Abstract
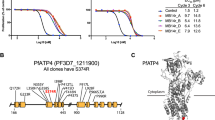
Antimalarial compounds with dual therapeutic and transmission-blocking activity are desired as high-value partners for combination therapies. Here, we report the identification and characterization of hexahydroquinolines (HHQs) that show low nanomolar potency against both pathogenic and transmissible intra-erythrocytic forms of the malaria parasite Plasmodium falciparum. This activity translates into potent transmission-blocking potential, as shown by in vitro male gamete formation assays and reduced oocyst infection and prevalence in Anopheles mosquitoes. In vivo studies illustrated the ability of lead HHQs to suppress Plasmodium berghei blood-stage parasite proliferation. Resistance selection studies, confirmed by CRISPR–Cas9-based gene editing, identified the digestive vacuole membrane-spanning transporter PfMDR1 (P. falciparum multidrug resistance gene-1) as a determinant of parasite resistance to HHQs. Haemoglobin and haem fractionation assays suggest a mode of action that results in reduced haemozoin levels and might involve inhibition of host haemoglobin uptake into intra-erythrocytic parasites. Furthermore, parasites resistant to HHQs displayed increased susceptibility to several first-line antimalarial drugs, including lumefantrine, confirming that HHQs have a different mode of action to other antimalarials drugs for which PfMDR1 is known to confer resistance. This work evokes therapeutic strategies that combine opposing selective pressures on this parasite transporter as an approach to countering the emergence and transmission of multidrug-resistant P. falciparum malaria.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
World Malaria Report 2016 (WHO, 2016); http://www.who.int/malaria/publications/world-malaria-report-2016
Wells, T. N. C., van Huijsduijnen, R. H. & Van Voorhis, W. C. Malaria medicines: a glass half full? Nat. Rev. Drug Discov. 14, 424–442 (2015).
Sinden, R. E. Targeting the parasite to suppress malaria transmission. Adv. Parasitol. 97, 147–185 (2017).
Josling, G. A. & Llinás, M. Sexual development in Plasmodium parasites: knowing when it’s time to commit. Nat. Rev. Microbiol. 13, 573–587 (2015).
Bousema, T. & Drakeley, C. Epidemiology and infectivity of Plasmodium falciparum and Plasmodium vivax gametocytes in relation to malaria control and elimination. Clin. Microbiol. Rev. 24, 377–410 (2011).
Cowman, A. F., Healer, J., Marapana, D. & Marsh, K. Malaria: biology and disease. Cell 167, 610–624 (2016).
Dicko, A. et al. Primaquine to reduce transmission of Plasmodium falciparum malaria in Mali: a single-blind, dose-ranging, adaptive randomised phase 2 trial. Lancet Infect. Dis. 16, 674–684 (2016).
White, N. J. Primaquine to prevent transmission of falciparum malaria. Lancet Infect. Dis. 13, 175–181 (2013).
Tanaka, T. Q. et al. A quantitative high throughput assay for identifying gametocytocidal compounds. Mol. Biochem. Parasitol. 188, 20–25 (2013).
Cevenini, L. et al. Multicolor bioluminescence boosts malaria research: quantitative dual-color assay and single-cell imaging in Plasmodium falciparum parasites. Anal. Chem. 86, 8814–8821 (2014).
Sanders, N. G., Sullivan, D. J., Mlambo, G., Dimopoulos, G. & Tripathi, A. K. Gametocytocidal screen identifies novel chemical classes with Plasmodium falciparum transmission blocking activity. PLoS ONE 9, e105817 (2014).
Bolscher, J. M. et al. A combination of new screening assays for prioritization of transmission-blocking antimalarials reveals distinct dynamics of marketed and experimental drugs. J. Antimicrob. Chemother. 70, 1357–1366 (2014).
Lucantoni, L., Duffy, S., Adjalley, S. H., Fidock, D. A. & Avery, V. M. Identification of MMV malaria box inhibitors of Plasmodium falciparum early-stage gametocytes using a luciferase-based high-throughput assay. Antimicrob. Agents Chemother. 57, 6050–6062 (2013).
Sun, W. et al. Chemical signatures and new drug targets for gametocytocidal drug development. Sci. Rep. 4, 3743 (2014).
Bowman, J. D. et al. Antiapicoplast and gametocytocidal screening to identify the mechanisms of action of compounds within the malaria box. Antimicrob. Agents Chemother. 58, 811–819 (2014).
Plouffe, D. M. et al. High-throughput assay and discovery of small molecules that interrupt malaria transmission. Cell Host Microbe 19, 114–126 (2016).
Miguel-Blanco, C. et al. Imaging-based high-throughput screening assay to identify new molecules with transmission-blocking potential against Plasmodium falciparum female gamete formation. Antimicrob. Agents Chemother. 59, 3298–3305 (2015).
Lucantoni, L., Loganathan, S. & Avery, V. M. The need to compare: assessing the level of agreement of three high-throughput assays against Plasmodium falciparum mature gametocytes. Sci. Rep. 7, 45992 (2017).
Delves, M. J. et al. Routine in vitro culture of P. falciparum gametocytes to evaluate novel transmission-blocking interventions. Nat. Protoc. 11, 1668–1680 (2016).
de Koning-Ward, T. F., Gilson, P. R. & Crabb, B. S. Advances in molecular genetic systems in malaria. Nat. Rev. Microbiol. 13, 373–387 (2015).
Flannery, E. L., Fidock, D. A. & Winzeler, E. A. Using genetic methods to define the targets of compounds with antimalarial activity. J. Med. Chem. 56, 7761–7771 (2013).
Plouffe, D. et al. In silico activity profiling reveals the mechanism of action of antimalarials discovered in a high-throughput screen. Proc. Natl Acad. Sci. USA 105, 9059–9064 (2008).
Adjalley, S. H. et al. Quantitative assessment of Plasmodium falciparum sexual development reveals potent transmission-blocking activity by methylene blue. Proc. Natl Acad. Sci. USA 108, E1214–E1223 (2011).
Coulibaly, B. et al. Efficacy and safety of triple combination therapy with artesunate-amodiaquine-methylene blue for falciparum malaria in children: a randomized controlled trial in Burkina Faso. J. Infect. Dis. 211, 689–697 (2015).
Fidock, D. A., Rosenthal, P. J., Croft, S. L., Brun, R. & Nwaka, S. Antimalarial drug discovery: efficacy models for compound screening. Nat. Rev. Drug Discov. 3, 509–520 (2004).
Corey, V. C. et al. A broad analysis of resistance development in the malaria parasite. Nat. Commun. 7, 11901 (2016).
Ng, C. L. et al. CRISPR-Cas9-modified pfmdr1 protects Plasmodium falciparum asexual blood stages and gametocytes against a class of piperazine-containing compounds but potentiates artemisinin-based combination therapy partner drugs. Mol. Microbiol. 101, 381–393 (2016).
Ferreira, P. E. et al. PfMDR1: mechanisms of transport modulation by functional polymorphisms. PLoS ONE 6, 3–10 (2011).
Sidhu, A. B. S., Valderramos, S. G. & Fidock, D. A. pfmdr1 mutations contribute to quinine resistance and enhance mefloquine and artemisinin sensitivity in Plasmodium falciparum. Mol. Microbiol. 57, 913–926 (2005).
Veiga, M. I. et al. Globally prevalent PfMDR1 mutations modulate Plasmodium falciparum susceptibility to artemisinin-based combination therapies. Nat. Commun. 7, 11553 (2016).
Sá, J. M. et al. Geographic patterns of Plasmodium falciparum drug resistance distinguished by differential responses to amodiaquine and chloroquine. Proc. Natl Acad. Sci. USA 106, 18883–18889 (2009).
Combrinck, J. M. et al. Optimization of a multi-well colorimetric assay to determine haem species in Plasmodium falciparum in the presence of anti-malarials. Malar. J. 14, 253 (2015).
Combrinck, J. M. et al. Insights into the role of heme in the mechanism of action of antimalarials. ACS Chem. Biol. 8, 133–137 (2013).
Fukami, T. & Yokoi, T. The emerging role of human esterases. Drug Metab. Pharmacokinet. 27, 466–477 (2012).
Blagborough, A. M. et al. Transmission-blocking interventions eliminate malaria from laboratory populations. Nat. Commun. 4, 1812–1817 (2013).
De Mello, C. X. et al. Comparison of the sensitivity of imprint and scraping techniques in the diagnosis of American tegumentary leishmaniasis in a referral centre in Rio de Janeiro, Brazil. Parasitol. Res. 109, 927–933 (2011).
Gil, J. P. & Krishna, S. pfmdr1 (Plasmodium falciparum multidrug drug resistance gene 1): a pivotal factor in malaria resistance to artemisinin combination therapies. Expert Rev. Anti. Infect. Ther. 15, 527–543 (2017).
Sigala, P. A. & Goldberg, D. E. The peculiarities and paradoxes of Plasmodium heme metabolism. Annu. Rev. Microbiol. 68, 259–278 (2014).
Tilley, L., Straimer, J., Gnädig, N. F., Ralph, S. A. & Fidock, D. A. Artemisinin action and resistance in Plasmodium falciparum. Trends Parasitol. 32, 682–696 (2016).
Venkatesan, M. et al. Polymorphisms in Plasmodium falciparum chloroquine resistance transporter and multidrug resistance 1 genes: parasite risk factors that affect treatment outcomes for P. falciparum malaria after artemether-lumefantrine and artesunate-amodiaquine. Am. J. Trop. Med. Hyg. 91, 833–843 (2014).
Petersen, I., Eastman, R. & Lanzer, M. Drug-resistant malaria: molecular mechanisms and implications for public health. FEBS Lett. 585, 1551–1562 (2011).
Petersen, I. et al. Balancing drug resistance and growth rates via compensatory mutations in the Plasmodium falciparum chloroquine resistance transporter. Mol. Microbiol. 97, 381–395 (2015).
Roberts, L., Egan, T. J., Joiner, K. A. & Hoppe, H. C. Differential effects of quinoline antimalarials on endocytosis in Plasmodium falciparum. Antimicrob. Agents Chemother. 52, 1840–1842 (2008).
Wunderlich, J., Rohrbach, P. & Dalton, J. P. The malaria digestive vacuole. Front. Biosci. 1, 1424–1448 (2012).
Rohrbach, P. et al. Genetic linkage of pfmdr1 with food vacuolar solute import in Plasmodium falciparum. EMBO J. 25, 3000–3011 (2006).
Reiling, S. J. & Rohrbach, P. Monitoring PfMDR1 transport in Plasmodium falciparum. Malar. J. 14, 270 (2015).
Brunner, R. et al. UV-triggered affinity capture identifies interactions between the Plasmodium falciparum multidrug resistance protein 1 (PfMDR1) and antimalarial agents in live parasitized cells. J. Biol. Chem. 288, 22576–22583 (2013).
Pleeter, P., Lekostaj, J. K. & Roepe, P. D. Purified Plasmodium falciparum multi-drug resistance protein (PfMDR 1) binds a high affinity chloroquine analogue. Mol. Biochem. Parasitol. 173, 158–161 (2010).
Cowman, A. F., Karcz, S., Galatis, D. & Culvenor, J. G. A P-glycoprotein homologue of Plasmodium falciparum is localized on the digestive vacuole. J. Cell Biol. 113, 1033–1042 (1991).
Tumwebaze, P. et al. Changing antimalarial drug resistance patterns identified by surveillance at three sites in Uganda. J. Infect. Dis. 215, 631–635 (2017).
Rottmann, M. et al. Spiroindolones, a potent compound class for the treatment of malaria. Science 329, 1175–1180 (2010).
Sidhu, S. B. A., Uhlemann, A., Valderramos, S. G., Krishna, S. & Fidock, D. A. Decreasing pfmdr1 copy number in Plasmodium falciparum malaria heightens susceptibility to mefloquine, lumefantrine, halofantrine, quinine and artemisinin. J. Infect. Dis. 194, 528–535 (2006).
ChEMBL-NTD (EMBL-EBI); https://www.ebi.ac.uk/chemblntd
Duffy, S. & Avery, V. M. Identification of inhibitors of Plasmodium falciparum gametocyte development. Malar. J. 12, 408 (2013).
Lucantoni, L., Fidock, D. A. & Avery, V. M. A luciferase-based, high-throughput assay for screening and profiling transmission-blocking compounds against Plasmodium falciparum gametocytes. Antimicrob. Agents Chemother. 60, 2097–2107 (2016).
Ruecker, A. et al. A male and female gametocyte functional viability assay to identify biologically relevant malaria transmission-blocking drugs. Antimicrob. Agents Chemother. 58, 7292–7304 (2014).
Duffy, S. & Avery, V. M. Development and optimization of a novel 384-well anti-malarial imaging assay validated for high-throughput screening. Am. J. Trop. Med. Hyg. 86, 84–92 (2012).
Fletcher, S. & Avery, V. M. A novel approach for the discovery of chemically diverse anti-malarial compounds targeting the Plasmodium falciparum coenzyme A synthesis pathway. Malar. J. 13, 343 (2014).
Delves, M. J. et al. Male and female Plasmodium falciparum mature gametocytes show different responses to antimalarial drugs. Antimicrob. Agents Chemother. 57, 3268–3274 (2013).
McNamara, C. W. et al. Targeting Plasmodium PI(4)K to eliminate malaria. Nature 504, 248–253 (2013).
Pereira, M. R. et al. In vivo and in vitro antimalarial properties of azithromycin–chloroquine combinations that include the resistance reversal agent amlodipine. Antimicrob. Agents Chemother. 55, 3115–3124 (2011).
Goodyer, I. D. & Taraschi, T. F. Plasmodium falciparum: a simple, rapid method for detecting parasite clones in microtiter plates. Exp. Parasitol. 86, 158–160 (1997).
Manary, M. J. et al. Identification of pathogen genomic variants through an integrated pipeline. BMC Bioinformatics 15, 63–76 (2014).
Hameed, P. S. et al. Triaminopyrimidine is a fast-killing and long-acting antimalarial clinical candidate. Nat. Commun. 6, 6715 (2015).
Ekland, E. H., Schneider, J. & Fidock, D. A. Identifying apicoplast-targeting antimalarials using high-throughput compatible approaches. FASEB J. 25, 3583–3593 (2011).
D’Alessandro, S. et al. A Plasmodium falciparum screening assay for anti-gametocyte drugs based on parasite lactate dehydrogenase detection. J. Antimicrob. Chemother. 68, 2048–2058 (2013).
Tsirigos, K. D., Peters, C., Shu, N., Lukas, K. & Elofsson, A. The TOPCONS web server for consensus prediction of membrane protein topology and signal peptides. Nucleic Acids Res. 43, W401–407 (2015).
Alva, V. et al. The MPI Bioinformatics Toolkit as an integrative platform for advanced protein sequence and structure. Nucleic Acids Res. 44, W410–415 (2016).
Sali, A., Potterton, L., Yuan, F., Van Vlijmen, H. & Karplus, M. Evaluation of comparative protein modeling by MODELLER. Proteins 23, 318–326 (1995).
Emsley, P. & Lohkamp, B. Features and development of Coot. Acta Crystallogr. D D66, 486–501 (2010).
Krieger, E. & Vriend, G. Models @ Home: distributed computing in bioinformatics using a screensaver based approach. Bioinformatics 18, 315–318 (2002).
Bikadi, Z. & Hazai, E. Application of the PM6 semi-empirical method to modeling proteins enhances docking accuracy of AutoDock. J. Cheminform 1, 15 (2009).
Acknowledgements
The authors thank T.T. Diagana (Novartis Institute for Tropical Diseases, Singapore) for provision of the compounds, the Red Cross (Australia and the USA) for the provision of human blood for cell cultures, and G. Stevenson for assistance with the triaging of compounds following screening. The authors acknowledge the Bill and Melinda Gates Foundation (grant OPP1040399 to D.A.F. and V.M.A. and grant OPP1054480 to E.A.W. and D.A.F.), the National Institutes of Health (grant R01 AI103058 to E.A.W. and D.A.F., grant R01 AI50234 to D.A.F, and R01 AI110329 to T.J.E.), the Australian Research Council (LP120200557 to V.M.A.) and the Medicines for Malaria Venture for their continued support. P.E.F. and M.I.V. are supported by the Northern Portugal Regional Operational Programme (NORTE 2020), under the Portugal 2020 Partnership Agreement, through the European Regional Development Fund (FEDER).
Author information
Authors and Affiliations
Contributions
L.L. and S.D. screened the Novartis-GNF Malaria Box against early- and late-stage GAMs using LUC or GFP-imaging technologies, assayed ABS parasites and mammalian cells and confirmed compound potency. L.L., S.D. and V.M.A. analysed screening data and selected the HHQs for further evaluation. T.L., K.L.S. and S.L.H. contributed in vitro membrane feeding and male gametocyte exflagellation assay data. A.R., R.E.S. and M.D. contributed the DGFA data. M.V. and T.R.S.K. performed in vivo efficacy studies. K.R., M.V. and T.R.S.K. performed in vivo transmission-blocking studies. S.G. performed selections for HHQ-resistant lines, which were cloned by O.L. and M.V. Whole-genome sequence analysis was performed by V.C.C., P.P.H. and E.A.W. Gene editing of pfmdr1 was performed by M.V. with help from C.L.N. and D.A.F. Susceptibility assays on resistant lines with clinical and experimental antimalarials were performed by M.V. with help from J.M.M. Early- and late-stage GAM susceptibility testing was performed by G.S. and P.A. Haem fractionation data were provided by J.M.C. and T.J.E. PfMDR1 modelling studies were performed by P.E.F and M.I.V. Data were compiled by M.V., L.L. and D.A.F., who wrote the manuscript with input from V.M.A. All authors approved of the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary Information
Supplementary Information
Supplementary Tables 1–17, Supplementary Figures 1–5.
Rights and permissions
About this article
Cite this article
Vanaerschot, M., Lucantoni, L., Li, T. et al. Hexahydroquinolines are antimalarial candidates with potent blood-stage and transmission-blocking activity. Nat Microbiol 2, 1403–1414 (2017). https://doi.org/10.1038/s41564-017-0007-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0007-4
This article is cited by
-
Catalyst-free synthesis of highly functionalized triazole hexahydroquinoline carbohydrazide scaffolds via four-component cyclocondensation reaction
Molecular Diversity (2024)
-
Identification and characterisation of the haemozoin of Haemonchus contortus
Parasites & Vectors (2023)
-
Isoliensinine from Cissampelos pariera rhizomes exhibits potential gametocytocidal and anti-malarial activities against Plasmodium falciparum clinical isolates
Malaria Journal (2023)
-
Novel magnetic nanoparticles with morpholine tags as multirole catalyst for synthesis of hexahydroquinolines and 2-amino-4,6-diphenylnicotinonitriles through vinylogous anomeric-based oxidation
Research on Chemical Intermediates (2019)
-
5-Oxo-hexahydroquinoline: an attractive scaffold with diverse biological activities
Molecular Diversity (2019)