Abstract
Complex polymicrobial biofilm communities are abundant in nature particularly in the human oral cavity where their composition and fitness can affect health. While the study of these communities during disease is essential and prevalent, little is known about interactions within the healthy plaque community. Here we describe interactions between two of the most abundant species in this healthy microbiome, Haemophilus parainfluenzae and Streptococcus mitis. We discovered that H. parainfluenzae typically exists adjacent to mitis group streptococci in vivo with which it is also positively correlated based on microbiome data. By comparing in vitro coculture data to ex vivo microscopy we revealed that this co-occurrence is density dependent and further influenced by H2O2 production. We discovered that H. parainfluenzae utilizes a more redundant, multifactorial response to H2O2 than related microorganisms and that this system’s integrity enhances streptococcal fitness. Our results indicate that mitis group streptococci are likely the in vivo source of NAD for H. parainfluenzae and also evoke patterns of carbon utilization in vitro for H. parainfluenzae similar to those observed in vivo. Our findings describe mechanistic interactions between two of the most abundant and prevalent members of healthy supragingival plaque that contribute to their in vivo survival.
Similar content being viewed by others
Introduction
Taxon-taxon interactions are of great importance for understanding the structure and function of natural biofilms. Many bacteria occur in nature as part of complex polymicrobial biofilms [1]. The behavior of individual species within these biofilms is elaborate and likely influenced by the spatial organization between physical and chemical substrates [1, 2] produced by distinct species [3,4,5]. Metagenomic and metatranscriptome techniques give us an unprecedented ability to observe group composition and behaviors, but cannot resolve which species might be influencing one another nor describe their spatial interactions. Metabolite-based interactions can influence community composition through crossfeeding or trophic interactions [6, 7], complementation of auxotrophies [8] as well as competition for nutrients and production of inhibitory substances [9,10,11,12]. These types of polymicrobial associations have been shown to enhance the resiliency, fitness, and stability of these communities [4, 13, 14]. Maintaining stable oral microbial communities can preserve oral health, thus there is a clear need to identify the interactions between prominent species in host-associated polymicrobial communities and how they might shape not only constituency but physical structure.
To delve into mechanisms underlying taxon-taxon interactions we chose to study the naturally occurring biofilm of human dental plaque. The human supragingival plaque biofilm has been long studied, dating to the very 1st microscopy observations of Antony von Leeuwenhoek in 1683 (ref. [15]). Hypotheses about direct interaction of bacteria and assembly of plaque structures have been developed from work visualizing the spatial structure of plaque, both on extracted teeth [16,17,18,19] and on epoxy resin crowns or enamel chips worn in the mouth to allow plaque to develop [16, 20,21,22,23,24]. Recently, more detailed biogeography of plaque structure was observed [25]. One notable association was between Streptococcus and other genera at the margins of biofilm structures. Streptococci can influence community interactions due to their ability to rapidly consume carbohydrates, produce large amounts of acidic fermentation products, and excrete antimicrobial substances including reactive oxygen species. These properties can support the growth of some species while excluding others [9, 26,27,28]. Many oral streptococci including mitis group members are known to produce various antimicrobial substances, including hydrogen peroxide (H2O2), in aerobic conditions. Therefore, any bacterium adjacent to these Streptococcus spp. aerobically would need to have adapted the ability to withstand H2O2 (refs. [7, 21]). Alongside Streptococcus species within the mitis group (S. mitis, S. oralis, S. australis, S. infantis, and others), Haemophilus parainfluenzae is one of the most abundant and prevalent species in the dental plaque of healthy individuals [29,30,31,32,33]. Whilst frequently characterized as an opportunistic pathogen and implicated in diseases including endocarditis [34], in the oral cavity H. parainfluenzae is a commensal and is associated with beneficial immunomodulatory effects [35].
The strains identified as S. mitis encompass a broader range of diversity than many bacterial species but appear to be members of a genomically coherent species. Like many species within the mitis group, S. mitis strains share less than 95% average nucleotide identity (ANI), the standard threshold used to delineate bacterial species identity [30]. This may be due to the high rate of recombination among some members of the mitis group [36, 37]. The strains identified as S. mitis form several distinct clades [38,39,40]. One clade previously identified as “S. mitis biovar 2” has been reclassified as S. oralis subsp. dentisani [30]. The ANI values indicate the remaining S. mitis clades are made up of strains that constitute a continuum of lineages [30]. A more recent analysis indicated that while the remaining clades represent potential subspecies, they are not separate species [40]. For these reasons and because we were primarily interested in S. mitis as a representative of the broader mitis group, we elected to treat S. mitis as a single species and we selected its type strain as a representative of the species for in vitro experiments.
To identify mechanisms of interaction that take place within this healthy polymicrobial biofilm we investigated interactions between two of its most abundant members, H. parainfluenzae and S. mitis. Despite the frequent co-occurrence of these species in proximity in vivo, little is known about both the spatial relationship and metabolic mechanisms of interactions between these species. In this study, we observed that these taxa exist in intimate proximity in dental plaque and exhibit localized density dependences that also exist in vitro which are driven by streptococcal H2O2 production. We also discovered a highly redundant H2O2 resistance system in H. parainfluenzae different from that in other Haemophilus species. We then observed multiple mechanisms of H. parainfluenzae interaction including NAD auxotrophy complementation by mitis group streptococci and the upregulation of alternative carbon and energy pathways during in vitro coculture that are also observed in in vivo gene expression data. These results provide a robust characterization of H. parainfluenzae’s role in the oral microbiota and reveal ways it has evolved to exist alongside streptococci in the oral cavity and likely beyond. This study details interactions between two prominent members of a complex natural biofilm community and allows us to demonstrate mechanisms of interaction that likely help drive micron-scale arrangements between these microorganisms.
Results
H. parainfluenzae co-occurs with S. mitis and related streptococci in human supragingival plaque
To determine which species H. parainfluenzae was most likely to interact with we used microbiome data from 117 subjects sampled by the Human Microbiome Project (HMP). Analysis of HMP species-assigned metagenomic data indicated that H. parainfluenzae is an abundant and prevalent member of supragingival plaque detected in all 117 subjects averaging 7.6% relative abundance based on sequence reads (Fig. 1, Table S1). We compared the upper quartile (n = 29) of these subjects ranked by highest H. parainfluenzae abundance to the remainder of subjects (n = 88) via LEfSe analysis [41] to determine which species were significantly likely to co-occur with H. parainfluenzae (Fig. 1, Fig. S1). Interestingly, this indicated that individuals enriched in H. parainfluenzae have abundant Streptococcus sp. especially those in the mitis group. These data were what led us to investigate H. parainfluenzae interactions with S. mitis as a mitis group representative in order to examine the mechanisms of taxon-taxon interactions.
Human Microbiome Project (HMP) metagenome data of supragingival plaque was used to plot the relative abundance and prevalence of species of interest including Haemophilus parainfluenzae (blue), several mitis group streptococci and all remaining Streptococcus spp. (dark gray). (Top Right) The top 25% of subjects based on H. parainfluenzae abundance were compared to the remainder via LEfSe analysis. Shown are species enriched in this comparison with an LDA score ≤ −3.0. Full datasets in Table S1 and Fig. S1.
Species-specific FISH demonstrates frequent Haemophilus spp. co-proximity with a subset of the mitis group and suggests a density-dependent relationship
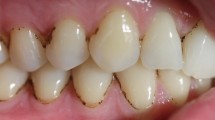
To determine whether the large-scale co-occurrence of H. parainfluenzae and S. mitis was reflected in micrometer-scale spatial structure, we analyzed the organization of these two taxa using fluorescence in situ hybridization (FISH) and imaging. Within the supragingival plaque, S. mitis and related streptococci are frequent features of H. parainfluenzae’s micron-scale environment. We used FISH probes targeting H. parainfluenzae and other Haemophilus spp. and S. mitis and its close relatives (Fig. S2), referred to hereafter as mitis group species (including S. infantis, S. oralis, S. peroris, and S. pneumoniae; see Table S2 for details of probe specificity). Visualizing these species showed that 92% of Haemophilus spp. cells in supragingival plaque are located within 10 μm of mitis group species (Fig. 2A, Table S2). The median distance separating a Haemophilus spp. cell from the nearest mitis group cell was 1.14 μm. Thus, most cells of Haemophilus spp. co-occur with mitis group species at micrometer scales.
A Histogram of the distance to the nearest S. mitis group cell, measured edge-to-edge, from each of the 37,591 Haemophilus spp. cells in 41 fields of view. B Pair correlations between Haemophilus spp. and S. mitis group cells. The lighter lines represent the bounds of the 95% confidence interval for the correlation values. The dashed line represents the null hypothesis: the pair correlation equals one. n = 41 fields of view. C Plaque with sparsely distributed Haemophilus spp. (cyan). D Plaque with high abundances of both S. mitis group (magenta) and Haemophilus spp. (i) Most Haemophilus spp. cells are within a few microns of the nearest S. mitis group cell. Generally, Haemophilus spp. cells are randomly distributed with respect to S. mitis group. (ii) Haemophilus spp. avoids the highest densities of S. mitis group. Scale bars indicate 10 µm. Images show hybridization signal from probes Smit651 (mitis group streptococci) and Hpar441 (Haemophilus spp.); probe sequences are given in Table S2 and additional probe images are shown in Fig. S2.
In addition to proximity, images suggested a density-dependent relationship between these two taxa, with reduced densities of Haemophilus spp. in areas with the highest densities of the mitis group species (Fig. 2C). We quantified this density effect by dividing each image into 1024 blocks of ~6 μm on a side (Fig. 2D) and measuring the density of mitis group species, Haemophilus spp., and total bacteria within each block. The mean density of Haemophilus spp. increased as the percent of a block covered by the mitis group increased from 0 to around 20% (Fig. S3). This trend is likely due, at least in part, to variation in the overall quantity of plaque in each block because the density of each taxon has a positive linear relationship with the overall plaque density. As the density of the mitis group increased above 20%, the mean Haemophilus spp. density decreased, suggesting a density-dependent inhibitory effect of the mitis group on Haemophilus spp. growth. Given that both microbiome sequencing and microscopy imaging of in vivo supragingival plaque samples indicate significant co-occurrence and co-proximity between Haemophilus spp. and the mitis group (Figs. 1 and 2), it is important to determine the mechanisms that underlie these observations.
S. mitis eliminates H. parainfluenzae via production of H2O2
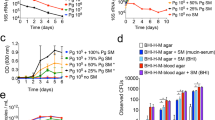
Culturing H. parainfluenzae and S. mitis together under controlled conditions in vitro revealed a dose-dependent interaction: H. parainfluenzae was killed by S. mitis, but only when abundance of S. mitis was high relative to that of H. parainfluenzae. We used a colony biofilm model [42, 43] in which polycarbonate membranes were inoculated with S. mitis and H. parainfluenzae, either separately or together, and permitted to grow for 24 h; cell abundance was then assessed by resuspending the cells and plating for colony counts (see “Methods”). Cocultures inoculated at equal densities revealed that S. mitis reduced H. parainfluenzae numbers (Fig. 3) nearly 100-fold below inoculum density, indicating active killing. This effect is dose-dependent as we observed a significant reduction in the growth yield of H. parainfluenzae compared to monoculture when inoculums of S. mitis were either equivalent or threefold greater than H. parainfluenzae. However, when S. mitis inoculum was tenfold lower than H. parainfluenzae, there was no significant change in the growth yield of H. parainfluenzae compared to monoculture.
H. parainfluenzae (Hp) CFU counts in mono and coculture with wildtype (Sm) or a pyruvate oxidase mutant (ΔspxB) of S. mitis. Hp had an initial inoculum using 10 µL at an OD600 of 1, which corresponds to 4.65 × 106 CFU/ml. Wildtype (Sm) and S. mitis ΔspxB with initial inoculums using 10 µL at an OD600 of either 0.1, 1 or 3 which corresponds to an average of 2.45 × 105, 1.55 × 106 or 3.45 × 106 CFU/ml. Data are mean CFU counts with error bars indicating standard deviation for n ≥ 3. * denotes p < 0.001 using a Student’s t test compared to monoculture.
The mechanism of dose-dependent killing of H. parainfluenzae involves H2O2 production by S. mitis. The reduction of H. parainfluenzae numbers in coculture was abolished if the cocultured strain of S. mitis lacked pyruvate oxidase (ΔspxB) and thus unable to produce H2O2. Figure 3 demonstrates that H. parainfluenzae growth yield when cocultured with S. mitis ΔspxB is not significantly different from H. parainfluenzae monoculture at any ratio tested, indicating that S. mitis-produced H2O2 is responsible for H. parainfluenzae inhibition. Additional coculture experiments indicate that H. parainfluenzae was unable to grow in pH neutralized supernatants of S. mitis unless they were pre-treated with exogenous catalase (data not shown), further supporting the finding that H2O2 production limits H. parainfluenzae growth in these conditions and that acid production is not sufficient to inhibit H. parainfluenzae. Thus, our observed density-dependent exclusion of H. parainfluenzae by S. mitis in ex vivo samples (Fig. 2) can be explained by H2O2 toxicity.
Although the density-dependent killing of H. parainfluenzae by S. mitis implies an antagonistic interaction, the presence of H. parainfluenzae enhances S. mitis growth yield in biofilm coculture (Fig. S4). This enhancement was abolished when S. mitis was cocultured with H. parainfluenzae ΔoxyR but not with other catalase or cytochrome peroxidase mutants either individually or combined (Fig. S4A). The addition of exogenous catalase elevated S. mitis monoculture yields near to those observed in coculture without catalase which suggests that H2O2 detoxification is primarily responsible for this enhanced yield (Fig. S4B). However, a > 10-fold yield enhancement was still observed in coculture with H. parainfluenzae supplemented with exogenous catalase indicating further interactions that benefit S. mitis fitness. The role of H2O2 decomposition in S. mitis yield increase was further confirmed when comparing mono vs coculture of S. mitis vs its ΔspxB mutant where the mutant demonstrated greater yield in monoculture and coculture vs the wild-type with a modest increase coculture yield (Fig. S4C) reminiscent of the wild-type coculture with exogenous catalase.
Individual H2O2 sensitive genes do not affect the fitness of H. parainfluenzae in coculture with S. mitis
The presence of S. mitis within a few micrometers of most H. parainfluenzae cells in natural plaque suggests that H2O2 and other inhibitory or promotive compounds excreted by S. mitis are reasonably expected to perfuse the substrate in which most H. parainfluenzae grow [44]. We investigated the growth effects of known H2O2 relevant gene products in H. parainfluenzae by constructing gene deletions and assessing the fitness of the deletion strains after H2O2 exposure. Genes in H. parainfluenzae relevant to peroxide metabolism include oxyR whose gene product is a global transcriptional regulator responsive to H2O2. Also present is a single catalase (katA) essential for H2O2 resistance in H. influenzae [45, 46]. In addition, H. parainfluenzae possesses a cytochrome c peroxidase (ccp), orthologs of which are important for peroxide resistance in other Pasteurellaceae [47], Campylobacter jejuni [48], and H. influenzae [49]. We also made deletions of the peroxiredoxin (prx), peroxiredoxin-glutaredoxin (pdgX) and glucose-6-phosphate dehydrogenase (g6p) genes known to contribute to oxidative stress protection in H. influenzae and other species [46, 50,51,52].
We quantified changes in H2O2 resistance by a zone of inhibition assay (Fig. 4A), in which an agar plate inoculated with a lawn of wild-type or deletion-strain H. parainfluenzae is exposed to a small filter saturated with H2O2 and sensitivity is measured as the diameter of the zone of inhibition. As expected, the largest increase in sensitivity was observed in H. parainfluenzae with deletions of the global H2O2-responsive transcriptional regulator ΔoxyR, while all other individual gene deletion mutants were less sensitive to H2O2. A ΔkatA + Δccp double mutant demonstrated a significant increase in sensitivity compared to individual mutants, indicating a combinatorial effect of these gene products. Surprisingly, MIC concentrations for many mutants were identical to the wild type (Table S3) indicating that individual contributions of these genes to H2O2 tolerance are minimal. We next used the colony biofilm model to test the fitness of each mutant in coculture with H2O2-producing S. mitis (Fig. 4B). OxyR was shown to be essential for H. parainfluenzae survival in coculture, while deletion of individual genes typically controlled by OxyR showed no significant difference compared to wildtype which contrasts greatly to similar mutants in H. influenzae (katA) [45, 46] and other species [53,54,55]. However, the ΔkatA + Δccp mutants’ growth yield was significantly inhibited in S. mitis coculture again indicating an additive effect of these gene products on H2O2 resistance. ΔkatA and ΔoxyR mutants were also generated in another oral isolate strain of H. parainfluenzae and showed similar trends in regard to H2O2 sensitivity (Fig. S5). These data suggest that the mechanism of H. parainfluenzae resistance to H2O2 involves a complex multifactorial system unlike other Haemophilus spp. yet characterized.
We assessed the sensitivity of wildtype H. parainfluenzae (WT) to H2O2 or coculture yield with S. mitis vs mutants lacking oxyR, catalase (katA), cytochrome c peroxidase (ccp), peroxiredoxin (prx), glutaredoxin-peroxiredoxin (pdgX), DNA-binding protein from starved cells (dps) and glucose-6-phosphate dehydrogenase (g6p). A Zones of inhibition by 30% H2O2 exposure (areas of no visible H. parainfluenzae growth) were measured after 24 h and fold change calculated compared to wildtype. Data are the mean fold change relative to WT; error bars indicate standard deviation for n ≥ 3. All strains were significantly different from WT, while ΔkatA-Δccp was significantly different from ΔkatA (padj < 0.05), based on t test with Bonferroni correction. B WT and mutant H. parainfluenzae CFU following 24 h coculture with S. mitis. H. parainfluenzae inoculums were 4.65 × 106 CFU/ml. S. mitis inoculum was 2.45 × 105 CFU/ml. Data are represented as mean CFU, error bars indicate standard deviation for n ≥ 3. *denotes p < 0.05 by Student’s t test compared to monoculture.
S. mitis and other Streptococcus sp. support H. parainfluenzae growth
Like other Haemophilus sp., H. parainfluenzae is a NAD auxotroph and must obtain NAD, nicotinamide mononucleotide (NMN) or nicotinamide riboside (NR) from the host or other microorganisms [56]. We found that human saliva was unable to support H. parainfluenzae growth unless NAD was added (data not shown) suggesting that adjacent microbes are the source of NAD for H. parainfluenzae in the oral cavity. As Corynebacterium and Streptococcus are two of the most abundant genera in supragingival plaque, we tested whether species from these genera could complement NAD auxotrophy. Spot assays on H. parainfluenzae lawns on medium lacking NAD showed robust growth of H. parainfluenzae adjacent to S. mitis and S. sanguinis but no other taxa (Fig. 5). These data demonstrate that H. parainfluenzae obtains NAD from these taxa when they are in close proximity. It is notable that multiple species that significantly correlate with H. parainfluenzae in microbiome data (Fig. 1) also enhance its growth most strongly in the absence of NAD compared to other Streptococcus spp. Similar results were also observed in two other oral isolates of H. parainfluenzae (Fig. S6).
Cultures of S. mitis, S. sanguinis, S. gordonii, S. cristatus, Corynebacterium matruchotii and C. durum were grown overnight, normalized based on optical density, and spotted onto paper discs over lawns of H. parainfluenzae spread on solid agar medium lacking NAD. Plates were incubated for 48 h before observation. Rings near the disc indicate H. parainfluenzae growth. NAD was added a positive control (NAD+).
In vitro transcriptional responses of H. parainfluenzae to S. mitis
Transcriptome analysis of H. parainfluenzae in aerobic coculture with S. mitis indicated a multifaceted response to oxidative stress. We observed that H. parainfluenzae significantly differentially expressed 387 genes greater than twofold in coculture (Appendix 1, Fig. S7), compared to monoculture. Surprisingly, the differentially expressed genes did not include catalase; however, based on transcript abundance catalase is well expressed under both conditions. Among genes typically involved in oxidative stress responses, dps, which encodes an iron sequestration protein, had a 2.2-fold increase in coculture suggesting that H. parainfluenzae is sequestering free intracellular Fe2+, to prevent oxidative damage. Surprisingly, however, several other genes involved in H2O2 were repressed in coculture, including ccp, pdgX, thioredoxin (trxA), glutaredoxin (grx) and thiol peroxidase (tsa), demonstrating the complex nature of the oxidative stress response in this species. H. parainfluenzae stress responses were induced in coculture, likely due to H2O2–related damage by S. mitis. We observed modest but significant upregulation of the hfq chaperone encoding gene, universal stress protein E (uspE) and lexA. LexA is involved in the repression of genes involved in the SOS response of E. coli [56]. Hfq is known to be involved in the stress responses of many species [57]. Paralogs of the Universal stress proteins including uspE, are known to be involved in response to DNA damage [58]. Genes likely involved in DNA repair are also induced in coculture including those encoding DNA ligase, exodeoxyribonuclease V beta chain and endonuclease V.
Streptococcus spp. are known to rapidly consume carbohydrates, and transcriptional data suggest that H. parainfluenzae in coculture switches from carbohydrate consumption to alternative sources of carbon and energy. There was increased expression of genes suggesting the breakdown of glycerophospholipids resulting in the uptake and utilization of glycerol, including the predicted extracellular patatin-like phospholipase (2.5-fold), lysophospholipase L2 (3.8-fold), glycerol uptake facilitator protein (5.3-fold), glycerol kinase (3.4-fold), and a fatty acid degradation regulator (2.1-fold). In addition, there was an increase in expression of fructose 1,6 bisphosphatase (2.8-fold), indicating active gluconeogenesis, consistent with H. parainfluenzae growth on 3-carbon intermediates such as glycerol. There was also evidence of the uptake and catabolism of the sialic acid, N-acetylneuraminic acid as suggested by an increase in expression of SHS family sialic acid transporter (twofold), sialic acid utilization regulator (3.6-fold), N-acetylneuraminate lyase (twofold), N-acetylmannosamine kinase (3.8-fold), and N-acetylmannosamine-6-phosphate 2-epimerase (3.1-fold). Last, there was an increase in the expression of genes involved in the oligopeptide transport system, oppA (threefold), oppB (2.2-fold), oppC (2.3-fold), oppD (2.7-fold), and oppF (2.4-fold). These data together suggest uptake of alternate carbon and energy sources in coculture. S. mitis transcriptional responses to H. parainfluenzae are the focus of a separate study.
In vivo transcriptional responses of H. parainfluenzae vs in vitro
We hypothesized that in vivo gene expression of H. parainfluenzae is largely influenced by S. mitis due to their in vivo proximity (Fig. 2) and that our in vitro coculture data may be predictive of H. parainfluenzae behavior in vivo. To obtain quantitative data, we compared in vitro H. parainfluenzae transcriptomes to two separate in vivo metatranscriptome datasets from healthy dental plaque [59, 60]. We aligned metatranscriptomes to the H. parainfluenzae genome generating a dataset of H. parainfluenzae transcription within the complex plaque biofilm. We then compared these samples to our in vitro monoculture and then determined which genes differentially expressed in vivo are also expressed in in vitro coculture.
Comparing significantly differentially expressed gene patterns shared between all 3 datasets (in vitro coculture and both in vivo metatranscriptomes) we observed that 18 genes were mutually upregulated including genes involved in glycerol catabolism, gluconeogenesis, and pili biogenesis (Table S4). We also observed 22 genes that were mutually downregulated including those involved in stringent response, methionine metabolism, and fur (Table S5, Fig. 6B). In individual metatranscriptome comparisons, we saw even more genes that were significantly differentially expressed in the presence of S. mitis in vitro. This included repression of ccp, genes involved in methionine metabolism, stringent response, metal transport genes, fur, and induction of genes involved in glycerol catabolism, gluconeogenesis, cell stress, DNA damage/repair related pathways, and peptide/oligopeptide transport (Tables S6-S9).
Mono vs coculture RNASeq results (in vitro) for H. parainfluenzae indicated 387 significantly differentially expressed genes (DEG) above twofold. By comparing in vitro monoculture to in vivo metatranscriptome reads from Benítez-Páez 2014 (Meta 1) and Jorth 2014 (Meta 2) we were able to generate two sets of in vivo-specific DEGs to determine differences in (A) significantly upregulated or (B) significantly downregulated genes shared between in vitro coculture and in vivo conditions.
These data suggest that aspects of H. parainfluenzae transcriptional responses in plaque can be recapitulated in in vitro coculture with S. mitis. One notably absent overlap between in vitro cocultures vs in vivo datasets was the upregulation of genes involved in lactate oxidation observed in vivo. Since most Streptococcus spp. produce lactate as a metabolic end product [26], many oral taxa have evolved the ability to utilize lactate as a carbon source. Metatranscriptome data suggests that H. parainfluenzae catabolizes lactate in vivo, which it did not do in coculture indicating that further carbon source competition and crossfeeding is occurring in vivo that did not occur in vitro on complex medium. Overall, these data indicate numerous overlaps of in vitro H. parainfluenzae transcriptional responses to S. mitis that also occur in vivo where they exist in close proximity.
Discussion
Our findings reveal multiple factors that may dictate the spatial organization and behavior of two abundant oral commensal bacteria in vivo. The availability and distribution of oxygen has been hypothesized to dictate the spatial organization of bacteria within oral biofilm structures and in mixed communities of soil bacteria, where bacterial respiration creates a reduced environment enabling the growth of strict anaerobes in the biofilm interior [2, 25]. An important facet of oxygen availability is the production of H2O2 by some microbes, which can be an important mechanism of protection against invading or competing species in a variety of environments from the oral cavity [61] to the surface of the oceans [62]. Also, the exchange of nutrients including electron donors and cofactors or nutrients between taxa can determine growth and interaction between them [8, 63]. Here we demonstrate that streptococcal metabolism can potentially dictate both spatial organization and constituency within a naturally occurring multispecies biofilm.
The production of H2O2 by early colonizing Streptococcus spp. is regarded as an important mechanism of protecting commensals from invading species and is believed to play a role in determining species composition [61]. Our in vitro findings show that S. mitis can inhibit or kill H. parainfluenzae in a dose-dependent manner via the production of H2O2. This appears to be mirrored in our in vivo findings where streptococci seemingly exclude H. parainfluenzae above a certain density. We assert that this density dependent interaction between these taxa is important for driving heterogeneity and spatial organization within this complex community.
Given the close association in plaque samples between Haemophilus spp. and H2O2 producing streptococci we investigated mechanisms of H2O2 resistance in H. parainfluenzae and demonstrated that in the strains tested it possesses a highly redundant, multifactorial oxidative stress response that substantially differs from other closely related species, including H. influenzae. We demonstrated that while loss of OxyR caused a significant increase in H2O2 sensitivity, loss of catalase or other individual gene products that provide H2O2 resistance did not (Fig. 4A, Table S3, Fig. S5) which directly contrasts with the importance of catalase in H. influenzae, whose deletion leads to its inability to survive high concentrations of H2O2 (refs. [32, 36]). Streptococcal H2O2 production is thought to provide a competitive advantage in vivo and presents a source of stress that coexisting bacterial species must tolerate. This ability to deal with streptococci-produced H2O2 has been demonstrated in multiple species [10, 44, 61, 64]. Given that H. parainfluenzae is found closely associated with S. mitis in plaque and that they co-occur together in other sites of the oral cavity [65], an intriguing hypothesis is that this redundant system is born from H. parainfluenzae’s constant proximity to a H2O2 producer in multispecies biofilms. Evaluating the regulation of this redundant system is under current investigation by our group. We also noted that the fitness of S. mitis in vitro was reduced when cocultured with H. parainfluenzae strains impaired in H2O2 resistance (Fig. S4) and that when exogenous H2O2 detoxification occurred this fitness was restored (Fig. S4B). However, we also observed that H. parainfluenzae still further increased S. mitis yield when H2O2 was not a factor indicating further mutual benefit beyond H2O2 responses.
Paradoxically, H. parainfluenzae can be inhibited by but is found adjacent to S. mitis. This could be due to the ability of S. mitis to complement the NAD auxotrophy of H. parainfluenzae. We observed that abundant plaque Corynebacterium spp. were unable to restore H. parainfluenzae’s growth, but some oral streptococci and S. mitis in particular could (Fig. 5) while host saliva could not. Perhaps this explains why 92% of H. parainfluenzae cells lie within 10 µm of S. mitis in vivo (Fig. 2A). It is interesting to note that both H. parainfluenzae and S. mitis are found as commensals not just in the same sites of the human oral cavity [65], but also in other sites in the nasopharynx [66, 67]. Thus, Streptococcus sp.-produced NAD could therefore be an important determinant of H. parainfluenzae’s ability to survive and colonize various sites of the human body. While this may appear to be disadvantageous to H. parainfluenzae, it is also possible given the ubiquitous nature of mitis group streptococci in the oral cavity, that H. parainfluenzae evolved this auxotrophy due to the constant availability of extracellular NAD. Interestingly NADP also complements this auxotrophy as H. parainfluenzae possesses an NAD kinase that can convert NADP to NAD and vice versa. This kinase was not differentially expressed in our coculture conditions. However, an NADP-specific glutamate hydrogenase was upregulated in coculture 3.1-fold (Supplemental) which suggests that it may play a role in both low pH defense as well as a response to oxidative stress which is under further investigation. These data are reminiscent of other cases in which auxotrophy development confers a fitness advantage demonstrated in broad genome-scale metabolic models [68] as well as direct testing of artificial auxotrophs generated in E. coli and Acinetobacter baylyi [69] where classes of auxotrophs showed a fitness advantage vs non-auxotrophs when substrates were available. Further, cross-feeding synthetically induced by artificial auxotrophy in E. coli conferred a fitness advantage to both strains [70]. Thus, H. parainfluenzae may have adapted to exist in close proximity to certain Streptococcus spp. due to their NAD/P production whose manner of export is as yet unknown.
When comparing the H. parainfluenzae coculture in vitro transcriptome to in vivo supragingival plaque metatranscriptomes (Fig. 6) we observed similar patterns of carbon catabolism including the shift from utilization of sugars to small organic acids and alcohols while downregulating genes in the stringent response suggesting greater access to peptides. Given their in vivo proximity, it is extremely likely that mitis group streptococci are what induce these same H. parainfluenzae behaviors in vivo, highlighting a facet of our reductionist approach that can uniquely explain behaviors in a complex multispecies in vivo biofilm. One notable pathway absent from our in vitro cocultures was lactate oxidation which has previously been shown to be critical for co-infection between organisms with Streptococcus spp. by cross-feeding on this fermentation product [7]. In vivo, metatranscriptome data indicated that H. parainfluenzae was also upregulating lactate oxidation gene products which would be expected in the more diverse, competitive natural plaque environment compared to in vitro coculture alone. H. parainfluenzae differentially expressed many other genes in coculture with S. mitis that suggest additional stress responses and changes in nutrient availability that have not yet been investigated.
H. parainfluenzae and S. mitis are found abundantly not just in supragingival plaque but also in other sites of the human oral cavity [33, 65]. The abundance of these two taxa has been associated with health and their reduction is associated with the development of oral squamous cell carcinomas [31, 71]. Both taxa have also been shown to prevent the adhesion and attachment of the oral pathogen Porphyromonas gingivalis [72] and their mutual proximity has been observed at the genus level [25]. Therefore, further investigation of the mechanistic interactions that take place between these two species could provide insights into maintaining their presence and thus prevention of disease. Overall, there is a wealth of information and techniques available to study the composition, structure, and gene expression of complex multispecies communities. However, specific mechanisms that exist between individual members of these communities can be hard to discern from broad observational methods. Using a reductionist approach on two highly abundant and prevalent species in the oral cavity we determined how these taxa can support growth of each other and dictate their micron-scale distribution within this environment while suggesting additional mechanisms in use in the greater in vivo community.
Materials and methods
Strains and media
Strains and plasmids used in this study are listed in Table S10. Unless indicated, Streptococcus mitis was cultured using Brain Heart Infusion (BHI) broth or solid agar supplemented with Yeast extract (YE), Haemophilus parainfluenzae had additional supplementation with 15 µg/ml Nicotinamide Adenine Dinucleotide (Sigma-Aldrich) and 15 µg/ml Hemin (Sigma-Aldrich)—(BHI-YE-HP). Escherichia coli was grown on Luria Broth (LB). H. parainfluenzae and S. mitis were grown at 37 °C and 5% CO2 for aerobic conditions and in 5% H2, 10% CO2 and 85% N2 in anaerobic conditions. E. coli was grown at 37 °C in standard atmospheric conditions with liquid cultures shaken at 200 RPM. Antibiotics were used at the following concentrations: kanamycin 40 µg/ml, vancomycin 5 µg/ml, and spectinomycin 50 µg/ml for E. coli and H. parainfluenzae. In total, 200 µg/ml spectinomycin was used to select for H. parainfluenzae transformants.
Genomic and plasmid DNA isolation
H. parainfluenzae Genomic DNA was isolated using the DNeasy Blood & Tissue kit (Qiagen) according to the manufacturer’s instructions. Plasmid isolations were performed using QIAprep spin miniprep kits (Qiagen).
Genetic manipulation of H. parainfluenzae
Gene deletions were generated using derivatives of a suicide vector pMRKO [7] as listed in Table S10. H. parainfluenzae was transformed via conjugation and mutants were screened via PCR and Sanger sequencing. Procedures involved in plasmid construction and H. parainfluenzae transformation are outlined in the supplementary methods section (Table S11).
Read abundance data
MetaPhlAn [73] species-assigned metagenomic sequence data from the Human Microbiome Project (Human Microbiome Project Consortium 2012) for the “Supragingival Plaque” oral site was 1st sorted based on predicted read abundance for Haemophilus parainfluenzae. Using the top quartile (highest 25% of samples enriched for H. parainfluenzae) we compared these samples to the bottom 75% and performed LEfSe analysis [41] to predict species significantly encountered at higher H. parainfluenzae abundance. LDA of a log10 score ≥3 were deemed significant.
Plaque collection, fixation, and storage
Supragingival plaque was collected according to a protocol approved by New England IRB; all donors provided informed consent. Donors were instructed to refrain from practicing oral hygiene for 24 h before plaque collection. Plaque samples were collected from 7 donors by supervised self-sampling using a toothpick to scrape the supragingival surface of the teeth, avoiding the gingival margin. We fixed the plaque in a solution of 2% paraformaldehyde (PFA) in PBS buffer on ice for 2–6 h. The PFA was removed by three washes with 10 mM Tris HCl buffer (pH 7.5). The samples were stored in a 1:1 (vol/vol) solution of 10 mM Tris HCl (pH 7.5) and 100% ethanol at –20 °C.
DNA FISH and mounting
Aliquots of plaque were dried on UltraStick Slides (Thermo Scientific) for 10 min at 46 °C. The plaque was hybridized with 2 μM of each probe in a 900 mM NaCl, 20 mM Tris HCl (pH 7.5), 0.01% SDS, and 20% formamide hybridization buffer for 3 h in a humid chamber at 46 °C. Non-hybridized probe was removed by washing the slides in prewarmed 215 mM NaCl, 20 mM Tris HCl (pH 7.5), and 5 mM EDTA wash buffer for 15 min at 48 °C. The slides were rocked once during the wash incubation. The slides were rinsed in chilled deionized water and allowed to mostly air-dry before the samples were mounted in ProLong Gold Antifade mounting medium (ThermoFisher) under a #1.5 coverslip. The slides were dried flat in the dark.
Imaging
We imaged the hybridized plaque with an LSM 780 Confocal Microscope (Zeiss) with a Plan-Apochromat 40x/1.4 NA objective. Each field of view was simultaneously excited by linear scanning with 405, 488, 561, and 633 nm laser lines. The emission spectra were decomposed by linear unmixing using ZEN software (Zeiss) using reference spectra recorded from pure cultures of reference cells (Leptotrichia buccalis) hybridized with the Eub338 probe labeled with the appropriate fluorophore as described above. To obtain a random sample of the masses of plaque large enough to permit spatial analysis, we scanned transects spaced every 5 μm along the coverslip at ×40 magnification and imaged every mass of plaque that was at least 70 μm in diameter and 250 μm away from the previously imaged fields of view. For each donor, we imaged the first 20 fields of view that satisfied these criteria or as many fields of view as were present on the slide. To maximize the number of bacteria captured in each image, we imaged the focal plane closest to the surface of the slide.
Image analysis
To allow quantitative analysis of the spatial distribution of the taxa of interest, we used FIJI [74] to create binarized images of the subset of S. mitis group species, H. parainfluenzae, and total bacteria from mitis group, H. parainfluenzae, Streptococcus spp., Pasteurellaceae spp., and Eubacteria probe images as described in the supplementary methods.
We evaluated the pair correlations between S. mitis and H. parainfluenzae over different distances using a linear dipole analysis performed in Daime 2.2 [75, 76]. For this analysis, the reference space in each image was restricted to the area in the binary bacterial biomass image. We used all possible dipoles with lengths ranging from 0.15 to 99.90 μm in steps of 0.45 μm.
We evaluated trends between the local densities of both taxa, by dividing each field of view into 1024 6.64 μm by 6.64 μm blocks, discarding blocks that did not contain any of the binary bacterial mass image, and calculating the fraction of each block that was covered by the S. mitis and H. parainfluenzae binary images.
Mono and coculture assays
Colony biofilm assays were carried out as described previously [43]. Briefly, equal volumes of H. parainfluenzae and/or S. mitis were spotted either in mono or coculture on sterile 25 mm 0.2 µm polycarbonate membranes (MilliporeSigma) and placed on BHIYE HP agar plates. H. parainfluenzae was spotted at an OD600 of 1 and S. mitis at OD600 of either 0.1, 1 or 3. The plates were then incubated for 24 h at 37 °C in 5% CO2. The membranes were then transferred to sterile media, mixed to ensure complete resuspension of the colony into the media; serially diluted and plated for CFU enumeration. S. mitis was enumerated by counting CFUs on BHI-YE and H. parainfluenzae on BHI-YEHP with 5 µg/ml vancomycin.
Disk diffusion assays
Cultures of H. parainfluenzae were grown anaerobically in BHI-YEHP overnight. All strains were then adjusted to an OD600 of 1 and 100 µl was spread plated on BHIYE HP plates and incubated aerobically for 2 hours at 37 °C in 5% CO2. 5 µl of 30% H2O2 was then added to a sterile 5 mm paper disk and plates were incubated for 24 hours at 37 °C in 5% CO2. The diameters of the zones of inhibition were then measured using a caliper in at least 3 axes.
Coculture transcriptome sample preparation
RNASeq analyses were carried out on mono and coculture samples following the colony biofilm assays described above. Briefly, H. parainfluenzae was spotted on the polycarbonate membranes at an OD600 of 1 and S. mitis at OD600 0.1. The plates were then incubated for 22 h at 37 °C in 5% CO2. The membranes were then transferred onto fresh media for 4 h and immediately placed in RNAlater solution (Ambion). Experiments were carried out in biological duplicates. RNA extraction, library preparation and sequencing were then carried out by the Microbial ‘Omics Core facility at the Broad Institute. Sequences are submitted to the NIH SRA Gene Expression Omnibus (GEO) database and can be found under submission GSE158845.
Transcriptome analyses
The H. parainfluenzae ATCC 33392 genome (GCA_000191405.1) was obtained from NCBI and annotations were generated using RAST under default settings [77,78,79]. RNASeq reads were aligned, mapped and differentially expressed genes were analyzed using bowtie2 (ref. [80]), HTSeq [81] and DESeq2 (ref. [82]) with a Unix based pipeline, generated in this paper (available at https://github.com/dasithperera-hub/RNASeq-analysis-toolkit). This pipeline was also used to carry out a pathway analysis by mapping DEGs to KEGG orthology (https://www.genome.jp/kegg-bin/get_htext?ko00001). This allowed for improved annotations and the identification of potential pathways that are involved in coculture. The same pipeline was used to analyze H. parainfluenzae gene expression in published metatranscriptome datasets [59, 60].
Complementing nicotinamide adenine dinucleotide (NAD) auxotrophy of H. parainfluenzae
Overnight cultures of H. parainfluenzae were washed 3 times in 1x Phosphate buffered saline (PBS) and diluted to an OD600 of 0.1 and spread plated on a plate containing BHI-YE supplemented with Hemin and 20 units/ml catalase. 5 µL of bacteria at an OD600 of 1 was added to a sterile paper disk and incubated for 48 h. Strains used for spotting include C. matruchotii, C. durum, S. mitis, S. sanguinis, S. cristatus, and S. gordonii.
References
Stoodley P, Sauer K, Davies DG, Costerton JW. Biofilms as complex differentiated communities. Annu Rev Microbiol. 2002;56:187–209.
Borer B, Tecon R, Or D. Spatial organization of bacterial populations in response to oxygen and carbon counter-gradients in pore networks. Nat Commun. 2018;9:769.
Liu W, Jacquiod S, Brejnrod A, Russel J, Burmølle M, Sørensen SJ. Deciphering links between bacterial interactions and spatial organization in multispecies biofilms. ISME J. 2019;13:3054–66.
Liu W, Russel J, Burmølle M, Sørensen SJ, Madsen JS. Micro-scale intermixing: a requisite for stable and synergistic co-establishment in a four-species biofilm. ISME J. 2018;12:1940–51.
Shrivastava A, Patel VK, Tang Y, Yost SC, Dewhirst FE, Berg HC. Cargo transport shapes the spatial organization of a microbial community. Proc Natl Acad Sci USA. 2018;115:8633–8.
Gralka M, Szabo R, Stocker R, Cordero OX. Trophic Interactions and the Drivers of Microbial Community Assembly. Curr Biol. 2020;30:R1176–88.
Ramsey MM, Rumbaugh KP, Whiteley M. Metabolite cross-feeding enhances virulence in a model polymicrobial infection. PLoS Pathog. 2011;7:e1002012.
Zengler K, Zaramela LS. The social network of microorganisms—how auxotrophies shape complex communities. Nat Rev Microbiol. 2018;16:383–90.
Mashimo PA, Yamamoto Y, Nakamura M, Reynolds HS, Genco RJ. Lactic acid production by oral Streptococcus mitis inhibits the growth of oral Capnocytophaga. J Periodontol. 1985;56:548–52.
Redanz S, Cheng X, Giacaman RA, Pfeifer CS, Merritt J, Kreth J. Live and let die: Hydrogen peroxide production by the commensal flora and its role in maintaining a symbiotic microbiome. Mol Oral Microbiol. 2018;33:337–52.
Kreth J, Zhang Y, Herzberg MC. Streptococcal antagonism in oral biofilms: Streptococcus sanguinis and Streptococcus gordonii interference with Streptococcus mutans. J Bacteriol. 2008;190:4632–40.
Liu W, Røder HL, Madsen JS, Bjarnsholt T, Sørensen SJ, Burmølle M. Interspecific bacterial interactions are reflected in multispecies biofilm spatial organization. Front Microbiol. 2016;7:1366.
Ren D, Madsen JS, Sørensen SJ, Burmølle M. High prevalence of biofilm synergy among bacterial soil isolates in cocultures indicates bacterial interspecific cooperation. ISME J. 2015;9:81–9.
Lee KWK, Periasamy S, Mukherjee M, Xie C, Kjelleberg S, Rice SA. Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J. 2014;8:894–907.
Mikx FH. [The study of dental plaque by Antoni van Leeuwenhoek in 1683]. Ned Tijdschr Tandheelkd. 1983;90:421–4.
Listgarten MA, Mayo HE, Tremblay R. Development of dental plaque on epoxy resin crowns in man. A light and electron microscopic study. J Periodontol. 1975;46:10–26.
Noiri Y, Ozaki K, Nakae H, Matsuo T, Ebisu S. An immunohistochemical study on the localization of Porphyromonas gingivalis, Campylobacter rectus and Actinomyces viscosus in human periodontal pockets. J Periodontal Res. 1997;32:598–607.
Zijnge V, van Leeuwen MBM, Degener JE, Abbas F, Thurnheer T, Gmür R, et al. Oral biofilm architecture on natural teeth. PloS One. 2010;5:e9321.
Zijnge V, Ammann T, Thurnheer T, Gmür R Subgingival Biofilm Structure. In: Kinane DF, Mombelli A, editors. Frontiers of Oral Biology [Internet]. Basel: KARGER; 2011 [cited 2021 Jul 28]. p. 1–16. Available from: https://www.karger.com/Article/FullText/329667
Nyvad B, Fejerskov O. Transmission electron microscopy of early microbial colonization of human enamel and root surfaces in vivo. Scand J Dent Res. 1987;95:297–307.
Nyvad B, Fejerskov O. Scanning electron microscopy of early microbial colonization of human enamel and root surfaces in vivo. Scand J Dent Res. 1987 95:287–96.
Palmer RJ, Gordon SM, Cisar JO, Kolenbrander PE. Coaggregation-mediated interactions of streptococci and actinomyces detected in initial human dental plaque. J Bacteriol. 2003;185:3400–9.
Chalmers NI, Palmer RJ, Cisar JO, Kolenbrander PE. Characterization of a Streptococcus sp.-Veillonella sp. community micromanipulated from dental plaque. J Bacteriol. 2008;190:8145–54.
Palmer RJ, Shah N, Valm A, Paster B, Dewhirst F, Inui T, et al. Interbacterial adhesion networks within early oral biofilms of single human hosts. Appl Environ Microbiol. 2017;83:e00407–17.
Mark Welch JL, Rossetti BJ, Rieken CW, Dewhirst FE, Borisy GG. Biogeography of a human oral microbiome at the micron scale. Proc Natl Acad Sci USA. 2016;113:E791–800. Feb 9.
Kreth J, Merritt J, Qi F. Bacterial and host interactions of oral streptococci. DNA Cell Biol. 2009;28:397–403.
van der Hoeven JS, Toorop AI, Mikx RH. Symbiotic relationship of Veillonella alcalescens and Streptococcus mutans in dental plaque in gnotobiotic rats. Caries Res. 1978;12:142–7.
Mikx FH, Van der Hoeven JS. Symbiosis of Streptococcus mutans and Veillonella alcalescens in mixed continuous cultures. Arch Oral Biol. 1975;20:407–10.
Zheng W, Tan TK, Paterson IC, Mutha NVR, Siow CC, Tan SY, et al. Streptobase: an oral streptococcus mitis group genomic resource and analysis platform. PloS One. 2016;11:e0151908.
Jensen A, Scholz CFP, Kilian M. Re-evaluation of the taxonomy of the Mitis group of the genus Streptococcus based on whole genome phylogenetic analyses, and proposed reclassification of Streptococcus dentisani as Streptococcus oralis subsp. dentisani comb. nov., Streptococcus tigurinus as Streptococcus oralis subsp. tigurinus comb. nov., and Streptococcus oligofermentans as a later synonym of Streptococcus cristatus. Int J Syst Evol Microbiol. 2016;66:4803–20.
Liljemark WF, Bloomquist CG, Uhl LA, Schaffer EM, Wolff LF, Pihlstrom BL, et al. Distribution of oral Haemophilus species in dental plaque from a large adult population. Infect Immun. 1984;46:778–86.
Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner ACR, Yu W-H, et al. The human oral microbiome. J Bacteriol. 2010;192:5002–17.
Eren AM, Borisy GG, Huse SM, Mark Welch JL. Oligotyping analysis of the human oral microbiome. Proc Natl Acad Sci USA. 2014;111:E2875–2884.
Sen Yew H, Chambers ST, Roberts SA, Holland DJ, Julian KA, Raymond NJ, et al. Association between HACEK bacteraemia and endocarditis. J Med Microbiol. 2014;63:892–5.
Tseng Y-C, Yang H-Y, Lin W-T, Chang C-B, Chien H-C, Wang H-P, et al. Salivary dysbiosis in Sjögren’s syndrome and a commensal-mediated immunomodulatory effect of salivary gland epithelial cells. NPJ Biofilms Microbiomes. 2021;7:21.
Chi F, Nolte O, Bergmann C, Ip M, Hakenbeck R. Crossing the barrier: evolution and spread of a major class of mosaic pbp2x in Streptococcus pneumoniae, S. mitis and S. oralis. Int J Med Microbiol. 2007;297:503–12.
Donati C, Hiller NL, Tettelin H, Muzzi A, Croucher NJ, Angiuoli SV, et al. Structure and dynamics of the pan-genome of Streptococcus pneumoniae and closely related species. Genome Biol. 2010;11:R107.
Bishop CJ, Aanensen DM, Jordan GE, Kilian M, Hanage WP, Spratt BG. Assigning strains to bacterial species via the internet. BMC Biol. 2009;7:3.
Kilian M, Tettelin H. Identification of Virulence-Associated Properties by Comparative Genome Analysis of Streptococcus pneumoniae, S. pseudopneumoniae, S. mitis, Three S. oralis Subspecies, and S. infantis. mBio. 2019;10:e01985–19.
Velsko IM, Perez MS, Richards VP. Resolving phylogenetic relationships for streptococcus mitis and streptococcus oralis through core- and pan-genome analyses. Genome Biol Evol. 2019;11:1077–87.
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12:R60.
Merritt JH, Kadouri DE, O’Toole GA Growing and analyzing static biofilms. Curr Protoc Microbiol. 2005 Jul;Chapter 1:Unit 1B.1.
Ramsey MM, Freire MO, Gabrilska RA, Rumbaugh KP, Lemon KP. Staphylococcus aureus shifts toward commensalism in response to corynebacterium species. Front Microbiol. 2016;7:1230.
Liu X, Ramsey MM, Chen X, Koley D, Whiteley M, Bard AJ. Real-time mapping of a hydrogen peroxide concentration profile across a polymicrobial bacterial biofilm using scanning electrochemical microscopy. Proc Natl Acad Sci USA. 2011;108:2668–73.
Bishai WR, Howard NS, Winkelstein JA, Smith HO. Characterization and virulence analysis of catalase mutants of Haemophilus influenzae. Infect Immun. 1994;62:4855–60.
Juneau RA, Pang B, Armbruster CE, Murrah KA, Perez AC, Swords WE. Peroxiredoxin-glutaredoxin and catalase promote resistance of nontypeable Haemophilus influenzae 86-028NP to oxidants and survival within neutrophil extracellular traps. Infect Immun. 2015;83:239–46.
Takashima E, Konishi K. Characterization of a quinol peroxidase mutant in Aggregatibacter actinomycetemcomitans. FEMS Microbiol Lett. 2008;286:66–70.
Ishikawa T, Mizunoe Y, Kawabata S, Takade A, Harada M, Wai SN, et al. The iron-binding protein Dps confers hydrogen peroxide stress resistance to Campylobacter jejuni. J Bacteriol. 2003;185:1010–7.
Wong SMS, Alugupalli KR, Ram S, Akerley BJ. The ArcA regulon and oxidative stress resistance in Haemophilus influenzae. Mol Microbiol. 2007;64:1375–90.
Izawa S, Maeda K, Miki T, Mano J, Inoue Y, Kimura A. Importance of glucose-6-phosphate dehydrogenase in the adaptive response to hydrogen peroxide in Saccharomyces cerevisiae. Biochem J. 1998;330:811–7.
Lundberg BE, Wolf RE, Dinauer MC, Xu Y, Fang FC. Glucose 6-phosphate dehydrogenase is required for Salmonella typhimurium virulence and resistance to reactive oxygen and nitrogen intermediates. Infect Immun. 1999;67:436–8.
Perkins A, Nelson KJ, Parsonage D, Poole LB, Karplus PA. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem Sci. 2015;40:435–45.
Panek HR, O’Brian MR. KatG is the primary detoxifier of hydrogen peroxide produced by aerobic metabolism in Bradyrhizobium japonicum. J Bacteriol. 2004;186:7874–80.
Kim J-S, Holmes RK. Characterization of OxyR as a negative transcriptional regulator that represses catalase production in Corynebacterium diphtheriae. PloS One. 2012;7:e31709.
Park B, Nizet V, Liu GY. Role of Staphylococcus aureus catalase in niche competition against Streptococcus pneumoniae. J Bacteriol. 2008;190:2275–8.
Kamenšek S, Podlesek Z, Gillor O, Zgur-Bertok D. Genes regulated by the Escherichia coli SOS repressor LexA exhibit heterogeneous expression. BMC Microbiol. 2010;10:283.
Deng Y, Chen C, Zhao Z, Zhao J, Jacq A, Huang X, et al. The RNA chaperone Hfq is involved in colony morphology, nutrient utilization and oxidative and envelope stress response in vibrio alginolyticus. PloS One. 2016;11:e0163689.
Gustavsson N, Diez A, Nyström T. The universal stress protein paralogues of Escherichia coli are co-ordinately regulated and co-operate in the defence against DNA damage. Mol Microbiol. 2002;43:107–17.
Jorth P, Turner KH, Gumus P, Nizam N, Buduneli N, Whiteley M. Metatranscriptomics of the human oral microbiome during health and disease. mBio 2014;5:e01012–01014.
Benítez-Páez A, Belda-Ferre P, Simón-Soro A, Mira A. Microbiota diversity and gene expression dynamics in human oral biofilms. BMC Genomics. 2014;15:311.
Zhu L, Kreth J. The role of hydrogen peroxide in environmental adaptation of oral microbial communities. Oxid Med Cell Longev. 2012;2012:717843.
Ma L, Calfee BC, Morris JJ, Johnson ZI, Zinser ER. Degradation of hydrogen peroxide at the ocean’s surface: the influence of the microbial community on the realized thermal niche of Prochlorococcus. ISME J. 2017;12:473–84.
Seth EC, Taga ME. Nutrient cross-feeding in the microbial world. Front Microbiol. 2014;5:350.
Zhu B, Macleod LC, Newsome E, Liu J, Xu P. Aggregatibacter actinomycetemcomitans mediates protection of Porphyromonas gingivalis from Streptococcus sanguinis hydrogen peroxide production in multi-species biofilms. Sci Rep. 2019;9:4944.
Mark Welch JL, Dewhirst FE, Borisy GG. Biogeography of the oral microbiome: the site-specialist hypothesis. Annu Rev Microbiol. 2019;73:335–58.
Kosikowska U, Biernasiuk A, Rybojad P, Łoś R, Malm A. Haemophilus parainfluenzae as a marker of the upper respiratory tract microbiota changes under the influence of preoperative prophylaxis with or without postoperative treatment in patients with lung cancer. BMC Microbiol. 2016;16:62.
Könönen E, Jousimies-Somer H, Bryk A, Kilpi T, Kilian M. Establishment of streptococci in the upper respiratory tract: longitudinal changes in the mouth and nasopharynx up to 2 years of age. J Med Microbiol. 2002;51:723–30.
Seif Y, Choudhary KS, Hefner Y, Anand A, Yang L, Palsson BO. Metabolic and genetic basis for auxotrophies in Gram-negative species. Proc Natl Acad Sci USA. 2020;117:6264–73.
D’Souza G, Waschina S, Pande S, Bohl K, Kaleta C, Kost C. Less is more: selective advantages can explain the prevalent loss of biosynthetic genes in bacteria. Evol Int J Org Evol. 2014;68:2559–70.
Pande S, Merker H, Bohl K, Reichelt M, Schuster S, de Figueiredo LF, et al. Fitness and stability of obligate cross-feeding interactions that emerge upon gene loss in bacteria. ISME J. 2014;8:953–62.
Yang C-Y, Yeh Y-M, Yu H-Y, Chin C-Y, Hsu C-W, Liu H, et al. Oral microbiota community dynamics associated with oral squamous cell carcinoma staging. Front Microbiol. 2018;9:862.
Van Hoogmoed CG, Geertsema-Doornbusch GI, Teughels W, Quirynen M, Busscher HJ, Van der Mei HC. Reduction of periodontal pathogens adhesion by antagonistic strains. Oral Microbiol Immunol. 2008;23:43–8.
Segata N, Waldron L, Ballarini A, Narasimhan V, Jousson O, Huttenhower C. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods. 2012;9:811–4.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Daims H, Wagner M. In situ techniques and digital image analysis methods for quantifying spatial localization patterns of nitrifiers and other microorganisms in biofilm and flocs. Methods Enzymol. 2011;496:185–215.
Daims H, Lücker S, Wagner M. daime, a novel image analysis program for microbial ecology and biofilm research. Environ Microbiol. 2006;8:200–13.
Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S, Olsen GJ, et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep. 2015;5:8365.
Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T, et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2014;42:D206–14.
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Anders S, Pyl PT, Huber W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Acknowledgements
We thank Janet Atoyan and the RI-EPSCOR sequencing facility at URI for sequence generation, Jonathan Livny and the Microbial ‘Omics Core and Genomics Platform for their help with RNASeq library sequencing and guidance on experimental design, the RI-INBRE/SURF program for supporting summer undergraduate research for KCL, the URI Science and Engineering fellows program for supporting summer undergraduate research for KC and the Annual Mark Wilson conference attendees for many valuable suggestions and discussion.
Funding
This work was funded by the NIDCR/NIH (R01DE027958—MR, JMW), (R01DE022586—JMW), NIGMS/RI-INBRE early career development award (P20GM103430—MR) and the Rhode Island Foundation Medical Research Fund (20164348—MR). This work was supported by the USDA National Institute of Food and Agriculture, Hatch Formula project accession number 1017848.
Author information
Authors and Affiliations
Contributions
MR and JMW designed research; DP, AM, VML, KCL, EA, and KC performed research, JMW, AM, DP and MR wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Perera, D., McLean, A., Morillo-López, V. et al. Mechanisms underlying interactions between two abundant oral commensal bacteria. ISME J 16, 948–957 (2022). https://doi.org/10.1038/s41396-021-01141-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01141-3
This article is cited by
-
The oral microbiome: diversity, biogeography and human health
Nature Reviews Microbiology (2024)
-
Corncob structures in dental plaque reveal microhabitat taxon specificity
Microbiome (2022)