Abstract
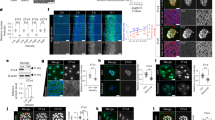
Integrin-mediated cell–matrix adhesions are key to sensing the geometry and rigidity of extracellular environments and influence vital cellular processes. In vivo, the extracellular matrix is composed of fibrous arrays. To understand the fibre geometries that are required for adhesion formation, we patterned nanolines of various line widths and arrangements in single, crossing or paired arrays with the integrin-binding peptide Arg-Gly-Asp. Single thin lines (width ≤30 nm) did not support cell spreading or formation of focal adhesions, despite the presence of a high density of Arg-Gly-Asp, but wide lines (>40 nm) did. Using super-resolution microscopy, we observed stable, dense integrin clusters formed on parallel (within 110 nm) or crossing thin lines (mimicking a matrix mesh) similar to those on continuous substrates. These dense clusters bridged the line pairs by recruiting activated but unliganded integrins, as verified by integrin mutants unable to bind ligands that coclustered with ligand-bound integrins when present in an active extended conformation. Thus, in a fibrous extracellular matrix mesh, stable integrin nanoclusters bridge between thin (≤30 nm) matrix fibres and bring about downstream consequences of cell motility and growth.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the article (and its Supplementary Information files), and from the corresponding author upon reasonable request.
References
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Geiger, B., Spatz, J.P. & Bershadsky, A.D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell. Biol. 10, 21–33 (2009).
Humphries, J. D., Byron, A. & Humphries, M. J. Integrin ligands at a glance. J. Cell Sci. 119, 3901–3903 (2006).
Cukierman, E., Pankov, R., Stevens, D. R. & Yamada, K. M. Taking cell-matrix adhesions to the third dimension. Science 294, 1708–1712 (2001).
Doyle, A. D. & Yamada, K. M. Mechanosensing via cell-matrix adhesions in 3D microenvironments. Exp. Cell Res. 343, 60–66 (2016).
Cui, H., Webber, M. J. & Stupp, S. I. Self-assembly of peptide amphiphiles: from molecules to nanostructures to biomaterials. Biopolymers 94, 1–18 (2010).
Hartgerink, J. D., Beniash, E. & Stupp, S. I. Self-assembly and mineralization of peptide-amphiphile nanofibers. Science 294, 1684–1688 (2001).
Cavalcanti-Adam, E. A. et al. Cell spreading and focal adhesion dynamics are regulated by spacing of integrin ligands. Biophys. J. 92, 2964–2974 (2007).
Schvartzman, M. et al. Nanolithographic control of the spatial organization of cellular adhesion receptors at the single-molecule level. Nano Lett. 11, 1306–1312 (2011).
Lehnert, D. et al. Cell behaviour on micropatterned substrata: limits of extracellular matrix geometry for spreading and adhesion. J. Cell Sci. 117, 41–52 (2004).
Hirschfeld-Warneken, V. C. et al. Cell adhesion and polarisation on molecularly defined spacing gradient surfaces of cyclic RGDfK peptide patches. Eur. J. Cell Biol. 87, 743–750 (2008).
Oria, R. et al. Force loading explains spatial sensing of ligands by cells. Nature 552, 219–224 (2017).
Hu, S. et al. Structured illumination microscopy reveals focal adhesions are composed of linear subunits. Cytoskeleton 72, 235–245 (2015).
Shroff, H. et al. Dual-color superresolution imaging of genetically expressed probes within individual adhesion complexes. Proc. Natl Acad. Sci. USA 104, 20308–20313 (2007).
Bray, D., Levin, M. D. & Morton-Firth, C. J. Receptor clustering as a cellular mechanism to control sensitivity. Nature 393, 85–88 (1998).
Pageon, S. V. et al. Functional role of T-cell receptor nanoclusters in signal initiation and antigen discrimination. Proc. Natl Acad. Sci. USA 113, E5454–E5463 (2016).
Manz, B. N., Jackson, B. L., Petit, R. S., Dustin, M. L. & Groves, J. T-cell triggering thresholds are modulated by the number of antigen within individual T-cell receptor clusters. Proc. Natl Acad. Sci. USA 108, 9089–9094 (2011).
Chang, A. C. et al. Single molecule force measurements in living cells reveal a minimally tensioned integrin state. ACS Nano 10, 10745–10752 (2016).
Choi, C. K. et al. Actin and alpha-actinin orchestrate the assembly and maturation of nascent adhesions in a myosin II motor-independent manner. Nat. Cell Biol. 10, 1039–1050 (2008).
Alexandrova, A. Y. et al. Comparative dynamics of retrograde actin flow and focal adhesions: formation of nascent adhesions triggers transition from fast to slow flow. PLoS ONE 3, e3234 (2008).
Changede, R. & Sheetz, M. Integrin and cadherin clusters: a robust way to organize adhesions for cell mechanics. Bioessays 39, 1–12 (2017).
Changede, R., Xu, X., Margadant, F. & Sheetz, M. P. Nascent integrin adhesions form on all matrix rigidities after integrin activation. Dev. Cell 35, 614–621 (2015).
Bachir, A. I. et al. Integrin-associated complexes form hierarchically with variable stoichiometry in nascent adhesions. Curr. Biol. 24, 1845–1853 (2014).
Wolfenson, H. et al. Tropomyosin controls sarcomere-like contractions for rigidity sensing and suppressing growth on soft matrices. Nat. Cell Biol. 18, 33–42 (2016).
Saxena, M., Changede, R., Hone, J., Wolfenson, H. & Sheetz, M. P. Force-induced calpain cleavage of talin is critical for growth, adhesion development, and rigidity sensing. Nano Lett. 17, 7242–7251 (2017).
Saxena, M. et al. EGFR and HER2 activate rigidity sensing only on rigid matrices. Nat. Mater. 16, 775–781 (2017).
Geiger, B., Salomon, D., Takeichi, M. & Hynes, R. O. A chimeric N-cadherin/beta 1-integrin receptor which localizes to both cell-cell and cell-matrix adhesions. J. Cell Sci. 103, 943–951 (1992). Pt 4.
Smilenov, L., Briesewitz, R. & Marcantonio, E. E. Integrin beta-1 cytoplasmic domain dominant negative effects revealed by lysophosphatidic acid treatment. Mol. Biol. Cell 5, 1215–1223 (1994).
Smilenov, L. B., Mikhailov, A., Pelham, R. J., Marcantonio, E. E. & Gundersen, G. G. Focal adhesion motility revealed in stationary fibroblasts. Science 286, 1172–1174 (1999).
Roca-Cusachs, P., Gauthier, N. C., Del Rio, A. & Sheetz, M. P. Clustering of alpha(5)beta(1) integrins determines adhesion strength whereas alpha(v)beta(3) and talin enable mechanotransduction. Proc. Natl Acad. Sci. USA 106, 16245–16250 (2009).
Luo, B. H., Springer, T. A. & Takagi, J. Stabilizing the open conformation of the integrin headpiece with a glycan wedge increases affinity for ligand. Proc. Natl Acad. Sci. USA 100, 2403–2408 (2003).
Xiong, J. P. et al. Crystal structure of the extracellular segment of integrin alpha Vbeta3. Science 294, 339–345 (2001).
Xiong, J. P. et al. Crystal structure of the extracellular segment of integrin alpha Vbeta3 in complex with an Arg-Gly-Asp ligand. Science 296, 151–155 (2002).
Humphries, M. J. Integrin structure. Biochem. Soc. Trans. 28, 311–339 (2000).
Cai, H. et al. Molecular occupancy of nanodot arrays. ACS Nano 10, 4173–4183 (2016).
Jurchenko, C., Chang, Y., Narui, Y., Zhang, Y. & Salaita, K. S. Integrin-generated forces lead to streptavidin-biotin unbinding in cellular adhesions. Biophys. J. 106, 1436–1446 (2014).
Cai, H. et al. Spatial control of biological ligands on surfaces applied to T cell activation. Methods Mol. Biol. 1584, 307–331 (2017).
Cai, H. et al. Full control of ligand positioning reveals spatial thresholds for T cell receptor triggering. Nat. Nanotechnol. 13, 610–617 (2018).
Cluzel, C. et al. The mechanisms and dynamics of (alpha)v(beta)3 integrin clustering in living cells. J. Cell Biol. 171, 383–392 (2005).
Lippincott-Schwartz, J. & Patterson, G. H. Photoactivatable fluorescent proteins for diffraction-limited and super-resolution imaging. Trends Cell Biol. 19, 555–565 (2009).
Rossier, O. et al. Integrins beta1 and beta3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat. Cell Biol. 14, 1057–1067 (2012).
Campbell, I. D. & Humphries, M. J. Integrin structure, activation, and interactions. Cold Spring Harb. Perspect. Biol. 3, 3 (2011).
Yu, C. H., Law, J. B., Suryana, M., Low, H. Y. & Sheetz, M. P. Early integrin binding to Arg-Gly-Asp peptide activates actin polymerization and contractile movement that stimulates outward translocation. Proc. Natl Acad. Sci. USA 108, 20585–20590 (2011).
Schreiner, C. L. et al. Isolation and characterization of Chinese hamster ovary cell variants deficient in the expression of fibronectin receptor. J. Cell Biol. 109, 3157–3167 (1989).
Liu, Y. et al. Nanoparticle tension probes patterned at the nanoscale: impact of integrin clustering on force transmission. Nano Lett. 14, 5539–5546 (2014).
Legate, K. R. & Fassler, R. Mechanisms that regulate adaptor binding to beta-integrin cytoplasmic tails. J. Cell Sci. 122(2), 187–198 (2009).
Kuo, J. C., Han, X., Hsiao, C. T., Yates, J. R. 3rd & Waterman, C. M. Analysis of the myosin-II-responsive focal adhesion proteome reveals a role for beta-Pix in negative regulation of focal adhesion maturation. Nat. Cell Biol. 13, 383–393 (2011).
Schiller, H. B., Friedel, C. C., Boulegue, C. & Fassler, R. Quantitative proteomics of the integrin adhesome show a myosin II-dependent recruitment of LIM domain proteins. EMBO Rep. 12, 259–266 (2011).
Geiger, T. & Zaidel-Bar, R. Opening the floodgates: proteomics and the integrin adhesome. Curr. Opin. Cell Biol. 24, 562–568 (2012).
Hu, X. et al. Cooperative vinculin binding to talin mapped by time-resolved super resolution microscopy. Nano Lett. 16, 4062–4068 (2016).
Golji, J. & Mofrad, M. R. K. The talin dimer structure orientation is mechanically regulated. Biophys. J. 107, 1802–1809 (2014).
Deeg, J. A. et al. Impact of local versus global ligand density on cellular adhesion. Nano Lett. 11, 1469–1476 (2011).
Loftus, J. C. et al. A beta 3 integrin mutation abolishes ligand binding and alters divalent cation-dependent conformation. Science 249, 915–918 (1990).
Zhang, X. et al. Talin depletion reveals independence of initial cell spreading from integrin activation and traction. Nat. Cell Biol. 10, 1062 (2008).
Wu, H. P., Cheng, T. L. & Tseng, W. L. Phosphate-modified TiO2 nanoparticles for selective detection of dopamine, levodopa, adrenaline, and catechol based on fluorescence quenching. Langmuir 23, 7880–7885 (2007).
Holzmeister, P. et al. Quantum yield and excitation rate of single molecules close to metallic nanostructures. Nat. Commun. 5, 5356 (2014).
Breshike, C. J., Riskowski, R. A. & Strouse, G. F. Leaving Forster resonance energy transfer behind: nanometal surface energy transfer predicts the size-enhanced energy coupling between a metal nanoparticle and an emitting dipole. J. Phys. Chem. C 117, 23942–23949 (2013).
Dulkeith, E. et al. Fluorescence quenching of dye molecules near gold nanoparticles: radiative and nonradiative effects. Phys. Rev. Lett. 89, 203002 (2002).
Cai, H. et al. Molecular occupancy of nanodot arrays. ACS Nano 10, 4173–4183 (2016).
Rajh, T. et al. Surface restructuring of nanoparticles: an efficient route for ligand-metal oxide crosstalk. J. Phys. Chem. B 106, 10543–10552 (2002).
Acknowledgements
We thank G. Giannone, Neurosciences Bordeaux, France and the Michael W. Davidson group, The Florida State University, Tallahassee, FL, USA for DNA constructs. We thank H. Wolfenson for his help with initial experiments, P. Kathirvel for cloning the double-mutant β3 construct and M. Lee for help with illustrations. This work was supported by intramural funds from the Mechanobiology Institute. R.C. is supported by Singapore National Research Foundation’s CRP grant (No. NRF2012NRF-CRP001-084), and M.P.S. received National Institutes of Health (NIH) grant support related to this project (no. RO1-GM113022). S.J.W. and H.C. were supported by the National Science Foundation under award no. CMMI-1300590 and NIH Common Fund Nanomedicine program grant no. PN2 EY016586. The Columbia Nano Initiative provided cleanroom and processing facilities. This work was performed in part at the Center for Nanoscale Materials, a US Department of Energy Office of Science User Facility, and was supported by the US Department of Energy, Office of Science under contract no. DE-AC02-06CH11357.
Author information
Authors and Affiliations
Contributions
R.C. and M.P.S conceived and designed the experiments. H.C. and S.J.W. designed and prepared the AuPd and Ti nanopatterned substrates. R.C. and H.C. standardized the functionalization of AuPd substrates with RGD. R.C. standardized the functionalization of Ti substrates and performed the experiments, analysed the data and wrote the manuscript. R.C., M.P.S., S.J.W. and H.C., prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9, Supplementary Video Legends 1–3 and Supplementary Table 1
Supplementary Video 1
MEFs do not spread well on single-line 1D geometry
Supplementary Video 2
MEFs spread well and form large adhesions on line pair 2D geometry
Supplementary Video 3
Dynamics of b3GFP on single lines and line pairs
Rights and permissions
About this article
Cite this article
Changede, R., Cai, H., Wind, S.J. et al. Integrin nanoclusters can bridge thin matrix fibres to form cell–matrix adhesions. Nat. Mater. 18, 1366–1375 (2019). https://doi.org/10.1038/s41563-019-0460-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-019-0460-y
This article is cited by
-
Organization, dynamics and mechanoregulation of integrin-mediated cell–ECM adhesions
Nature Reviews Molecular Cell Biology (2023)
-
The glycocalyx affects the mechanotransductive perception of the topographical microenvironment
Journal of Nanobiotechnology (2022)
-
Control cell migration by engineering integrin ligand assembly
Nature Communications (2022)
-
Ligand functionalization of titanium nanopattern enables the analysis of cell–ligand interactions by super-resolution microscopy
Nature Protocols (2022)
-
Current international research into cellulose as a functional nanomaterial for advanced applications
Journal of Materials Science (2022)