Abstract
Environmental DNA surveys reveal that most fungal diversity represents uncultured species. We sequenced the genomes of eight uncultured species across the fungal tree of life using a new single-cell genomics pipeline. We show that, despite a large variation in genome and gene space recovery from each single amplified genome (SAG), ≥90% can be recovered by combining multiple SAGs. SAGs provide robust placement for early-diverging lineages and infer a diploid ancestor of fungi. Early-diverging fungi share metabolic deficiencies and show unique gene expansions correlated with parasitism and unculturability. Single-cell genomics holds great promise in exploring fungal diversity, life cycles and metabolic potential.
Similar content being viewed by others
Main
Genomics research enables a well-resolved phylogenetic backbone for the fungal tree of life and describes how the fungal nutritional toolkit has evolved over a billion years1. A complete grasp of these evolutionary patterns requires a thorough understanding of the enzymes and metabolites utilized by all fungi for accessing diverse sources of nutrition. However, much of what we know about the evolution of the fungal metabolic repertoire is biased towards fungi that are model systems, enzyme factories or important human or plant pathogens2. This emphasis results in an underrepresentation of uncultured lineages in the fungal genomic tree and the inaccessibility of these clades to comparative genomic analysis.
Environmental DNA amplicon surveys provide ample evidence for a high diversity of uncultured fungi that are unable to complete their life cycle in axenic conditions. Most studies recover species that are not represented in genomic databases3,4,5. This demonstrates a collectively limited knowledge of the true diversity of fungi, a problem exacerbated when considering early-diverging lineages, which are primarily microscopic and diverse in biotrophic groups1,6. Among these uncultured fungal lineages are entire hyperdiverse phyla, such as the Cryptomycota7. Even among the culturable early-diverging fungi (EDF), such as the Chytridiomycota, environmental DNA studies show almost no overlap with known and sampled species8,9. Because genome sequencing has thus far been limited to cultured fungi, the phylogeny of EDF remains poorly resolved. A better understanding of their genomes will provide clues regarding evolutionary history, basic biology and metabolism (for example, ref. 10).
Cultivation-independent methods for sequencing environmental microbial taxa, driven largely by shotgun metagenomics, have been applied for over a decade. Owing in part to the complexity of metagenomic data, the application of single-cell genomics has sharply increased over the past 5 years11,12,13. These methods rely on isolation and lysis of individual cells with subsequent whole-genome amplification and sequencing14. Most of the current environmental work has focused primarily on bacterial and archaeal systems15 (see ref. 13 for a review); however, an increasing number of eukaryotic genomes have been reported from single-cell sequencing16,17,18,19. Although recent efforts successfully demonstrated both single-nucleus de novo genome sequencing of the endomycorrhizal fungus Rhizophagus irregularis20 and high-throughput microfluidics single-cell sequencing21, these methods are neither generalizable nor easily adoptable in other laboratories. Fungi pose several challenges for scaling up single-cell genomics. These include structural challenges, such as the presence of a cell wall and diverse cellular morphologies, as well as genomic challenges, such as multiple chromosomes and higher ploidy, mitochondrial genomes, wide GC variation and transposable elements. To develop methods for robust capture and de novo assembly of environmental fungal genomes, we designed a project using known target species to explore the technical challenges prior to isolating and sequencing more complex environmental samples.
Here, we applied single-cell genomics methods for the first time to uncultured mycoparasitic EDF from the Cryptomycota, Chytridiomycota and Zoopagomycota22: Rozella allomycis, Caulochytrium protostelioides, Dimargaris cristalligena, Piptocephalis cylindrospora, Syncephalis pseudoplumigaleata and Thamnocephalis sphaerospora. We focused primarily on biotrophic mycoparasites because they are an understudied group of fungi that are widely represented among EDF23 and they sporulate abundantly when cultured with their hosts. We also included the pollen saprotroph Blyttiomyces helicus (Chytridiomycota) and Daphnia parasite Metschnikowia bicuspidata (Ascomycota) because samples of these fungi also contain DNA from numerous non-target species, similar to environmental samples. In this study, we provide genomic insights into the biology and evolutionary histories of these uncultivated species. We illustrated robust placement of novel lineages among EDF, demonstrated higher than haploid ploidy as a common characteristic of these lineages and revealed interesting gene family evolution patterns outside the Dikarya. We also highlighted common metabolic deficiencies among uncultured lineages and tested whether these deficiencies could be overcome through culturing efforts with addition of limiting reagents. Collectively, these approaches will facilitate further study on diverse and uncultured environmental eukaryotes.
Results
Single-cell pipeline successfully captures fungal genomes with high completeness
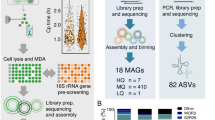
We developed and applied our single-cell pipeline (Supplementary Fig. 1) to eight diverse target species. We recovered genomes from individual cells (‘1-cell’), as well as pools of multiple cells sequenced as one library (‘10-/30-/50-/100-cells’). These individual libraries (Supplementary Table 1) were combined in separate co-assemblies (a technique routinely used in microbial single-cell projects15,24,25) to maximize completeness.
Figure 1b summarizes genome completeness (per cent CEGMA (Core Eukaryotic Genes Mapping Approach)26), assembly size and total gene content of co-assemblies and individual assemblies. Generally, CEGMA percentages increased when more cells were incorporated into a library, ranging from 2.8% (1 cell, C. protostelioides) to 96.7% (100 cell, M. bicuspidata). For all species, co-assemblies were most complete (relative to the 1-cell or 50-cell/100-cell libraries), ranging from 73.36% (P. cylindrospora) to 99.34% (D. cristalligena). Assembly sizes of the 1-cell libraries range from 0.5 Mb (C. protostelioides) to 21.1 Mb (B. helicus). The largest single-library assembly was 30.1 Mb (100 cell, B. helicus). Co-assembly sizes ranged from 10.6 Mb (C. protostelioides) to 46.5 Mb (B. helicus). Predicted gene counts ranged from 111 (1 cell, C. protostelioides) to 6,941 (100 cell, D. cristalligena) for single-library assemblies. Co-assembly gene counts ranged from 3,328 (C. protostelioides) to 12,167 (B. helicus). Overall, there is a strong positive correlation between estimated genome completeness and assembly size (Fig. 2a). Within a single taxon, the trend suggests that more cells sorted per library results in more complete libraries. With the exception of R. allomycis, >75% of reads of a given library were incorporated into an assembly (Fig. 2b).
a, RAxML tree constructed from MCL clustering from across the fungi and deep-branching eukaryote outgroups. Support values are based on 1,000 bootstrap replicates. Bootstrap values (<100%) are indicated on branches. Fungal species are annotated with simplified lifestyle icons. Species sequenced using single-cell methods are coloured in magenta. Enrichment cultures for R. allomycis and C. protostelioides and a related species of M. bicuspidata are denoted with an asterisk. b, Estimated completeness (CEGMA (%)), assembly size and gene model statistics for fungal genomes. Metrics for the most complete individual libraries are indicated on the respective graphs, either as ‘single’ (referring to 1-cell libraries), ‘multiple’ (referring to 100-cell libraries in all cases except for T. sphaerospora, in which it refers to a 50-cell library) or ‘CoASM’ (referring to co-assemblies). ‘Isolate’ refers to assemblies derived from genomic material sequenced using non-single-cell methods.
a, Scatter plot showing the correlation between estimated completeness and assembly size for individual libraries and co-assemblies for a given species. b, Scatter plot showing the correlation between estimated completeness and the percentage of reads from a given library mapped to the co-assembly for that organism. Individual library sizes are reflected in the size of the points.
Single-cell assemblies provide robust fungal phylogeny
To help resolve fungal phylogeny, a maximum likelihood tree was built from whole-genome clustering data from the eight co-assemblies along with major fungal and deep-branching eukaryotic representatives (Supplementary Table 2). The phylogenetic placement of our co-assembled genomes agrees with current taxonomic understanding: D. cristalligena groups with Kickxellomycotina; B. helicus and C. protostelioides with Chytridiomycota; R. allomycis with Microsporidia as sister to the rest of the fungi; and P. cylindrospora, S. pseudoplumigaleata and T. sphaerospora form a monophyletic group sister to the Kickxellomycotina within the Zoopagomycota (Fig. 1a).
Because single-cell libraries will only recover a partial genome, we determined the effect of genome completeness and input cell number on generating an accurate phylogeny. We annotated all libraries from D. cristalligena, C. protostelioides, R. allomycis and M. bicuspidata and constructed individual phylogenetic trees using each library (Supplementary Data). These topologies were compared to the co-assembly-based tree in Fig. 1a. Only three libraries (7.7%) were incongruent for that particular taxon, despite libraries having as few as 15 genes belonging to orthologous gene clusters (median: 434): one R. allomycis (1 cell, 1,095 clusters) and two C. protostelioides (100 cell and 10 cell, 1,091 and 1,090 clusters, respectively) (Supplementary Table 3). These results show that less-complete single-celled libraries can be used to generate accurate phylogenies.
Single-cell genomes indicate that higher ploidy is common in EDF
Single-cell genomes provide a unique opportunity to separate polymorphisms among cells from polymorphisms within cells. The ploidy of most EDF species is largely unknown but a few have been shown to be higher than haploid27,28,29. Analyses with k-mer graphs and allele frequency spectra can successfully distinguish between haploid and non-haploid organisms29,30,31. For C. protostelioides and R. allomycis, our results indicate haploid and triploid patterns, respectively (Fig. 3a–d). We were able to identify putative single-nucleotide polymorphisms (SNPs) in single-cell libraries of both species, but only for R. allomycis do single-cell SNPs match those identified in isolate sequencing (Fig. 3e).
a–g, Analysis of diversity in population sequencing (a–d) and single-cell sequencing (e–g). R. allomycis (a,b) and C. protostelioides (c,d) population variability is shown in the k-mer distribution plot (a,c) and in the allele frequency spectra (b,d). In panel e, recovery of isolate SNPs from a single cell (1) and multiple cells (1–100) are shown. In panel f, the amount of heterozygous SNPs (allele frequencies above 25%) in single-cell and multiple-cell libraries is indicative of higher ploidy. In panel g, high numbers of shared heterozygous SNPs in non-haploid organisms are shown. For e–g, n = 4 biologically independent samples, in which each dot corresponds to one library. The horizontal black lines indicate the mean values.
Single-cell sequencing can also identify heterozygous SNPs in diploid cells32. By using C. protostelioides and R. allomycis as haploid and non-haploid models, respectively, we identified five higher-than-haploid species (Fig. 3f). Moreover, because we are able to show that most variants identified in our five non-haploid species are present in multiple libraries (Fig. 3g), our patterns are consistent with heterozygous SNPs expected in all cells of an isolate. These results show that single-cell sequencing can elucidate the ploidy of fungi and suggest that the majority of EDF are non-haploid.
We also identified cell-to-cell polymorphisms among one-cell libraries of D. cristalligena (Supplementary Fig. 2). We determined that six SNPs for a particular gene were exactly shared between two libraries (AHPZW and AHPZP), whereas one was unique to a third library (AHSAA).
Single-cell genomes highlight common deficiencies in primary metabolism of uncultured fungi
Genome sequencing of uncultured EDF allows us to explore how metabolic content has changed during the course of fungal evolution. Compared to the common ancestor of fungi (Supplementary Fig. 3), D. cristalligena shows gains in high-level metabolic categories, whereas B. helicus has mainly losses.
Examination of the primary metabolism of genomes can reveal deficiencies responsible for unculturability, ideally resulting in successful culturing through supplementing media with missing metabolites33. One challenge with predicting missing function using single-cell genomes is the presence of false negatives from missing data, which we address by searching for commonalities in missing pathways across our target taxa. The conserved pathways found in 75% of ‘free-living’ fungi, overlaid with those missing from ≥5 target fungi, reflect consistent enzymatic losses (Fig. 4a).
a, Metabolic pathway map with enzymes found in ~70% of ‘free-living’ fungi and absent in at least one of the single-cell fungi shown as a gradient: darker red shades indicate more single-cell species missing the same enzyme in a given pathway. b–f, Certain pathways with a high degree of such common losses: assimilatory sulfate reduction (sulfate to sulfide) (b), thiamine biosynthesis (HMP and TZE to thiamine phosphate) (c), spermidine synthesis (S-adenosylmethionine to spermidine) (d), biotin metabolism (dethiobiotin to biotin (and biotinyl-5′-AMP)) (e), and citrate synthesis (citrate to/from oxaloacetate and acetyl-CoA) (f). APS, adenylyl sulfate; HMP, 4-amino-5-hydroxymethyl-2-methylpyrimidine; HMP-P, HMP phosphate; HMP-PP, HMP diphosphate; PAPS, phosphoadenylyl sulfate; TZE, 5-(2-hydroxyethyl)-4-methylthiazole; TZP, TZE phosphate. The coloured boxes for b–f indicate the presence (solid) or the absence (shaded) of a given enzyme in a genome: Ra, R. allomycis (red); Bh, B. helicus (blue); Cp, C. protostelioides (green); Dc, D. cristalligena (purple); Pc, P. cylindrospora (orange); Ts, T. sphaerospora (yellow); Sp, S. pseudoplumigaleata (brown); Mb, M. bicuspidata (pink).
The absence of spermidine synthase (enzyme classification (EC) 2.5.1.16) and S-adenosylmethionine decarboxylase (EC 4.1.1.50) suggests that almost all target taxa are unable to make spermidine and/or homospermidine (Fig. 4d). Spermidine is involved in the regulation of processes such as virulence and sporulation34,35, and deficiency in polyamines leads to auxotrophy and attenuation of virulence36,37.
Another deficiency common to almost all of the target genomes is in the assimilatory sulfate reduction pathway38 (Fig. 4b). ATP sulfurylase (EC 2.7.7.4) and adenylyl-sulfate kinase (EC 2.7.1.25) are missing from all except R. allomycis and a culturable M. bicuspidata relative (hereafter: NRRL YB-4993). Phosphoadenylyl-sulfate reductase (EC 1.8.4.8) and sulfite reductase (NADPH) (EC 1.8.1.2) are missing from all target genomes except M. bicuspidata NRRL YB-4993.
Metabolism of biotin is another common deficiency among target genomes (Fig. 4e). In plants, it is synthesized from dethiobiotin by biotin synthase (EC 2.8.1.6)39. This enzyme is only found in M. bicuspidata and B. helicus. Similarly, biotin–(acetyl-CoA-carboxylase) ligase (EC 6.3.4.15) is only found in M. bicuspidata, R. allomycis and S. pseudoplumigaleata.
Biosynthesis of thiamine phosphate is accomplished via thiamine phosphate synthase (EC 2.5.1.3) and hydroxyethylthiazole kinase (EC 2.7.1.50). These enzymes are absent from all target fungi except B. helicus. Both enzymes are absent from M. bicuspidata from Daphnia but are found in M. bicuspidata NRRL YB-4993 (Fig. 4c).
Entrance into the tricarboxylic acid cycle is accomplished through citrate synthase (EC 2.3.3.1), which is found in all free-living and target fungi. This enzyme facilitates the conversion of acetyl-CoA to citrate. ATP citrate synthase (EC 2.3.3.8) facilitates the same reaction but also generates ATP in the process. It is absent from all target fungi except for the chytrids C. protostelioides and B. helicus (Fig. 4f).
Based on our primary metabolism results, we attempted preliminary media-supplementing axenic culturing experiments for three mycoparasitic taxa: D. cristalligena, S. pseudoplumigaleata and P. cylindrospora. We tested the efficacy of five supplements to produce axenic growth of these fungi and obtained mixed results. S. pseudoplumigaleata either did not grow or was contaminated by the host fungus. P. cylindrospora grew axenically but weakly on all media treatments, including the control plates, but was not able to complete its life cycle on any of the media formulations. D. cristalligena responded best to our experiments; this fungus grew faster and more abundantly on the media that included all five supplements when compared to all other treatments. However, even on the fully supplemented axenic media, D. cristalligena was not able to complete its life cycle (Supplementary Fig. 4). Further experiments will be necessary to fully characterize axenic growth of these and related mycoparasitic EDF (see the Supplementary Information for complete experimental details).
Proteinase and CAZymes reveal ecology of uncultured fungi
To explore parasitism strategies, we focused broadly on abundance patterns of enzymatic protein domains. Comparing carbohydrate-active enzymes (CAZymes; http://www.cazy.org)40 and proteases (MEROPS; https://www.ebi.ac.uk/merops/)41, we found that facultative mycoparasites have a CAZyme-to-peptidase ratio most similar to plant pathogens, whereas that for obligate mycoparasites is most similar to animal pathogens (Fig. 5). B. helicus, the only saprotroph in this data set, has the most CAZymes among our single-cell genomes, with expansions of families that target pollen polysaccharides (Supplementary Table 4). To complement these broad observations, we focused narrowly on families often associated with mycoparasitism: subtilases, metallopeptidases and chitinases.
CAZymes and proteases (defined as in the CAZY database and the MEROPS database) were predicted for each species in the taxon list. Total counts of each were plotted to describe the overall ratio. Correlation was determined using the linear model function (lm) in R, which implements QR decomposition. A trend line for saprobe species (n = 19) was omitted owing to a low r2 value. Similarly, a trend line for mycorrhizal species was omitted owing to a low species count (n = 2).
Subtilases in Zoopagomycota
Subtilases, one of the largest clans of serine endopeptidases, are found in fungal entomopathogens, mycoparasites and plant pathogens42,43. Recent work has characterized novel categories of serine proteases in fungi44,45 in the Dikarya. We explored subtilase abundance in EDF mycoparasites and found that D. cristalligena subtilase sequences form a group distinct from others, whereas the Zoopagomycotina (P. cylindrospora, T. sphaerospora and S. pseudoplumigaleata) sequences split roughly equally between known proteinase K-like proteins and the group formed by D. cristalligena (Fig. 6a). Figure 6b shows the domain architecture of orthologous subtilase proteins predicted among the Zoopagomycota and illustrates expansions specific to D. cristalligena, T. sphaerospora and S. pseudoplumigaleata.
a, Network figure highlighting the relationships between fungal subtilases. The distance reflects sequence similarity. Groups marked as Li-1 through to Li-4 refer to clusters previously reported by Li et al.45, from which the data set originated. Bh, B. helicus; Cp, C. protostelioides; Dc, D. cristalligena; Mb, M. bicuspidata; Pc, P. cylindrospora; Ra, R. allomycis; Sp, S. pseudoplumigaleata; Ts, T. sphaerospora. b, Subtilase domain architecture in Zoopagomycota genomes, for proteins in a given homologous cluster. Mycoparasites sequenced using single-cell methods from this study (labelled with *) have generally more subtilase proteins. For S. pseudoplumigaleata and T. sphaerospora, these are predominately single-domain (PF00082) proteins. Domain architecture colours: blue, PF00082; light grey, PF02225; dark grey, PF06280. Lifestyles are indicated with simplified icons. Species abbreviations are consistent with panel a and also include: Cc, Conidiobolus coronatus; Cr, C. reversa; Lp, L. pennispora.
Metallopeptidases in Zoopagomycota
Class M36 metallopeptidases, also known as fungalysins, hydrolyze laminins, elastin, collagen and keratin46. The amphibian pathogen Batrachochytrium dendrobatidis has an expansion of this metallopeptidase47, which is presumably used in the degradation of keratin-rich amphibian skin. Orthologous metallopeptidase proteins were identified in the three species of Zoopagomycotina and D. cristalligena. A phylogenetic tree (Supplementary Fig. 5) highlights their relationship to other fungalysin proteins and illustrates a unique expansion among the Zoopagomycotina.
Chitinases in EDF
Chitin is a definitive component of the fungal cell wall48,49,50. It is broken down by chitinases of the glycoside hydrolase family 18 (GH18) and GH19 families and a recently described family (AA11)51. The two glycoside hydrolase families do not share similarities in protein sequence, structure or mechanism of action52. Furthermore, GH18 chitinases are widely distributed, whereas GH19 chitinases are described mainly from plants and act as defences to insect or fungal invaders.
There are a few known fungal representatives of GH19 chitinases, exclusively from the Cryptomycota and the Microsporidia. We also found GH19 chitinases for the first time in the Chytridiomycota and the Zoopagomycota. A small expansion was found in Rhizoclosmatium globosum, a saprotrophic chytridiomycete. Putative GH19 chitinases were also identified in D. cristalligena, Linderina pennispora and Coemansia reversa (Kickxellomycotina), and in P. cylindrospora (Zoopagomycotina). Only one protein from D. cristalligena had an additional CBM19 domain (Supplementary Fig. 6). D. cristalligena also has a unique expansion of the non-catalytic CBM18 family. Finally, the mycoparasites exclusively have AA11 genes, with clear expansions in D. cristalligena and T. sphaerospora (Supplementary Table 4).
Secondary metabolite expansion in D. cristalligena
Secondary metabolites are non-essential compounds produced by fungi for various purposes, including antagonism of other microorganisms, pathogenesis and iron chelation. Few secondary metabolites have been identified from EDF53. Our results support this observation (Supplementary Table 5), with most of the species containing one or fewer non-ribosomal peptide synthetases (NRPSs) or NRPS-like proteins and two or fewer polyketide synthases or polyketide synthase-like proteins. However, D. cristalligena possesses 27 NRPS genes divided between two lineage-specific expansions in two specific clades (Supplementary Fig. 7). Several of these homologous NRPS proteins also show modular synteny, suggesting multiple duplication events within D. cristalligena. Based on phylogenetic analysis of adenylation domains (Supplementary Fig. 7), these secondary metabolites are most closely related to epipolythiodioxopiperazine toxins that disrupt cell membranes. Trichoderma mycoparasites are known to produce several epipolythiodioxopiperazine toxins, including gliotoxin54. Future work is necessary to determine whether these epipolythiodioxopiperazine-like expansions are related to mycoparasitism in D. cristalligena. Most of the NRPS genes reside alone on relatively short contigs (Supplementary Table 6), precluding identification of a traditional fungal secondary metabolite gene cluster.
Hydrophobins in C. protostelioides
Hydrophobins are small cysteine-rich proteins involved in the development of aerial hyphae in certain filamentous fungi55. These proteins are currently only described in the Dikarya, with no evidence of their presence in EDF56. Caulochytrium is the only zoosporic genus known to produce aerial stalks and sporangia reminiscent of filamentous fungi57. We found 14 putative hydrophobins in the single-cell C. protostelioides proteome, all of which contained the highly conserved 8-cysteine marker region. Phylogeny and hydropathy profiles both suggest that the C. protostelioides proteins are more closely related to the group 1 proteins found in the Dikarya (Supplementary Fig. 8). Furthermore, we found one putative group 2 hydrophobin in the Mortierella elongata proteome. These findings represent the first examples of hydrophobins outside the Dikarya.
Discussion
In this study, we used single-cell genomics to create near-complete assemblies of uncultured fungi. This approach allowed us to capture an estimated 73–99% of the genome in multiple-cell co-assemblies ranging from 478 to 8,398 scaffolds. Our analysis shows that genome completeness from a single cell ranges from 6% to 88%, and that combining multiple cells can considerably increase assembly completeness from 6% to 80% (worst case) and from 88% to 96.7% (best case). Co-assemblies of genome data from different single cells further increase genome recovery while losing single-cell resolution for other analyses (for example, heterozygosity).
The co-assemblies allowed annotation of 3,328–12,167 proteins in each species, and common orthologous proteins were used to create a phylogenetic tree, placing these uncultured lineages among their cultured counterparts with robust bootstrap support. We placed Zoopagomycotina as a sister branch to Kickxellomycotina, although with minimal (69% bootstrap) support. A characteristic uniting these subphyla is that certain taxa in both groups produce merosporangia, that is, linear sporangia with few spores58. Although Dimargaris has been difficult to confidently place in ribosomal RNA gene phylogenies owing to a rapid rate of sequence evolution59, here, D. cristalligena is placed with strong (100%) support as a sister to the Kickxellales. An rRNA-based phylogeny placed B. helicus with the order Spizellomycetales and Rhizophlyctidiales but without statistical support60. In our maximum likelihood tree, B helicus groups with the Spizellomycetales representative Spizellomyces punctatus. Caulochytrium was placed in the Spizellomycetales by Barr61 based on ultrastructural data. However, Barr’s concept of the order has been radically reshaped by molecular phylogenetics60. Until this study, no molecular phylogenetic analyses have included Caulochytrium; thus, there was no clear null hypothesis for this chytrid. Although C. protostelioides grouped with the Chytridiales representative R. globosum (97% support), the precise relationship of C. protostelioides to other chytrids will require additional genome sequencing.
Single-cell sequencing is uniquely capable of addressing questions in fungal genetics regarding the organization of genetic variation. Analysis of single cells allows testing of whether detected SNPs are specific to an individual or due to variation among multiple genotypes in a variable population. The species targeted by single-cell sequencing were found to correspond to two different groups: non-haploid with high levels of heterozygosity and haploid with negligible heterozygosity (Fig. 3f). Importantly, taxa that we suspected of being diploid had >10,000 SNPs shared across libraries, whereas taxa that were haploid typically had <2,000 SNPs shared across libraries. One caveat is that fungi could be genetically diploid but show low heterozygosity, as is the case for Saccharomyces spp.62. Surprisingly, we found that R. allomycis is triploid, which is rare in fungi (for example, in Epichloë63) and usually reflects a sexually sterile condition. This observation of greater than diploidy has precedent in the microsporidia (for example, the tetraploid Nosema29), and the chytrid B. dendrobatidis has diploid to tetraploid nuclei64,65. Four of the six basal fungi are not haploid, including three of four Zoopagomycota. The preponderance of heterozygous species at the base of the fungal tree implies a diploid (or higher ploidy) ancestor of the fungi. Beyond estimating ploidy, the single-cell methods presented here could be applied broadly to other fungal taxa where culturing has been unsuccessful (for example, certain rust fungi and ectomycorrhizae), to facilitate genetic mapping, test for genetic segregation and establish mating systems using cohorts of meiotically produced spores.
The uncultured fungi in this study have diverse phylogenetic backgrounds and nutritional strategies. Most are parasites, presumably dependent on hosts for certain nutrients, which may explain their unculturability. By mapping the proteome to primary metabolism pathways, we observed major deficiencies in the ability to synthesize the full set of amino acids and polyamines, which probably result in auxotrophies. Overall, there is a mosaic of losses of essential metabolism genes across the uncultured taxa, consistent with patterns seen in the parasitic Cryptomycota66. The deficiencies in the biotin and thiamine pathways in mycoparasites are intriguing given that Syncephalis species have been successfully grown using beef liver67,68, which contains measurable quantities of both biotin and thiamine69. However, Syncephalis do not sporulate nor complete their life cycle under these conditions68. Most chytrid pollen saprotrophs are culturable, yet the unculturability of B. helicus may be explained, as it shares deficiencies in spermidine and sulfur metabolism. M. bicuspidata from Daphnia is similar to strain NRRL YB-4993 from brine shrimp (although they are not conspecific70) and lacks 15 enzymes that are broadly linked to aspects of urea, sulfate and thiamine metabolism. Given that the majority are also missing from various other target species, this may help to explain its unculturability. R. allomycis is unlikely to be axenically cultured because it is missing a large number of genes for critical pathways, losses that potentially cannot be corrected for as seen in microsporidia71.
Mycoparasitic fungi face unique challenges because they must parasitize a fungal host using fungal enzymes without disrupting their own cells. However, the specificity of this antagonism is not fully characterized72. Genomic analysis of the facultative mycoparasite Trichoderma spp. revealed several key features, including gene expansions and numerous antifungal-producing secondary metabolite gene clusters73. Nevertheless, many fungal lineages contain mycoparasites and there is no generalized analysis of the traits that are selected for in this ecological transition. We did not find any orthologous genes that were a signal of mycoparasitism across all taxa. In obligate mycoparasites, such as those studied here, we found a CAZyme-to-protease ratio more similar to that of animal pathogens rather than plant pathogens. An expansion of subtilases was observed among mycoparasitic Zoopagomycota, forming a group that was distinct from known subtilase families and among subtilases from other fungi. The GH19 chitinases are exclusively restricted to EDF lineages, which suggests that this gene family is ancestral but has been lost.
Having genomes from these EDF enabled us to uncover traits that were previously only described in Dikarya. We observed a vast expansion of secondary metabolism NRPS genes unique to D. cristalligena among other EDF. Similarly, we provide evidence for multiple hydrophobins in C. protostelioides, the only member of the Chytridiomycota known to form aerial hyphal-like spore-producing structures.
Here, we show that single-cell methods can dramatically expand our understanding of fungal biology. Given the large numbers of as-yet-unsequenced EDF genomes, many of which have never been cultured, these methods help to clarify the expectations for future assembly and functional annotation of genomes from those lineages. Furthermore, given the large numbers of uncultured Dikarya, these methods can be effectively applied across the phylogeny. As the scale of single-cell genomics increases13, we will be able to generate a full picture of the fungal tree of life that precisely reveals the complete extent of fungal diversity, regardless of the culturability of the individual taxa.
Methods
Strains and sample preparation
A dual culture of parasite R. allomycis CSF55 with its host fungus Allomyces sp. was established and used previously to sequence the genome of the parasite74. For this study, spore suspensions of R. allomycis were obtained under optimal conditions by washing the plates with a dilute Tween solution. An estimated 106–107 spores of the parasite with up to 5% of host spores were obtained. The sample was preserved in 10% sterile glycerol solution, shipped on dry ice and stored at −80 °C.
A dual culture of C. protostelioides ATCC 52028 with its host Sordaria was used to isolate parasitic zoospores at 2.5 × 106 per ml. The zoospore suspension was preserved in 10% dimethylsulfoxide with 10% FBS, shipped on dry ice and stored at −80 °C.
B. helicus was grown to a high density of cells through enrichment methods using spruce pollen in bog water. The sample was obtained from Perch Pond Fen near Old Town, Penobscot County, Maine, USA, in June 2014. This enrichment culture was filtered through a 40-μm mesh (removing pollen and sporangia) and concentrated by centrifugation to about 5 × 104 zoospores per ml. The sample was preserved in 10% glycerol, shipped on ice and stored at −80 °C.
M. bicuspidata was isolated from an infected population of the water flea Daphnia dentifera grown under laboratory conditions. D. dentifera samples were rinsed repeatedly with deionized water. Then, insect pins were used to puncture the carapace and a micropipette was used to collect haemolymph, which contained a mixture of M. bicuspidata yeast cells and ascospores. Cells were preserved in 10% glycerol at a concentration of 105 spores per ml and stored at −80 °C.
D. cristalligena RSA 468 was grown on V8 juice agar (1 small can of original V8 juice (5.5 oz, 163 ml), diluted to 1 l with deionized water, 3 g CaCO3 and 20 g agar) and cultured with Cokeromyces recurvatus. Spores were shipped in 10% sterile glycerol.
S. pseudoplumigaleata Benny S71-1 was grown on Mucor moelleri on 10% wheat germ agar (Wg10 (ref. 75)). Parasite hyphae and spores were shipped in 50% glycerol.
T. sphaerospora RSA 1356 was grown on V8 juice agar in dual culture with C. recurvatus and harvested from petri plates. The sample was stored in 50% glycerol at −80 °C.
P. cylindrospora RSA 2659 was cultivated on potato dextrose agar with the host Umbelopsis isabellina. The culture was grown on many petri dishes and the spores of both the fungus and the host were removed from the culture by washing the plates with 0.2% Triton X-100. An estimated 2.5 × 107 spores per ml of parasite with host were obtained and preserved in 10% glycerol at −80 °C.
All mycoparasites described above are considered obligate mycoparasites. For benchmarking purposes, we used non-single-cell approaches to sequence material gathered from genomic DNA extracted from enrichment cultures of C. protostelioides. We also took advantage of genomic resources from M. bicuspidata NRRL YB-4993, a parasite of brine shrimp70 related to but not considered conspecific with the Daphnia parasitic M. bicuspidata targeted in this study, and the published genome of R. allomycis74. These three genomic DNA isolates were used to compare with their respective single-cell genomes.
Single-cell genomics pipeline
The pipeline schema is shown in Supplementary Fig. 1. After sample collection, as described in the preceding section, individual cells were isolated using a one-step or two-step FACS (BD Influx Cell sorter) procedure. Target population enrichment in the original sample, determined by microscopy, dictated the use of one-step or two-step FACS. Cells were sorted into 384-well plates at counts of 1, 10, 30, 50 or 100 cells per well. Sorting accuracy was verified microscopically (Zeiss Axio Observer D1) and according to morphology.
Verified target cells were lysed using a combination of enzymatic and alkaline solutions. Briefly, cell lysis solutions were prepared with the following reagents: KOH dry pellets (reconstituted to 500 mM with nuclease-free water), 1 M dithiothreitol, and HCl (Stop Buffer) were obtained from the REPLI-g WGA Single-Cell Kit (150345, Qiagen); Tween 20 (P9416-100 ml, Sigma)); 0.5 M EDTA (AM92606, Ambion); proteinase K (P8107S, NEB); phenylmethylsulfonyl fluoride (532789-5 g, Sigma); and EGTA (E3889-10 g, Sigma). Genome amplification via multiple displacement amplification was achieved using either the REPLI-g WGA Single-Cell Kit (150345, Qiagen) according to the manufacturer’s instructions, or in-house using 10 mM dNTP (N0447L, NEB), 500 mM hexamers (37617009, IDT), phi29 polymerase (M029L, NEB), DMSO (D8418-50 ml, Sigma) and 10× buffer (400 mM Tris-HCl, pH 7.5 (AM9855, Ambion), 500 mM KCL (AM9640G, Ambion), 100 mM MgCl2 (AM9530, Ambion), 50 mM (NH4)2SO4 (AA4418-100 g, Sigma) and 20 mM dithiothreitol (P2325, Invitrogen)). We determined that, although the in-house approach is cheaper and yields better coverage, it is less practical for large-scale production as it requires several days of reagent cleaning prior to the reaction, whereas REPLI-g comes pre-purified and requires only a few hours of additional cleaning. Furthermore, downstream co-assembly of libraries reduces the problem of random bias.
As a quality-control step, amplified cells were 18S rDNA screened using the following primers: M13CRYPTO2-2F (5′-GTTTTCCCAGTCACGACCACAGGGAGGTAGTGACAG-3′), M13AU4v2 (5′-CAGGAAAAGCTATGACGCCTCACTAAGCCATTC-3′), M13DPD360FE (5′-GTTTTCCCAGTCACGACCGGAGARGGMGCMTGAGA-3′), M13DPD1492RE (5′-CAGGAAACAGCTATGACACCTTGTTACGRCTT-3′), M13SR1RFor (5′-GTTTTCCCAGTCACGACTACCTGGTTGATYCTGCCAGT-3′) and M13NS4Rev (5′-CAGGAAACAGCTATGACCTTCCGTCAATTCCTTTAAG-3′). Amplified 18S rDNA fragments were treated with ExoSap-IT (Thermo Fisher Scientific) and Sanger sequenced. Sequences were queried using BLAST against the National Center for Biotechnology Information non-redundant (NCBI nr) and Assembling the Fungal Tree Of Life (AFTOL)76 databases. Verified target wells were prepared using the TruSeq library method. Between three and seven libraries were pooled together and sequenced on one lane of Illumina HiSeq 1T v4-v6 2 × 150 bp. All pooled libraries belonged to the same species. Prior to establishing this protocol, we tested for crosstalk between TruSeq libraries on the MiSeq platform during pre-assembly library screening and found no such occurrence for the fungal single-cell pipeline. However, owing to previous observations of library crosstalk in the Joint Genome Institute (JGI) bacterial single-cell pipeline, the fungal single-cell pipeline decided on the cautionary same-species pooling until further examination was completed. Fungal single-cell pipeline TruSeq libraries were prepared on eight tube strips with individual caps using automated multichannel pipettes, which excludes plate-based robotic potential cross-contamination between samples.
In the fungal single-cell pipeline, there are several steps of rigorous filtering of potential contaminating sequences, at both the read level (pre-assembly) and the contig level (post-assembly). All Illumina reads were run through a filtering pipeline prior to assembly. BBDuk (https://sourceforge.net/projects/bbmap/) was used with the settings filterk = 27 and trimk = 27 to remove Illumina adapters, known Illumina artefacts, phiX and quality trim both ends to Q12. Resulting reads containing more than one ‘N’, with quality scores (before trimming) averaging <8 over the read or length under 40 bp after trimming were discarded. Remaining reads were mapped to a masked version of human hg19 with BBMap, discarding all hits over 93% identity. Reads mapping to mouse were also similarly discarded. Reads were also normalized to make the coverage more uniform, including removing reads that have very high coverage (>100). Individual libraries from each species were assembled with SPAdes77 (v2.6 or v3.0), using the ‘careful’ and ‘single-cell’ options, and k-mer sizes of 21, 33 or 55. After assembly, scaffolds of <2 kb in length were removed as, in our experience, they are phylogenetically ambiguous. Assembled contigs were also compared using BLAST to a set of contaminant databases: bacterial, mouse, human, feline and canine. Finally, we performed tetramer principle component analysis and removed any outlier contigs. This extra cautious read usage for assembly probably leads to reduced completeness and removes symbiotic occurrences but guarantees one species genome instead.
Supplementary Table 1 presents summaries of HiSeq libraries of variable number of cells for each organism. Bold libraries denote inclusion in co-assemblies. Underlined libraries denote individual annotation. To generate co-assemblies, reads from each library were extracted, pooled and co-assembled, again with SPAdes. The final co-assemblies were annotated using the JGI Annotation Pipeline78. For D. cristalligena, C. protostelioides, R. allomycis and M. bicuspidata, all individual libraries were additionally annotated. For P. cylindrospora, T. sphaerospora, S. pseudoplumigaleata and B. helicus, only the most complete (highest CEGMA score) 1-cell and 100-cell (1 cell and 50 cell for T. sphaerospora) individual libraries were additionally annotated. All final annotations were loaded into MycoCosm, the JGI Fungal Genomics Resource78, for public presentation and comparative analysis.
Annotation completeness
CEGMA26 as a general measure of completeness has been deprecated in favour of BUSCO79. However, because the underlying data for BUSCO relies heavily on Dikarya fungi, we have observed (Supplementary Fig. 9) that BUSCO dramatically underestimates the coverage of EDF lineages, particularly within the Chytridiomycota, Blastocladiomycota and Mucoromycota. As such, we continue to use CEGMA metrics for this study as it focuses primarily on EDF.
Heterozygous polymorphism discovery
Paired-end reads were aligned to assembled draft genomes using BWA80 (v0.7.12-r1044) using default parameters. Variants were identified using FreeBayes81 (v1.0.2-58-g054b257) with parameters ‘–pooled-continuous –min-coverage 5’. As non-haploid organisms have multiple genome copies per cell, increased counts of higher-frequency SNPs (allele frequency > 0.25) in their single cells are indicative of heterozygosity. Only SNPs with an allele frequency >25% were kept for further analysis. Comparisons of ploidy levels between species were performed on four single-cell libraries with the highest completeness and quality. K-mer graphs were plotted using the kmercountexact.sh script of the BBTools package (http://jgi.doe.gov/data-and-tools/bbtools/).
For intraspecific variation, reads from the eight D. cristalligena one-cell libraries were aligned to the D. cristalligena co-assembled genome using BBMap (v37.76) (https://sourceforge.net/projects/bbmap/). Variants were identified using FreeBayes81 (v1.1.0-54-g49413aa) with parameters ‘–-min-coverage 5 –F 0.01 –C 2 –no-mnps –no-complex’. SNPs with 100% disagreement with reference and 100% frequency were kept and mapped to genes for which all libraries had >90% coverage. A set visualization plot (Supplementary Fig. 2) was generated using the R implementation of the UpSet set visualization technique82 (https://cran.r-project.org/package=UpSetR).
Functional analyses
Subtilase cluster analysis was achieved using CLANS83 and a list of identified protein sequences described in Li et al.45. Metallopeptidase and hydrophobin trees were constructed using RAxML84 (v8.2.2; GAMMA + WAGF substitution model; 100 bootstrap replicates) and PFAM seed set sequences: PF02128 and PF01185 + PF06766, respectively. Trees were visualized using the Interactive Tree of Life (iTOL) website85 (http://itol.embl.de).
For phylogenetic analysis of identified NRPS genes, adenylation domains were mined from proteomes using HMMER86 (v3.1b2) with a hidden Markov model based on the fungal and bacterial domains identified by Bushley and Turgeon53. MUSCLE87 (v3.8.31) was used to align extracted domains with those used for the hidden Markov model creation and gaps were removed manually. RAxML84 (v8.2.8) was used for phylogenetic reconstruction using the gamma model of rate heterogeneity and the RTREV substitution matrix with 100 bootstrap replicates.
Phylogenetic analysis
All-v-all blastp88 (v2.2.26) was run using a cut-off of 1 × 10−5 on the predicted proteomes of the representative set of fungal and outgroup species provided in Supplementary Table 2. Clusters were predicted using MCL89 (v1.008) with an inflation value of 2. A python script identified clusters containing at most one gene copy per genome, allowing for up to 8 missing taxa per cluster, which resulted in 805 total clusters selected. Each cluster was aligned using MAFFT90 (v7.047) on each cluster, with a multiple sequence alignment algorithm detected automatically (using the –auto flag). Alignments were trimmed with Gblocks91 (v0.91b) and concatenated, followed by rapid bootstrap maximum likelihood tree building with RAxML84, 27,751 distinct alignment patterns, 100 bootstrap replicates and using the GAMMA + WAGF protein model. For all 29,255 positions in the alignment, the target genomes had a median of 14% missing, compared to the median of 2.5% missing of others.
Orthogroup reconstruction
Using MCL clustering89, orthologues were collected using a minimum of three taxa per orthogroup, ignoring single-cell C. protostelioides and R. allomycis in favour of the respective co-assemblies. The R package APE92 was used for ancestral-state reconstruction. Protein functions predicted using PRIAM93 were mapped to each internal node of the phylogenetic tree presented in Fig. 1. Gains and losses in single-cell lineages were considered as relative to the ancestral fungal node (Supplementary Fig. 3): D. cristalligena shows noticeable gains in all high-level metabolic categories, whereas B. helicus has mainly losses.
Metabolic reconstruction
Enzyme classifications were predicted for all 60 proteomes in Supplementary Table 2 using PRIAM93. A subset of 24 fungal, non-single-cell proteomes were defined as ‘free living’ based on culturability. The presence/absence counts were converted to a binary presence/absence matrix for each enzyme classification found in each species and filtered according to ‘found in free-living’ threshold of present in 18 out of 24 (75%) free-living species and absent in at least 5 single-cell species. ‘Single cell’ includes C. protostelioides and R. allomycis culture isolate genomes in place of those respective single-cell genomes. The resulting pattern of conserved losses among ‘core’ metabolic pathways was summarized and displayed using IPath94.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The co-assembled genomes and annotations of the target species are available through MycoCosm (https://genome.jgi.doe.gov/fungi) and Genbank using the following MycoCosm URLs and NCBI accessions, respectively: R. allomycis CSF55 single cell (https://genome.jgi.doe.gov/Rozal_SC1; QUVT00000000), B. helicus Perch Fen single cell (https://genome.jgi.doe.gov/Blyhe1; QPFV00000000), C. protostelioides ATCC 52028 single cell (https://genome.jgi.doe.gov/Caupr_SCcomb; QUVS00000000), D. cristalligena RSA 468 single cell (https://genome.jgi.doe.gov/DimcrSC1; QRFA00000000), P. cylindrospora RSA 2659 single cell (https://genome.jgi.doe.gov/Pipcy3_1; QPFT00000000), T. sphaerospora RSA 1356 single cell (https://genome.jgi.doe.gov/Thasp1; QUVU00000000), S. pseudoplumigaleata Benny S71-1 single cell (https://genome.jgi.doe.gov/Synps1; QUVV00000000) and M. bicuspidata single cell (https://genome.jgi.doe.gov/Metbi_SCcomb; QUVR00000000). The whole-genome sequence for the non-single-cell isolate C. protostelioides ATCC 52028 is available through MycoCosm (https://genome.jgi.doe.gov/Caupr1) and Genbank (QAJV00000000). The whole-genome sequences for the non-single-cell isolate of R. allomycis CSF55 was not determined in this study and is available through MycoCosm (https://genome.jgi.doe.gov/Rozal1_1) and Genbank (ATJD00000000). The genome sequence for the non-single-cell M. bicuspidata NRRL YB-4993 was also not determined in this study and is available through MycoCosm (https://genome.jgi.doe.gov/Metbi1) and Genbank (LXTC00000000).
References
Berbee, M. L., James, T. Y. & Strullu-Derrien, C. Early diverging fungi: diversity and impact at the dawn of terrestrial life. Annu. Rev. Microbiol. 71, 41–59 (2017).
Grigoriev, I. V. et al. Fueling the future with fungal genomics. Mycology 2, 192–209 (2011).
Hibbett, D. S. et al. Progress in molecular and morphological taxon discovery in fungi and options for formal classification of environmental sequences. Fungal Biol. Rev. 25, 38–47 (2011).
Shirouzu, T., Uno, K., Hosaka, K. & Hosoya, T. Early-diverging wood-decaying fungi detected using three complementary sampling methods. Mol. Phylogenet. Evol. 98, 11–20 (2016).
Tedersoo, L., Bahram, M., Puusepp, R., Nilsson, R. H. & James, T. Y. Novel soil-inhabiting clades fill gaps in the fungal tree of life. Microbiome 5, 42 (2017).
Grossart, H.-P., Wurzbacher, C., James, T. Y. & Kagami, M. Discovery of dark matter fungi in aquatic ecosystems demands a reappraisal of the phylogeny and ecology of zoosporic fungi. Fungal Ecol. 19, 28–38 (2016).
Jones, M. D. M. et al. Discovery of novel intermediate forms redefines the fungal tree of life. Nature 474, 200–203 (2011).
Freeman, K. R. et al. Evidence that chytrids dominate fungal communities in high-elevation soils. Proc. Natl Acad. Sci. USA 106, 18315–18320 (2009).
Jobard, M., Rasconi, S., Solinhac, L., Cauchie, H.-M. & Sime-Ngando, T. Molecular and morphological diversity of fungi and the associated functions in three European nearby lakes. Environ. Microbiol. 14, 2480–2494 (2012).
Chang, Y. et al. Phylogenomic analyses indicate that early fungi evolved digesting cell walls of algal ancestors of land plants. Genome Biol. Evol. 7, 1590–1601 (2015).
Blainey, P. C. The future is now: single-cell genomics of bacteria and archaea. FEMS Microbiol. Rev. 37, 407–427 (2013).
Gawad, C., Koh, W. & Quake, S. R. Single-cell genome sequencing: current state of the science. Nat. Rev. Genet. 17, 175–188 (2016).
Woyke, T., Doud, D. F. R. & Schulz, F. The trajectory of microbial single-cell sequencing. Nat. Methods 14, 1045–1054 (2017).
Rinke, C. et al. Obtaining genomes from uncultivated environmental microorganisms using FACS-based single-cell genomics. Nat. Protoc. 9, 1038–1048 (2014).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Yoon, H. S. et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists. Science 332, 714–717 (2011).
Roy, R. S. et al. Single cell genome analysis of an uncultured heterotrophic stramenopile. Sci. Rep. 4, 4780 (2014).
López-Escardó, D. et al. Evaluation of single-cell genomics to address evolutionary questions using three SAGs of the choanoflagellate Monosiga brevicollis. Sci. Rep. 7, 11025 (2017).
Strassert, J. F. H. et al. Single cell genomics of uncultured marine alveolates shows paraphyly of basal dinoflagellates. ISME J. 12, 304–308 (2018).
Lin, K. et al. Single nucleus genome sequencing reveals high similarity among nuclei of an endomycorrhizal fungus. PLoS Genet. 10, e1004078 (2014).
Lan, F., Demaree, B., Ahmed, N. & Abate, A. R. Single-cell genome sequencing at ultra-high-throughput with microfluidic droplet barcoding. Nat. Biotechnol. 35, 640–646 (2017).
Spatafora, J. W. et al. A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108, 1028–1046 (2016).
Jeffries, P. & Young, T. W. K. Interfungal Parasitic Relationships (CAB International, Wallingford, 1994).
Blainey, P. C., Mosier, A. C., Potanina, A., Francis, C. A. & Quake, S. R. Genome of a low-salinity ammonia-oxidizing archaeon determined by single-cell and metagenomic analysis. PLoS ONE 6, e16626 (2011).
Dodsworth, J. A. et al. Single-cell and metagenomic analyses indicate a fermentative and saccharolytic lifestyle for members of the OP9 lineage. Nat. Commun. 4, 1854 (2013).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Emerson, R. Current trends of experimental research on the aquatic Phycomycetes. Annu. Rev. Microbiol. 4, 169–200 (1950).
Morehouse, E. A. et al. Multilocus sequence typing suggests the chytrid pathogen of amphibians is a recently emerged clone. Mol. Ecol. 12, 395–403 (2003).
Pelin, A., Selman, M., Aris-Brosou, S., Farinelli, L. & Corradi, N. Genome analyses suggest the presence of polyploidy and recent human-driven expansions in eight global populations of the honeybee pathogen Nosema ceranae. Environ. Microbiol. 17, 4443–4458 (2015).
Rogers, M. B. et al. Chromosome and gene copy number variation allow major structural change between species and strains of Leishmania. Genome Res. 21, 2129–2142 (2011).
Ropars, J. et al. Evidence for the sexual origin of heterokaryosis in arbuscular mycorrhizal fungi. Nat. Microbiol. 1, 16033 (2016).
Lodato, M. A. et al. Somatic mutation in single human neurons tracks developmental and transcriptional history. Science 350, 94–98 (2015).
Omsland, A., Hackstadt, T. & Heinzen, R. A. Bringing culture to the uncultured: Coxiella burnetii and lessons for obligate intracellular bacterial pathogens. PLoS Pathog. 9, e1003540 (2013).
Stevens, L. & Winther, M. D. Spermine, spermidine and putrescine in fungal development. Adv. Microb. Physiol. 19, 63–148 (1979).
Valdés-Santiago, L., Cervantes-Chávez, J. A., León-Ramírez, C. G. & Ruiz-Herrera, J. Polyamine metabolism in fungi with emphasis on phytopathogenic species. J. Amino Acids 2012, 837932 (2012).
Jiménez-Bremont, J. F., Ruiz-Herrera, J. & Dominguez, A. Disruption of gene YlODC reveals absolute requirement of polyamines for mycelial development in Yarrowia lipolytica. FEMS Yeast Res. 1, 195–204 (2001).
Valdés-Santiago, L., Cervantes-Chávez, J. A. & Ruiz-Herrera, J. Ustilago maydis spermidine synthase is encoded by a chimeric gene, required for morphogenesis, and indispensable for survival in the host. FEMS Yeast Res. 9, 923–935 (2009).
Paietta, J. V. in Biochemistry and Molecular Biology. The Mycota (A Comprehensive Treatise on Fungi as Experimental Systems for Basic and Applied Research) Vol. 3 (eds Brambl, R. & Marzluf, G. A.) 369–383 (Springer, Berlin, Heidelberg, 2004).
Maruyama, J.-I. & Kitamoto, K. Expanding functional repertoires of fungal peroxisomes: contribution to growth and survival processes. Front. Physiol. 4, 177 (2013).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Rawlings, N. D., Barrett, A. J. & Finn, R. Twenty years of the MEROPS database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 44, D343–D350 (2016).
Siezen, R. J. & Leunissen, J. A. Subtilases: the superfamily of subtilisin-like serine proteases. Protein Sci. 6, 501–523 (1997).
Hu, G. & Leger, R. J. S. A phylogenomic approach to reconstructing the diversification of serine proteases in fungi. J. Evol. Biol. 17, 1204–1214 (2004).
Muszewska, A., Taylor, J. W., Szczesny, P. & Grynberg, M. Independent subtilases expansions in fungi associated with animals. Mol. Biol. Evol. 28, 3395–3404 (2011).
Li, J., Gu, F., Wu, R., Yang, J. & Zhang, K.-Q. Phylogenomic evolutionary surveys of subtilase superfamily genes in fungi. Sci. Rep. 7, 45456 (2017).
Brouta, F. et al. Purification and characterization of a 43.5 kDa keratinolytic metalloprotease from Microsporum canis. Med. Mycol. 39, 269–275 (2001).
Rosenblum, E. B., Stajich, J. E., Maddox, N. & Eisen, M. B. Global gene expression profiles for life stages of the deadly amphibian pathogen Batrachochytrium dendrobatidis. Proc. Natl Acad. Sci. USA 105, 17034–17039 (2008).
Flach, J., Pilet, P. E. & Jollès, P. What’s new in chitinase research? Experientia 48, 701–716 (1992).
Duo-Chuan, L. Review of fungal chitinases. Mycopathologia 161, 345–360 (2006).
Latgé, J.-P. The cell wall: a carbohydrate armour for the fungal cell. Mol. Microbiol. 66, 279–290 (2007).
Hemsworth, G. R., Henrissat, B., Davies, G. J. & Walton, P. H. Discovery and characterization of a new family of lytic polysaccharide monooxygenases. Nat. Chem. Biol. 10, 122–126 (2014).
Henrissat, B. Classification of chitinases modules. EXS 87, 137–156 (1999).
Bushley, K. E. & Turgeon, B. G. Phylogenomics reveals subfamilies of fungal nonribosomal peptide synthetases and their evolutionary relationships. BMC Evol. Biol. 10, 26 (2010).
Howell, C. R., Stipanovic, R. D. & Lumsden, R. D. Antibiotic production by strains of Gliocladium virens and its relation to the biocontrol of cotton seedling diseases. Biocontrol Sci. Technol. 3, 435–441 (1993).
Wösten, H. A. Hydrophobins: multipurpose proteins. Annu. Rev. Microbiol. 55, 625–646 (2001).
de Vries, O. M. H., Peter Fekkes, M., Wösten, H. A. B. & Wessels, J. G. H. Insoluble hydrophobin complexes in the walls of Schizophyllum commune and other filamentous fungi. Arch. Microbiol. 159, 330–335 (1993).
Olive, L. S. Caulochytrium protostelioides sp. nov., a new chytrid with aerial spporangia. Am. J. Bot. 67, 568–574 (1980).
Benjamin, R. K. The Merosporangium. Mycologia 58, 1–42 (1966).
White, M. M. et al. Phylogeny of the Zygomycota based on nuclear ribosomal sequence data. Mycologia 98, 872–884 (2006).
James, T. Y. et al. A molecular phylogeny of the flagellated fungi (Chytridiomycota) and description of a new phylum (Blastocladiomycota). Mycologia 98, 860–871 (2006).
Barr, D. J. S. An outline for the reclassification of the Chytridiales, and for a new order, the Spizellomycetales. Can. J. Bot. 58, 2380–2394 (1980).
Tsai, I. J., Bensasson, D., Burt, A. & Koufopanou, V. Population genomics of the wild yeast Saccharomyces paradoxus: quantifying the life cycle. Proc. Natl Acad. Sci. USA 105, 4957–4962 (2008).
Saikkonen, K., Young, C. A., Helander, M. & Schardl, C. L. Endophytic Epichloë species and their grass hosts: from evolution to applications. Plant Mol. Biol. 90, 665–675 (2016).
Schloegel, L. M. et al. Novel, panzootic and hybrid genotypes of amphibian chytridiomycosis associated with the bullfrog trade. Mol. Ecol. 21, 5162–5177 (2012).
Rosenblum, E. B. et al. Complex history of the amphibian-killing chytrid fungus revealed with genome resequencing data. Proc. Natl Acad. Sci. USA 110, 9385–9390 (2013).
Quandt, C. A.et al. The genome of an intranuclear parasite, Paramicrosporidium saccamoebae, reveals alternative adaptations to obligate intracellular parasitism. eLife 6, e29594 (2017).
Ellis, J. J. On growing Syncephalis in pure culture. Mycologia 58, 465–469 (1966).
Lazarus, K. L., Benny, G. L., Ho, H.-M. & Smith, M. E. Phylogenetic systematics of Syncephalis (Zoopagales, Zoopagomycotina), a genus of ubiquitous mycoparasites. Mycologia 109, 333–349 (2017).
Staggs, C. G., Sealey, W. M., McCabe, B. J., Teague, A. M. & Mock, D. M. Determination of the biotin content of select foods using accurate and sensitive HPLC/avidin binding. J. Food. Compost. Anal. 17, 767–776 (2004).
Riley, R. et al. Comparative genomics of biotechnologically important yeasts. Proc. Natl Acad. Sci. USA 113, 9882–9887 (2016).
Moran, N. A. & Bennett, G. M. The tiniest tiny genomes. Annu. Rev. Microbiol. 68, 195–215 (2014).
Druzhinina, I. S. et al. Trichoderma: the genomics of opportunistic success. Nat. Rev. Microbiol. 9, 749–759 (2011).
Mukherjee, P. K., Horwitz, B. A. & Kenerley, C. M. Secondary metabolism in Trichoderma—a genomic perspective. Microbiology 158, 35–45 (2012).
James, T. Y. et al. Shared signatures of parasitism and phylogenomics unite Cryptomycota and Microsporidia. Curr. Biol. 23, 1548–1553 (2013).
Benny, G. L., Ho, H.-M., Lazarus, K. & Smith, M. E. Five new species of the obligate mycoparasite Syncephalis (Zoopagales, Zoopagomycotina) from soil. Mycologia 108, 1114–1129 (2016).
Lutzoni, F. et al. Assembling the fungal tree of life: progress, classification, and evolution of subcellular traits. Am. J. Bot. 91, 1446–1480 (2004).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42, D699–D704 (2014).
Simão, F. A. et al. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/abs/1207.3907 (2012).
Lex, A., Gehlenborg, N., Strobelt, H., Vuillemot, R. & Pfister, H. UpSet: visualization of intersecting sets. IEEE Trans. Vis. Comput. Graph. 20, 1983–1992 (2014).
Frickey, T. & Lupas, A. CLANS: a Java application for visualizing protein families based on pairwise similarity. Bioinformatics 20, 3702–3704 (2004).
Stamatakis, A. et al. Exploring new search algorithms and hardware for phylogenetics: RAxML meets the IBM cell. J. VLSI Signal Process. Syst. Signal Image Video Technol. 48, 271–286 (2007).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Edgar, R. C., Drive, R. M. & Valley, M. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic Local Alignment Search Tool. J. Mol. Biol. 215, 403–410 (1990).
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Claudel-Renard, C., Chevalet, C., Faraut, T. & Kahn, D. Enzyme-specific profiles for genome annotation: PRIAM. Nucleic Acids Res. 31, 6633–6639 (2003).
Yamada, T., Letunic, I., Okuda, S., Kanehisa, M. & Bork, P. iPath2.0: interactive pathway explorer. Nucleic Acids Res. 39, W412–W415 (2011).
Acknowledgements
The work conducted by the US Department of Energy JGI, a DOE Office of Science User Facility, is supported by the Office of Science of the US Department of Energy under contract no. DE-AC02-05CH11231. The authors thank M. Duffy (University of Michigan, MI, USA) for providing the sample of M. bicuspidata, J. Longcore (University of Maine, ME, USA) for B. helicus and N. Ivanova for metabolic biochemistry insights. S.R.A., I.V.G., T.Y.J. and C.A.Q. were supported by the NSF grant DEB-1354625. M.E.S., T.Y.J., N.K.R. and G.L.B. were supported by the NSF grant DEB-1441677 (Genealogy of Life (ZyGoLife) the conundrum of Kingdom Fungi).
Author information
Authors and Affiliations
Contributions
S.R.A., D.C., C.A.Q., T.Y.J. and A.P. wrote the manuscript with input from T.W., I.V.G., M.E.S. and N.K.R. D.C. performed the wet-bench protocol optimization and sequencing under supervision of J.-F.C. A.C. and B.A. performed sequencing and assembly, quality-control protocols and provided substantial technical input. S.R.A. and A.S. annotated the genomes. C.A.Q., S.R.A., A.P., A.S. and B.H. performed the comparative analyses. N.K.R. performed the media supplement axenic culture experiments. G.L.B., M.E.S. and T.Y.J. provided the biological samples. T.Y.J. and I.V.G. designed and coordinated the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Notes, Supplementary References, Supplementary Tables 5–8, Supplementary Figures 1–9 and Supplementary Dataset legends.
Supplementary Table 1
Select statistics for all sequenced libraries from each target species.
Supplementary Table 2
Descriptions and sources for fungal and outgroup proteomes used in comparative analyses.
Supplementary Table 3
Summary of tree phylogenies generated using select individual libraries.
Supplementary Table 4
Abundance counts of different chitinase and related CAZyme families among the target genomes.
Dataset 1
Contains 6 phylogenetic trees, each using a different library of Caulochytrium protostelioides: one 1-cell, one 10-cell, and four 100-cell libraries.
Dataset 2
Contains 14 phylogenetic trees, each using a different library of Dimargaris cristalligena: eight 1-cell, one 50-cell, and five 100-cell libraries.
Dataset 3
Contains 5 phylogenetic trees, each using a different library of Metschnikowia bicuspidata: three 1-cell, one 10-cell, and one 100-cell libraries.
Dataset 4
Contains 14 phylogenetic trees, each using a different library of Rozella allomycis: eleven 1-cell and three 100-cell libraries.
Dataset 5
Contains 1 phylogenetic tree, built using the co-assemblies and presented graphically as Figure 1.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahrendt, S.R., Quandt, C.A., Ciobanu, D. et al. Leveraging single-cell genomics to expand the fungal tree of life. Nat Microbiol 3, 1417–1428 (2018). https://doi.org/10.1038/s41564-018-0261-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0261-0
This article is cited by
-
Tools for microbial single-cell genomics for obtaining uncultured microbial genomes
Biophysical Reviews (2024)
-
Large-scale identification of genes involved in septal pore plugging in multicellular fungi
Nature Communications (2023)
-
Transitions of foliar mycobiota community and transcriptome in response to pathogenic conifer needle interactions
Scientific Reports (2022)
-
Genomic insights into the phylogeny and biomass-degrading enzymes of rumen ciliates
The ISME Journal (2022)
-
Ciliary transition zone evolution and the root of the eukaryote tree: implications for opisthokont origin and classification of kingdoms Protozoa, Plantae, and Fungi
Protoplasma (2022)