Abstract
Coronavirus disease 2019 (COVID-19) is caused by infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)1,2,3 and individuals with COVID-19 have symptoms that can be asymptomatic, mild, moderate or severe4,5. In the early phase of infection, T- and B-cell counts are substantially decreased6,7; however, IgM8,9,10,11 and IgG12,13,14 are detectable within 14 d after symptom onset. In COVID-19-convalescent individuals, spike-specific neutralizing antibodies are variable3,15,16. No specific drug or vaccine is available for COVID-19 at the time of writing; however, patients benefit from treatment with serum from COVID-19-convalescent individuals17,18. Nevertheless, antibody responses and cross-reactivity with other coronaviruses in COVID-19-convalescent individuals are largely unknown. Here, we show that the majority of COVID-19-convalescent individuals maintained SARS-CoV-2 spike S1- and S2-specific antibodies with neutralizing activity against the SARS-CoV-2 pseudotyped virus, and that some of the antibodies cross-neutralized SARS-CoV, Middle East respiratory syndrome coronavirus or both pseudotyped viruses. Convalescent individuals who experienced severe COVID-19 showed higher neutralizing antibody titres, a faster increase in lymphocyte counts and a higher frequency of CXCR3+ T follicular help (TFH) cells compared with COVID-19-convalescent individuals who experienced non-severe disease. Circulating TFH cells were spike specific and functional, and the frequencies of CXCR3+ TFH cells were positively associated with neutralizing antibody titres in COVID-19-convalescent individuals. No individuals had detectable autoantibodies. These findings provide insights into neutralizing antibody responses in COVID-19-convalescent individuals and facilitate the treatment and vaccine development for SARS-CoV-2 infection.
Similar content being viewed by others
Main
To investigate the antibody response after recovery from coronavirus disease 2019 (COVID-19), 67 convalescent individuals were recruited for this study, and blood was drawn on day 28 after discharge. The baseline clinical characteristics and laboratory findings on admission were retrospectively analysed (Extended Data Figs. 1 and 2). The binding and avidity of an antibody to a specific antigen reflect the overall strength and quality of the antibody. To assess the reactivity of COVID-19 serum antibodies, the binding activity and avidity of the serum to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike subunits S1 and S2 were examined using a human-IgG-specific enzyme-linked immunosorbent assay (ELISA). The results showed that all 67 sera contained anti-S1 (median titre = 4.61; interquartile range (IQR) = 4.01–4.61) and anti-S2 (median titre = 4.91; IQR = 4.61–5.52) IgG antibodies (Fig. 1a and Extended Data Fig. 3). Furthermore, the end-point titres of anti-S1 and anti-S2 antibodies were positively correlated (Extended Data Fig. 4a). To evaluate binding specificity, 61 sera collected from healthy control individuals were examined in parallel, and only one serum sample (1/61, 1.6%) was found to have binding activity against SARS-CoV-2 spike S1 but not S2 (Extended Data Fig. 5). Similar patterns were observed for antibody avidity, in which 64 out of 67 sera (3 sera were excluded as they did not meet the requirements for the avidity assay) exhibited medium avidity to S1 (median = 44.50; IQR = 34.50–51.75) and S2 (median = 58.00; IQR = 49.00–67.00; Fig. 1b and Extended Data Fig. 3). These data indicate that the majority of the COVID-19-convalescent individuals elicited and maintained robust SARS-CoV-2 spike-specific IgG antibodies with a medium avidity.
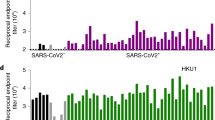
a, End-point dilution titre of serum IgG antibodies from individuals who recovered from COVID-19 (n = 67), determined by antibody binding to SARS-CoV-2 spike subunits S1 and S2 and shown as final dilution titres. b, The avidity of serum IgG antibodies to SARS-CoV-2 spike subunits S1 and S2 (n = 64), shown as avidity index. c, The percentage of sera derived from COVID-19-convalescent individuals with cross-binding activity to spike subunits S1 and S2 from both SARS-CoV and MERS-CoV. d, The percentage of sera derived from COVID-19-convalescent individuals that neutralizes SARS-CoV-2 pseudotyped virus (ID50 > 30) (n = 65/67, 97.01%). e, Neutralization titre of sera derived from COVID-19-convalescent individuals against SARS-CoV-2 pseudotyped virus (ID50, n = 65). f, The percentage of the SARS-CoV-2-neutralizing sera (n = 65) that shows cross-neutralization against pseudotyped viruses of SARS-CoV and MERS-CoV. The plus and minus symbols indicate with and without cross-neutralization, respectively. For a and e, end-point titres and neutralization titres were log10-transformed. For a, b, and e, data are median ± IQR (25–75%).
The cross-reactivity of antibodies has been reported for coronaviruses, as sera from recovered SARS individuals showed cross-reaction with Middle East respiratory syndrome coronavirus (MERS-CoV)19. SARS-CoV-2 shows high sequence homology with SARS-CoV and less similarity to MERS-CoV2,3. To examine whether the serum antibodies derived from COVID-19-convalescent individuals cross-react with other coronaviruses, a binding assay was performed using spike subunits S1 and S2 from both SARS-CoV and MERS-CoV. The results showed that 38.81% (26/67) and 89.55% (60/67) of the sera bound to SARS-CoV S1 and S2, respectively (Fig. 1c and Extended Data Fig. 3). For MERS-CoV, no binding was detected for S1, whereas 73.13% (49/67) of the sera bound to S2 (Fig. 1c and Extended Data Fig. 3). As negative controls, 61 healthy sera were included in each experiment, and no cross-binding was found for SARS-CoV S1. Only 1.67% (1/61), 6.66% (4/61) and 1.67% (1/61) of the sera showed binding activity to SARS-CoV S2, MERS-CoV S1 and MERS-CoV S2, respectively (Extended Data Fig. 5). These results suggest that COVID-19-convalescent individuals elicited antibodies with cross-binding activity to SARS-CoV and MERS-CoV.
Next, sera neutralization of SARS-CoV-2 and its cross-neutralization of SARS-CoV and MERS-CoV were determined. SARS-CoV-2 pseudotyped virus was constructed and used for neutralization20, and previously developed SARS-CoV and MERS-CoV pseudotyped viruses were used for cross-neutralization21. The results showed that 97.01% (65/67) of the sera from individuals who had recovered from COVID-19 contained neutralizing antibodies that efficiently neutralized SARS-CoV-2 pseudotyped virus (50% inhibitory dilution (ID50) = 2.75; IQR = 2.34–3.08; Fig. 1d,e and Extended Data Fig. 3), and the neutralization titres were positively correlated with the end-point binding titres of anti-S1 and anti-S2 antibodies (Extended Data Fig. 4b,c). Cross-neutralization was observed and variable for different viruses; 65.67% (44/67), 7.46% (5/67) and 10.45% (7/67) of the sera cross-neutralized SARS-CoV, MERS-CoV and both viruses, respectively. Furthermore, 16.42% (11/67) of the sera showed no cross-neutralization activity (Fig. 1f and Extended Data Fig. 3). However, having cross-neutralization activity did not seem to affect the ability of a serum to neutralize SARS-CoV-2, as the serum groups with and without cross-neutralization activity showed no significant difference in the neutralization titres for SARS-CoV-2 (Extended Data Fig. 6a). Similarly, neutralization titres for SARS-CoV-2 were also comparable between those sera that cross-neutralized SARS-CoV, MERS-CoV or both viruses (Extended Data Fig. 6b). Nevertheless, cross-neutralization titres for SARS-CoV and MERS-CoV were significantly lower than their neutralization titres for SARS-CoV-2 in general (Extended Data Fig. 6c–e). Furthermore, autoantibodies were not detectable in the tested sera derived from COVID-19-convalescent individuals (Extended Data Fig. 7). These findings demonstrate that the majority of patients with COVID-19 elicited and maintained robust neutralizing antibody responses to SARS-CoV-2 after recovery; the serum of some convalescent individuals also contained antibodies with cross-binding and neutralizing activities to SARS-CoV, MERS-CoV or both coronaviruses.
It has been reported that patients with COVID-19 with severe and mild symptoms showed distinct clinical and immunological presentations5,17,18. To study whether the severity of COVID-19 symptoms that patients experienced affects the neutralizing antibody responses after disease recovery, 67 convalescent individuals were categorized into severe (17/67) and non-severe (50/67) groups, according to the severity of disease that the patients had (Extended Data Fig. 1). Compared with the non-severe group, the severe group exhibited higher end-point titres of antibodies, anti-S1 (P = 0.011) and anti-S2 (P = 0.010; Fig. 2a) antibodies, and neutralization titres (P < 0.001) to SARS-CoV-2 (Fig. 2b). However, in the severe and non-severe groups, there was no difference in the avidity indexes of either anti-S1 (P = 0.609) or anti-S2 (P = 0.857; Fig. 2c) antibodies. Although the severe group had an older age (P < 0.001), a longer course of disease (P = 0.007), more comorbidities (P = 0.044) and a higher total of underlying diseases (P < 0.001) compared with the non-severe group (Extended Data Fig. 1), multivariable analysis of the factors that may be associated with neutralizing antibody responses revealed that neutralizing antibody titres were correlated with the severity of disease (odds ratio (OR) = 5.04, 95% confidence interval (CI) = 1.003–25.30; Table 1).
a, Comparison of end-point titres of SARS-CoV-2 spike S1- and S2-specific IgG antibodies from COVID-19-convalescent individuals who had experienced severe (n = 17) and non-severe (n = 50) symptoms. b, Comparison of neutralization titres to SARS-CoV-2 pseudotyped virus for the sera from convalescent individuals who had severe (n = 17) and non-severe (n = 50) COVID-19. c, Comparison of avidity indexes of SARS-CoV-2 spike S1- and S2-specific IgG antibodies derived from convalescent individuals who had severe (n = 16) and non-severe (n = 48) COVID-19. d,e, The kinetics of lymphocyte counts (d) and the fold change in lymphocytes relative to admission for the time points as indicated (e) for the severe (red, circle) and non-severe (blue, square) groups of convalescent individuals, in which the number of patients included in the analysis was as follows (severe versus non-severe): on admission (n = 17 versus n = 50), mid-course of disease (n = 17 versus n = 41), discharge (n = 16 versus n = 30), 14 d after discharge (n = 13 versus n = 40) and 28 d after discharge (n = 13 versus n = 40). For a and b, end-point titres and neutralization titres were logarithmically transformed. In d and e, the dotted lines indicate the bottom line of the normal range of lymphocyte count and the baseline of the fold change in lymphocytes related to admission, respectively. For a–e, data are median ± IQR (25–75%). Mann–Whitney U-tests were used to analyse the difference between two groups; P < 0.05 was considered to be a two-tailed significant difference.
As the neutralizing antibody responses are essentially related to the functional lymphocytes, the lymphocyte kinetics in severe and non-severe groups were therefore investigated from admission to recovery phase at the following five time points: admission, mid-course, discharge, day 14 and day 28 after discharge (Fig. 2d). The results revealed that the majority of the severe group (16/17, 94.12%) had lower lymphocyte counts at the first four time-points (P ≤ 0.05). This observation is consistent with those of a recent report, in which patients with severe COVID-19 showed decreased T- and B-cell counts4, and lymphopaenia was often observed on admission, especially for those with severe symptoms4,7. Here, lymphocyte counts of the non-severe group on admission were close to the bottom line of the normal range (Fig. 2d). Lymphocyte counts of both groups increased gradually, and the severe group was restored to a normal level on discharge (median, from 0.65 to 2.28 × 109 per litre) and maintained this level for two weeks. From day 14 to day 28 after discharge, lymphocyte counts of the severe group increased rapidly and reached a mean value that was higher than that of the non-severe group (P = 0.08; Fig. 2d). Although lymphocyte counts of both groups increased, the severe group showed higher fold changes at all time points, related to admission, compared with the non-severe group (P < 0.01; Fig. 2e), suggesting that convalescent individuals who had severe COVID-19 have higher restoration rates of lymphocytes, which may contribute to antibody maintenance and neutralizing antibody responses.
TFH cells, a T-cell subset, have been identified as professional B helper T cells in past decades and are required for T-dependent antibody production22,23. Circulating TFH cells are representative of germinal centre TFH cells and have an important role in T-dependent B-cell maturation and antibody production24. As the severe group had higher antibody titres in binding and neutralization (Fig. 2a,b), as well as a faster increase in lymphocyte counts (Fig. 2d), it is interesting to investigate whether the TFH cells of lymphocytes are correlated with the neutralization effect. Circulating TFH cells were analysed by gating on PD-1+CXCR5+CD4+ T cells (Extended Data Fig. 8a). Analysis of the frequency of total circulating TFH cells showed that COVID-19-convalescent individuals and healthy controls had no difference in the frequency of total TFH cells (Extended Data Fig. 8b); however, the frequency of CXCR3+ TFH-cell subsets in total TFH cells of COVID-19-convalescent individuals (both severe and non-severe groups) was lower compared with that of healthy controls (Extended Data Fig. 8c–e). When analysed by severe, non-severe and healthy control groups, total TFH cells were comparable among three groups (Fig. 3a); however, similar to the healthy control group, the severe group showed a higher frequency of CXCR3+ TFH cells compared with the non-severe group (P = 0.006). Accordingly, the frequency of CXCR3− TFH cells (P = 0.001) and the ratio of CXCR3+/CXCR3− TFH cells (P = 0.013) were lower and higher compared with the non-severe group, respectively (Fig. 3b–d). Correlation coefficient analysis revealed that the frequency of total TFH cells had no correlation with neutralization titres (ID50; r = −0.004, P = 0.985), but CXCR3+ TFH cells were positively correlated with neutralization titres (r = 0.486, P = 0.012; Fig. 3e,f). As expected, CXCR3− TFH cells (r = −0.435, P = 0.025) and the ratio of CXCR3+/CXCR3− TFH cells (r = 0.467, P = 0.016) were negatively and positively correlated with neutralization titres, respectively (Fig. 3g,h). These findings suggest that CXCR3+ TFH cells may have a dominant role in the early initiation and/or maintenance of neutralizing antibody responses in individuals who have recovered from COVID-19. The relatively high frequency of CXCR3+ TFH-cell subsets may contribute to the higher level of neutralizing antibody responses in the severe group compared with that in non-severe group.
a–d, The frequency of circulating total TFH cells (a), CXCR3+ TFH cells (b) and CXCR3− TFH cells (c), as well as the ratio of CXCR3+/CXCR3− TFH cells (d) in healthy controls (HC) (n = 17), severe groups (n = 10) and non-severe groups (n = 16). The gating strategy for the TFH cells (PD-1+CXCR5+CD4+ T cells) is presented in Extended Data Fig. 8a, and a comparison of TFH-cell subsets between healthy controls (n = 17) and individuals who recovered from COVID-19 (n = 26) is shown in Extended Data Fig. 8b. NS, not significant. e–h, Correlation analysis of the neutralization titres of antibodies and the frequencies of total TFH cells (e), CXCR3+ TFH cells (f) and CXCR3− TFH cells (g), as well as the neutralization titres of antibodies and the ratio of CXCR3+/CXCR3− TFH cells (h) in individuals who recovered from COVID-19 (n = 26). Neutralization titres were log10-transformed. i–l, Spike-specific circulating TFH-cell response after antigen stimulation. Spike-specific CD154+ TFH cells (n = 5) (i) and OX40+CD25+ TFH cells (n = 6) (j), and intracellular IL-21 (n = 5) (k) and IFNγ (n = 6) (l) were measured using flow cytometry after stimulation. For a–d, data are median ± IQR (25–75%). One-way analysis of variance was used to compare the difference between multiple groups, and Tukey’s multiple-comparisons test was used to compare differences within the groups. For e–h, Spearman’s rank correlation coefficient was used to describe the association between the frequency of TFH-cell subsets and the neutralization titres. For i–l, paired t-tests were used to analyse the difference in the frequency of spike-specific TFH-cell subsets from COVID-19-convalescent individuals and healthy controls after stimulation. P < 0.05 was considered to be a two-tailed significant difference.
To further examine whether the TFH cells were SARS-CoV-2 spike specific, peripheral blood mononuclear cells (PBMCs) from COVID-19-convalescent individuals and healthy controls were stimulated with SARS-CoV-2 spike (S1 + S2) proteins or bovine serum albumin (BSA), and the antigen-specific TFH-cell populations were analysed for the expression of surface marker CD154, the frequency of the OX40+CD25+ population, and the release of cytokines IL-21 and IFNγ (Extended Data Fig. 9a–d). As expected, CD154 expression and the CD25+OX40+ population were significantly increased in spike-stimulated total TFH cells and CXCR3+ and CXCR3− TFH-cell subsets compared with BSA-stimulated controls (Fig. 3i,j). By contrast, SARS-CoV-2 spike stimulation did not significantly increase CD154 expression and the frequency of the CD25+OX40+ population in TFH cells or their subsets of PBMCs from healthy controls (Fig. 3i,j). These results suggest that the responsive TFH cells were SARS-CoV-2 spike specific. IL-21 is a signature marker cytokine of TFH cells and is critical for TFH-cell function25,26. Similarly, the release of IL-21 was significantly increased in total TFH cells and TFH-cell subsets after spike stimulation compared with the BSA-stimulated controls (Fig. 3k). IFNγ was produced mainly by CXCR3+ TFH cells after spike stimulation (Fig. 3l); for the healthy control group, a very low level of IFNγ was detected after stimulation with spike or BSA (Fig. 3l). Together, these data demonstrate that circulating TFH cells in individuals who have recovered from COVID-19 are spike specific and functional—this is critical for the initiation and/or maintenance of neutralizing antibodies after SARS-CoV-2 infection.
Here, we systematically investigated neutralizing antibody responses in individuals who had recovered from COVID-19. The majority of COVID-19-convalescent individuals maintained antibodies with neutralization activity against SARS-CoV-2 (Fig. 1d), and some of these antibodies cross-neutralized SARS-CoV and/or MERS-CoV (Fig. 1f). The cross-neutralization to SARS-CoV and MERS-CoV may be explained, at least in part, by the sequence homology of spike proteins of these three coronaviruses. SARS-CoV-2 spike amino acid sequences share ~76% with SARS-CoV spike but have only ~35% identity with MERS-CoV spike27, which may explain the finding that COVID-19 sera showed cross-neutralization more efficiently to SARS-CoV than to MERS-CoV (Fig. 1f). Neutralizing antibodies in COVID-19-convalescent individuals were reported but only for those with mild symptoms3,15. The cross-binding activity of COVID-19 sera was recently tested against SARS-CoV (spike and S1) and MERS-CoV spike, but included only three serum samples28. A report published during the revision of this manuscript also supported the finding that sera from patients who recovered from COVID-19 cross-reacted with SARS-CoV and MERS-CoV29. Patient-derived SARS-CoV monoclonal and receptor binding domain (RBD)-specific S309 antibodies could cross-neutralize SARS-CoV-2 (refs. 30,31,32); however, a low titre of SARS-CoV serum was found without cross-neutralization for SARS-CoV-2 (ref. 16). Indeed, with up to 67 participants, our study showed that antibody binding titres correlate positively with neutralization titres (Extended Data Fig. 4a–c), and neutralizing antibodies derived from COVID-19-convalescent individuals efficiently neutralized SARS-CoV-2 but were less efficient at neutralizing SARS-CoV and MERS-CoV (Extended Data Fig. 6c–e). It should be noted that cross-neutralization with MERS-CoV was most likely mediated by S2 binding only, as there was no detectable binding to S1 (Fig. 1c and Extended Data Fig. 3), which is consistent with recent reports19,28. The existence of cross-neutralizing antibodies in COVID-19-convalescent individuals provides the possibility to isolate broad neutralizing antibodies and to design pan-coronavirus vaccines.
Interestingly, individuals who recovered from severe COVID-19 elicited and maintained higher antibody and neutralization titres compared with the non-severe group (Fig. 2). Furthermore, neutralizing antibody titres were positively correlated with the severity of COVID-19 rather than other factors (Table 1 and Extended Data Fig. 1). These findings were similar to those from a report for patients who had recovered from MERS-CoV, in which the levels of neutralizing antibodies were positively associated with the number of days in ICU, viral shedding and the need for ventilation—several characteristics of critical conditions33. In individuals who recovered from MERS-CoV and SARS-CoV-2 infections, neutralizing antibody titres were positively correlated with antigen-specific CD4+ T cells15,33, and neutralizing antibody responses were more stable and lasted longer in those individuals who had recovered from severe symptoms of MERS-CoV infection34. Here, a rapid increase in lymphocyte counts in the severe COVID-19 convalescent group may be responsible for the production of the high levels of neutralizing antibodies (Fig. 2d–e). Indeed, this hypothesis is supported by the fact that a higher frequency of CXCR3+ TFH-cell subsets existed in the severe group compared with in non-severe group and was positively correlated with the neutralizing antibody titres (Fig. 3b–h). The correlation of CXCR3+ TFH cells with neutralizing antibody responses has been reported in other virus infections. Our previous study showed that, in patients with hepatitis C, CXCR3+ TFH cells are positively correlated with hepatitis C virus-neutralizing antibody titres and contributed to memory-B-cell activation and differentiation35. CXCR3+ICOS+CXCR5+CD4+ T cells36 and circulating T-helper-1-biased helper cells37 were also reported to be positively correlated with influenza virus- and HIV-specific antibody responses, respectively. TFH cells, as professional B helper cells, were differentiated, activated and progressively increased from day 7 after onset of SARS-CoV-2 infection38. Antigen stimulation substantially increased the frequency of the spike-specific CD154+ and OX40+CD25+ populations, as well as IL-21- and IFNγ-releasing cells in total TFH cells and CXCR3+ and CXCR3− TFH-cell subsets (Fig. 3i–l), which strongly supports the notion that TFH cells are involved in neutralizing antibody responses in the SARS-CoV-2 infection. However, CXCR3+ TFH cells and CXCR3− TFH cells may have different roles in antibody initiation and/or maintenance. Similar findings were published during the revision of this manuscript, in which CCR6+CXCR3− cells were dominant spike-specific circulating TFH cells that were negatively correlated with neutralizing antibody titres39. However, it remains unknown whether these circulating TFH cells originate from germinal centres and are able to support long-lived plasma cells that are critical for the development of a SARS-CoV-2 vaccine.
Taken together, this study provides insights into the understanding of antibody responses in COVID-19-convalescent individuals and evidence facilitating the development of broad neutralizing antibodies and pan-coronavirus vaccines.
Methods
Individuals and sample collection
A total of 67 individuals who had recovered from COVID-19 were enrolled in this study, and diagnosis of COVID-19 was made according to WHO interim guidance. All patients presenting as outpatients showed fever or respiratory symptoms; chest computed tomography scans identified abnormal pulmonary nodules, and SARS-CoV-2 infection was further confirmed using quantitative PCR at the local health authority. All patients were hospitalized in the Department of Infectious Disease, The Centre Hospital of Shaoyang, Hunan Province, China, between 23 January 2020 and 2 March 2020. Patients with COVID-19 were classified into either the severe group or the non-severe group according to the Guidelines for the Diagnosis and Treatment of New Coronavirus Pneumonia (v.5) published by the National Health Commission of China. Patients were classified into the severe group once they met one of the following criteria: (1) shortness of breath with a respiratory rate ≥30 times per minute; (2) oxygen saturation (resting state) ≤93%; (3) PaO2/FiO2 ≤ 300 mmHg. Otherwise, they were assigned to the non-severe group. Of the 67 patients, 17 were categorized with severe conditions and 50 had mild to moderate symptoms (referred to here as non-severe). The medical histories and the results of physical, haematological, biochemical, radiological and microbiological analyses were retrospectively evaluated and analysed. Peripheral blood of the recovered individuals was collected on day 28 after discharge, corresponding to 44–52 d after the onset of symptoms. PBMCs and serum were isolated and frozen in liquid nitrogen and in ultra-low temperature freezers, respectively.
End-point titre assay of serum antibodies
The end-point titre of serum antibodies was determined by measuring the binding activity of serum to the SARS-CoV-2 spike subunits S1 and S2 using an IgG-specific ELISA. In brief, 96-well plates (Corning) were coated with S1 or S2 at 200 ng per well in PBS at 4 °C overnight. The plates were washed five times with PBS-T (0.05% Tween-20 in PBS) and then blocked with blocking buffer (2% FBS and 2% BSA in PBS-T) for 30 min. Twofold serial dilutions of serum, starting from 1:20 dilution, were added to the 96-well plates in triplicate (100 µl per well) and incubated for 1 h at room temperature. Spike S1- and S2-specific antibodies were detected using horseradish peroxidase (HRP)-conjugated anti-human IgG and 3,3′,5,5′-tetramethylbenzidine substrate (Thermo Fisher Scientific). Sera from healthy individuals were used as negative controls, and monoclonal antibodies specific for the RBD of SARS-CoV-2 spike protein (anti-RBD/SARS-CoV-2; generated in the laboratory; unpublished data) was used as a positive control. Optical density at 450 nm (OD450) was measured for each reaction, and an OD450 of threefold above the cutoff-OD450 value was considered to be a positive readout. The cross-reactivity of the serum with subunits S1 and/or S2 of either SARS-CoV or MERS-CoV spike was examined using an optimized serum dilution of 1:1,000. All antigens were His-tagged (SARS-CoV-2 spike S1 and S2; SARS-CoV spike S1 and S2; MERS-CoV spike S1 and S2) and were purchased (Sino Biological).
Avidity assay of serum antibodies
The avidity of IgG antibodies specific to SARS-CoV-2 spike S1 and S2 was measured using a modified two-step approach that was described previously40,41. In the first step, serum dilutions were optimized to obtain an OD450 value within the range of 0.5–1.5, such that it ensured a linear measurement of the antibody avidity. The second step was an ELISA, but included an elution procedure of 1 M sodium thiocyanate (NaSCN). These measurements were performed in triplicate. The avidity index of an antibody was calculated as ODNaSCN 1 M/ODNaSCN 0 M × 100%.
Neutralization assay of serum antibodies
The neutralization activity of serum was determined by the decrease in luciferase expression after pseudotyped virus infection of Huh7 cells, as described previously for the neutralization assay of HIV pseudotyped virus20,21,42. The ID50 was defined as the serum dilution, at which the relative light unit (RLU) value was reduced by 50% compared with the control solution wells (virus + cells). The ID50 values were calculated using nonlinear regression, that is, log10-transformed inhibitor versus response (four parameters). In brief, SARS-CoV-2 pseudotyped virus was incubated in duplicate with the serial dilutions of sera samples (six dilutions: 1:30, 1:90, 1:270, 1:810, 1:2,430, 1:7,290) at 37 °C for 1 h. Freshly trypsinized cells were then added and incubated at 5% CO2 and 37 °C for 24 h, and the luminescence was measured. In parallel, control wells with only virus or cells were included in six replicates. The background RLU value (cells only) was subtracted from each determination. The cross-neutralization of serum to SARS-CoV- and MERS-CoV-pseudotyped viruses was performed using similar procedures as to SARS-CoV-2-pseudotyped viruses. Healthy control sera were used as negative controls. Sera from guinea pigs immunized with the spike protein from SARS-CoV-2, SARS-CoV or MERS-CoV were used as positive controls in the neutralization experiments for SARS-CoV-2, SARS-CoV or MERS-CoV, respectively. The cut-off value was defined as ID50 = 30, and ID50 > 30 was considered to have a neutralization effect. Neutralization titres were logarithmically transformed.
Autoantibody detection
Sixteen autoimmune antibodies were tested to assess whether the autoimmunity had occurred after recovery from COVID-19. Anti-double-stranded DNA and anti-ANA antibodies were detected using ELISA (Zeus Scientific); anti-nucleosome, anti-histone, anti-SmD1, anti-U1-SnRNP, anti-SS-A/Ro 60KD, anti-SS-A/Ro 52KD, anti-SS-B/La, anti-Sc1-70, anti-CENP-B, anti-Jo-1 and anti-PO/38KD antibodies were examined using Line Immuno Assay (LIA), according to the manufacturer’s protocols (HUMAN).
Flow cytometry
To analyse the circulating TFH cells, cryopreserved PBMCs were thawed in a 37 °C water bath and cultured immediately in RPMI 1640 medium supplemented with 10% FBS in 5% CO2 at 37 °C overnight. For cell-surface staining, 1 × 106 PBMCs were first labelled using the LIVE/DEAD Fixable Blue Dead Cell Stain Kit (Thermo Fisher Scientific) to distinguish live and dead cells, and then treated with Fc Block (BioLegend) to block non-specific binding. The treated PBMCs were stained with antibodies, which were pretitrated to an optimized dilution and fluorescently labelled, in 96-well V-bottom plates at 4 °C for 30 min. The fluorescently labelled antibodies used were as follows: BUV737 mouse anti-human CD4 (SK3) and PE mouse anti-human CXCR3 (1C6; BD Biosciences), FITC mouse anti-human PD-1 (EH12.2H7; BioLegend) and PE-eFluor 610 mouse anti-human CXCR5 (MU5UBEE; Thermo Fisher Scientific). Samples were loaded onto a MoFlo XDP Flow Cytometer (Beckman Coulter) immediately after antibody staining. The gating of cell populations was based on the mean fluorescence intensity minus one and unstained controls. All data were analysed using FlowJo v.10.0 (Tree Star).
Antigen-specific TFH-cell assay
A CD154 assay and a cytokine-independent activation-induced marker assay (AIMs, OX40/CD25) were used to assess spike-specific (S1 and S2) circulating-TFH-cell responses after stimulation43,44. In brief, cryopreserved PBMCs were thawed and kept resting in complete RPMI 1640 medium in 5% CO2 at 37 °C overnight. Cells (1 × 106) were stimulated with SARS-CoV-2 spike protein (S1 + S2, 5 μg ml−1) or BSA (5 μg ml−1, Sigma-Aldrich) in 5% CO2 at 37 °C for 24 h. Concanavalin A (5 μg ml−1, Sigma-Aldrich) was used as a CD154 assay positive control, and staphylococcal enterotoxin B (1 μg ml−1, Toxin Technology) was used as the OX40/CD25 assay positive control. PBMCs from healthy controls were also stimulated in the same conditions. Furthermore, to test the spike-specific cytokine secretion of TFH cells, a conventional intracellular cytokine staining assay was used45,46. Intracellular secretion of IL-21 and IFNγ was measured after antigen stimulation of TFH cells. In brief, PBMCs were stimulated with SARS-CoV-2 spike protein (S1 + S2, 5 μg ml−1) or BSA (5 μg ml−1) in the presence of anti-CD28 (1 μg ml−1) and anti-CD49d (1 μg ml−1) (BioLegend) antibodies for 6 h, and brefeldin A (1:1,000, BioLegend) was added 1 h after stimulation. Anti-CD3 antibodies (1 μg ml−1, BioLegend) were included as a positive control. TFH cells were gated as live CD4+PD-1+CXCR5+ T cells. PE mouse anti-human CD154 (24-31), PE-Cy7 mouse anti-human OX40 (ACT35), APC-mouse anti-human IL-21 (3A3-N2), PE-Cy7 mouse anti-human IFNγ (4S.B3) (BioLegend) and PE-Cy5 mouse anti-human CD25 (M-A251) (BD Biosciences) were purchased.
Statistical analysis and reproducibility
Baseline clinical characteristics were non-normally distributed, continuous variables were expressed as median ± IQR. Rank variables were expressed as constituent ratios. Mann–Whitney U-tests were used to analyse two independent variables, and paired t-tests were used to compare two paired samples. One-way analysis of variance was used to compare the differences between multiple groups, and Tukey’s multiple-comparisons test was used to compare within the groups. χ2 tests or Fisher’s exact tests were used to analyse different distributions between groups. Spearman’s rank correlation coefficient was used to measure the correlation between two different variables. A univariate and multivariable binary logistic regression model was used to rank the factors affecting the production of neutralization antibodies. OR values and 95% two-sided CIs were generalized by equation models to describe the factors contributing to antibody responses. Analyses of the data were performed using SPSS v.26 and GraphPad Prism v.8.0. Unless otherwise stated, all numeral data shown in this study were collected from at least three independent experiments. Thus, the results shown are representative of the biological replicates.
Ethics approval
This study was performed in accordance with the Good Clinical Practice and the Declaration of Helsinki principles for ethical research. The study protocol was approved by the Institutional Review Board of The Center Hospital of Shaoyang (V.1.0, 20200301), Hunan Province, China. Each participant provided informed consent by signing a written consent form. Medical data were collected from electronic records of the hospitals using standardized Data Collection Forms recommended by the International Severe Acute Respiratory and Emerging Infection Consortium.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Source data are provided with this paper and are available via Figshare (https://doi.org/10.6084/m9.figshare.13139423.v2). Any other data that support the findings of this study are available from the corresponding authors on request.
References
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 382, 727–733 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Guan, W. J. et al. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 382, 1708–1720 (2020).
Hu, Z. et al. Clinical characteristics of 24 asymptomatic infections with COVID-19 screened among close contacts in Nanjing, China. Sci. China Life Sci. 63, 706–711 (2020).
Wang, F. et al. Characteristics of peripheral lymphocyte subset alteration in COVID-19 pneumonia. J. Infect. Dis. 221, 1762–1769 (2020).
Chen, G. et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J. Clin. Invest. 130, 2620–2629 (2020).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 26, 1033–1036 (2020).
Chen, X. et al. Human monoclonal antibodies block the binding of SARS-CoV-2 spike protein to angiotensin converting enzyme 2 receptor. Cell Mol. Immunol. 17, 647–649 (2020).
Qin, C. et al. Dysregulation of immune response in patients with coronavirus 2019 (COVID-19) in Wuhan, China. Clin. Infect. Dis. 71, 762–768 (2020).
Wang, F. et al. The laboratory tests and host immunity of COVID-19 patients with different severity of illness. JCI Insight 5, e137799 (2020).
Guo, L. et al. Profiling early humoral response to diagnose novel coronavirus disease (COVID-19). Clin. Infect. Dis. 71, 778–785 (2020).
Zhao, J. et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa344 (2020).
Long, Q. X. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 26, 845–848 (2020).
Ni, L. et al. Detection of SARS-CoV-2-specific humoral and cellular immunity in COVID-19 convalescent individuals. Immunity 52, 971–977 (2020).
Anderson, D. E. et al. Lack of cross-neutralization by SARS patient sera towards SARS-CoV-2. Emerg. Microbes Infect. 9, 900–902 (2020).
Duan, K. et al. Effectiveness of convalescent plasma therapy in severe COVID-19 patients. Proc. Natl Acad. Sci. USA 117, 9490–9496 (2020).
Shen, C. et al. Treatment of 5 critically Ill patients with COVID-19 with convalescent plasma. JAMA 323, 1582–1589 (2020).
Chan, K. H. et al. Cross-reactive antibodies in convalescent SARS patients’ sera against the emerging novel human coronavirus EMC (2012) by both immunofluorescent and neutralizing antibody tests. J. Infect. 67, 130–140 (2013).
Nie, J. et al. Establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2. Emerg. Microbes Infect. 9, 680–686 (2020).
Ma, J. et al. Development of SARS and MERS neutralization methods based on pseudoviruses. Chin. J. Virol. 35, 189–195 (2019).
Breitfeld, D. et al. Follicular B helper T cells express CXC chemokine receptor 5, localize to B cell follicles, and support immunoglobulin production. J. Exp. Med. 192, 1545–1552 (2000).
Schaerli, P. et al. CXC chemokine receptor 5 expression defines follicular homing T cells with B cell helper function. J. Exp. Med. 192, 1553–1562 (2000).
Morita, R. et al. Human blood CXCR5+CD4+ T cells are counterparts of T follicular cells and contain specific subsets that differentially support antibody secretion. Immunity 34, 108–121 (2011).
Nurieva, R. I. et al. Generation of T follicular helper cells is mediated by interleukin-21 but independent of T helper 1, 2, or 17 cell lineages. Immunity 29, 138–149 (2008).
Vogelzang, A. et al. A fundamental role for interleukin-21 in the generation of T follicular helper cells. Immunity 29, 127–137 (2008).
Grifoni, A. et al. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host Microbe 27, 671–680 (2020).
Okba, N. M. A. et al. Severe acute respiratory syndrome coronavirus 2-specific antibody responses in coronavirus disease patients. Emerg. Infect. Dis. 26, 1478–1488 (2020).
Wang, Y. et al. Kinetics of viral load and antibody response in relation to COVID-19 severity. J. Clin. Invest. 130, 5235–5244 (2020).
Wang, C. et al. A human monoclonal antibody blocking SARS-CoV-2 infection. Nat. Commun. 11, 2251 (2020).
Tian, X. et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 9, 382–385 (2020).
Pinto, D. et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 583, 290–295 (2020).
Zhao, J. et al. Recovery from the Middle East respiratory syndrome is associated with antibody and T-cell responses. Sci. Immunol. 2, eaan5393 (2017).
Alshukairi, A. N. et al. Antibody response and disease severity in healthcare worker MERS survivors. Emerg. Infect. Dis. 22, 1113–1115 (2016).
Zhang, J. et al. Circulating CXCR3+ Tfh cells positively correlate with neutralizing antibody responses in HCV-infected patients. Sci. Rep. 9, 10090 (2019).
Bentebibel, S. E. et al. Induction of ICOS+CXCR3+CXCR5+ TH cells correlates with antibody responses to influenza vaccination. Sci. Transl. Med. 5, 176ra132 (2013).
Baiyegunhi, O. et al. Frequencies of Circulating Th1-Biased T Follicular Helper Cells in Acute HIV-1 Infection Correlate with the Development of HIV-Specific Antibody Responses and Lower Set Point Viral Load. J. Virol. 92, e00659-618 (2018).
Thevarajan, I. et al. Breadth of concomitant immune responses prior to patient recovery: a case report of non-severe COVID-19. Nat. Med. 26, 453–455 (2020).
Juno, J. A. et al. Humoral and circulating follicular helper T cell responses in recovered patients with COVID-19. Nat. Med. 26, 1428–1434 (2020).
Welten, S. P. M., Redeker, A., Toes, R. E. M. & Arens, R. Viral persistence induces antibody inflation without altering antibody avidity. J. Virol. 90, 4402–4411 (2016).
Ray, R. et al. Characterization of antibodies induced by vaccination with hepatitis C virus envelope glycoproteins. J. Infect. Dis. 202, 862–866 (2010).
Chen, Q. et al. Development and optimization of a sensitive pseudovirus-based assay for HIV-1 neutralizing antibodies detection using A3R5 cells. Hum. Vaccin. Immunother. 14, 199–208 (2018).
Dan, J. M. et al. A cytokine-independent approach to identify antigen-specific human germinal center T follicular helper cells and rare antigen-specific CD4+ T cells in blood. J. Immunol. 197, 983–993 (2016).
Bentebibel, S. E. et al. ICOS+PD-1+CXCR3+ T follicular helper cells contribute to the generation of high-avidity antibodies following influenza vaccination. Sci. Rep. 6, 26494 (2016).
Gauduin, M. C. Intracellular cytokine staining for the characterization and quantitation of antigen-specific T lymphocyte responses. Methods 38, 263–273 (2006).
Gauduin, M. C. et al. Optimization of intracellular cytokine staining for the quantitation of antigen-specific CD4+ T cell responses in rhesus macaques. J. Immunol. Methods 288, 61–79 (2004).
Acknowledgements
We thank all of the participants. This work was supported by the COVID-19 Emergency Special Program of Hunan Province (no. 2020SK3052, to W. Liu), Key Research and Development Project of Chenzhou City, Hunan Province (no. ZDYF2020010 and ZDYF2020007) and by the SC1-PHE-CORONAVIRUS-2020: Advancing knowledge for the clinical and public health response to the 2019-nCoV epidemic’ from the European Commission (CORONADX, no. 101003562, to Y.-P.L.).
Author information
Authors and Affiliations
Contributions
X. Qu, W. Liu, Y.-P.L. and Y.W. contributed to the study design and data interpretation. Q. Wang, T.X., M.H., D.P., G.L., X.X., S.H., Y.D., J.L., W. Li, Z. Lu, H.C., T.Z. and Q.L. contributed to clinical management, patient recruitment and data collection. J.Z., Q. Wu, Z. Liu, J.W., Y.H., T.B., T.W., W.H., K.J., L.N., W.G., D. Luo and Y.-P.L. contributed to sample processing, assay development and performing experiments. J.Z., Q. Wu, Z. Liu, T.W., D. Lei and R.H. contributed to statistical analysis and data visualization. X. Qu, W. Liu, Y.-P.L. and J.Z. drafted the manuscript. Y.W., X. Qu and Y.-P.L. contributed to revision of the manuscript for important intellectual content. X. Qu and W. Liu provided supervision. All of the authors met authorship criteria and approved the publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Microbiology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Baseline characteristics of COVID-19 patients included in this study.
Data of continuous variable are expressed as median (Interquartile range, IQR), rank variables are expressed as positive cases (percentage). Mann-Whitney U test was used to continuous variable companion analysis, χ² or Fisher’s exact tests was used to rank variables data analysis. *p < 0.05 was considered to be a two-tailed significant difference between severe and non-severe groups.
Extended Data Fig. 2 Laboratory findings of COVID-19 patients on admission.
Data were collected on admission and presented as median, IQR (25%-75%). Mann-Whitney U test was used to analysis of difference between two groups. *p < 0.05 was considered to be a two-tailed significant difference.
Extended Data Fig. 3 Reactivity of anti-SARS-CoV-2 antibody to the spike subunits S1 and S2 and neutralization to SARS-CoV-2, SARS-CoV, and MERS-CoV viruses.
Antibody binding titers to S1 and S2 of SARS-CoV-2 spike were performed by endpoint dilution assay and expressed as endpoint dilution titers. The avidity of antibody is expressed as avidity index (%). Cross-binding to spike S1 and S2 from both SARS-CoV and MERS-CoV were expressed as (+) (with binding) and (-) (without binding). Neutralization to SARS-CoV-2 and cross-neutralization to SARS-CoV or MERS-CoV were expressed as ID50, and ID50 > 30 was defined as positive, according to the cutoff value of neutralization assay. ND, not detectable; NA, no applicable. See Methods for the details of each assay.
Extended Data Fig. 4 Correlation analysis of the neutralization titers and the endpoint titers of anti-S1 and anti-S2 antibodies.
Correlation analysis of endpoint titers of anti-S1 and anti-S2 antibodies (a) (n = 67), neutralization titers and anti-S1 titers (b) (n = 65), and neutralization titers and anti-S2 titers (c) (n = 65). Endpoint titers and neutralization titers are log10-transformed. Spearman’s rank coefficient of correlation was used for the correlation analysis, and p < 0.05 was considered to be significant difference.
Extended Data Fig. 5 Binding analysis of healthy control sera against spike S1 and S2 from SARS-CoV-2, SARS-CoV, and MERS-CoV.
Sixty-one healthy control sera were examined for its binding to the spike S1 and S2 from SARS-CoV-2, SARS-CoV, and MERS-CoV. The binding activity is expressed as (+) (with binding) and (-) (without binding). See Methods for the details of the assay.
Extended Data Fig. 6 Analysis of neutralization titers for the sera with cross-neutralization activity.
a, Neutralization titers to SARS-CoV-2 pseudotyped virus for the sera with (n = 56) and without (n = 11) cross-neutralization activity to SARS-CoV and/or MERS-CoV. b, Comparison of neutralization titers to SARS-CoV-2 (n = 11) for the sera without cross-neutralization and with cross-neutralization to SARS-CoV (n = 44), MERS-CoV (n = 5), and both viruses (n = 7). c-e, Comparison of neutralization titer (specific and cross neutralization) for each serum that shows cross-neutralization activity. c, Neutralization titers to SARS-CoV and SARS-CoV-2 for those sera cross-neutralizing SARS-CoV (n = 44). d, Neutralization titers to MERS-CoV and SARS-CoV-2 for those sera cross-neutralizing MERS-CoV (n = 5). e, Neutralization titers to SARS-CoV, MERS-CoV, and SARS-CoV-2 for those sera cross-neutralizing both SARS-CoV and MERS-CoV (n = 7). Neutralization titers are log10-transformed. In a and b, data presented as median, interquartile range (25%-75%). Mann-Whitney U test is used to compare the difference between two groups (a). One-way ANOVA is used to compare the difference among multi-groups, and Tukey’s multiple comparisons test is used for intra-groups (b). In c-e, paired t test is used to analysis the difference between groups. p < 0.05 was considered to be significant difference between the groups.
Extended Data Fig. 7 Analysis of autoimmune antibodies in recovered COVID-19 individuals.
ANA: Antinuclear antibodies; Anti-ds-DNA: Anti-double-stranded DNA; SmD1: Core small nuclear ribonucleoprotein particle splicing factor; SS-A/Ro: Anti-Sjogren’s Syndrome antigen B antibody; SS-B/La: Anti-Sjogren’s Syndrome B antibody; Scl-70: Autoantibodies to topoisomerase I; CENP-B: The major human centromere autoantigen; Jo-1: Histidyl-transfer RNA synthetase; Anti-P0: Anti-P antibodies react against acidic phosphorylated ribosomal proteins P0, P1, and P2 (with molecular mass of 38, 19, and 17 kDa, respectively). ANA titer≥20 and Anti-ds-DNA ≥ 180 are defined as positive (+) or less are negative (-).
Extended Data Fig. 8 The gating strategy and the frequency of circulating Tfh cell subsets in recovered COVID-19 individuals.
a, Representative gating strategy for circulating Tfh (PD-1+CXCR5+ CD4+ T) cells, CXCR3+ and CXCR3− Tfh cells. Gating of cell populations is based on the mean fluorescence intensity ‘minus one’ (FMO). b–e, Comparison of between healthy controls (HC) (n = 17) percentage of Tfh cells (b), CXCR3+ Tfh cells (c), CXCR3− Tfh cells (d), and the ratio of CXCR3+/CXCR3− Tfh cells (e) and recovered COVID-19 individuals (n = 26). In b-e, data presented as median, interquartile range (25%-75%). Mann-Whitney U test was used to compare the difference between two groups, and p < 0.05 was considered to be a two-tailed significant difference.
Extended Data Fig. 9 SARS-CoV-2 spike-specific Tfh cell response after antigen stimulation.
Representative flow plots of the antigen specific Tfh cells after stimulation. To determine the expression of CD154 (a) and OX40 and CD25 (b), PBMCs from COVID-19 individuals and healthy controls were stimulated with SARS-CoV-2 spike subunits S1 and S2, or BSA. Con A and SEB were used as positive controls. After stimulation, surface CD154 and both OX40 and CD25 were measured by flow cytometry gating on PD-1+ CXCR5+ CD4+ T cells. For intracellular cytokine secretion, PBMCs from COVID-19 individuals and healthy controls were stimulated with SARS-CoV-2 spike subunits S1 and S2, or BSA, in the presence of anti-CD28, anti-CD49d. Anti-CD3 was as positive control. After stimulation, intracellular IL-21 (c) and IFN-γ (d) were measured by flow cytometry gating on PD-1+ CXCR5+ CD4+ T cells. The gating of cell populations was based on the mean fluorescence intensity ‘minus one’ (FMO).
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Zhang, J., Wu, Q., Liu, Z. et al. Spike-specific circulating T follicular helper cell and cross-neutralizing antibody responses in COVID-19-convalescent individuals. Nat Microbiol 6, 51–58 (2021). https://doi.org/10.1038/s41564-020-00824-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-020-00824-5
This article is cited by
-
A chimeric porcine reproductive and respiratory syndrome virus 1 strain containing synthetic ORF2-6 genes can trigger T follicular helper cell and heterologous neutralizing antibody responses and confer enhanced cross-protection
Veterinary Research (2024)
-
Neutralizing immunity against coronaviruses in Tanzanian health care workers
Scientific Reports (2024)
-
Humoral immunity for durable control of SARS-CoV-2 and its variants
Inflammation and Regeneration (2023)
-
The immune response to SARS-CoV-2 in people with HIV
Cellular & Molecular Immunology (2023)
-
B lymphocytes in COVID-19: a tale of harmony and discordance
Archives of Virology (2023)