Abstract
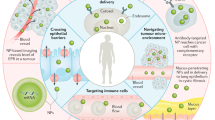
Development of targeted nanoparticle drug carriers often requires complex synthetic schemes involving both supramolecular self-assembly and chemical modification. These processes are generally difficult to predict, execute, and control. We describe herein a targeted drug delivery system that is accurately and quantitatively predicted to self-assemble into nanoparticles based on the molecular structures of precursor molecules, which are the drugs themselves. The drugs assemble with the aid of sulfated indocyanines into particles with ultrahigh drug loadings of up to 90%. We devised quantitative structure-nanoparticle assembly prediction (QSNAP) models to identify and validate electrotopological molecular descriptors as highly predictive indicators of nano-assembly and nanoparticle size. The resulting nanoparticles selectively targeted kinase inhibitors to caveolin-1-expressing human colon cancer and autochthonous liver cancer models to yield striking therapeutic effects while avoiding pERK inhibition in healthy skin. This finding enables the computational design of nanomedicines based on quantitative models for drug payload selection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Peer, D. et al. Nanocarriers as an emerging platform for cancer therapy. Nat. Nanotech. 2, 751–760 (2007).
Schroeder, A. et al. Treating metastatic cancer with nanotechnology. Nat. Rev. Cancer 12, 39–50 (2012).
Yaari, Z. et al. Theranostic barcoded nanoparticles for personalized cancer medicine. Nat. Commun. 7, 13325 (2016).
Wilhelm, S. et al. Analysis of nanoparticle delivery to tumours. Nat. Rev. Mater. 1, 16014 (2016).
Cheng, Z., Al Zaki, A., Hui, J. Z., Muzykantov, V. R. & Tsourkas, A. Multifunctional nanoparticles: Cost versus benefit of adding targeting and imaging capabilities. Science 338, 903–910 (2012).
Lammers, T., Kiessling, F., Hennink, W. E. & Storm, G. Drug targeting to tumors: Principles, pitfalls and (pre-) clinical progress. J. Control. Release 161, 175–187 (2012).
Shamay, Y. et al. P-Selectin is a nanotherapeutic delivery target in the tumor microenvironment. Sci. Transl. Med. 8, 345ra87 (2016).
Mizrachi, A. et al. Tumour-specific Pi3k inhibition via nanoparticle-targeted delivery in head and neck squamous cell carcinoma. Nat. Commun. 8, 14292 (2017).
Maojo, V. et al. Nanoinformatics: A new area of research in nanomedicine. Int. J. Nanomed. 7, 3867–3890 (2012).
Irwin, J. J. et al. An aggregation advisor for ligand discovery. J. Med. Chem. 58, 7076–7087 (2015).
Seidler, J., McGovern, S. L., Doman, T. N. & Shoichet, B. K. Identification and prediction of promiscuous aggregating inhibitors among known drugs. J. Med. Chem. 46, 4477–4486 (2003).
Feng, B. Y., Shelat, A., Doman, T. N., Guy, R. K. & Shoichet, B. K. High-throughput assays for promiscuous inhibitors. Nat. Chem. Biol. 1, 146–148 (2005).
Alskar, L. C., Porter, C. J. & Bergstrom, C. A. Tools for early prediction of drug loading in lipid-based formulations. Mol. Pharm. 13, 251–261 (2016).
Fourches, D. et al. Quantitative nanostructure-activity relationship modeling. ACS Nano 4, 5703–5712 (2010).
Puzyn, T. et al. Using nano-QSAR to predict the cytotoxicity of metal oxide nanoparticles. Nat. Nanotech. 6, 175–178 (2011).
Zhang, Y. et al. Lipid-modified aminoglycoside derivatives for in vivo siRNA delivery. Adv. Mater. 25, 4641–4645 (2013).
Roxbury, D., Jagota, A. & Mittal, J. Sequence-specific self-stitching motif of short single-stranded DNA on a single-walled carbon nanotube. J. Am. Chem. Soc. 133, 13545–13550 (2011).
Lee, O. S., Stupp, S. I. & Schatz, G. C. Atomistic molecular dynamics simulations of peptide amphiphile self-assembly into cylindrical nanofibers. J. Am. Chem. Soc. 133, 3677–3683 (2011).
Frederix, P. W. et al. Exploring the sequence space for (tri-)peptide self-assembly to design and discover new hydrogels. Nat. Chem. 7, 30–37 (2015).
Shi, C. et al. Drug-specific nanocarrier design for efficient anticancer therapy. Nat. Commun. 6, 7449 (2015).
Alphandery, E., Grand-Dewyse, P., Lefevre, R., Mandawala, C. & Durand-Dubief, M. Cancer therapy using nanoformulated substances: Scientific, regulatory and financial Aspects. Expert. Rev. Anticancer. Ther. 15, 1233–1255 (2015).
Agarwal, A., Lvov, Y., Sawant, R. & Torchilin, V. Stable nanocolloids of poorly soluble drugs with high drug content prepared using the combination of sonication and layer-by-layer lechnology. J. Control. Release 128, 255–260 (2008).
Muller, R. H. & Keck, C. M. Challenges and solutions for the delivery of biotech drugs—a review of drug nanocrystal technology and lipid nanoparticles. J. Biotech. 113, 151–170 (2004).
McLaughlin, C. K. et al. Stable colloidal drug aggregates catch and release active enzymes. ACS Chem. Biol. 11, 992–1000 (2016).
Shi, C., Wu, J. B. & Pan, D. Review on near-infrared heptamethine cyanine dyes as theranostic agents for tumor imaging, targeting, and photodynamic therapy. J. Biomed. Opt. 21, 50901 (2016).
Yang, X. et al. Near Ir heptamethine cyanine dye-mediated cancer imaging. Clin. Cancer Res. 16, 2833–2844 (2010).
McArthur, E. A., Godbe, J. M., Tice, D. B. & Weiss, E. A. A study of the binding of cyanine dyes to colloidal quantum dots using spectral signatures of dye aggregation. J. Phys. Chem. 116, 6136–6142 (2012).
Slavnova, T. D., Gorner, H. & Chibisov, A. K. Cyanine-based J-aggregates as a chirality-sensing supramolecular system. J. Phys. Chem. B 115, 3379–3384 (2011).
Fofang, N. T., Grady, N. K., Fan, Z., Govorov, A. O. & Halas, N. J. Plexciton dynamics: exciton–plasmon coupling in a J-aggregate–Au nanoshell complex provides a mechanism for nonlinearity. Nano Lett. 11, 1556–1560 (2011).
Todeschini, R. & Consonni, V. Handbook of Molecular Descriptors Vol. 11 (eds Mannhold, R., Kubinyi, H. & Timmerman, H.) 285–286 (Wiley, New York, NY, 2008).
Kier, L. B. & Hall, L. H. An electrotopological-state index for atoms in molecules. Pharm. Res. 7, 801–807 (1990).
Consonni, V., Todeschini, R. & Pavan, M. Structure/response correlations and similarity/diversity analysis by Getaway descriptors. 1. Theory of the novel 3d molecular descriptors. J. Chem., Inf. Comput. Sci. 42, 682–692 (2002).
Wishart, D. S. et al. Drugbank: A comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 34, D668–672 (2006).
Rhee, Y. M. & Pande, V. S. Multiplexed-replica exchange molecular dynamics method for protein folding simulation. Biophys. J. 84, 775–786 (2003).
Zhou, R. in Protein Folding Protocols (eds Bai, Y. & Nussinov, R.) 205–223 (Springer, New York, NY, 2007).
Voigt, J., Christensen, J. & Shastri, V. P. Differential uptake of nanoparticles by endothelial cells through polyelectrolytes with affinity for caveolae. Proc. Natl Acad. Sci. USA 111, 2942–2947 (2014).
Jena, P. V. et al. Photoluminescent carbon nanotubes interrogate the permeability of multicellular tumor spheroids. Carbon 97, 99–109 (2016).
Wang, Z., Tiruppathi, C., Cho, J., Minshall, R. D. & Malik, A. B. Delivery of nanoparticle: Complexed drugs across the vascular endothelial barrier via caveolae. IUBMB Life 63, 659–667 (2011).
Chrastina, A., Massey, K. A. & Schnitzer, J. E. Overcoming in vivo barriers to targeted nanodelivery. WIREs. Nanomed. Nanobiotechnol. 3, 421–437 (2011).
Chen, X. & Calvisi, D. F. Hydrodynamic transfection for generation of novel mouse models for liver cancer research. Am. J. Path. 184, 912–923 (2014).
O’Donnell, K. A. et al. A Sleeping Beauty mutagenesis screen reveals a tumor suppressor role for Ncoa2/Src-2 in liver cancer. Proc. Natl Acad. Sci. USA 109, E1377–1386 (2012).
Nicholls, A. Confidence limits, error bars and method comparison in molecular modeling. Part 1: the calculation of confidence intervals. J. Comput. Aided Mol. Des. 28, 887–918 (2014).
Schomburg, K. T., Wetzer, L. & Rarey, M. Interactive design of generic chemical patterns. Drug Discovery Today 18, 651–658 (2013).
Acknowledgements
This work was supported in part by the NIH New Innovator Award (DP2-HD075698), National Cancer Institute (CA 013106), and Cancer Center Support Grant (P30 CA008748), the Expect Miracles Foundation - Financial Services Against Cancer, the Anna Fuller Fund, the Louis V. Gerstner Jr. Young Investigator’s Fund, the Frank A. Howard Scholars Program, Cycle for Survival, the Alan and Sandra Gerry Metastasis Research Initiative, Mr. William H. Goodwin and Mrs. Alice Goodwin and the Commonwealth Foundation for Cancer Research, the Experimental Therapeutics Center, the Imaging & Radiation Sciences Program, and the Center for Molecular Imaging and Nanotechnology of Memorial Sloan Kettering Cancer. This work is supported in part by a New York State Department of Health Fixed Term Agreement (Contract# DOH01-C30315GG-3450000). The opinions, results, findings and/or interpretations of data contained therein are the responsibility of the contractor and do not necessarily represent the opinions, interpretations or policy of the state or, if funded with federal funds, the applicable federal funding agency. Y.S. was supported by the Center for Metastasis Research (CMR) Scholars Fellowship Program. D.R. was supported by an American Cancer Society – Roaring Fork Valley Postdoctoral Fellowship. S.W.L. is a Howard Hughes Medical Institute Investigator and the Geoffrey Beene Chair for Cancer Biology (MSKCC). J.B. was supported by the Tow Foundation Postdoctoral Fellowship, Center for Molecular Imaging and Nanotechnology at MSKCC. We would like to thank the following facilities at MSKCC: Molecular Cytology Core Facility, Small Animal Imaging, Anti-tumor Assessment, and Electron Microscopy. The molecular dynamics work used the Extreme Science and Engineering Discovery Environment (XSEDE), supported by National Science Foundation grant number TG-MCB-130013. We would like to thank N. Lampen for assistance with electron microscopy. We would also like to thank P.V. Jena, R. Williams, J. Harvey, T. Galassi, J. Humm and C. Horoszko for helpful discussions.
Author information
Authors and Affiliations
Contributions
Y.S. and D.A.H. conceived the project and designed experiments. Y.S. analysed data, designed, and conducted the self-assembly experiments. D.R. performed MD experiments and analysis. D.F.T. and J.L. performed the in vivo liver cancer model. Y.S., V.K.R., A.M. and J.S. performed all other in vivo experiments, electron microscopy and tissue staining. Y.S., V.K.R., K.N., J.L.S., M.R.N., K.C. and J.S. performed in vitro experiments. Y.S. developed the tumour spheroid model. E.B., J.L.S., K.N., R.S., M.R.N., K.C., K.S.G., M.D. and D.C.J. performed experiments for the computational drug screening and nanoparticle self-assembly validation experiments. J.B. performed F-NMR studies for drug biodistribution. M.I. performed self-assembly categorization of DrugBank small molecule drugs. M.I. and J.D.C. conducted statistical analysis of QSNAP descriptors. Y.S. and D.A.H. wrote the paper. D.A.H., S.W.L. and J.D.C. supervised the research.
Corresponding author
Ethics declarations
Competing interests
J. D. C. is a member of the Scientific Advisory Board for Schrödinger, LLC.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Discussion, Supplementary Figures 1–24, Supplementary Tables 1–6
Rights and permissions
About this article
Cite this article
Shamay, Y., Shah, J., Işık, M. et al. Quantitative self-assembly prediction yields targeted nanomedicines. Nature Mater 17, 361–368 (2018). https://doi.org/10.1038/s41563-017-0007-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41563-017-0007-z
This article is cited by
-
Application of molecular dynamics simulation in self-assembled cancer nanomedicine
Biomaterials Research (2023)
-
A multifunctional integrated biomimetic spore nanoplatform for successively overcoming oral biological barriers
Journal of Nanobiotechnology (2023)
-
Fragment-based drug nanoaggregation reveals drivers of self-assembly
Nature Communications (2023)
-
Staggered-layer-boosted flexible Bi2Te3 films with high thermoelectric performance
Nature Nanotechnology (2023)
-
Emerging nanobiotechnology for precise theranostics of hepatocellular carcinoma
Journal of Nanobiotechnology (2022)