Abstract
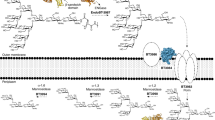
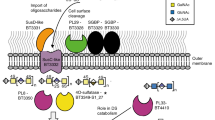
Glycans are the major carbon sources available to the human colonic microbiota. Numerous N-glycosylated proteins are found in the human gut, from both dietary and host sources, including immunoglobulins such as IgA that are secreted into the intestine at high levels. Here, we show that many mutualistic gut Bacteroides spp. have the capacity to utilize complex N-glycans (CNGs) as nutrients, including those from immunoglobulins. Detailed mechanistic studies using transcriptomic, biochemical, structural and genetic techniques reveal the pathway employed by Bacteroides thetaiotaomicron (Bt) for CNG degradation. The breakdown process involves an extensive enzymatic apparatus encoded by multiple non-adjacent loci and comprises 19 different carbohydrate-active enzymes from different families, including a CNG-specific endo-glycosidase activity. Furthermore, CNG degradation involves the activity of carbohydrate-active enzymes that have previously been implicated in the degradation of other classes of glycan. This complex and diverse apparatus provides Bt with the capacity to access the myriad different structural variants of CNGs likely to be found in the intestinal niche.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The full RNA-Seq data are provided in Supplementary Table 1 and have also been submitted to https://www.ncbi.nlm.nih.gov/geo/ with the accession number GSE129572. The crystal structures are deposited in the Protein Data Bank under the accession numbers 6Q63 and 6Q64. The other data that support the findings in this paper are available upon request from the corresponding authors.
References
McNeil, N. I. The contribution of the large intestine to energy supplies in man. Am. J. Clin. Nutr. 39, 338–342 (1984).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Khoruts, A., Dicksved, J., Jansson, J. K. & Sadowsky, M. J. Changes in the composition of the human fecal microbiome after bacteriotherapy for recurrent Clostridium difficile-associated diarrhea. J. Clin. Gastroenterol. 44, 354–360 (2010).
O’Keefe, S. J. et al. Products of the colonic microbiota mediate the effects of diet on colon cancer risk. J. Nutr. 139, 2044–2048 (2009).
Koropatkin, N. M., Cameron, E. A. & Martens, E. C. How glycan metabolism shapes the human gut microbiota. Nat. Rev. Microbiol. 10, 323–335 (2012).
Koropatkin, N. M., Martens, E. C., Gordon, J. I. & Smith, T. J. Starch catabolism by a prominent human gut symbiont is directed by the recognition of amylose helices. Structure 16, 1105–1115 (2008).
Luis, A. S. et al. Dietary pectic glycans are degraded by coordinated enzyme pathways in human colonic Bacteroides. Nat. Microbiol. 3, 210–219 (2018).
Cuskin, F. et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 517, 165–169 (2015).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e21 (2016).
Chung, C. Y., Majewska, N. I., Wang, Q., Paul, J. T. & Betenbaugh, M. J. SnapShot: N-glycosylation processing pathways across kingdoms. Cell 171, 258–258 (2017).
Mathias, A. & Corthesy, B. N-Glycans on secretory component: mediators of the interaction between secretory IgA and gram-positive commensals sustaining intestinal homeostasis. Gut Microbes 2, 287–293 (2011).
Corfield, A. P. The interaction of the gut microbiota with the mucus barrier in health and disease in human. Microorganisms 6, 78 (2018).
Mestecky, J., Russell, M. W., Jackson, S. & Brown, T. A. The human IgA system: a reassessment. Clin. Immunol. Immunopathol. 40, 105–114 (1986).
Hughes, G. J., Reason, A. J., Savoy, L., Jaton, J. & Frutiger-Hughes, S. Carbohydrate moieties in human secretory component. Biochim. Biophys. Acta 1434, 86–93 (1999).
Garrido, D. et al. Endo-beta-N-acetylglucosaminidases from infant gut-associated bifidobacteria release complex N-glycans from human milk glycoproteins. Mol. Cell. Proteomics 11, 775–785 (2012).
Sebaihia, M. et al. The multidrug-resistant human pathogen Clostridium difficile has a highly mobile, mosaic genome. Nat. Genet. 38, 779–786 (2006).
Bohle, L. A., Mathiesen, G., Vaaje-Kolstad, G. & Eijsink, V. G. An endo-beta-N-acetylglucosaminidase from Enterococcus faecalis V583 responsible for the hydrolysis of high-mannose and hybrid-type N-linked glycans. FEMS Microbiol. Lett. 325, 123–129 (2011).
Renzi, F. et al. The N-glycan glycoprotein deglycosylation complex (Gpd) from Capnocytophaga canimorsus deglycosylates human IgG. PLoS Pathog. 7, e1002118 (2011).
Collin, M. & Fischetti, V. A. A novel secreted endoglycosidase from Enterococcus faecalis with activity on human immunoglobulin G and ribonuclease B. J. Biol. Chem. 279, 22558–22570 (2004).
Cao, Y., Rocha, E. R. & Smith, C. J. Efficient utilization of complex N-linked glycans is a selective advantage for Bacteroides fragilis in extraintestinal infections. Proc. Natl Acad. Sci. USA 111, 12901–12906 (2014).
Robb, M. et al. Molecular characterization of N-glycan degradation and transport in Streptococcus pneumoniae and its contribution to virulence. PLoS Pathog. 13, e1006090 (2017).
Sjogren, J. et al. EndoS2 is a unique and conserved enzyme of serotype M49 group A Streptococcus that hydrolyses N-linked glycans on IgG and alpha1-acid glycoprotein. Biochem. J. 455, 107–118 (2013).
Collin, M. & Olsen, A. EndoS, a novel secreted protein from Streptococcus pyogenes with endoglycosidase activity on human IgG. EMBO J. 20, 3046–3055 (2001).
Dupoiron, S. et al. The N-glycan cluster from Xanthomonas campestris pv. campestris: a toolbox for sequential plant N-glycan processing. J. Biol. Chem. 290, 6022–6036 (2015).
Forster, S. C. et al. A human gut bacterial genome and culture collection for improved metagenomic analyses. Nat. Biotechnol. 37, 186–192 (2019).
Martens, E. C. et al. Recognition and degradation of plant cell wall polysaccharides by two human gut symbionts. PLoS Biol. 9, e1001221 (2011).
Rogowski, A. et al. Glycan complexity dictates microbial resource allocation in the large intestine. Nat. Commun. 6, 7481 (2015).
Ndeh, D. et al. Complex pectin metabolism by gut bacteria reveals novel catalytic functions. Nature 544, 65–70 (2017).
Bagenholm, V. et al. Galactomannan catabolism onferred by a polysaccharide utilization locus of Bacteroides ovatus: enzyme synergy and crystal structure of a beta-mannanase. J. Biol. Chem. 292, 229–243 (2017).
Tamura, K. et al. Molecular mechanism by which prominent human gut bacteroidetes utilize mixed-linkage beta-glucans, major health-promoting cereal polysaccharides. Cell Rep. 21, 417–430 (2017).
Temple, M. J. et al. A Bacteroidetes locus dedicated to fungal 1,6-beta-glucan degradation: unique substrate conformation drives specificity of the key endo-1,6-beta-glucanase. J. Biol. Chem. 292, 10639–10650 (2017).
Larsbrink, J. et al. A discrete genetic locus confers xyloglucan metabolism in select human gut Bacteroidetes. Nature 506, 498–502 (2014).
Martens, E. C., Koropatkin, N. M., Smith, T. J. & Gordon, J. I. Complex glycan catabolism by the human gut microbiota: the Bacteroidetes Sus-like paradigm. J. Biol. Chem. 284, 24673–24677 (2009).
Reeves, A. R., Wang, G. R. & Salyers, A. A. Characterization of four outer membrane proteins that play a role in utilization of starch by Bacteroides thetaiotaomicron. J. Bacteriol. 179, 643–649 (1997).
Thomas, F. et al. Characterization of the first alginolytic operons in a marine bacterium: from their emergence in marine Flavobacteriia to their independent transfers to marine Proteobacteria and human gut Bacteroides. Environ. Microbiol. 14, 2379–2394 (2012).
Muchmore, E. A., Diaz, S. & Varki, A. A structural difference between the cell surfaces of humans and the great apes. Am. J. Phys. Anthropol. 107, 187–198 (1998).
Park, K. H. et al. Structural and biochemical characterization of the broad substrate specificity of Bacteroides thetaiotaomicron commensal sialidase. Biochim. Biophys. Acta 1834, 1510–1519 (2013).
Almagro-Moreno, S. & Boyd, E. F. Insights into the evolution of sialic acid catabolism among bacteria. BMC Evolut. Biol. 9, 118 (2009).
Phansopa, C. et al. Characterization of a sialate-O-acetylesterase (NanS) from the oral pathogen Tannerella forsythia that enhances sialic acid release by NanH, its cognate sialidase. Biochem. J. 472, 157–167 (2015).
Garbe, J. et al. EndoE from Enterococcus faecalis hydrolyzes the glycans of the biofilm inhibiting protein lactoferrin and mediates growth. PloS One 9, e91035 (2014).
Waddling, C. A., Plummer, T. H. Jr., Tarentino, A. L. & Van Roey, P. Structural basis for the substrate specificity of endo-beta-N-acetylglucosaminidase F(3). Biochemistry 39, 7878–7885 (2000).
Green, E. D., Adelt, G., Baenziger, J. U., Wilson, S. & Van Halbeek, H. The asparagine-linked oligosaccharides on bovine fetuin. Structural analysis of N-glycanase-released oligosaccharides by 500-megahertz 1H NMR spectroscopy. J. Biol. Chem. 263, 18253–18268 (1988).
Tailford, L. E. et al. Mannose foraging by Bacteroides thetaiotaomicron: structure and specificity of the beta-mannosidase, BtMan2A. J. Biol. Chem. 282, 11291–11299 (2007).
Pluvinage, B. et al. Inhibition of the pneumococcal virulence factor StrH and molecular insights into N-glycan recognition and hydrolysis. Structure 19, 1603–1614 (2011).
Liu, T. et al. Structural determinants of an insect beta-N-acetyl-D-hexosaminidase specialized as a chitinolytic enzyme. J. Biol. Chem. 286, 4049–4058 (2011).
Theodoratou, E. et al. Glycosylation of plasma IgG in colorectal cancer prognosis. Sci. Rep. 6, 28098 (2016).
Martens, E. C., Chiang, H. C. & Gordon, J. I. Mucosal glycan foraging enhances fitness and transmission of a saccharolytic human gut bacterial symbiont. Cell Host Microbe 4, 447–457 (2008).
Martens, E. C., Roth, R., Heuser, J. E. & Gordon, J. I. Coordinate regulation of glycan degradation and polysaccharide capsule biosynthesis by a prominent human gut symbiont. J. Biol. Chem. 284, 18445–18457 (2009).
Kubinak, J. L. et al. MyD88 signaling in T cells directs IgA-mediated control of the microbiota to promote health. Cell Host Microbe 17, 153–163 (2015).
Lawrence, R. M. & Pane, C. A. Human breast milk: current concepts of immunology and infectious diseases. Curr. Probl. Pediatr. Adolesc. Health Care 37, 7–36 (2007).
Charnock, S. J. et al. Key residues in subsite F play a critical role in the activity of Pseudomonas fluorescens subspecies cellulosa xylanase A against xylooligosaccharides but not against highly polymeric substrates such as xylan. J. Biol. Chem. 272, 2942–2951 (1997).
Gasteiger E. et al. in The Proteomics Protocols Handbook (ed. Walker, J. M.) 571–607 (Humana Press, 2015).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006).
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007).
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010).
Cowtan, K. Recent developments in classical density modification. Acta Crystallogr. D 66, 470–478 (2010).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D 62, 1002–1011 (2006).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Keegan, R. M. & Winn, M. D. Automated search-model discovery and preparation for structure solution by molecular replacement. Acta Crystallogr. D 63, 447–457 (2007).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
The PyMOL Molecular Graphics System v.2.0 (Schrödinger LLC, 2017).
Collaborative Computational Project, Number 4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Hehemann, J. H., Kelly, A. G., Pudlo, N. A., Martens, E. C. & Boraston, A. B. Bacteria of the human gut microbiome catabolize red seaweed glycans with carbohydrate-active enzyme updates from extrinsic microbes. Proc. Natl Acad. Sci. USA 109, 19786–19791 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zerbino, D. R. et al. Ensembl 2018. Nucleic Acids Res. 46, D754–d761 (2018).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, GBIF, 2018).
Zhang, Z., Xie, J., Zhang, F. & Linhardt, R. J. Thin-layer chromatography for the analysis of glycosaminoglycan oligosaccharides. Anal. Biochem. 371, 118–120 (2007).
Ceroni, A. et al. GlycoWorkbench: a tool for the computer-assisted annotation of mass spectra of glycans. J. Proteome Res. 7, 1650–1659 (2008).
Juncker, A. S. et al. Prediction of lipoprotein signal peptides in Gram-negative bacteria. Protein Sci. 12, 1652–1662 (2003).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Markowitz, V. M. et al. IMG: the Integrated Microbial Genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–D122 (2012).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Holm, L. & Laakso, L. M. Dali server update. Nucleic Acids Res. 44, W351–W355 (2016).
Krissinal, E. & Hanrick, K. PDBeFold (European Bioinformatics Institute, 2009).
Dereeper, A., Audic, S., Claverie, J. M. & Blanc, G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evolut. Biol. 10, 8 (2010).
Dereeper, A. et al. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 36, W465–W469 (2008).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Letunic, I. & Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 46, D493–D496 (2018).
Letunic, I., Doerks, T. & Bork, P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 43, D257–D260 (2015).
Acknowledgements
We thank C. Morland (Newcastle University, UK) for his expert technical assistance, and F. Cuskin (Newcastle University, UK) and L. Royle (Ludger, UK) for insightful conversations about the data. We would like to thank Diamond Light Source (Oxfordshire, UK) for beamtime (proposal mx13587 and mx18598) and the staff of beamline I03 and 104-1 for assistance with crystal testing and data collection. We thank J. Casement (Bioinformatics Support Unit, Newcastle University, UK) for analysing the raw RNA-Seq data. We thank J. Sonnenberg (Stanford, USA) for the ΔBT0455 Bt strain, R. Lewis (Newcastle University, UK) for his guidance and advice in looking for structural homologues and R. Hirt (Newcastle University, UK) for his advice on phylogenetics. The work was funded by the BBSRC/Innovate UK IB catalyst award to D.N.B. ‘Glycoenzymes for Bioindustries’ (BB/M029018/1).
Author information
Authors and Affiliations
Contributions
J.B., A.S.L. and L.I.C. carried out enzyme kinetics. J.B., P.A.U., A.S.L., O.R. and L.I.C. carried out enzymes assays. J.H. made substrates. J.B. and L.I.C. carried out Bacteroides growth. L.I.C. carried out the whole-cell assays. P.A.U. and O.R. carried out the LC–MS. P.A.U. and L.I.C. analysed the LC–MS data. J.B. and L.I.C. purified proteins and set up crystal trays. A.B. collected crystals and data. A.B. and N.P. solved crystal structures. J.B., D.N. and L.I.C. produced Bacteroides gene deletion strains. L.I.C. carried out the bioinformatic analysis. L.I.C., D.N.B., P.A.U., J.B., A.S.L., D.I.R.S. and E.C.M. designed experiments. L.I.C., E.C.L. and D.N.B wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Results and Discussion, legends for Supplementary Tables 1–13, Supplementary Figures 1–24, and Supplementary References.
Supplementary Data 1
RNA seq data table showing average normalized base counts in glucose and bovine α1-acid glycoprotein (α1AGP) samples.
Rights and permissions
About this article
Cite this article
Briliūtė, J., Urbanowicz, P.A., Luis, A.S. et al. Complex N-glycan breakdown by gut Bacteroides involves an extensive enzymatic apparatus encoded by multiple co-regulated genetic loci. Nat Microbiol 4, 1571–1581 (2019). https://doi.org/10.1038/s41564-019-0466-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0466-x
This article is cited by
-
Particle-attached bacteria act as gatekeepers in the decomposition of complex phytoplankton polysaccharides
Microbiome (2024)
-
An expanded transcriptome atlas for Bacteroides thetaiotaomicron reveals a small RNA that modulates tetracycline sensitivity
Nature Microbiology (2024)
-
The gut microbiota and its biogeography
Nature Reviews Microbiology (2024)
-
Outer membrane utilisomes mediate glycan uptake in gut Bacteroidetes
Nature (2023)
-
Crystal structure of the CoV-Y domain of SARS-CoV-2 nonstructural protein 3
Scientific Reports (2023)