Abstract
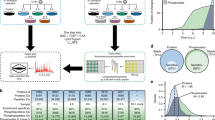
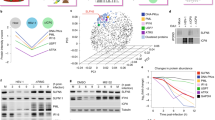
To orchestrate context-dependent signalling programmes, poxviruses encode two dual-specificity enzymes, the F10 kinase and the H1 phosphatase. These signalling mediators are essential for poxvirus production, yet their substrate profiles and systems-level functions remain enigmatic. Using a phosphoproteomic screen of cells infected with wild-type, F10 and H1 mutant vaccinia viruses, we systematically defined the viral signalling network controlled by these enzymes. Quantitative cross-comparison revealed 33 F10 and/or H1 phosphosites within 17 viral proteins. Using this proteotype dataset to inform genotype–phenotype relationships, we found that H1-deficient virions harbour a hidden hypercleavage phenotype driven by reversible phosphorylation of the virus protease I7 (S134). Quantitative phosphoproteomic profiling further revealed that the phosphorylation-dependent activity of the viral early transcription factor, A7 (Y367), underlies the transcription-deficient phenotype of H1 mutant virions. Together, these results highlight the utility of combining quantitative proteotype screens with mutant viruses to uncover proteotype–phenotype–genotype relationships that are masked by classical genetic studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Biedenkopf, N., Lier, C. & Becker, S. Dynamic phosphorylation of VP30 is essential for Ebola virus life cycle. J. Virol. 90, 4914–4925 (2016).
Mondal, A., Potts, G. K., Dawson, A. R., Coon, J. J. & Mehle, A. Phosphorylation at the homotypic interface regulates nucleoprotein oligomerization and assembly of the influenza virus replication machinery. PLoS Pathog. 11, e1004826 (2015).
Zhao, X. et al. Phosphorylation of Beet black scorch virus coat protein by PKA is required for assembly and stability of virus particles. Sci. Rep. 5, 11585 (2015).
Sharma, K. et al. Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep. 8, 1583–1594 (2014).
Wojcechowskyj, J. A. et al. Quantitative phosphoproteomics reveals extensive cellular reprogramming during HIV-1 entry. Cell Host Microbe 13, 613–623 (2013).
Kato, A. et al. Herpes simplex virus 1 protein kinase Us3 phosphorylates viral dUTPase and regulates its catalytic activity in infected cells. J. Virol. 88, 655–666 (2014).
Humphrey, S. J., Azimifar, S. B. & Mann, M. High-throughput phosphoproteomics reveals in vivo insulin signaling dynamics. Nat. Biotechnol. 33, 990–995 (2015).
Moss, B. in Fields Virology Vol. 2 (eds Estes, M. K. et al.) 2906–2945 (Lippincott Williams & Wilkins, Philadelphia, 2007).
Resch, W., Hixson, K. K., Moore, R. J., Lipton, M. S. & Moss, B. Protein composition of the vaccinia virus mature virion. Virology 358, 233–247 (2007).
Chung, C.-S. et al. Vaccinia virus proteome: identification of proteins in vaccinia virus intracellular mature virion particles. J. Virol. 80, 2127–2140 (2006).
Condit, R. C., Moussatche, N. & Traktman, P. In a nutshell: structure and assembly of the vaccinia virion. Adv. Virus Res. 66, 31–124 (2006).
Lin, S. & Broyles, S. S. Vaccinia protein kinase 2: a second essential serine/threonine protein kinase encoded by vaccinia virus. Proc. Natl Acad. Sci. USA. 91, 7653–7657 (1994).
Wang, S. & Shuman, S. Vaccinia virus morphogenesis is blocked by temperature-sensitive mutations in the F10 gene, which encodes protein kinase 2. J. Virol. 69, 6376–6388 (1995).
Byrd, C. M., Bolken, T. C. & Hruby, D. E. The vaccinia virus I7L gene product is the core protein proteinase. J. Virol. 76, 8973–8976 (2002).
Ericsson, M. et al. Characterization of ts 16, a temperature-sensitive mutant of vaccinia virus. J. Virol. 69, 7072–7086 (1995).
Traktman, P., Caligiuri, A., Jesty, S. A., Liu, K. & Sankar, U. Temperature-sensitive mutants with lesions in the vaccinia virus F10 kinase undergo arrest at the earliest stage of virion morphogenesis. J. Virol. 69, 6581–6587 (1995).
Hedengren-Olcott, M., Byrd, C. M., Watson, J. & Hruby, D. E. The vaccinia virus G1L putative metalloproteinase is essential for viral replication in vivo. J. Virol. 78, 9947–9953 (2004).
Ansarah-Sobrinho, C. & Moss, B. Role of the I7 protein in proteolytic processing of vaccinia virus membrane and core components. J. Virol. 78, 6335–6343 (2004).
Ansarah-Sobrinho, C. & Moss, B. Vaccinia virus G1 protein, a predicted metalloprotease, is essential for morphogenesis of infectious virions but not for cleavage of major core proteins. J. Virol. 78, 6855–6863 (2004).
Senkevich, T. G., White, C. L., Koonin, E. V. & Moss, B. Complete pathway for protein disulfide bond formation encoded by poxviruses. Proc. Natl Acad. Sci. USA. 99, 6667–6672 (2002).
Guan, K. L., Broyles, S. S. & Dixon, J. E. A Tyr/Ser protein phosphatase encoded by vaccinia virus. Nature 350, 359–362 (1991).
Liu, K., Lemon, B. & Traktman, P. The dual-specificity phosphatase encoded by vaccinia virus, VH1, is essential for viral transcription in vivo and in vitro. J. Virol. 69, 7823–7834 (1995).
Mercer, J. & Traktman, P. Genetic and cell biological characterization of the vaccinia virus A30 and G7 phosphoproteins. J. Virol. 79, 7146–7161 (2005).
Derrien, M., Punjabi, A., Khanna, M., Grubisha, O. & Traktman, P. Tyrosine phosphorylation of A17 during vaccinia virus infection: Involvement of the H1 phosphatase and the F10 kinase. J. Virol. 73, 7287–7296 (1999).
Betakova, T., Wolffe, E. J. & Moss, B. Regulation of vaccinia virus morphogenesis: phosphorylation of the A14L and A17L membrane proteins and C-terminal truncation of the A17L protein are dependent on the F10L kinase. J. Virol. 73, 3534–3543 (1999).
Szajner, P., Weisberg, A. S. & Moss, B. Evidence for an essential catalytic role of the F10 protein kinase in vaccinia virus morphogenesis. J. Virol. 78, 257–265 (2004).
Assarsson, E. et al. Kinetic analysis of a complete poxvirus transcriptome reveals an immediate-early class of genes. Proc. Natl Acad. Sci. USA. 105, 2140–2145 (2008).
Traktman, P. et al. Elucidating the essential role of the A14 phosphoprotein in vaccinia virus morphogenesis: construction and characterization of a tetracycline-inducible recombinant. J. Virol. 74, 3682–3695 (2000).
Mercer, J. & Traktman, P. Investigation of structural and functional motifs within the vaccinia virus A14 phosphoprotein, an essential component of the virion membrane. J. Virol. 77, 8857–8871 (2003).
Wu, Y. et al. Multilayered genetic and omics dissection of mitochondrial activity in a mouse reference population. Cell 158, 1415–1430 (2014).
Ulaeto, D., Grosenbach, D. & Hruby, D. E. The vaccinia virus 4c and A-type inclusion proteins are specific markers for the intracellular mature virus particle. J. Virol. 70, 3372–3377 (1996).
Matson, J., Chou, W., Ngo, T. & Gershon, P. D. Static and dynamic protein phosphorylation in the vaccinia virion. Virology 452-453, 310–323 (2014).
Byrd, C. M., Bolken, T. C. & Hruby, D. E. Molecular dissection of the vaccinia virus I7L core protein proteinase. J. Virol. 77, 11279–11283 (2003).
Condit, R. C., Motyczka, A. & Spizz, G. Isolation, characterization, and physical mapping of temperature-sensitive mutants of vaccinia virus. Virology 128, 429–443 (1983).
Pengue, G., Caputo, A., Rossi, C., Barbanti-Brodano, G. & Lania, L. Transcriptional silencing of human immunodeficiency virus type 1 long terminal repeat-driven gene expression by the Krüppel-associated box repressor domain targeted to the transactivating response element. J. Virol. 69, 6577–6580 (1995).
Punjabi, A. & Traktman, P. Cell biological and functional characterization of the vaccinia virus F10 kinase: implications for the mechanism of virion morphogenesis. J. Virol. 79, 2171–2190 (2005).
Grimley, P. M., Rosenblum, E. N., Mims, S. J. & Moss, B. Interruption by rifampin of an early stage in vaccinia virus morphogenesis: accumulation of membranes which are precursors of virus envelopes. J. Virol. 6, 519–533 (1970).
Unger, B. & Traktman, P. Vaccinia virus morphogenesis: A13 phosphoprotein is required for assembly of mature virions. J. Virol. 78, 8885–8901 (2004).
Rodriguez, D., Rodriguez, J. R. & Esteban, M. The vaccinia virus 14-kilodalton fusion protein forms a stable complex with the processed protein encoded by the vaccinia virus A17L gene. J. Virol. 67, 3435–3440 (1993).
Vanslyke, J. K., Whitehead, S. S., Wilson, E. M. & Hruby, D. E. The multistep proteolytic maturation pathway utilized by vaccinia virus P4a protein: a degenerate conserved cleavage motif within core proteins. Virology 183, 467–478 (1991).
Sarov, I. & Joklik, W. K. Studies on the nature and location of the capsid polypeptides of vaccinia virions. Virology 50, 579–592 (1972).
VanSlyke, J. K., Franke, C. A. & Hruby, D. E. Proteolytic maturation of vaccinia virus core proteins: identification of a conserved motif at the N termini of the 4b and 25K virion proteins. J. Gen. Virol. 72, 411–416 (1991).
Whitehead, S. S. & Hruby, D. E. Differential utilization of a conserved motif for the proteolytic maturation of vaccinia virus proteins. Virology 200, 154–161 (1994).
Yang, W. P., Kao, S. Y. & Bauer, W. R. Biosynthesis and post-translational cleavage of vaccinia virus structural protein VP8. Virology 167, 585–590 (1988).
Bisht, H., Weisberg, A. S., Szajner, P. & Moss, B. Assembly and disassembly of the capsid-like external scaffold of immature virions during vaccinia virus morphogenesis. J. Virol. 83, 9140–9150 (2009).
Szajner, P., Weisberg, A. S., Lebowitz, J., Heuser, J. & Moss, B. External scaffold of spherical immature poxvirus particles is made of protein trimers, forming a honeycomb lattice. J. Cell Biol. 170, 971–981 (2005).
Unger, B., Mercer, J., Boyle, K. A. & Traktman, P. Biogenesis of the vaccinia virus membrane: genetic and ultrastructural analysis of the contributions of the A14 and A17 proteins. J. Virol. 87, 1083–1097 (2013).
Gross, C. H. & Shuman, S. Vaccinia virions lacking the RNA helicase nucleoside triphosphate phosphohydrolase II are defective in early transcription. J. Virol. 70, 8549–8557 (1996).
Broyles, S. S., Yuen, L., Shuman, S. & Moss, B. Purification of a factor required for transcription of vaccinia virus early genes. J. Biol. Chem. 263, 10754–10760 (1988).
Hu, X., Wolffe, E. J., Weisberg, A. S., Carroll, L. J. & Moss, B. Repression of the A8L gene, encoding the early transcription factor 82-kilodalton subunit, inhibits morphogenesis of vaccinia virions. J. Virol. 72, 104–112 (1998).
Resch, W. & Moss, B. The conserved poxvirus L3 virion protein is required for transcription of vaccinia virus early genes. J. Virol. 79, 14719–14729 (2005).
Yang, Z. & Moss, B. Interaction of the vaccinia virus RNA polymerase-associated 94-kilodalton protein with the early transcription factor. J. Virol. 83, 12018–12026 (2009).
Hagen, C. J., Titong, A., Sarnoski, E. A. & Verardi, P. H. Antibiotic-dependent expression of early transcription factor subunits leads to stringent control of vaccinia virus replication. Virus Res. 181, 43–52 (2014).
Hu, X., Carroll, L. J., Wolffe, E. J. & Moss, B. De novo synthesis of the early transcription factor 70-kilodalton subunit is required for morphogenesis of vaccinia virions. J. Virol. 70, 7669–7677 (1996).
Hutchinson, E. C. et al. Mapping the phosphoproteome of influenza A and B viruses by mass spectrometry. PLoS Pathog. 8, e1002993 (2012).
Bergström Lind, S. et al. The phosphoproteome of the adenovirus type 2 virion. Virology 433, 253–261 (2012).
Ngo, T., Mirzakhanyan, Y., Moussatche, N. & Gershon, P. D. Protein primary structure of the vaccinia virion at increased resolution. J. Virol. 90, 9905–9919 (2016).
Schmidt, F. I., Bleck, C. K. E., Helenius, A. & Mercer, J. Vaccinia extracellular virions enter cells by macropinocytosis and acid-activated membrane rupture. EMBO J. 30, 3647–3661 (2011).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Duan, G., Li, X. & Köhn, M. The human DEPhOsphorylation database DEPOD: a 2015 update. Nucleic Acids Res. 43, D531–D535 (2015).
Schmidt, F. I. et al. Vaccinia virus entry is followed by core activation and proteasome-mediated release of the immunomodulatory effector VH1 from lateral bodies. Cell Rep. 4, 464–476 (2013).
Kilcher, S. et al. siRNA screen of early poxvirus genes identifies the AAA + ATPase D5 as the virus genome-uncoating factor. Cell Host Microbe 15, 103–112 (2014).
Mercer, J. & Helenius, A. Vaccinia virus uses macropinocytosis and apoptotic mimicry to enter host cells. Science 320, 531–535 (2008).
Bodenmiller, B. & Aebersold, R. Quantitative analysis of protein phosphorylation on a system-wide scale by mass spectrometry-based proteomics. Methods Enzymol. 470, 317–334 (2010).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Choi, M. et al. MSstats: an R package for statistical analysis of quantitative mass spectrometry-based proteomic experiments. Bioinformatics 30, 2524–2526 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57, 289–300 (1995).
UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015).
Deutsch, E. W. et al. Trans-Proteomic Pipeline, a standardized data processing pipeline for large-scale reproducible proteomics informatics. Proteomics Clin. Appl. 9, 745–754 (2015).
Schilling, B. et al. Platform-independent and label-free quantitation of proteomic data using MS1 extracted ion chromatograms in Skyline: application to protein acetylation and phosphorylation. Mol. Cell. Proteom. 11, 202–214 (2012).
Bruderer, R. et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen treated 3D liver microtissues. Mol. Cell. Proteom. 14, 1400–1410 (2015).
Bruderer, R. et al. Optimization of experimental parameters in data-independent mass spectrometry significantly increases depth and reproducibility of results. Mol. Cell. Proteom. 16, 2296–2309 (2017).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 16, 2296–72 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Fathi, Z. & Condit, R. C. Phenotypic characterization of a vaccinia virus temperature-sensitive complementation group affecting a virion component. Virology 181, 273–276 (1991).
Acknowledgements
We would like to thank P. Traktman, R. Condit, B. Moss and P. H. Verardi for generously donating VACV mutants for this study. We greatly acknowledge A. Frei, S. Götze and A. Leitner for maintenance of the mass spectrometers. We are grateful to all members of the Mercer and Wollscheid laboratories for critical comments and suggestions throughout this project. This work was supported by the Swiss National Science Foundation (31003A_160259 to B.W.) and the InfectX project from the Swiss Initiative in Systems Biology SystemsX.ch (to B.W.), the MRC Programme Grant (MC_UU12018/7) (J.M.), the European Research Council (649101 UbiProPox) (J.M.) and the Swiss National Foundation Ambizione (PZ00P3_131988) (J.M.).
Author information
Authors and Affiliations
Contributions
K.N., S.K., J.M. and B.W. designed the project and wrote the manuscript. K.N. performed the proteomic experiments. K.N., U.O. and J.V. analysed the proteomics data and U.O. performed the phosphorylation site relocalization analysis. M.S. produced the viruses. S.K., C.B. and C.K.M. designed and performed the biochemical validations. S.K, C.K.E.B. and I.W. performed the EM analysis. A.M. contributed ideas and gave assistance for the phosphoproteomic workflows. All authors discussed the results and implications of the findings and provided comments on the manuscript at all stages.
Corresponding authors
Ethics declarations
Competing interests
J.V. is an employee of Biognosys AG and helped with the Spectronaut DIA analysis in the revised version of the manuscript. K.N. joined Biognosys AG during the revision process of the manuscript upon finishing his PhD at ETH Zurich.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–8, Supplementary Reference.
Supplementary Table 1
Label-free quantification phosphoproteome data of HeLa cells infected with VACV WT, H1 and F10, and uninfected cells.
Supplementary Table 2
Label-free quantification of MV WT versus H1 proteomes.
Supplementary Table 3
Label-free quantification phosphoproteome data of MV WT versus H1.
Supplementary Table 4
Label-free quantification data of MV WT versus H1 phosphotyrosine enrichment.
Supplementary Table 5
Primers used in this study.
Rights and permissions
About this article
Cite this article
Novy, K., Kilcher, S., Omasits, U. et al. Proteotype profiling unmasks a viral signalling network essential for poxvirus assembly and transcriptional competence. Nat Microbiol 3, 588–599 (2018). https://doi.org/10.1038/s41564-018-0142-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-018-0142-6