Abstract
Amplicon sequencing (for example, of the 16S rRNA gene) identifies the presence and relative abundance of microbial community members. However, metagenomic sequencing is needed to identify the genetic content and functional potential of a community. Metagenomics is challenging in samples dominated by host DNA, such as those from the skin, tissue and respiratory tract. Here, we combine advances in amplicon and metagenomic sequencing with culture-enriched molecular profiling to study the human microbiota. Using the cystic fibrosis lung as an example, we cultured an average of 82.13% of the operational taxonomic units representing 99.3% of the relative abundance identified in direct sequencing of sputum samples; importantly, culture enrichment identified 63.3% more operational taxonomic units than direct sequencing. We developed the PLate Coverage Algorithm (PLCA) to determine a representative subset of culture plates on which to conduct culture-enriched metagenomics, resulting in the recovery of greater taxonomic diversity—including of low-abundance taxa—with better metagenome-assembled genomes, longer contigs and better functional annotations when compared to culture-independent methods. The PLCA is also applied as a proof of principle to a previously published gut microbiota dataset. Culture-enriched molecular profiling can be used to better understand the role of the human microbiota in health and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing results are publicly available (BioProject ID PRJNA503799). The PLCA algorithm is available from https://github.com/fwhelan/PLCA.
Code availability
All code developed by the authors is available under a GNU licence at http://github.com/fwhelan/PLCA and https://github.com/shekas3/BinTaxaAssigner.
References
Van Leeuwenhoek, A. Microscopical observations about animals in the scurf of the teeth. Philos. Trans. R Soc. Lond. B Biol. Sci. 14, 568–574 (1683).
Turnbaugh, P. J. et al. The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 1, 6ra14 (2009).
Huttenhower, C. et al. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Spor, A., Koren, O. & Ley, R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nat. Rev. Microbiol. 9, 279–290 (2011).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Olesen, S. W. & Alm, E. J. Dysbiosis is not an answer. Nat. Microbiol. 1, 16228 (2016).
Shade, A. Diversity is the question, not the answer. ISME J. 11, 1–6 (2017).
Finegold, S. M., Attebery, H. R. & Sutter, V. L. Effect of diet on human fecal flora: comparison of Japanese and American diets. Am. J. Clin. Nutr. 27, 1456–1469 (1974).
Goodman, A. L. et al. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl Acad. Sci. USA 108, 6252–6257 (2011).
Lagier, J.-C. et al. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin. Microbiol. Infect. 18, 1185–1193 (2012).
Rettedal, E. A., Gumpert, H. & Sommer, M. O. A. Cultivation based multiplex phenotyping of human gut microbiota allows targeted recovery of previously uncultured bacteria. Nat. Commun. 5, 4714 (2014).
Lau, J. T. et al. Capturing the diversity of the human gut microbiota through culture-enriched molecular profiling. Genome Med. 8, 72 (2016).
Hilt, E. E. et al. Urine is not sterile: use of enhanced urine culture techniques to detect resident bacterial flora in the adult female bladder. J. Clin. Microbiol. 52, 871–876 (2014).
Myles, I. A. et al. A method for culturing Gram-negative skin microbiota. BMC Microbiol. 16, 60 (2016).
Thompson, H., Rybalka, A., Moazzez, R., Dewhirst, F. E. & Wade, W. G. In-vitro culture of previously uncultured oral bacterial phylotypes. Appl. Environ. Microbiol. 81, 8307–8314 (2015).
Sibley, C. D. et al. Culture enriched molecular profiling of the cystic fibrosis airway microbiome. PLoS ONE 6, e22702 (2011).
Oh, J. et al. Biogeography and individuality shape function in the human skin metagenome. Nature 514, 59–64 (2014).
Wang, W.-L. et al. Application of metagenomics in the human gut microbiome. World J. Gastroenterol. 21, 803–814 (2015).
Zhang, C. et al. Identification of low abundance microbiome in clinical samples using whole genome sequencing. Genome Biol. 16, 265 (2015).
Lim, Y. W. et al. Metagenomics and metatranscriptomics: windows on CF-associated viral and microbial communities. J. Cyst. Fibros. 12, 154–164 (2013).
Huang, Y. J. & LiPuma, J. J. The microbiome in cystic fibrosis. Clin. Chest Med. 37, 59–67 (2015).
Zhao, J. et al. Decade-long bacterial community dynamics in cystic fibrosis airways. Proc. Natl Acad. Sci. USA 109, 5809–5814 (2012).
Whelan, F. J. et al. Longitudinal sampling of the lung microbiota in individuals with cystic fibrosis. PLoS ONE 12, e0172811 (2017).
Lagier, J.-C. et al. Culture of previously uncultured members of the human gut microbiota by culturomics. Nat. Microbiol. 1, 16203 (2016).
Surette, M. G. The cystic fibrosis lung microbiome. Ann. Am. Thorac. Soc. 11(Suppl. 1), S61–S65 (2014).
Field, T. R., Sibley, C. D., Parkins, M. D., Rabin, H. R. & Surette, M. G. The genus Prevotella in cystic fibrosis airways. Anaerobe 16, 337–344 (2010).
van der Gast, C. J. et al. Partitioning core and satellite taxa from within cystic fibrosis lung bacterial communities. ISME J. 5, 780–791 (2011).
Tunney, M. M. et al. Detection of anaerobic bacteria in high numbers in sputum from patients with cystic fibrosis. Am. J. Respir. Crit. Care Med. 177, 995–1001 (2008).
Parkins, M. D. & Floto, R. A. Emerging bacterial pathogens and changing concepts of bacterial pathogenesis in cystic fibrosis. J. Cyst. Fibros. 14, 293–304 (2015).
Pop, M. et al. Individual-specific changes in the human gut microbiota after challenge with enterotoxigenic Escherichia coli and subsequent ciprofloxacin treatment. BMC Genomics 17, 440 (2016).
Coleman, F. T. et al. Hypersusceptibility of cystic fibrosis mice to chronic Pseudomonas aeruginosa oropharyngeal colonization and lung infection. Proc. Natl Acad. Sci. USA 100, 1949–1954 (2003).
Jorth, P. et al. Regional isolation drives bacterial diversification within cystic fibrosis lungs. Cell Host Microbe 18, 307–319 (2015).
Lieberman, T. D. et al. Genetic variation of a bacterial pathogen within individuals with cystic fibrosis provides a record of selective pressures. Nat. Genet. 46, 82–87 (2014).
Pompilio, A. et al. Stenotrophomonas maltophilia phenotypic and genotypic diversity during a 10-year colonization in the lungs of a cystic fibrosis patient. Front. Microbiol. 7, 1551 (2016).
Ferretti, P. et al. Mother-to-infant microbial transmission from different body sites shapes the developing infant gut microbiome. Cell Host Microbe 24, 133–145 (2018).
Li, S. S. et al. Durable coexistence of donor and recipient strains after fecal microbiota transplantation. Science 352, 586–589 (2016).
Nicholls, S. M. et al. Probabilistic recovery of cryptic haplotypes from metagenomic data. Preprint at https://www.biorxiv.org/content/10.1101/117838v1 (2017).
Creevey, C. J., Doerks, T., Fitzpatrick, D. A., Raes, J. & Bork, P. Universally distributed single-copy genes indicate a constant rate of horizontal transfer. PLoS ONE 6, e22099 (2011).
Frank, D. N. et al. Molecular–phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007).
Collins, S. M. A role for the gut microbiota in IBS. Nat. Rev. Gastroenterol. Hepatol. 11, 497–505 (2014).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Gohir, W., Whelan, F. J., Surette, M. G., Moore, C. & Jonathan, D. Pregnancy-related changes in the maternal gut microbiota are dependent upon the mother’s periconceptional diet. Gut Microbes 6, 310–320 (2015).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1095 (2015).
Naseribafrouei, A. et al. Correlation between the human fecal microbiota and depression. Neurogastroenterol. Motil. 26, 1155–1162 (2014).
Grice, E. A. & Segre, J. A. The skin microbiome. Nat. Rev. Microbiol. 9, 244–253 (2011).
Wang, J. et al. Metagenomic sequencing reveals microbiota and its functional potential associated with periodontal disease. Sci. Rep. 3, 1843 (2013).
Dickson, R. P., Erb-Downward, J. R., Martinez, F. J. & Huffnagle, G. B. The microbiome and the respiratory tract. Annu. Rev. Physiol. 78, 481–504 (2015).
Chi, B., Chauhan, S. & Kuramitsu, H. Development of a system for expressing heterologous genes in the oral spirochete treponema denticola and its use in expression of the treponema pallidum flaA gene. Infect. Immun. 67, 3653–3656 (1999).
Camanocha, A. & Dewhirst, F. E. Host-associated bacterial taxa from Chlorobi, Chloroflexi, GN02, Synergistetes, SR1, TM7 and WPS-2 phyla/candidate divisions. J. Oral Microbiol. 6, 25468 (2014).
Marcy, Y. et al. Dissecting biological ‘dark matter’ with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc. Natl Acad. Sci. USA 104, 11889–11894 (2007).
Meyer, K. C., Sharma, A., Rosenthal, N. S., Peterson, K. & Brennan, L. Regional variability of lung inflammation in cystic fibrosis. Am. J. Respir. Crit. Care Med. 156, 1536–1540 (1997).
Stressmann, F. A. et al. Does bacterial density in cystic fibrosis sputum increase prior to pulmonary exacerbation? J. Cyst. Fibros. 10, 357–365 (2011).
Quigley, E. M. Gut bacteria in health and disease. Gastroenterol. Hepatol. 9, 560–569 (2013).
Whelan, F. J. & Surette, M. G. A comprehensive evaluation of the sl1p pipeline for 16S rRNA gene sequencing analysis. Microbiome 5, 100 (2017).
Fuchs, H. J. et al. Effect of aerosolized recombinant human DNase on exacerbations of respiratory symptoms and on pulmonary function in patients with cystic fibrosis. The Pulmozyme Study Group. N. Engl. J. Med. 331, 637–642 (1994).
Sibley, C. D. et al. McKay agar enables routine quantification of the ‘Streptococcus milleri’ group in cystic fibrosis patients. J. Med. Microbiol. 59, 534–540 (2010).
Whelan, F. J., Rossi, L., Stearns, J. C. & Surette, M. G. Culture and molecular profiling of the respiratory tract microbiota. Methods Mol. Biol. 1894, 49–61 (2018).
Whelan, F. J. et al. The loss of topography in the microbial communities of the upper respiratory tract in the elderly. Ann. Am. Thorac. Soc. 11, 513–521 (2014).
Bartram, A. K., Lynch, M. D. J., Stearns, J. C., Moreno-Hagelsieb, G. & Neufeld, J. D. Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired end illumina reads. Appl. Environ. Microbiol. 77, 3846–3852 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G. & Neufeld, J. D. PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics 13, 31 (2012).
Ye, Y. Identification and quantification of abundant species from pyrosequences of 16S rRNA by consensus alignment. In Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine 153–157 (IEEE, 2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10, e1003531 (2014).
Asnicar, F., Weingart, G., Tickle, T. L., Huttenhower, C. & Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 3, e1029 (2015).
Pheatmap: pretty heatmaps. R Package v.1.0.12 (CRAN, 2012).
Denton, M., Hall, M., Todd, N., Kerr, K. & Littlewood, J. Improved isolation of Stenotrophomonas maltophilia from the sputa of patients with cystic fibrosis using a selective medium. Clin. Microbiol. Infect. 6, 395–396 (2000).
Schmieder, R. & Edwards, R. Fast identification and removal of sequence contamination from genomic and metagenomic datasets. PLoS ONE 6, e17288 (2011).
Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Wu, Y.-W., Tang, Y.-H., Tringe, S. G., Simmons, B. A. & Singer, S. W. MaxBin: an automated binning method to recover individual genomes from metagenomes using an expectation-maximization algorithm. Microbiome 2, 26 (2014).
Sczyrba, A. et al. Critical assessment of metagenome interpretation—a benchmark of metagenomics software. Nat. Methods 14, 1063–1071 (2017).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells and metagenomes. Genome Res. 25, 1043–1055 (2015).
Breitwieser, F. P. & Salzberg, S. L. KrakenHLL: confident and fast metagenomics classification using unique k-mer counts. Genome Biol. 19, 198 (2018).
Lee, S. T. M. et al. Tracking microbial colonization in fecal microbiota transplantation experiments via genome-resolved metagenomics. Microbiome 5, 50 (2017).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Mao, C. et al. Curation, integration and visualization of bacterial virulence factors in PATRIC. Bioinformatics 31, 252–258 (2015).
Huerta-Cepas, J. et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol. Biol. Evol. 34, 2115–2122 (2017).
Huerta-Cepas, J. et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 44, D286–D293 (2016).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Jia, B. et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 45, D566–D573 (2017).
Skinnider, M. A. et al. Genomes to natural products PRediction Informatics for Secondary Metabolomes (PRISM). Nucleic Acids Res. 43, 9645–9662 (2015).
Bland, C. et al. CRISPR Recognition Tool (CRT): a tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinformatics 8, 209 (2007).
Acknowledgements
This research was funded in part by a Canadian Institutes of Health Research (CIHR) Doctoral Scholarship, a Cystic Fibrosis Canada (CFC) Studentship and a Marie Skłodowska-Curie Individual Fellowship (GA no. 793818) awarded to F.J.W., and grants from CIHR, CFC and a Tier 1 Canada Research Chair to M.G.S. We thank the patients and healthcare professionals at the Calgary Adult Cystic Fibrosis Clinic for their participation and assistance with this study. We acknowledge critical intellectual conversations with J.T. Lau and J.C. Szamosi. We wish to acknowledge that this research was conducted on traditional territory shared by the Haudenosaunee confederacy and the Anishinaabe nations as well as the peoples of the Treaty 7 region in Southern Alberta.
Author information
Authors and Affiliations
Contributions
F.J.W. is the primary author of this prepared manuscript. H.R.R. and M.D.P. collected patient information and enrolled willing participants for this study. B.W. collected, processed and cultured all sputum samples in addition to all biological and technical controls. F.J.W. and S.A.S. isolated DNA from culture/sputum material, and ran PCR reactions to amplify the 16S rRNA gene variable 3 region. S.A.S. performed the enrichment of Stenotrophomonas. F.J.W. prepared DNA for metagenomic sequencing. F.J.W. processed and analysed all 16S rRNA gene and metagenomic sequencing results. S.S. provided code for the taxonomic assignment of metagenomic bins. F.J.W., M.D.P. and M.G.S. conceptualized the experimental outline. F.J.W. conducted all data analyses and wrote the manuscript. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Media used for culture enrichment.
Extended Data Fig. 2 Using more stringent abundance thresholds, the vast majority of the cystic fibrosis lung microbiota is still captured by culture-enriched methods. Culture-enriched 16S rRNA gene sequencing increases OTU recovery.
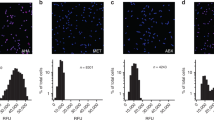
Using a cutoff of >= 0.1% relative abundance, 65.6% of the lung microbiota was identified only by culture across samples (a), with only 4.3% of OTUs not being cultured (b). c. In each sample, more OTUs were recovered via culture-enriched sequencing than by direct sequencing. d. Overall, an average of 65.5% of all OTUs were identified only via culture. e. The α-diversity of the original samples subject to culture enrichment.
Extended Data Fig. 3 Culture enriches for low abundance taxa.
The first sample in our dataset was re-sequenced to a depth of 972,834 reads. A rank order curve, and associated quantification of the recovered OTUs (inset), illustrates the number of OTUs present in the original (41,199) and re-sequenced sample as well as at rarefied depths, indicating the increased recovery of cultured OTUs by direct sequencing as sequencing depth increases. The inset shows, at two different read cutoffs, that with increasing sequencing depth, the number of OTUs seen only in culture decreases.
Extended Data Fig. 4 No specific culture condition consistently recapitulates the originating sputum sample.
Principle coordinate analyses based on the Bray Curtis β-diversity metric indicates that, across all 20 samples, no culture condition consistently recapitulates the microbial community of the originating sputum sample. For a list of media abbreviations see Extended Data Table 1; Aer = aerobic, Ana = anaerobic.
Extended Data Fig. 5 The importance of selective and non-selective culture conditions in capturing the genus-level diversity of the cystic fibrosis microbiome.
This Figure is a labelled version of Fig. 3c.
Extended Data Fig. 6 The importance of selective and non-selective culture conditions in capturing the OTU-level diversity of the cystic fibrosis microbiome.
A heatmap indicates the breadth of culture conditions necessary to culture this community at the OTU level.
Extended Data Fig. 7
The optimal plate sets needed to sequence all samples (S) in this dataset, and the number of OTUs which would be obtained.
Extended Data Fig. 8 Culture-enriched metagenomic sequencing finds similar communities to 16S rRNA gene sequencing.
Comparisons of the bacterial composition of 16S rRNA gene sequencing to the 16S rRNA gene sequences obtained via whole-genome metagenomics reveal similar communities in the culture conditions amplified as part of the de novo PLCA (a-b) and adjusted PLCA plate sets. (c-d), Communities are compared visually using taxonomic summaries (a,c) and quantitatively using PCoA of the Bray Curtis β-diversity metric (b,d). 16S=16S rRNA gene sequencing; MG=shotgun metagenomic sequencing.
Extended Data Fig. 9 The PLCA consistently recovers targeted OTUs and is not specific to the cystic fibrosis lung microbiota.
a, When the adjusted PLCA was applied to the 20 samples in this cystic fibrosis dataset, it consistently recovered the targeted OTUs (orange), though some (gray) were not recovered as metagenomic bins due to inadequate sequencing depth, or the inability to separate species into separate bins. The overlaying numbers represent the number of OTUs in each category. Additional species obtained as a consequence of being present on a plate with a targeted OTU are shown as yellow dots. b, The PLCA is not specific to the cystic fibrosis lung microbiota or to a particular set of culture conditions; here, we apply the PLCA to previously published culture-enriched gut microbiota data (reference 12). Even through the culture conditions used by Lau et al. differ from those used in this study, the PLCA still predicts successful recovery of almost all species at abundances above the PLCA thresholds (dotted line).
Extended Data Fig. 10 Biological (a-b) and technical (c-d) replicates of sputum and culture-enriched 16S rRNA gene sequencing.
PCoA plots of the Bray Curtis distance between sputum profiles show close clustering of 3 sputum biological replicates prior to plating, n=6 (a), and cultured replicates (technical replicates after plating of 3 sputum samples x 2 culture replicates of 6 representative media types, n=108, c). Polygons/lines connect the replicates in each plot; specifically between duplicate sputum sampling of 3 individuals (labelled A-C) in a and between triplicate platings of sputums in c. In c, the legend labels refer to the media type (for example BHI), environmental condition (Aer, aerobic or Ana, anaerobic), and replicate number (1–6). Taxonomic summaries of biological samples prior to plating (b) and 3 technical replicates each of 2 sputum samples from 3 patients (d) show the consistency of these techniques. Labelling of biological (A-C) and technical (1–6) replicates is consistent with parts a and c. In cases of no visible growth (that is MAC_Aer_3 and MAC_Aer_4) no samples were collected. In some cases, there was some difference in abundance of organisms between replicates (for example KVLB 5,6 and McKay 5,6) on which there were low bacterial counts and only at the lowest dilution plated (the cultured organisms representing ~104 CFU/ml and less than 1% in the original sputum sample). These will be subject to more variability in plating.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Supplementary Tables 1–7.
Source data
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 9
Statistical Source Data.
Rights and permissions
About this article
Cite this article
Whelan, F.J., Waddell, B., Syed, S.A. et al. Culture-enriched metagenomic sequencing enables in-depth profiling of the cystic fibrosis lung microbiota. Nat Microbiol 5, 379–390 (2020). https://doi.org/10.1038/s41564-019-0643-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0643-y
This article is cited by
-
Capturing the microbial dark matter in desert soils using culturomics-based metagenomics and high-resolution analysis
npj Biofilms and Microbiomes (2023)
-
Microbiota in mesenteric adipose tissue from Crohn’s disease promote colitis in mice
Microbiome (2021)
-
A prevalent and culturable microbiota links ecological balance to clinical stability of the human lung after transplantation
Nature Communications (2021)
-
A complete and flexible workflow for metaproteomics data analysis based on MetaProteomeAnalyzer and Prophane
Nature Protocols (2020)