Abstract
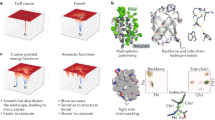
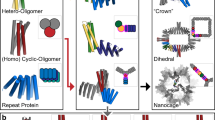
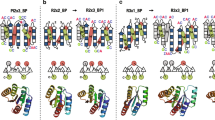
Protein crystallization plays a central role in structural biology. Despite this, the process of crystallization remains poorly understood and highly empirical, with crystal contacts, lattice packing arrangements and space group preferences being largely unpredictable. Programming protein crystallization through precisely engineered side-chain–side-chain interactions across protein–protein interfaces is an outstanding challenge. Here we develop a general computational approach for designing three-dimensional protein crystals with prespecified lattice architectures at atomic accuracy that hierarchically constrains the overall number of degrees of freedom of the system. We design three pairs of oligomers that can be individually purified, and upon mixing, spontaneously self-assemble into >100 µm three-dimensional crystals. The structures of these crystals are nearly identical to the computational design models, closely corresponding in both overall architecture and the specific protein–protein interactions. The dimensions of the crystal unit cell can be systematically redesigned while retaining the space group symmetry and overall architecture, and the crystals are extremely porous and highly stable. Our approach enables the computational design of protein crystals with high accuracy, and the designed protein crystals, which have both structural and assembly information encoded in their primary sequences, provide a powerful platform for biological materials engineering.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available in the main text or as supplementary materials. Design scripts examples and design models (cages including T32-15, T32-15-6H, O43-2, T33-ECY54, T33-ECY55, T33-ECY59, T33-ECY66, T33-ECY67, O43-UWN453, O43-ZL1, O43-ZL7 and O32-ZL4, and crystal contact dihedrals of F4132-1-0, F4132-2, I432-1, F4132-2-ex1, F4132-2-ex2 and F4132-2-ex3) are available in the supplementary information files and through Zenodo79. SAXS and hyperspectral data are available as source data files. Crystallographic datasets have been deposited in the PDB (accession codes 8CUU, 8CUV, 8CUW, 8CUX, 8CWS, 8CUS, 8CUT, 8CWZ and 8FAR). CryoEM maps and corresponding atomic models have been deposited in the PDB (accession codes 8CWY and 8SZZ) and the Electron Microscopy Data Bank (accession codes EMD-27031 and EMD-40926). Source data are provided with this paper.
References
Berman, H. M. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Rupp, B. Biomolecular Crystallography: Principles, Practice, and Application to Structural Biology (Garland Science, 2010).
Mcpherson, A. Introduction to protein crystallization. Methods 34, 254–265 (2004).
Desiraju, G. R. Crystal engineering: a holistic view. Angew. Chem. Int. Ed. 46, 8342–8356 (2007).
Sontz, P. A., Bailey, J. B., Ahn, S. & Tezcan, F. A. A metal organic framework with spherical protein nodes: rational chemical design of 3D protein crystals. J. Am. Chem. Soc. 137, 11598–11601 (2015).
Subramanian, R. H. et al. Design of metal-mediated protein assemblies via hydroxamic acid functionalities. Nat. Protoc. 16, 3264–3297 (2021).
Kostiainen, M. A. et al. Electrostatic assembly of binary nanoparticle superlattices using protein cages. Nat. Nanotechnol. 8, 52–56 (2013).
Liljeström, V., Mikkilä, J. & Kostiainen, M. A. Self-assembly and modular functionalization of three-dimensional crystals from oppositely charged proteins. Nat. Commun. 5, 4445 (2014).
Brodin, J. D., Auyeung, E. & Mirkin, C. A. DNA-mediated engineering of multicomponent enzyme crystals. Proc. Natl Acad. Sci. USA 112, 4564–4569 (2015).
Partridge, B. E., Winegar, P. H., Han, Z. & Mirkin, C. A. Redefining protein interfaces within protein single crystals with DNA. J. Am. Chem. Soc. https://doi.org/10.1021/jacs.1c04191 (2021).
Zhou, K. et al. On-axis alignment of protein nanocage assemblies from 2D to 3D through the aromatic stacking interactions of amino acid residues. ACS Nano 12, 11323–11332 (2018).
Lanci, C. J. et al. Computational design of a protein crystal. Proc. Natl Acad. Sci. USA 109, 7304–7309 (2012).
Huang, P.-S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320–327 (2016).
Bai, Y., Luo, Q. & Liu, J. Protein self-assembly via supramolecular strategies. Chem. Soc. Rev. 45, 2756–2767 (2016).
Luo, Q., Hou, C., Bai, Y., Wang, R. & Liu, J. Protein assembly: versatile approaches to construct highly ordered nanostructures. Chem. Rev. 116, 13571–13632 (2016).
Zhu, J. et al. Protein assembly by design. Chem. Rev. 121, 13701–13796 (2021).
Hartje, L. F. & Snow, C. D. Protein crystal based materials for nanoscale applications in medicine and biotechnology. WIREs Nanomed. Nanobiotechnol. 11, e1547 (2019).
Heater, B. S., Yang, Z., Lee, M. M. & Chan, M. K. In vivo enzyme entrapment in a protein crystal. J. Am. Chem. Soc. 142, 9879–9883 (2020).
Conejero-Muriel, M., Rodríguez-Ruiz, I., Verdugo-Escamilla, C., Llobera, A. & Gavira, J. A. Continuous sensing photonic lab-on-a-chip platform based on cross-linked enzyme crystals. Anal. Chem. 88, 11919–11923 (2016).
Vilenchik, L. Z., Griffith, J. P., St. Clair, N., Navia, M. A. & Margolin, A. L. Protein crystals as novel microporous materials. J. Am. Chem. Soc. 120, 4290–4294 (1998).
Basu, S. K., Govardhan, C. P., Jung, C. W. & Margolin, A. L. Protein crystals for the delivery of biopharmaceuticals. Expert Opin. Biol. Ther. 4, 301–317 (2004).
Cotton, F. A. Chemical Applications of Group Theory (Wiley, 1990).
Yeates, T. O., Liu, Y. & Laniado, J. The design of symmetric protein nanomaterials comes of age in theory and practice. Curr. Opin. Struct. Biol. 39, 134–143 (2016).
Laniado, J. & Yeates, T. O. A complete rule set for designing symmetry combination materials from protein molecules. Proc. Natl Acad. Sci. USA 117, 31817–31823 (2020).
King, N. P. et al. Computational design of self-assembling protein nanomaterials with atomic level accuracy. Science 336, 1171–1174 (2012).
King, N. P. et al. Accurate design of co-assembling multi-component protein nanomaterials. Nature 510, 103–108 (2014).
Bale, J. B. et al. Structure of a designed tetrahedral protein assembly variant engineered to have improved soluble expression: designed protein tetrahedron. Protein Sci. 24, 1695–1701 (2015).
Ueda, G. et al. Tailored design of protein nanoparticle scaffolds for multivalent presentation of viral glycoprotein antigens. eLife 9, e57659 (2020).
Wang, J. Y. et al. Improving the secretion of designed protein assemblies through negative design of cryptic transmembrane domains. Proc. Natl Acad. Sci. USA 120, e2214556120 (2023).
Sheffler, W. et al. Fast and versatile sequence-independent protein docking for nanomaterials design using RPXDock. PLoS Comput. Biol. 19, e1010680 (2023).
Brunette, T. et al. Exploring the repeat protein universe through computational protein design. Nature 528, 580–584 (2015).
Boyken, S. E. et al. De novo design of protein homo-oligomers with modular hydrogen-bond network–mediated specificity. Science 352, 680–687 (2016).
Fallas, J. A. et al. Computational design of self-assembling cyclic protein homo-oligomers. Nat. Chem. 9, 353–360 (2017).
Boyken, S. E. et al. De novo design of tunable, pH-driven conformational changes. Science 364, 658–664 (2019).
Hsia, Y. et al. Design of multi-scale protein complexes by hierarchical building block fusion. Nat. Commun. 12, 2294 (2021).
Leman, J. K. et al. Macromolecular modeling and design in Rosetta: recent methods and frameworks. Nat. Methods 17, 665–680 (2020).
Wulff, G. On the question of speed of growth and dissolution of crystal surfaces. Z. Kristallogr. 34, 449–530 (1901).
Jeliazkov, J. R., Robinson, A. C., García-Moreno, E. B., Berger, J. M. & Gray, J. J. Toward the computational design of protein crystals with improved resolution. Acta Crystallogr. D 75, 1015–1027 (2019).
Lai, Y. T. et al. Structure of a designed protein cage that self-assembles into a highly porous cube. Nat. Chem. 6, 1065–1071 (2014).
Yan, E.-K. et al. Cross-linked protein crystals by glutaraldehyde and their applications. RSC Adv. 5, 26163–26174 (2015).
Lee, S. et al. Shape memory in self-adapting colloidal crystals. Nature 610, 674–679 (2022).
Künzle, M., Eckert, T. & Beck, T. Binary protein crystals for the assembly of inorganic nanoparticle superlattices. J. Am. Chem. Soc. 138, 12731–12734 (2016).
Tian, Y. et al. Ordered three-dimensional nanomaterials using DNA-prescribed and valence-controlled material voxels. Nat. Mater. 19, 789–796 (2020).
Sun, J. et al. Core-controlled polymorphism in virus-like particles. Proc. Natl Acad. Sci. USA 104, 1354–1359 (2007).
Lach, M., Strelow, C., Meyer, A., Mews, A. & Beck, T. Encapsulation of gold nanoparticles into redesigned ferritin nanocages for the assembly of binary superlattices composed of fluorophores and gold nanoparticles. ACS Appl. Mater. Interfaces 14, 10656–10668 (2022).
Schulz, F. et al. Structural order in plasmonic superlattices. Nat. Commun. 11, 3821 (2020).
Junker, N. O. et al. Optical properties of metacrystals based on protein nanocages. Adv. Funct. Mater. https://doi.org/10.1002/adfm.202303260 (2023).
Ross, M. B., Mirkin, C. A. & Schatz, G. C. Optical properties of one‑, two‑, and three-dimensional arrays of plasmonic nanostructures. J. Phys. Chem. C 120, 816–830 (2016).
Ross, M. B., Ku, J. C., Vaccarezza, V. M., Schatz, G. C. & Mirkin, C. A. Nanoscale form dictates mesoscale function in plasmonic DNA-nanoparticle superlattices. Nat. Nanotechnol. https://doi.org/10.1038/nnano.2015.68 (2015).
Fleishman, S. J. et al. RosettaScripts: a scripting language interface to the Rosetta macromolecular modeling suite. PLoS ONE 6, e20161 (2011).
The PyMOL Molecular Graphics System v.1.8 (Schrödinger, LLC, 2015).
Studier, F. W. Protein production by auto-induction in high-density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Schmitt, J., Hess, H. & Stunnenberg, H. G. Affinity purification of histidine-tagged proteins. Mol. Biol. Rep. 18, 223–230 (1993).
Tetter, S. & Hilvert, D. Enzyme encapsulation by a ferritin cage. Angew. Chem. 129, 15129–15132 (2017).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL -3000: the integration of data reduction and structure solution – from diffraction images to an initial model in minutes. Acta Crystallogr. D 62, 859–866 (2006).
Winn, M. D. et al. Overview of the CCP 4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Carragher, B. et al. Leginon: an automated system for acquisition of images from vitreous ice specimens. J. Struct. Biol. 132, 33–45 (2000).
Sun, M. et al. Practical considerations for using K3 cameras in CDS mode for high-resolution and high-throughput single particle cryo-EM. J. Struct. Biol. 213, 107745 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Wang, R. Y.-R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
DiMaio, F., Leaver-Fay, A., Bradley, P., Baker, D. & André, I. Modeling symmetric macromolecular structures in Rosetta3. PLoS ONE 6, e20450 (2011).
Barad, B. A. et al. EMRinger: side chain–directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Dyer, K. N. et al. High-throughput SAXS for the characterization of biomolecules in solution: a practical approach. Methods Mol. Biol. 1091, 245–258 (2013).
Classen, S. et al. Implementation and performance of SIBYLS: a dual endstation small-angle X-ray scattering and macromolecular crystallography beamline at the Advanced Light Source. J. Appl. Crystallogr. 46, 1–13 (2013).
Schneidman-Duhovny, D., Hammel, M. & Sali, A. FoXS: a web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Res. 38, W540–W544 (2010).
Wang, S. et al. The emergence of valency in colloidal crystals through electron equivalents. Nat. Mater. 21, 580–587 (2022).
Senesi, A. J. & Lee, B. Small-angle scattering of particle assemblies. J. Appl. Crystallogr. 48, 1172–1182 (2015).
Li, T., Senesi, A. J. & Lee, B. Small angle X-ray scattering for nanoparticle research. Chem. Rev. 116, 11128–11180 (2016).
Ross, M. B., Blaber, M. G. & Schatz, G. C. Using nanoscale and mesoscale anisotropy to engineer the optical response of three-dimensional plasmonic metamaterials. Nat. Commun. 5, 4090 (2014).
Coronado, E. A. & Schatz, G. C. Surface plasmon broadening for arbitrary shape nanoparticles: a geometrical probability approach. J. Chem. Phys. 119, 3926–3934 (2003).
Li, Z. et al. Data for: accurate computational design of 3D protein crystals. Zenodo https://doi.org/10.5281/zenodo.8299428 (2023).
Matthews, B. W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968).
Acknowledgements
We thank F. Busch and V. Wysocki at Ohio State University for support with native mass spectrometry experiments. We thank X. Zuo and T. Jun at Argonne National Laboratory for help with the SAXS measurements. We thank R. M. Haynes at the Pacific Northwest Cryo-EM Center for help with cryoEM data collection. We thank J. Du, S. Zhang and J. De Yoreo at the University of Washington for help with the crystallization measurements and for discussions. We thank I. Kopyeva and C. DeForest at the University of Washington for help with the mechanical measurements and for discussions. We thank D. Oberthür, V. Kremling, J. Sprenger, E. Scheer, B. Klopprogge and H. Chapman at the Center for Free-Electron Laser Science, Hamburg, Germany, for investigating the crystals on a free-electron laser. We thank S. Weaver, K. Patel and T. Gonen at the University of California, Los Angeles, for help with the microED. We thank S. Dickinson, N. Bethel and M. Wu at the University of Washington for help with cryoEM sample preparation and for screening. We thank S. Caldwell at the University of Washington for help with analysing the crystallographic data. We thank X. Wang at the University of Washington for suggestions on collecting X-ray crystallography data. We thank T. Huddy, R. Kibler, N. Bethel, A. Khmelinskaia, D. Zambrano and R. Haas at the University of Washington for providing potential protein building blocks. We thank H. Bai, R. Kibler, T. Huddy, H. Pyles, C. Xu and A. Ljubetic at the University of Washington for help with scripting and software. We thank R. Ravichandran for help with the bioreactor production of proteins. We thank J. Decarreau for help with the optical and fluorescence microscope imaging. We also thank members of the Baker lab and Institute for Protein Design, particularly J. Bale, N. King and F. Dimaio, for useful discussions. This work was supported with funds provided by the Howard Hughes Medical Institute (W.S. and D.B.), an Amgen gift (S.W.), Novo Nordisk (W.Y.), the Institute for Protein Design Directors Fund (M.J.B.), the Audacious Project at the Institute for Protein Design (Z.L., A.K.B., A.J.B., Q.D., R.D., A.F. and D.B.), the Open Philanthropy Project Improving Protein Design Fund (Y.H., H.N., N.I.E. and D.B.), the Synergistic Discovery and Design project HR001117S0003 of the Defense Advanced Research Projects Agency under contract FA8750-17-C-0219 (H.H. and D.B.), postdoctoral scholarships from the Washington Research Foundation (J.M.L. and H.H.), the Nordstrom Barrier Institute for Protein Design Directors Fund (A.E.), a Human Frontiers Science Program Long Term Fellowship (A.C.), a Public Health Service National Research Service Award (T32GM007270) from the National Institute of General Medical Sciences (NIGMS; U.N.), a Graduate Research Fellowship Program grant from the National Science Foundation (NSF DGE-1762114 to E.C.Y.) and a US Department of Energy (DOE), Office of Science, grant DE-SC0018940 (A.K. and D.B.). We thank the staff of the APS Northeastern Collaborative Access Team beamlines, APS beamline 24-ID-C, which are funded by the NIGMS from the National Institutes of Health (NIH; P30 GM124165), and the APS 12-ID-B SAXS beamline. This research used resources of APS, which is a DOE Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357. We also thank the ALS beamline 8.2.2/8.2.1 and SIBYLS Beamline 12.3.1 at Lawrence Berkeley National Laboratory. This research used ALS resources, which is a DOE Office of Science User Facility under contract no. DE-AC02-05CH11231 and the Integrated Diffraction Analysis Technologies grant. The ALS-ENABLE beamlines are supported by the NIH through NIGMS (Grant No. P30 GM124169-01). The Berkeley Center for Structural Biology is supported in part by the NIH, NIGMS and the Howard Hughes Medical Institute. A portion of this research was supported by NIH grant U24GM129547 and performed at the Pacific Northwest Center for Cryo-EM at Oregon Health & Science University and accessed through the Environmental Molecular Sciences Laboratory (grid.436923.9), which is a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research. This work (M.Y.Y. and D.S.G.) was supported with funds provided by the DOE, Office of Science, Office of Basic Energy Sciences, as part of the Energy Frontier Research Centers program of the Center for the Science of Synthesis Across Scales under Award Number DE-SC0019288, located at University of Washington.
Author information
Authors and Affiliations
Contributions
U.N., Z.L., S.W., E.C.Y. and D.B. were responsible for the conceptualization of the study. Z.L., S.W., U.N., W.S. and D.B. presented the methodology. Z.L., S.W., U.N., E.C.Y. and R.D. undertook the investigations. Z.L., S.W., A.J.B. and C.W. performed the CryoEM. Z.L., S.W., U.N., A.K.B., M.J.B., H.N., A.K. and B.S. performed the X-ray crystallography. S.W., B.L., S.S. and G.H. performed the SAXS. M.Y.Y. and D.S.G. performed the hyperspectra and SEM. M.B.R. carried out the simulations. Z.L., U.N., E.C.Y., J.M.L., Y.H., Q.D., M.M., A.F., B.N., N.I.E. and W.Y. prepared the building blocks. Z.L., S.W., U.N., W.S., Y.H., A.C., Q.D. and A.E. provided the design protocols. H.H. carried out the Rosetta score function calculations. Z.L., S.W., U.N., A.J.B., W.Y. and D.B. provided the visualization. D.B. was responsible for funding acquisition. D.B. supervised the study. Z.L., S.W., U.N. and D.B. wrote the original draft of the manuscript. Z.L., S.W., U.N., A.K.B., A.J.B., M.Y.Y., E.C.Y., H.H., M.B.R. and D.B. reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
Z.L., S.W., U.N., E.C.Y., W.S., J.M.L., Y.H., B.S. and D.B. are inventors on a provisional patent application submitted by the University of Washington for the design, composition and function of the proteins created in this study.
Peer review
Peer review information
Nature Materials thanks Mauri Kostiainen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Design rules of 3D protein crystals.
a, Constrained degree of freedom (DOF): The angle of rotation at the designed dihedral crystal interface (Fig. 1a, right panel) must be precisely specified by the design process, where the C2 axis of the dihedral needs to coincide with the C2 axis of the space group. In this example, the disruptive effect (highlighted in red) of a 15-degree error in alignment on crystal assembly is illustrated; similar crystal lattice breakdowns occur with all deviations from the target alignment angle. b, Accessible secondary structure (SS): Dihedral interfaces with helices perpendicular to the symmetry axis (docked from T33-15 cage) are more designable than those with helices parallel to the symmetry axis (docked from T33-21 cage26). Interacting secondary structures are highlighted in red. c, Affinity and Specificity: Working interfaces have sufficient hydrophobic packing with specific polar interactions at the boundary. Highly hydrophobic interfaces destruct the designed self-assembly, including insoluble components and off-target assemblies.
Extended Data Fig. 2 Characterizations of the constituent cages of designed crystals.
a-d, T33-15-D3-4, e–h, T32-15, i-l, O43-2. a,e,i, SEC chromatograms of two oligomeric components (green and orange) and cages assembled via in-vitro mixing of components (blue). b,f,j, nsEM images (scale bars, 50 nm). c,g,k, overlay of the design model with 3D reconstructed nsEM density map/ cryoEM model (scale bars, 5 nm). d,h,l, SAXS profile and simulation results of cages.
Extended Data Fig. 3 Characterizations of new tetrahedral cages for crystal design.
a–e, from left to right, computational model, SEC chromatogram, SAXS profile, and nsEM images.
Extended Data Fig. 4 Characterizations of new octahedral cages for crystal design.
a–d, from left to right, computational model, SEC chromatogram, SAXS profile, and nsEM images.
Extended Data Fig. 5 Symmetric dockings of tetrahedral and octahedral cages into crystal lattices.
a, Two tetrahedral cages are docked along their C3 axis for crystal contacts of D3 dihedrals, which allow them to crystallize in the F4132 space group. b, Two octahedral cages are docked along their C3 axis for crystal contacts of D3 dihedrals, which allow them to crystallize in the I432 space group. See methods for detailed docking protocol.
Extended Data Fig. 6 Optical microscopy and cryoEM characterization of designed protein crystals.
a, Optical micrograph of F4132-1-0 crystals. b, Optical micrograph of F4132-1 crystals. c, CryoEM image of F4132-1 crystals. d, Optical micrograph of F4132-2-6H crystals. e, Optical micrograph of F4132-2 crystals. f, CryoEM image of F4132-2 crystals. g, Optical micrograph of I432-1 crystals. h, Optical micrograph of I432-1-CC crystals. i, CryoEM image of I432-1 crystals.
Extended Data Fig. 7 CryoEM data of the T32-15 cage.
a, Representative 2D class averages of the T32-15 cage. b, CryoEM local resolution map of the T32-15 cage (top) and built atomic model (bottom). Local resolution estimates range from ~2.5 Å at the core to ~4 Å along the crystal-contact forming helices. c, Map-to-model comparison within a low-resolution region (top) and a high-resolution region (bottom). d, Global FSC. e, Orientational distribution plot demonstrating full angular sampling.
Extended Data Fig. 8 Tuning the crystallization behavior of designed crystals by mutagenesis.
a, Mutations to the F4132-1 crystals. b, Mutations of F4132-2 crystals. c, Mutations and redesigns (orange) of I432-1 crystals. Top panels, crystal interface models based on X-ray structure. Interface side chains are hypothetically placed to demonstrate mutation sites. Bottom panels: optical micrographs of representative crystallization results. Scale bars, 100 µm.
Extended Data Fig. 9 Design pipeline for engineering crystal unit cell dimension.
The crystal contact of the F4132-2 crystal was redesigned with different DHR arm fusion. See Methods for the details of step a-g.
Supplementary information
Supplementary Information
Supplementary Figs. 1–33 and Tables 1–7.
Supplementary Data 1
PDB models of cages and dihedrals.
Supplementary Data 2
Example scripts for computational dockings and designs.
Source data
Source Data for Fig. 1
SAXS data and simulation of three designed crystals.
Source Data for Fig. 2
Analysis of PDB entry resolution, space groups and solvent content distribution.
Source Data for Fig. 3
SAXS data of crystals of engineered lattice size.
Source Data for Fig. 4
SAXS data and simulation of crystals with AuNPs.
Source Data for Fig. 5
Hyperspectral data of different AuNP superlattices.
Source Data for Fig. 6
Optical microscope measurement results of crystal sizes.
Source Data for Fig. 7
SAXS data of all designed protein cages reported in the extended data figures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Z., Wang, S., Nattermann, U. et al. Accurate computational design of three-dimensional protein crystals. Nat. Mater. 22, 1556–1563 (2023). https://doi.org/10.1038/s41563-023-01683-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-023-01683-1
This article is cited by
-
Blueprinting extendable nanomaterials with standardized protein blocks
Nature (2024)
-
Genetically encoded protein crystals by hierarchical design
Nature Materials (2023)