Abstract
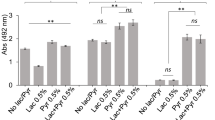
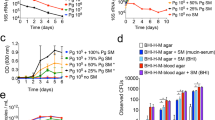
Many human infections are polymicrobial in origin, and interactions among community inhabitants shape colonization patterns and pathogenic potential1. Periodontitis, which is the sixth most prevalent infectious disease worldwide2, ensues from the action of dysbiotic polymicrobial communities3. The keystone pathogen Porphyromonas gingivalis and the accessory pathogen Streptococcus gordonii interact to form communities in vitro and exhibit increased fitness in vivo3,4. The mechanistic basis of this polymicrobial synergy, however, has not been fully elucidated. Here we show that streptococcal 4-aminobenzoate/para-amino benzoic acid (pABA) is required for maximal accumulation of P. gingivalis in dual-species communities. Metabolomic and proteomic data showed that exogenous pABA is used for folate biosynthesis, and leads to decreased stress and elevated expression of fimbrial adhesins. Moreover, pABA increased the colonization and survival of P. gingivalis in a murine oral infection model. However, pABA also caused a reduction in virulence in vivo and suppressed extracellular polysaccharide production by P. gingivalis. Collectively, these data reveal a multidimensional aspect to P. gingivalis–S. gordonii interactions and establish pABA as a critical cue produced by a partner species that enhances the fitness of P. gingivalis while diminishing its virulence.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Murray, J. L., Connell, J. L., Stacy, A., Turner, K. H. & Whiteley, M. Mechanisms of synergy in polymicrobial infections. J. Microbiol. 52, 188–199 (2014).
Kassebaum, N. J. et al. Global burden of severe periodontitis in 1990–2010: a systematic review and meta-regression. J. Dent. Res. 93, 1045–1053 (2014).
Lamont, R. J. & Hajishengallis, G. Polymicrobial synergy and dysbiosis in inflammatory disease. Trends Mol. Med. 21, 172–183 (2015).
Daep, C. A., Novak, E. A., Lamont, R. J. & Demuth, D. R. Structural dissection and in vivo effectiveness of a peptide inhibitor of Porphyromonas gingivalis adherence to Streptococcus gordonii. Infect. Immun. 79, 67–74 (2011).
Wright, C. J. et al. Microbial interactions in building of communities. Mol. Oral Microbiol. 28, 83–101 (2013).
Chawla, A. et al. Community signalling between Streptococcus gordonii and Porphyromonas gingivalis is controlled by the transcriptional regulator CdhR. Mol. Microbiol. 78, 1510–1522 (2010).
Kuboniwa, M. et al. Streptococcus gordonii utilizes several distinct gene functions to recruit Porphyromonas gingivalis into a mixed community. Mol. Microbiol. 60, 121–139 (2006).
Kubota, T. et al. Production of para-aminobenzoate by genetically engineered Corynebacterium glutamicum and non-biological formation of an N-glucosyl byproduct. Metab. Eng. 38, 322–330 (2016).
Maeda, K. et al. A Porphyromonas gingivalis tyrosine phosphatase is a multifunctional regulator of virulence attributes. Mol. Microbiol. 69, 1153–1164 (2008).
Kuboniwa, M. et al. Proteomics of Porphyromonas gingivalis within a model oral microbial community. BMC Microbiol. 9, 98 (2009).
Hendrickson, E. L. et al. Proteomics of Streptococcus gordonii within a model developing oral microbial community. BMC Microbiol. 12, 211 (2012).
Wegkamp, A., van Oorschot, W., de Vos, W. M. & Smid, E. J. Characterization of the role of para-aminobenzoic acid biosynthesis in folate production by Lactococcus lactis. Appl. Environ. Microbiol. 73, 2673–2681 (2007).
Orsomando, G. et al. Evidence for folate-salvage reactions in plants. Plant J. 46, 426–435 (2006).
Orsomando, G. et al. Plant gamma-glutamyl hydrolases and folate polyglutamates: characterization, compartmentation, and co-occurrence in vacuoles. J. Biol. Chem. 280, 28877–28884 (2005).
Pathirana, R. D., O’Brien-Simpson, N. M., Brammar, G. C., Slakeski, N. & Reynolds, E. C. Kgp and RgpB, but not RgpA, are important for Porphyromonas gingivalis virulence in the murine periodontitis model. Infect. Immun. 75, 1436–1442 (2007).
Wright, C. J. et al. Characterization of a bacterial tyrosine kinase in Porphyromonas gingivalis involved in polymicrobial synergy. MicrobiologyOpen 3, 383–394 (2014).
Whitmore, S. E. & Lamont, R. J. The pathogenic persona of community-associated oral streptococci. Mol. Microbiol 81, 305–314 (2011).
Valm, A. M. et al. Systems-level analysis of microbial community organization through combinatorial labeling and spectral imaging. Proc. Natl Acad. Sci. USA 108, 4152–4157 (2011).
Lamont, R. J. et al. Role of the Streptococcus gordonii SspB protein in the development of Porphyromonas gingivalis biofilms on streptococcal substrates. Microbiology 148, 1627–1636 (2002).
Li, Y. et al. Phylogenetic and functional gene structure shifts of the oral microbiomes in periodontitis patients. ISME J. 8, 1879–1891 (2014).
Hashino, E. et al. Erythritol alters microstructure and metabolomic profiles of biofilm composed of Streptococcus gordonii and Porphyromonas gingivalis. Mol. Oral Microbiol. 28, 435–451 (2013).
Kuboniwa, M. et al. Quantitative detection of periodontal pathogens using real-time polymerase chain reaction with TaqMan probes. Oral Microbiol. Immunol. 19, 168–176 (2004).
Houser, J. R. et al. Controlled measurement and comparative analysis of cellular components in E. coli reveals broad regulatory changes in response to glucose starvation. PLoS Comput. Biol. 11, e1004400 (2015).
Kwon, T., Choi, H., Vogel, C., Nesvizhskii, A. I. & Marcotte, E. M. MSblender: A probabilistic approach for integrating peptide identifications from multiple database search engines. J. Proteome Res. 10, 2949–2958 (2011).
Ohashi, Y. et al. Depiction of metabolome changes in histidine-starved Escherichia coli by CE-TOFMS. Mol. BioSys. 4, 135–147 (2008).
Yamamoto, H. et al. Statistical hypothesis testing of factor loading in principal component analysis and its application to metabolite set enrichment analysis. BMC Bioinformatics 15, 51 (2014).
Hirano, T., Beck, D. A., Demuth, D. R., Hackett, M. & Lamont, R. J. Deep sequencing of Porphyromonas gingivalis and comparative transcriptome analysis of a LuxS mutant. Front. Cell. Infect. Microbiol. 2, 79 (2012).
Wang, Q. et al. FOXO responses to Porphyromonas gingivalis in epithelial cells. Cell. Microbiol. 17, 1605–1617 (2015).
Park, Y. et al. Short fimbriae of Porphyromonas gingivalis and their role in coadhesion with Streptococcus gordonii. Infect. Immun. 73, 3983–3989 (2005).
Yilmaz, O., Watanabe, K. & Lamont, R. J. Involvement of integrins in fimbriae-mediated binding and invasion by Porphyromonas gingivalis. Cell. Microbiol. 4, 305–314 (2002).
Capestany, C. A., Tribble, G. D., Maeda, K., Demuth, D. R. & Lamont, R. J. Role of the Clp system in stress tolerance, biofilm formation, and intracellular invasion in Porphyromonas gingivalis. J. Bacteriol. 190, 1436–1446 (2008).
Moffatt-Jauregui, C. E. et al. Establishment and characterization of a telomerase immortalized human gingival epithelial cell line. J. Periodontal Res. 48, 713–721 (2013).
Sztukowska, M. et al. The C-terminal domains of the gingipain K polyprotein are necessary for assembly of the active enzyme and expression of associated activities. Mol. Microbiol. 54, 1393–1408 (2004).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–456 (2016).
Acknowledgements
Supported by AMED-CREST, AMED, MEXT/JSPS KAKENHI grant numbers 15H05057, and 15K20642 (M.K.), NIH grants DE012505 and DE011111 (R.J.L.), DE023193 (M.W.) DP1 OD009572 (E.M.M.) and DE014372 (M.H.), Army Research Office Grant W911NF-12–1–0390 (E.M.M.), and the Welch Foundation grant F1515 (E.M.M.).
Author information
Authors and Affiliations
Contributions
M.K., S.A.A. and A.S. performed metabolomics and community experiments. J.R.H., E.L.H., T.W. and D.A.C.B. performed proteomics experiments. Q.W. and D.P.M. performed PCR, blots, ELISAs, protease and attachment assays. Q.W., J.A.H. and H.W. performed animal experiments. M.K., M.W., A.A., H.W., E.M.M., M.H. and R.J.L. designed the study and interpreted data. M.K., M.W., A.A., M.H. and R.J.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–10, Supplementary Tables 1 and 2, Supplementary Table 6
Supplementary Table 1
MS/MS identification of P. gingivalis proteins differentially regulated by pABA
Supplementary Table 3
Metabolomic profiles in P. gingivalis treated with pABA
Supplementary Table 4
Metabolites decreased by pABA-mediated suppression of PLP-dependent enzymes in P. gingivalis
Supplementary Table 5
Metabolite classification pathways
Supplementary Table 7
Raw protein spectral counts
Supplementary Table 8
Raw peptide spectral counts
Supplementary Table 9
Spectral counts of P. gingivalis treated with or without pABA normalized such that their averages are identical for identical biological replicates
Supplementary Table 10
Outlier analysis of proteomic data
Rights and permissions
About this article
Cite this article
Kuboniwa, M., Houser, J.R., Hendrickson, E.L. et al. Metabolic crosstalk regulates Porphyromonas gingivalis colonization and virulence during oral polymicrobial infection. Nat Microbiol 2, 1493–1499 (2017). https://doi.org/10.1038/s41564-017-0021-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0021-6
This article is cited by
-
Polysaccharide-based antibacterial nanocomposite hydrogels with Janus structure for treatment of infected extraction socket
Science China Materials (2024)
-
An injectable and thermosensitive hydrogel with nano-aided NIR-II phototherapeutic and chemical effects for periodontal antibacteria and bone regeneration
Journal of Nanobiotechnology (2023)
-
Carvacrol combined with NIR light-responsive nano-drug delivery system with specific anti-bacteria, anti-inflammation, and immunomodulation for periodontitis
Nano Research (2023)
-
Genomic insights into the c-di-GMP signaling and biofilm development in the saprophytic spirochete Leptospira biflexa
Archives of Microbiology (2023)
-
Atypical cyclic di-AMP signaling is essential for Porphyromonas gingivalis growth and regulation of cell envelope homeostasis and virulence
npj Biofilms and Microbiomes (2022)