Abstract
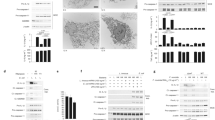
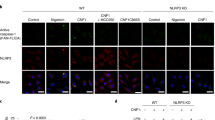
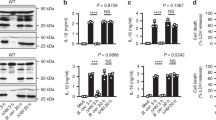
Group A Streptococcus (GAS) is among the top ten causes of infection-related mortality in humans. M protein is the most abundant GAS surface protein, and M1 serotype GAS strains are associated with invasive infections, including necrotizing fasciitis and toxic shock syndrome. Here, we report that released, soluble M1 protein triggers programmed cell death in macrophages (Mϕ). M1 served as a second signal for caspase-1-dependent NLRP3 inflammasome activation, inducing maturation and release of proinflammatory cytokine interleukin-1β (IL-1β) and macrophage pyroptosis. The structurally dynamic B-repeat domain of M1 was critical for inflammasome activation, which involved K+ efflux and M1 protein internalization by clathrin-mediated endocytosis. Mouse intraperitoneal challenge showed that soluble M1 was sufficient and specific for IL-1β activation, which may represent an early warning to activate host immunity against the pathogen. Conversely, in systemic infection, hyperinflammation associated with M1-mediated pyroptosis and IL-1β release could aggravate tissue injury.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Walker, M. J. et al. Disease manifestations and pathogenic mechanisms of group A Streptococcus. Clin. Microbiol. Rev. 27, 264–301 (2014).
Carapetis, J. R., Steer, A. C., Mulholland, E. K. & Weber, M. The global burden of group A streptococcal diseases. Lancet Infect. Dis. 5, 685–694 (2005).
Severin, A. et al. Proteomic analysis and identification of Streptococcus pyogenes surface-associated proteins. J. Bacteriol. 189, 1514–1522 (2007).
Phillips, G. N. Jr, Flicker, P. F., Cohen, C., Manjula, B. N. & Fischetti, V. A. Streptococcal M protein: α-helical coiled-coil structure and arrangement on the cell surface. Proc. Natl Acad. Sci. USA 78, 4689–4693 (1981).
Ghosh, P. The nonideal coiled coil of M protein and its multifarious functions in pathogenesis. Adv. Exp. Med. Biol. 715, 197–211 (2011).
Sanderson-Smith, M. et al. A systematic and functional classification of Streptococcus pyogenes that serves as a new tool for molecular typing and vaccine development. J. Infect. Dis. 210, 1325–1338 (2014).
Steer, A. C., Law, I., Matatolu, L., Beall, B. W. & Carapetis, J. R. Global emm type distribution of group A streptococci: systematic review and implications for vaccine development. Lancet Infect. Dis. 9, 611–616 (2009).
Zhu, L. et al. A molecular trigger for intercontinental epidemics of group A Streptococcus. J. Clin. Invest. 125, 3545–3559 (2015).
Okada, N., Liszewski, M. K., Atkinson, J. P. & Caparon, M. Membrane cofactor protein (CD46) is a keratinocyte receptor for the M protein of the group A Streptococcus. Proc. Natl Acad. Sci. USA 92, 2489–2493 (1995).
Oehmcke, S., Shannon, O., Morgelin, M. & Herwald, H. Streptococcal M proteins and their role as virulence determinants. Clin. Chim. Acta 411, 1172–1180 (2010).
LaRock, C. N. et al. Group A streptococcal M1 protein sequesters cathelicidin to evade innate immune killing. Cell Host Microbe 18, 471–477 (2015).
Dohrmann, S. et al. Group A streptococcal M1 protein provides resistance against the antimicrobial activity of histones. Sci. Rep. 7, 43039 (2017).
Herwald, H. et al. M protein, a classical bacterial virulence determinant, forms complexes with fibrinogen that induce vascular leakage. Cell 116, 367–379 (2004).
Macheboeuf, P. et al. Streptococcal M1 protein constructs a pathological host fibrinogen network. Nature 472, 64–68 (2011).
Soehnlein, O. et al. Neutrophil primary granule proteins HBP and HNP1-3 boost bacterial phagocytosis by human and murine macrophages. J. Clin. Invest. 118, 3491–3502 (2008).
Akesson, P., Schmidt, K. H., Cooney, J. & Bjorck, L. M1 protein and protein H: IgGFc- and albumin-binding streptococcal surface proteins encoded by adjacent genes. Biochem. J. 300, 877–886 (1994).
Kansal, R. G., McGeer, A., Low, D. E., Norrby-Teglund, A. & Kotb, M. Inverse relation between disease severity and expression of the streptococcal cysteine protease, SpeB, among clonal M1T1 isolates recovered from invasive group A streptococcal infection cases. Infect. Immun. 68, 6362–6369 (2000).
Sumby, P., Whitney, A. R., Graviss, E. A., DeLeo, F. R. & Musser, J. M. Genome-wide analysis of group A streptococci reveals a mutation that modulates global phenotype and disease specificity. PLoS Pathog. 2, e5 (2006).
Cole, J. N., Barnett, T. C., Nizet, V. & Walker, M. J. Molecular insight into invasive group A streptococcal disease. Nat. Rev. Microbiol. 9, 724–736 (2011).
Martinon, F., Burns, K. & Tschopp, J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-β. Mol. Cell 10, 417–426 (2002).
Miao, E. A., Rajan, J. V. & Aderem, A. Caspase-1-induced pyroptotic cell death. Immunol. Rev. 243, 206–214 (2011).
Franchi, L., Munoz-Planillo, R. & Nunez, G. Sensing and reacting to microbes through the inflammasomes. Nat. Immunol. 13, 325–332 (2012).
LaRock, C. N. & Nizet, V. Inflammasome/IL-1β responses to streptococcal pathogens. Front. Immunol. 6, 518 (2015).
Broderick, L., De Nardo, D., Franklin, B. S., Hoffman, H. M. & Latz, E. The inflammasomes and autoinflammatory syndromes. Annu. Rev. Pathol. 10, 395–424 (2015).
Wen, H., Ting, J. P. & O’Neill, L. A. A role for the NLRP3 inflammasome in metabolic diseases—did Warburg miss inflammation? Nat. Immunol. 13, 352–357 (2012).
Harder, J. et al. Activation of the Nlrp3 inflammasome by Streptococcus pyogenes requires streptolysin O and NF-κB activation but proceeds independently of TLR signaling and P2X7 receptor. J. Immunol. 183, 5823–5829 (2009).
Lin, A. E. et al. A group A Streptococcus ADP-ribosyltransferase toxin stimulates a protective interleukin 1β-dependent macrophage immune response. mBio 6, e00133 (2015).
Bauernfeind, F. G. et al. Cutting edge: NF-κB activating pattern recognition and cytokine receptors license NLRP3 inflammasome activation by regulating NLRP3 expression. J. Immunol. 183, 787–791 (2009).
Guo, H., Callaway, J. B. & Ting, J. P. Inflammasomes: mechanism of action, role in disease, and therapeutics. Nat. Med. 21, 677–687 (2015).
Gaidt, M. M. et al. Human monocytes engage an alternative inflammasome pathway. Immunity 44, 833–846 (2016).
Mariathasan, S. et al. Cryopyrin activates the inflammasome in response to toxins and ATP. Nature 440, 228–232 (2006).
Coll, R. C. et al. A small-molecule inhibitor of the NLRP3 inflammasome for the treatment of inflammatory diseases. Nat. Med. 21, 248–255 (2015).
Munoz-Planillo, R. et al. K+ efflux is the common trigger of NLRP3 inflammasome activation by bacterial toxins and particulate matter. Immunity 38, 1142–1153 (2013).
McNamara, C. et al. Coiled-coil irregularities and instabilities in group A Streptococcus M1 are required for virulence. Science 319, 1405–1408 (2008).
LaRock, C. N. et al. IL-1β is an innate immune sensor of microbial proteolysis. Sci. Immunol. 1, eaah3539 (2016).
Hsu, L. C. et al. IL-1β-driven neutrophilia preserves antibacterial defense in the absence of the kinase IKKβ. Nat. Immunol. 12, 144–150 (2011).
Persson, S. T., Wilk, L., Morgelin, M. & Herwald, H. Vigilant keratinocytes trigger pathogen-associated molecular pattern signaling in response to streptococcal M1 protein. Infect. Immun. 83, 4673–4681 (2015).
Pahlman, L. I. et al. Streptococcal M protein: a multipotent and powerful inducer of inflammation. J. Immunol. 177, 1221–1228 (2006).
Chattergoon, M. A. et al. HIV and HCV activate the inflammasome in monocytes and macrophages via endosomal Toll-like receptors without induction of type 1 interferon. PLoS Pathog. 10, e1004082 (2014).
Marina-Garcia, N. et al. Clathrin- and dynamin-dependent endocytic pathway regulates muramyl dipeptide internalization and NOD2 activation. J. Immunol. 182, 4321–4327 (2009).
Stewart, C. et al. Coiled-coil destabilizing residues in the group A Streptococcus M1 protein are required for functional interaction. Proc. Natl Acad. Sci. USA 113, 9515–9520 (2016).
Buffalo, C. Z. et al. Conserved patterns hidden within group A Streptococcus M protein hypervariability recognize human C4b-binding protein. Nat. Microbiol. 1, 16155 (2016).
Hornung, V. et al. Silica crystals and aluminum salts activate the NALP3 inflammasome through phagosomal destabilization. Nat. Immunol. 9, 847–856 (2008).
Chatellier, S. et al. Genetic relatedness and superantigen expression in group A Streptococcus serotype M1 isolates from patients with severe and nonsevere invasive diseases. Infect. Immun. 68, 3523–3534 (2000).
Lauth, X. et al. M1 protein allows Group A streptococcal survival in phagocyte extracellular traps through cathelicidin inhibition. J. Innate Immun. 1, 202–214 (2009).
Timmer, A. M. et al. Streptolysin O promotes group A Streptococcus immune evasion by accelerated macrophage apoptosis. J. Biol. Chem. 284, 862–871 (2009).
McCloy, R. A. et al. Partial inhibition of Cdk1 in G 2 phase overrides the SAC and decouples mitotic events. Cell Cycle 13, 1400–1412 (2014).
Acknowledgements
The authors thank the members of the P.G. and V.N. laboratories for their valuable input. This work was supported by NIH grants AI096837 (P.G. and V.N), AI077780 (V.N.) and AI52430 (H.M.H.) A.M.R. is a San Diego IRACDA Program fellow supported by NIGMS/NIH award K12GM068524, N.J.G. is supported by the UCSD Genetics Training Program (T32 GM008666) and the UCSD Global Health Institute and C.N.L. is a recipient of an A.P. Giannini Foundation Postdoctoral Research Fellowship.
Author information
Authors and Affiliations
Contributions
J.A.V., P.G. and V.N. formulated the original hypothesis, designed the study and analysed the results. J.A.V., A.M.R., N.J.G., C.N.L., N.G. and S.R.A. performed and optimized experiments. H.M.H. provided novel reagents. J.A.V., P.G. and V.N. wrote the manuscript and all authors reviewed the manuscript, data and conclusions before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Figures 1–10.
Rights and permissions
About this article
Cite this article
Valderrama, J.A., Riestra, A.M., Gao, N.J. et al. Group A streptococcal M protein activates the NLRP3 inflammasome. Nat Microbiol 2, 1425–1434 (2017). https://doi.org/10.1038/s41564-017-0005-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-017-0005-6
This article is cited by
-
Apoptosis in post-streptococcal glomerulonephritis and mechanisms for failed of inflammation resolution
Pediatric Nephrology (2024)
-
Pathogenesis, epidemiology and control of Group A Streptococcus infection
Nature Reviews Microbiology (2023)
-
Inflammasomes and Childhood Autoimmune Diseases: A Review of Current Knowledge
Clinical Reviews in Allergy & Immunology (2021)
-
NLRP3 inflammasome-mediated microglial pyroptosis is critically involved in the development of post-cardiac arrest brain injury
Journal of Neuroinflammation (2020)
-
NLRP3-dependent pyroptosis is required for HIV-1 gp120-induced neuropathology
Cellular & Molecular Immunology (2020)