developed a CRISPR interference platform for high-throughput gene repression that is compatible with most E. coli isolates and closely related species. We have applied it to assess the importance of ~3,400 nearly ubiquitous genes in three growth conditions in 18 representative E. coli strains spanning most common phylogroups and lifestyles of the species. Our screens revealed extensive variations in gene essentiality between strains and conditions. Investigation of the genetic determinants for these variations highlighted the importance of epistatic interactions with mobile genetic elements. In particular, we have shown how prophage-encoded defence systems against phage infection can trigger the essentiality of persistent genes that are usually non-essential. This study provides broad insights into the evolvability of gene essentiality and argues for the importance of studying various isolates from the same species under diverse conditions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw sequencing reads from CRISPRi screens are available at the European Nucleotide Archive (ENA) under accession no. PRJEB37847. Raw read counts for each CRISPRi screen are provided in Supplementary Table 11. Raw reads from RNA-seq experiments were deposited on ArrayExpress with accession no. E-MTAB-9036. Processed data are available in the Supplementary Tables. Genome sequences of K-12 MG1655, H120 and APEC O1 were deposited on the ENA under accession nos. GCA_904425475, GCA_902876715 and GCA_902880315, respectively. The accession numbers of the other strains used in the study are available in Supplementary Table 1. The sequence of pFR56 was deposited in GenBank under accession MT412099. Interpro (www.ebi.ac.uk/interpro/), Pfam (http://pfam.xfam.org/) and REBASE (http://rebase.neb.com/rebase/rebase.html) databases are available online. Source data are provided with this paper.
Code availability
Custom scripts used in the manuscript are available at https://gitlab.pasteur.fr/dbikard/ecocg.
Change history
08 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41564-021-00893-0
15 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41564-021-00889-w
References
Rancati, G., Moffat, J., Typas, A. & Pavelka, N. Emerging and evolving concepts in gene essentiality. Nat. Rev. Genet. 19, 34–49 (2017).
Jordan, I. K., Rogozin, I. B., Wolf, Y. I. & Koonin, E. V. Essential genes are more evolutionarily conserved than are nonessential genes in bacteria. Genome Res. 12, 962–968 (2002).
Zhang, J. & Yang, J.-R. Determinants of the rate of protein sequence evolution. Nat. Rev. Genet. 16, 409–420 (2015).
Turner, K. H., Wessel, A. K., Palmer, G. C., Murray, J. L. & Whiteley, M. Essential genome of Pseudomonas aeruginosa in cystic fibrosis sputum. Proc. Natl Acad. Sci. USA 112, 4110–4115 (2015).
Le Breton, Y. et al. Essential genes in the core genome of the human pathogen Streptococcus pyogenes. Sci. Rep. 5, 9838 (2015).
Freed, N. E., Bumann, D. & Silander, O. K. Combining Shigella Tn-seq data with gold-standard E. coli gene deletion data suggests rare transitions between essential and non-essential gene functionality. BMC Microbiol. 16, 203 (2016).
Poulsen, B. E. et al. Defining the core essential genome of Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 116, 10072–10080 (2019).
Galardini, M. et al. The impact of the genetic background on gene deletion phenotypes in S accharomyces cerevisiae. Mol. Syst. Biol. 15, e8831 (2019).
Dowell, R. D. et al. Genotype to phenotype: a complex problem. Science 328, 469 (2010).
van Opijnen, T., Dedrick, S. & Bento, J. Strain dependent genetic networks for antibiotic-sensitivity in a bacterial pathogen with a large pan-genome. PLOS Pathog. 12, e1005869 (2016).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Nichols, R. J. et al. Phenotypic landscape of a bacterial cell. Cell 144, 143–156 (2011).
Goodall, E. C. A. et al. The essential genome of Escherichia coli K-12. mBio 9, e02096 (2018).
Price, M. N. et al. Mutant phenotypes for thousands of bacterial genes of unknown function. Nature 557, 503–509 (2018).
Wetmore, K. M. et al. Rapid quantification of mutant fitness in diverse bacteria by sequencing randomly bar-coded transposons. mBio 6, e00306 (2015).
Rasko, D. A. et al. The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates. J. Bacteriol. 190, 6881–6893 (2008).
Touchon, M. et al. Organised genome dynamics in the Escherichia coli species results in highly diverse adaptive paths. PLoS Genet. 5, e1000344 (2009).
Touchon, M. et al. Phylogenetic background and habitat drive the genetic diversification of Escherichia coli. PLoS Genet. 16, e1008866 (2020).
Tenaillon, O., Skurnik, D., Picard, B. & Denamur, E. The population genetics of commensal Escherichia coli. Nat. Rev. Microbiol. 8, 207–217 (2010).
Denamur, E., Clermont, O., Bonacorsi, S. & Gordon, D. The population genetics of pathogenic Escherichia coli. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-020-0416-x (2020).
Subashchandrabose, S., Smith, S. N., Spurbeck, R. R., Kole, M. M. & Mobley, H. L. T. Genome-wide detection of fitness genes in uropathogenic Escherichia coli during systemic infection. PLoS Pathog. 9, e1003788 (2013).
Olson, M. A., Siebach, T. W., Griffitts, J. S., Wilson, E. & Erickson, D. L. Genome-wide identification of fitness factors in mastitis-associated Escherichia coli. Appl. Environ. Microbiol. 84, e02190 (2018).
Phan, M.-D. et al. The serum resistome of a globally disseminated multidrug resistant uropathogenic Escherichia coli clone. PLoS Genet. 9, e1003834 (2013).
Goh, K. G. K. et al. Genome-wide discovery of genes required for capsule production by uropathogenic Escherichia coli. mBio 8, e01558 (2017).
Shea, A. E. et al. Escherichia coli CFT073 fitness factors during urinary tract infection: identification using an ordered transposon library. Appl. Environ. Microbiol. 86, e00691–20 (2020).
Warr, A. R. et al. Transposon-insertion sequencing screens unveil requirements for EHEC growth and intestinal colonization. PLoS Pathog. 15, e1007652 (2019).
Bergmiller, T., Ackermann, M. & Silander, O. K. Patterns of evolutionary conservation of essential genes correlate with their compensability. PLoS Genet. 8, e1002803 (2012).
Patrick, W. M., Quandt, E. M., Swartzlander, D. B. & Matsumura, I. Multicopy suppression underpins metabolic evolvability. Mol. Biol. Evol. 24, 2716–2722 (2007).
Martínez-Carranza, E. et al. Variability of bacterial essential genes among closely related bacteria: the case of Escherichia coli. Front. Microbiol. 9, 1059 (2018).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Bikard, D. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 41, 7429–7437 (2013).
Vigouroux, A. & Bikard, D. CRISPR tools to control gene expression in bacteria. Microbiol. Mol. Biol. Rev. 84, e00077–19 (2020).
Rousset, F. & Bikard, D. CRISPR screens in the era of microbiomes. Curr. Opin. Microbiol. 57, 70–77 (2020).
Cui, L. et al. A CRISPRi screen in E. coli reveals sequence-specific toxicity of dCas9. Nat. Commun. 9, 1912 (2018).
Rousset, F. et al. Genome-wide CRISPR-dCas9 screens in E. coli identify essential genes and phage host factors. PLoS Genet. 14, e1007749 (2018).
Wang, T. et al. Pooled CRISPR interference screening enables genome-scale functional genomics study in bacteria with superior performance. Nat. Commun. 9, 2475 (2018).
Calvo-Villamañán, A. et al. On-target activity predictions enable improved CRISPR-dCas9 screens in bacteria. Nucleic Acids Res. 48, e64 (2020).
Li, S. et al. Genome-wide CRISPRi-based identification of targets for decoupling growth from production. ACS Synth. Biol. 9, 1030–1040 (2020).
Lee, H. H. et al. Functional genomics of the rapidly replicating bacterium Vibrio natriegens by CRISPRi. Nat. Microbiol. 4, 1105–1113 (2019).
Yao, L. et al. Pooled CRISPRi screening of the cyanobacterium Synechocystis sp PCC 6803 for enhanced industrial phenotypes. Nat. Commun. 11, 1666 (2020).
Liu, X. et al. Exploration of bacterial bottlenecks and Streptococcus pneumoniae pathogenesis by CRISPRi-seq. Cell Host Microbe https://doi.org/10.1016/j.chom.2020.10.001 (2020)
Schnider-Keel, U. et al. Autoinduction of 2,4-diacetylphloroglucinol biosynthesis in the biocontrol agent Pseudomonas fluorescens CHA0 and repression by the bacterial metabolites salicylate and pyoluteorin. J. Bacteriol. 182, 1215–1225 (2000).
Decrulle, A., Fernandez Rodriguez, J., Duportet, X. & Bikard, D. Optimized vector for delivery in microbial populations. International patent WO2018141907 (2018).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837–845 (1988).
Goodman, A. L. et al. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl Acad. Sci. USA 108, 6252–6257 (2011).
Rocha, E. P. C. & Danchin, A. An analysis of determinants of amino acids substitution rates in bacterial proteins. Mol. Biol. Evol. 21, 108–116 (2004).
Tian, W. & Skolnick, J. How well is enzyme function conserved as a function of pairwise sequence identity? J. Mol. Biol. 333, 863–882 (2003).
Tye, B.-K. & Lehman, I. R. Excision repair of uracil incorporated in DNA as a result of a defect in dUTPase. J. Mol. Biol. 117, 293–306 (1977).
Schaub, R. E. & Hayes, C. S. Deletion of the RluD pseudouridine synthase promotes SsrA peptide tagging of ribosomal protein S7. Mol. Microbiol. 79, 331–341 (2011).
Luo, P., He, X., Liu, Q. & Hu, C. Developing universal genetic tools for rapid and efficient deletion mutation in Vibrio species based on suicide T-vectors carrying a novel counterselectable marker, vmi480. PLoS ONE 10, e0144465 (2015).
Aakre, C. D., Phung, T. N., Huang, D. & Laub, M. T. A bacterial toxin inhibits DNA replication elongation through a direct interaction with the β sliding clamp. Mol. Cell 52, 617–628 (2013).
Harms, A., Brodersen, D. E., Mitarai, N. & Gerdes, K. Toxins, targets and triggers: an overview of toxin–antitoxin biology. Mol. Cell 70, 768–784 (2018).
Burroughs, A. M., Zhang, D., Schäffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015).
Bobonis, J. et al. Bacterial retrons encode tripartite toxin/antitoxin systems. Preprint at bioRxiv https://doi.org/10.1101/2020.06.22.160168 (2020).
Bobonis, J. et al. Phage proteins block and trigger retron toxin/antitoxin systems. Preprint at bioRxiv https://doi.org/10.1101/2020.06.22.160242 (2020).
Millman, A. et al. Bacterial retrons function in anti-phage defense. Cell https://doi.org/10.1016/j.cell.2020.09.065 (2020).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Ferrières, L. et al. Silent mischief: bacteriophage Mu insertions contaminate products of Escherichia coli random mutagenesis performed using suicidal transposon delivery plasmids mobilized by broad-host-range RP4 conjugative machinery. J. Bacteriol. 192, 6418–6427 (2010).
Engler, C., Gruetzner, R., Kandzia, R. & Marillonnet, S. Golden gate shuffling: a one-pot DNA shuffling method based on type IIs restriction enzymes. PLoS ONE 4, e5553 (2009).
Salis, H. M. in Methods in Enzymology Vol. 498 (Elsevier, 2011).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Hartley, J. L., Temple, G. F. & Brasch, M. A. DNA cloning using in vitro site-specific recombination. Genome Res. 10, 1788–1795 (2000).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 17, 132 (2016).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Steinegger, M. & Söding, J. Clustering huge protein sequence sets in linear time. Nat. Commun. 9, 2542 (2018).
St-Pierre, F. et al. One-step cloning and chromosomal integration of DNA. ACS Synth. Biol. 2, 537–541 (2013).
Clermont, O., Christenson, J. K., Denamur, E. & Gordon, D. M. The Clermont Escherichia coli phylo-typing method revisited: improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 5, 58–65 (2013).
Bouvet, O., Bourdelier, E., Glodt, J., Clermont, O. & Denamur, E. Diversity of the auxotrophic requirements in natural isolates of Escherichia coli. Microbiology 163, 891–899 (2017).
Roberts, R. J., Vincze, T., Posfai, J. & Macelis, D. REBASE—a database for DNA restriction and modification: enzymes, genes and genomes. Nucleic Acids Res. 43, D298–D299 (2015).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Treangen, T. J., Ondov, B. D., Koren, S. & Phillippy, A. M. The Harvest suite for rapid core-genome alignment and visualization of thousands of intraspecific microbial genomes. Genome Biol. 15, 524 (2014).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Mitchell, A. L. et al. InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res. 47, D351–D360 (2019).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Deatherage, D. E. & Barrick, J. E. Identification of mutations in laboratory-evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–188 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Acknowledgements
We thank B. Dupuy for sharing the use of the anaerobic chamber, as well as O. Tenaillon, P.-A. Kaminsky, G. Sezonov, A. Danchin and A. Calvo-Villamañan for useful discussions. We thank the P2M platform (Institut Pasteur, Paris, France) for genome sequencing and V. Briolat from the Biomics platform, C2RT, Institut Pasteur, Paris, France, supported by France Génomique (ANR-10-INBS-09-09) and IBISA. We also thank J. Rodríguez-Beltrán and S. Brisse for providing a strain of Citrobacter freundii and Klebsiella pneumoniae, respectively. This work was supported by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 677823), by the French Government’s Investissement d’Avenir programme and by Laboratoire d’Excellence ‘Integrative Biology of Emerging Infectious Diseases’ (ANR-10-LABX-62-IBEID). F.R. is supported by a doctoral scholarship from Ecole Normale Supérieure. E.D. was partially supported by the ‘Fondation pour la Recherche Médicale’ (Equipe FRM 2016, grant no. DEQ20161136698). E.P.C.R. was partially supported by the ‘Fondation pour la Recherche Médicale’ (Equipe FRM EQU201903007835).
Author information
Authors and Affiliations
Contributions
F.R. and D.B. designed the project. E.P.C.R. performed bioinformatic computation of the E. coli pangenome. E.D. and O.C. provided strains and genome sequences. F.R. performed experiments and analysed data. J.F.-R. and F.P.-F. participated in the design of pFR56. J.C.-C. provided experimental assistance. F.R., E.P.C.R. and D.B. wrote the manuscript. D.B. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Microbiology thanks Carol Gross and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The plasmid pFR56 is an optimized and transferrable platform for CRISPRi screening in Escherichia and closely-related genera.
a, In a previous study, we identified the ‘bad-seed’ effect in E. coli, a sequence-specific toxicity of dCas9 at high expression levels and whose mechanism remains to be elucidated8. We screened a ribosome-binding site (RBS) library to optimize the expression level of dCas9 in order to avoid this toxicity effect while maintaining efficient repression (see Methods).Ten-fold serial dilutions of cells were spotted on LB plates supplemented with chloramphenicol (Cm) and 50 µM DAPG. The nucleotide sequences of sgRNAs are described in Supplementary Table 8. b, The conjugation efficiency of pFR56 was assessed by measuring the proportion of chloramphenicol-resistant recipient cells in E. coli K-12 MG1655 and CFT073 and in Escherichia albertii, Escherichia fergusonii, Klebsiella pneumoniae and Citrobacter freundii (n = 3 independent replicates). c, Plasmid stability was assessed in these strains by measuring the proportion of chloramphenicol-resistant cells after 24 generation of growth in LB without chloramphenicol (n = 3 independent replicates). d, CRISPRi-mediated killing was measured in these strains after conjugation of an sgRNA targeting the essential gene rpsL. Ten-fold serial dilutions of cells were spotted on LB plates with chloramphenicol and with or without 50 µM DAPG. e, The EcoCG library was synthesized, cloned onto pFR56 and transferred to K-12 MG1655 by conjugation and the abundance of guides in the library was monitored by deep sequencing before and after 20 generations with dCas9 induction. f, The correlation of experimental replicates suggests an excellent reproducibility. Source data is available in Supplementary Table 11. g, The log2FC value was calculated for each sgRNA and the median log2FC value was used as a gene score to predict gene essentiality using the TraDIS dataset9 as ground truth. The plot shows the receiver operating characteristic (ROC) curve of the prediction model. The dashed black line marks the threshold chosen in further analysis (gene score < -3). h, For each gene, the standard-deviation of log2FC values of the different sgRNAs was calculated for the EcoCG library and for our previous genome-wide library8,10. In order to account for the difference in library size, we also calculated for each gene the mean standard-deviation of log2FC values obtained from all permutations of 3 sgRNAs from our previous library. i, Two examples of polar effects observed in our screen. Each dot represents a guide and its associated position and log2FC value. Genes from a same operon are displayed. In both cases, repression of a nonessential gene located upstream an essential gene induces a fitness defect. E: essential; NE: nonessential.
Extended Data Fig. 2 CRISPRi screening and RNA-seq experiments are highly reproducible.
(a-b) Histograms show the distribution of Pearson’s correlation coefficients between biological replicates. CRISPRi screening experiments were performed in two biological replicates (a) with 18 strains in LB (top) and GMM (bottom) and 14 strains in M9-glucose (middle). RNA-seq was performed in three biological replicates in 16 strains (b), representing 48 pairwise comparisons.
Extended Data Fig. 3 Comparison of CRISPRi screening results in different media reveals conditionally-essential genes.
(a,b) The median gene score across all strains was used to assess the most widespread genetic requirements in the E. coli species. a, Comparison between screening results in LB and M9-glucose medium highlights auxotrophy genes. Genes involved in amino acid and nucleotide biosynthesis and in sulfate assimilation are highlighted. b, Comparison of screening results in LB and in GMM reveal medium-specific essential genes. ATP synthase genes are highlighted in red, LB-specific essential genes are highlighted in blue and GMM-specific essential genes are highlighted in green.
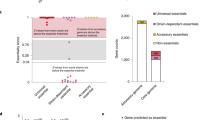
Extended Data Fig. 4 Evolution of the proportion of core- and pan-essential genes in the core genome (related to Fig. 1).
The fraction of core genes that are essential in all strains (a) or in at least one strain (b) was calculated for various sets of strains. Data are presented as mean values ± standard deviation of n = 250 random permutations.
Extended Data Fig. 5 The relationship between gene expression and the frequency of essentiality in an E. coli strain panel.
a, Spearman’s correlation between gene score in CRISPRi screens and gene expression for 2698 core genes is shown for the 16 strains assayed in both experiments. b, We selected 245 genes that are essential in all 16 strains (gene score < -3 in all strains), 87 genes that are variably essential (gene score < -3 in ≥1 strain and gene score > -1 in ≥1 strain), and 2056 genes that are never essential in the tested strains (gene score > -1 in all strains). Violin-plots show the distribution of the median gene expression level across all strains. Inside each distribution, the white dot shows the median and the extremities of the black bar show the 1st and 3rd quartiles of the distribution. p-values were calculated using a two-sided Mann-Whitney U test. c, For each ‘variably essential’ gene, we calculated Spearman’s correlation coefficient between gene expression level and gene score across the 16 strains. This plot shows the distribution of these correlation coefficients.
Extended Data Fig. 6 The influence of phylogeny on gene expression (related to Fig. 2).
For each pair of strains, we used Spearman’s correlation coefficient as a measure of the similarity in gene expression profile or gene essentiality profile. Each plot represents the relationship between the phylogenetic distance and the similarity in gene expression profile between a given strain and the 15 remaining strains assayed during RNA-seq experiments.
Extended Data Fig. 7 The influence of phylogeny on core gene essentiality (related to Fig. 2).
a, The Spearman correlation between CRISPRi fitness profile and phylogenetic distance was calculated for all pairs of strains (n = 153 in LB and GMM, 91 in M9) from the same or from different phylogroups, considering 3276 core protein-coding genes in LB (left), M9 (center) and GMM (right). Boxes show the 1st and 3rd quartiles with the median value at the center, while whiskers extend to the minimum and maximum of the distribution. P-values of Student’s two-tailed t-tests are shown. b, In each condition, we selected genes that are essential in at least 2 strains while being nonessential in at least 2 strains. On a case-by-case basis, we discarded genes in operons when the effect was due to a polar effect in the downstream gene to avoid redundancy (for instance we discarded ycaR whose fitness values are explained by a polar effect in kdsB). For each of the resulting gene (30 in LB, 41 in M9 and 56 in GMM), we calculated the Spearman correlation of the phylogenetic distance of pairs of strains with the absolute difference in their gene score. We then plotted the distribution of the correlation coefficients.
Extended Data Fig. 8 Functional redundancy between kdsB and kpsU.
a, A heatmap shows the fitness defect associated with the repression of kdsB. The gene score values are shown inside each box. Grey circles represent the presence of kpsU in the genome. b, A PCR using kpsU-specific primers (FR257 and FR258, Supplementary Table 7) followed by gel electrophoresis was performed to confirm the presence of kpsU in selected strains. A 469-bp product is expected when kpsU is present. This assay was performed once. c, Spot assays show the phenotype associated with the repression of kdsB, kpsU or both kdsB and kpsU simultaneously in strains carrying kpsU. Ten-fold serial dilutions were plated on LB supplemented with chloramphenicol and 50 µM DAPG. K-12-MG1655 does not have a copy of kpsU and was used as a control. Interestingly, CFT073 also carries kpsU but remain very sensitive to kdsB knockdown suggesting that kpsU might not be expressed or functional in these strains. The nucleotide sequences of sgRNAs are shown in Supplementary Table 8.
Extended Data Fig. 9 ECTA447_03166 expression level is insufficient in GMM to compensate the loss of dut in TA447 (related to Fig. 4).
a, Growth curves were performed in triplicates in LB or GMM with TA447 and TA447Δdut. b, The relative expression of TA447_03166 was measured by RT-qPCR in LB and in GMM. Horizontal bars show the mean of 3 biological replicates and 2 technical replicates. P-value is shown for a two-tailed t-test (t = 3.7391, 95% CI: [0.14,0.67], p = 0.009987).
Extended Data Fig. 10 ybaQ is a transcriptional repressor of the HigB-1 toxin.
a, A phylogenetic tree was built with the 18 strains assayed in CRISPRi screens. Grey squares indicate if ybaQ is essential in the corresponding strains. The genomic region of the ybaQ locus is shown on the right and varies according to three clades. (*) higB is interrupted by a stop codon in strain CFT073. b, CRISPRi screens results were validated by introducing ybaQ sgRNA in three strains where ybaQ repression has no effect (K-12-MG1655, E101 and TA447) and in five strains where ybaQ repression is toxic (UTI89, S88, JJ1886, APEC O1 and ROAR8). Ten-fold serial dilutions of cells were spotted on LB plates containing chloramphenicol with or without 50 µM DAPG to induce dCas9 expression. The sequence of the ybaQ sgRNA is provided in Supplementary Table 8. c, Silencing ybaQ in strain S88 induces the expression of higB-1. The horizontal bar shows the mean of 3 biological replicates and 2 technical replicates from RT-qPCR experiments. Primers used for qPCR are shown in Supplementary Table 9.
Supplementary information
Supplementary Information
Supplementary Figs. 1–2 and results.
Supplementary Tables 1–11.
Supplementary Tables 1–11.
Source data
Source Data Fig. 1
Source data for Fig. 1d.
Source Data Fig. 2
Source data for Fig. 2.
Source Data Fig. 3
Source data for Fig. 3.
Source Data Extended Data Fig. 1
Source data for Extended Data Fig. 1b,c.
Source Data Extended Data Fig. 2
Source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 9
Source data for Extended Data Fig. 9b.
Source Data Extended Data Fig. 10
Source data for Extended Data Fig. 10c.
Rights and permissions
About this article
Cite this article
Rousset, F., Cabezas-Caballero, J., Piastra-Facon, F. et al. The impact of genetic diversity on gene essentiality within the Escherichia coli species. Nat Microbiol 6, 301–312 (2021). https://doi.org/10.1038/s41564-020-00839-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00839-y
This article is cited by
-
(p)ppGpp modifies RNAP function to confer β-lactam resistance in a peptidoglycan-independent manner
Nature Microbiology (2024)
-
Metabolic cooperation between conspecific genotypic groups contributes to bacterial fitness
ISME Communications (2023)
-
Molecular mechanisms of antibiotic resistance revisited
Nature Reviews Microbiology (2023)
-
Characterization of antibiotic resistomes by reprogrammed bacteriophage-enabled functional metagenomics in clinical strains
Nature Microbiology (2023)
-
Transposon insertional mutagenesis of diverse yeast strains suggests coordinated gene essentiality polymorphisms
Nature Communications (2022)