Abstract
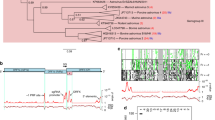
Enteroviruses comprise a large group of mammalian pathogens that includes poliovirus. Pathology in humans ranges from sub-clinical to acute flaccid paralysis, myocarditis and meningitis. Until now, all of the enteroviral proteins were thought to derive from the proteolytic processing of a polyprotein encoded in a single open reading frame. Here we report that many enterovirus genomes also harbour an upstream open reading frame (uORF) that is subject to strong purifying selection. Using echovirus 7 and poliovirus 1, we confirmed the expression of uORF protein in infected cells. Through ribosome profiling (a technique for the global footprinting of translating ribosomes), we also demonstrated translation of the uORF in representative members of the predominant human enterovirus species, namely Enterovirus A, B and C. In differentiated human intestinal organoids, uORF protein-knockout echoviruses are attenuated compared to the wild-type at late stages of infection where membrane-associated uORF protein facilitates virus release. Thus, we have identified a previously unknown enterovirus protein that facilitates virus growth in gut epithelial cells—the site of initial viral invasion into susceptible hosts. These findings overturn the 50-year-old dogma that enteroviruses use a single-polyprotein gene expression strategy and have important implications for the understanding of enterovirus pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The sequencing data reported in this paper have been deposited in ArrayExpress (http://www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-6180.
References
Suresh, S., Forgie, S. & Robinson, J. Non-polio Enterovirus detection with acute flaccid paralysis: a systematic review. J. Med. Virol. 90, 3–7 (2017).
Bedard, K. M. & Semler, B. L. Regulation of picornavirus gene expression. Microbes Infect. 6, 702–713 (2004).
Sweeney, T. R., Abaeva, I. S., Pestova, T. V. & Hellen, C. U. T. The mechanism of translation initiation on Type 1 picornavirus IRESs. EMBO J. 33, 76–92 (2014).
Pelletier, J., Flynn, M. E., Kaplan, G., Racaniello, V. & Sonenberg, N. Mutational analysis of upstream AUG codons of poliovirus RNA. J. Virol. 62, 4486–4492 (1988).
Hellen, C. U., Pestova, T. V. & Wimmer, E. Effect of mutations downstream of the internal ribosome entry site on initiation of poliovirus protein synthesis. J. Virol. 68, 6312–6322 (1994).
Kaminski, A., Poyry, T. A. A., Skene, P. J. & Jackson, R. J. Mechanism of Initiation site selection promoted by the human rhinovirus 2 internal ribosome entry site. J. Virol. 84, 6578–6589 (2010).
Pestova, T. V., Hellen, C. U. T. & Wimmer, E. A conserved AUG triplet in the 5′ nontranslated region of poliovirus can function as an initiation codon in vitro and in vivo. Virology 204, 729–737 (1994).
Meerovitch, K., Nicholson, R. & Sonenberg, N. In vitro mutational analysis of cis-acting RNA translational elements within the poliovirus type 2 5′ untranslated region. J. Virol. 65, 5895–5901 (1991).
Firth, A. & Brown, C. Detecting overlapping coding sequences in virus genomes. BMC Bioinform. 7, 75 (2006).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Irigoyen, N. et al. High-resolution analysis of coronavirus gene expression by RNA sequencing and ribosome profiling. PLoS Pathog. 12, e1005473 (2016).
Slobodskaya, O. R. et al. Poliovirus neurovirulence correlates with the presence of a cryptic AUG upstream of the initiator codon. Virology 221, 141–150 (1996).
Drummond, C. G. et al. Enteroviruses infect human enteroids and induce antiviral signaling in a cell lineage-specific manner. Proc. Natl Acad. Sci. USA 114, 1672–1677 (2017).
Ettayebi, K. et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 353, 1387–1393 (2016).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Kraiczy, J. et al. DNA methylation defines regional identity of human intestinal epithelial organoids and undergoes dynamic changes during development. Gut https://doi.org/10.1136/gutjnl-2017-314817 (2017).
Ward, T. et al. Decay-accelerating factor CD55 is identified as the receptor for echovirus 7 using CELICS, a rapid immuno-focal cloning method. EMBO J. 13, 5070–5074 (1994).
Racaniello, V. R. One hundred years of poliovirus pathogenesis. Virology 344, 9–16 (2006).
Kitamura, N. et al. Primary structure, gene organization and polypeptide expression of poliovirus RNA. Nature 291, 547–553 (1981).
Racaniello, V. R. & Baltimore, D. Molecular cloning of poliovirus cDNA and determination of the complete nucleotide sequence of the viral genome. Proc. Natl Acad. Sci. USA 78, 4887–4891 (1981).
Jacobson, M. F. & Baltimore, D. Polypeptide cleavages in the formation of poliovirus proteins. Proc. Natl Acad. Sci. USA 61, 77–84 (1968).
Royston, L. & Tapparel, C. Rhinoviruses and respiratory enteroviruses: not as simple as ABC. Viruses 8, 16 (2016).
Ohlmann, T. & Jackson, R. J. The properties of chimeric picornavirus IRESes show that discrimination between internal translation initiation sites is influenced by the identity of the IRES and not just the context of the AUG codon. RNA 5, 764–778 (1999).
Andreev, D. E. et al. Glycyl-tRNA synthetase specifically binds to the poliovirus IRES to activate translation initiation. Nucleic Acids Res. 40, 5602–5614 (2012).
Feng, Z. et al. A pathogenic picornavirus acquires an envelope by hijacking cellular membranes. Nature 496, 367–371 (2013).
Chen, Y.-H. et al. Phosphatidylserine vesicles enable efficient en bloc transmission of enteroviruses. Cell 160, 619–630 (2015).
Richards, A. L. & Jackson, W. T. Behind closed membranes: the secret lives of picornaviruses? PLoS Pathog. 9, e1003262 (2013).
Sin, J., McIntyre, L., Stotland, A., Feuer, R. & Gottlieb, R. A. Coxsackievirus B escapes the infected cell in ejected mitophagosomes. J. Virol. 91, e01347-17 (2017).
Cornell, C. T., Perera, R., Brunner, J. E. & Semler, B. L. Strand-specific RNA synthesis determinants in the RNA-dependent RNA polymerase of poliovirus. J. Virol. 78, 4397–4407 (2004).
Lulla, V. et al. Assembly of replication-incompetent African horse sickness virus particles: rational design of vaccines for all serotypes. J. Virol. 90, 7405–7414 (2016).
Loughran, G., Howard, M. T., Firth, A. E. & Atkins, J. F. Avoidance of reporter assay distortions from fused dual reporters. RNA 23, 1285–1289 (2017).
Vogt, D. A. & Ott, M. Membrane flotation assay. Bio-protocol 5, e1435 (2015).
Goodfellow, I. G. et al. Inhibition of coxsackie B virus infection by soluble forms of its receptors: binding affinities, altered particle formation, and competition with cellular receptors. J. Virol. 79, 12016–12024 (2005).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Simmonds, P. & Welch, J. Frequency and dynamics of recombination within different species of human enteroviruses. J. Virol. 80, 483–493 (2006).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 5, 113 (2004).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Stocsits, R. R., Hofacker, I. L., Fried, C. & Stadler, P. F. Multiple sequence alignments of partially coding nucleic acid sequences. BMC Bioinform. 6, 160 (2005).
Firth, A. E. Mapping overlapping functional elements embedded within the protein-coding regions of RNA viruses. Nucleic Acids Res. 42, 12425–12439 (2014).
McWilliam, H. et al. Analysis tool web services from the EMBL-EBI. Nucleic Acids Res. 41, W597–W600 (2013).
Acknowledgements
We thank the Cambridge NIHR BRC Cell Phenotyping Hub for assistance with confocal microscopy. We thank T. Sweeney, I. Brierley and E. Jan for stimulating discussions. This work was supported by Wellcome Trust grant no. 106207 and European Research Council grant no. 646891 to A.E.F and Wellcome Trust grant nos 097997/Z/11/Z and 207498/Z/17/Z to I.G.
Author information
Authors and Affiliations
Contributions
A.E.F. and V.L. conceived the project. V.L. performed the experiments. M.Z., K.M.N., M.H., Y.C. and I.G. established the organoid system, prepared and maintained the organoids and assisted with the organoid experiments. L.S. and N.J.S. established the poliovirus system and helped prepare poliovirus samples. N.I. advised and assisted with the Ribo-Seq experiments. A.E.F. performed the comparative genomic analyses. A.M.D. analysed the Ribo-Seq data. V.L. and A.E.F. wrote the manuscript. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–23, Supplementary Tables 1–4 and Supplementary References.
Supplementary Table 3
Statistical analysis of data from organoid-derived samples.
Supplementary Table 4
Statistical analysis of dual luciferase data.
Rights and permissions
About this article
Cite this article
Lulla, V., Dinan, A.M., Hosmillo, M. et al. An upstream protein-coding region in enteroviruses modulates virus infection in gut epithelial cells. Nat Microbiol 4, 280–292 (2019). https://doi.org/10.1038/s41564-018-0297-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0297-1
This article is cited by
-
Streamlined and sensitive mono- and di-ribosome profiling in yeast and human cells
Nature Methods (2023)
-
A novel approach to finding conserved features in low-variability gene alignments characterises RNA motifs in SARS-CoV and SARS-CoV-2
Scientific Reports (2023)
-
Persistent coxsackievirus B infection and pathogenesis of type 1 diabetes mellitus
Nature Reviews Endocrinology (2022)
-
A Single Mutation in the Cryptic AUG (cAUG) Affects In Vitro Translation and Replication Efficiencies and In Vivo Virulence of Coxsackievirus B3 (CVB3)
Current Microbiology (2022)
-
Unconventional viral gene expression mechanisms as therapeutic targets
Nature (2021)