Abstract
The cerebral ventricles are recognized as windows into brain development and disease, yet their genetic architectures, underlying neural mechanisms and utility in maintaining brain health remain elusive. Here we aggregated genetic and neuroimaging data from 61,974 participants (age range, 9 to 98 years) in five cohorts to elucidate the genetic basis of ventricular morphology and examined their overlap with neuropsychiatric traits. Genome-wide association analysis in a discovery sample of 31,880 individuals identified 62 unique loci and 785 candidate genes associated with ventricular morphology. We replicated over 80% of loci in a well-matched cohort of lateral ventricular volume. Gene set analysis revealed enrichment of ventricular-trait-associated genes in biological processes and disease pathogenesis during both early brain development and degeneration. We explored the age-dependent genetic associations in cohorts of different age groups to investigate the possible roles of ventricular-trait-associated loci in neurodevelopmental and neurodegenerative processes. We describe the genetic overlap between ventricular and neuropsychiatric traits through comprehensive integrative approaches under correlative and causal assumptions. We propose the volume of the inferior lateral ventricles as a heritable endophenotype to predict the risk of Alzheimer’s disease, which might be a consequence of prodromal Alzheimer’s disease. Our study provides an advance in understanding the genetics of the cerebral ventricles and demonstrates the potential utility of ventricular measurements in tracking brain disorders and maintaining brain health across the lifespan.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Our GWAS summary statistics for cerebral ventricles can be found at https://doi.org/10.6084/m9.figshare.21529491. The GWAS results are also available on the FUMA website (https://fuma.ctglab.nl/browse/; ID 619–625). The individual-level data used in the present study were obtained from UKB (https://www.ukbiobank.ac.uk/), ABCD (https://abcdstudy.org/), IMAGEN (http://imagen-project.org/), HCP (http://www.humanconnectome.org/) and ADNI (https://adni.loni.usc.edu/). The summary-level data for the CHARGE consortium were obtained from dbGaP (phs000930.v9.p1). The GWAS Catalog resource can be found at https://www.ebi.ac.uk/gwas/. The GWAS atlas can be found at https://atlas.ctglab.nl/PheWAS/. The DSigDB database can be found at http://dsigdb.tanlab.org/DSigDBv1.0/. Agora can be found at https://agora.adknowledgeportal.org/. The Human Protein Atlas can be found at https://www.proteinatlas.org/.
Code availability
This study used openly available software and code, specifically R (https://www.r-project.org/), PLINK (https://www.cog-genomics.org/plink/), GCTA (http://cnsgenomics.com/software/gcta/), IMPUTE (https://mathgen.stats.ox.ac.uk/impute/impute_v2.html), Michigan Imputation Server (https://imputationserver.sph.umich.edu/), MOSTest (https://github.com/precimed/mostest), METAL (http://csg.sph.umich.edu/abecasis/metal/), FUMA (https://fuma.ctglab.nl/), MAGMA (https://ctg.cncr.nl/software/magma/, also implemented in FUMA), SAIGE-GENE+ (https://saigegit.github.io/SAIGE-doc/), Enrichr (https://maayanlab.cloud/Enrichr/) and LDSC (https://github.com/bulik/ldsc/). Custom scripts for the analyses in this paper are available through GitHub (https://github.com/yjge/cerebral_ventricles).
References
Duy, P. Q. et al. Brain ventricles as windows into brain development and disease. Neuron 110, 12–15 (2022).
de Mélo Silva Júnior, M. L., Diniz, P. R. B., de Souza Vilanova, M. V., Basto, G. P. T. & Valença, M. M. Brain ventricles, CSF and cognition: a narrative review. Psychogeriatrics 22, 544–552 (2022).
Sapkota, S., McFall, G. P., Masellis, M., Dixon, R. A. & Black, S. E. Differential cognitive decline in Alzheimer’s disease is predicted by changes in ventricular size but moderated by apolipoprotein E and pulse pressure. J. Alzheimers Dis. 85, 545–560 (2022).
West, N. A. et al. Neuroimaging findings in midlife and risk of late-life dementia over 20 years of follow-up. Neurology 92, e917–e923 (2019).
Bethlehem, R. A. I. et al. Brain charts for the human lifespan. Nature 604, 525–533 (2022).
Lui, J. H., Hansen, D. V. & Kriegstein, A. R. Development and evolution of the human neocortex. Cell 146, 18–36 (2011).
Chojnacki, A. K., Mak, G. K. & Weiss, S. Identity crisis for adult periventricular neural stem cells: subventricular zone astrocytes, ependymal cells or both? Nat. Rev. Neurosci. 10, 153–163 (2009).
Duy, P. Q. et al. Impaired neurogenesis alters brain biomechanics in a neuroprogenitor-based genetic subtype of congenital hydrocephalus. Nat. Neurosci. 25, 458–473 (2022).
Richards, R. et al. Increased hippocampal shape asymmetry and volumetric ventricular asymmetry in autism spectrum disorder. NeuroImage Clin. 26, 102207 (2020).
Prigge, M. B. D. et al. A 16-year study of longitudinal volumetric brain development in males with autism. NeuroImage 236, 118067 (2021).
McWhinney, S. R. et al. Association between body mass index and subcortical brain volumes in bipolar disorders—ENIGMA study in 2735 individuals. Mol. Psychiatry 26, 6806–6819 (2021).
Schmaal, L. et al. Subcortical brain alterations in major depressive disorder: findings from the ENIGMA Major Depressive Disorder working group. Mol. Psychiatry 21, 806–812 (2016).
Okada, N. et al. Abnormal asymmetries in subcortical brain volume in schizophrenia. Mol. Psychiatry 21, 1460–1466 (2016).
Brugger, S. P. & Howes, O. D. Heterogeneity and homogeneity of regional brain structure in schizophrenia: a meta-analysis. JAMA Psychiatry 74, 1104–1111 (2017).
Vojinovic, D. et al. Genome-wide association study of 23,500 individuals identifies 7 loci associated with brain ventricular volume. Nat. Commun. 9, 3945 (2018).
Smith, S. M. et al. An expanded set of genome-wide association studies of brain imaging phenotypes in UK Biobank. Nat. Neurosci. 24, 737–745 (2021).
Scelsi, C. L. et al. The lateral ventricles: a detailed review of anatomy, development, and anatomic variations. AJNR Am. J. Neuroradiol. 41, 566–572 (2020).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
van der Meer, D. et al. Understanding the genetic determinants of the brain with MOSTest. Nat. Commun. 11, 3512 (2020).
Deming, Y. et al. Genome-wide association study identifies four novel loci associated with Alzheimer’s endophenotypes and disease modifiers. Acta Neuropathol. 133, 839–856 (2017).
Jansen, I. E. et al. Genome-wide meta-analysis for Alzheimer’s disease cerebrospinal fluid biomarkers. Acta Neuropathol. 144, 821–842 (2022).
Sha, Z., Schijven, D., Fisher, S. E. & Francks, C. Genetic architecture of the white matter connectome of the human brain. Sci. Adv. 9, eadd2870 (2023).
Bahrami, S. et al. Distributed genetic architecture across the hippocampal formation implies common neuropathology across brain disorders. Nat. Commun. 13, 3436 (2022).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Yoo, M. et al. DSigDB: drug signatures database for gene set analysis. Bioinformatics 31, 3069–3071 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Chen, Y. et al. Structural basis of ALDH1A2 inhibition by irreversible and reversible small molecule inhibitors. ACS Chem. Biol. 13, 582–590 (2018).
Piergiovanni, G. & Costanzo, V. GEMC1 is a novel TopBP1-interacting protein involved in chromosomal DNA replication. Cell Cycle 9, 3662–3666 (2010).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Burgess, S. et al. Guidelines for performing Mendelian randomization investigations [version 2; peer review: 2 approved]. Wellcome Open Research https://doi.org/10.12688/wellcomeopenres.15555.2 (2020).
Gorelick, P. B. et al. Defining optimal brain health in adults: a presidential advisory from the American Heart Association/American Stroke Association. Stroke 48, e284–e303 (2017).
Greenwood, A. K. et al. The AD Knowledge Portal: a repository for multi-omic data on Alzheimer’s disease and aging. Curr. Protoc. Hum. Genet. 108, e105 (2020).
Uhlén, M. et al. Proteomics: tissue-based map of the human proteome. Science 347, 1260419 (2015).
van der Meer, D. et al. The genetic architecture of human cortical folding. Sci. Adv. 7, eabj9446 (2021).
Makowski, C. et al. Discovery of genomic loci of the human cerebral cortex using genetically informed brain atlases. Science 375, 522–528 (2022).
Satizabal, C. L. et al. Genetic architecture of subcortical brain structures in 38,851 individuals. Nat. Genet. 51, 1624–1636 (2019).
Fame, R. M. & Lehtinen, M. K. Emergence and developmental roles of the cerebrospinal fluid system. Dev. Cell 52, 261–275 (2020).
Sha, Z. et al. The genetic architecture of structural left–right asymmetry of the human brain. Nat. Hum. Behav. 5, 1226–1239 (2021).
Wainschtein, P. et al. Assessing the contribution of rare variants to complex trait heritability from whole-genome sequence data. Nat. Genet. 54, 263–273 (2022).
Girirajan, S. Missing heritability and where to find it. Genome Biol. 18, 89 (2017).
Brouwer, R. M. et al. Genetic variants associated with longitudinal changes in brain structure across the lifespan. Nat. Neurosci. 25, 421–432 (2022).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Hansson, O. et al. The genetic regulation of protein expression in cerebrospinal fluid. EMBO Mol. Med. 15, e16359 (2023).
Zhang, X. et al. Bridging Integrator 1 (BIN1) genotype effects on working memory, hippocampal volume, and functional connectivity in young healthy individuals. Neuropsychopharmacology 40, 1794–1803 (2015).
Genon, S., Eickhoff, S. B. & Kharabian, S. Linking interindividual variability in brain structure to behaviour. Nat. Rev. Neurosci. 23, 307–318 (2022).
Jack, C. R. Jr. et al. NIA-AA Research Framework: toward a biological definition of Alzheimer’s disease. Alzheimers Dement. 14, 535–562 (2018).
Macdonald, K. E., Bartlett, J. W., Leung, K. K., Ourselin, S. & Barnes, J. The value of hippocampal and temporal horn volumes and rates of change in predicting future conversion to AD. Alzheimer Dis. Assoc. Disord. 27, 168–173 (2013).
Coupé, P. et al. Hippocampal-amygdalo-ventricular atrophy score: Alzheimer disease detection using normative and pathological lifespan models. Hum. Brain Mapp. 43, 3270–3282 (2022).
Lee Gregory, M., Burton, V. J. & Shapiro, B. K. in Neurobiology of Brain Disorders (eds Zigmond, M. J. et al.) 18–41 (Academic Press, 2015).
Coleman, J. Young brain fluid improves memory in old mice. Nature https://doi.org/10.1038/d41586-022-01282-1 (2022).
Sasabayashi, D. et al. Subcortical brain volume abnormalities in individuals with an at-risk mental state. Schizophr. Bull. 46, 834–845 (2020).
Lewis, M. M. et al. Asymmetrical lateral ventricular enlargement in Parkinson’s disease. Eur. J. Neurol. 16, 475–481 (2009).
Kuo, F. & Massoud, T. F. Structural asymmetries in normal brain anatomy: a brief overview. Ann. Anat. 241, 151894 (2022).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Zhou, W. et al. Efficiently controlling for case–control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet. 53, 1097–1103 (2021).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Psaty, B. M. et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circ. Cardiovasc. Genet. 2, 73–80 (2009).
Casey, B. J. et al. The Adolescent Brain Cognitive Development (ABCD) study: imaging acquisition across 21 sites. Dev. Cogn. Neurosci. 32, 43–54 (2018).
Lee, P. H. et al. Genetic association of attention-deficit/hyperactivity disorder and major depression with suicidal ideation and attempts in children: the Adolescent Brain Cognitive Development Study. Biol. Psychiatry 92, 236–245 (2022).
Schumann, G. et al. The IMAGEN study: reinforcement-related behaviour in normal brain function and psychopathology. Mol. Psychiatry 15, 1128–1139 (2010).
Van Essen, D. C. et al. The Human Connectome Project: a data acquisition perspective. NeuroImage 62, 2222–2231 (2012).
Hendrix, J. A. et al. The Worldwide Alzheimer’s Disease Neuroimaging Initiative: an update. Alzheimers Dement. 11, 850–859 (2015).
Alfaro-Almagro, F. et al. Image processing and quality control for the first 10,000 brain imaging datasets from UK Biobank. NeuroImage 166, 400–424 (2018).
Miller, K. L. et al. Multimodal population brain imaging in the UK Biobank prospective epidemiological study. Nat. Neurosci. 19, 1523–1536 (2016).
Kong, X. Z. et al. Mapping cortical brain asymmetry in 17,141 healthy individuals worldwide via the ENIGMA Consortium. Proc. Natl Acad. Sci. USA 115, E5154–e5163 (2018).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
O’Connell, J. et al. Haplotype estimation for biobank-scale data sets. Nat. Genet. 48, 817–820 (2016).
Chang et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, s13742-015-0047-8 (2015).
Yang, J., Zeng, J., Goddard, M. E., Wray, N. R. & Visscher, P. M. Concepts, estimation and interpretation of SNP-based heritability. Nat. Genet. 49, 1304–1310 (2017).
Yang, J., Hong Lee, S., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Weiner, D. J. et al. Polygenic architecture of rare coding variation across 394,783 exomes. Nature 614, 492–499 (2023).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2020).
Weiner, D. J. et al. Polygenic architecture of rare coding variation across 394783 exomes. Nature 614, 492–499 (2023).
Holst, K. K., Scheike, T. H. & Hjelmborg, J. B. The liability threshold model for censored twin data. Comput. Stat. Data Anal. 93, 324–335 (2016).
Zhou, W. et al. SAIGE-GENE+ improves the efficiency and accuracy of set-based rare variant association tests. Nat. Genet. 54, 1466–1469 (2022).
Aschard, H., Vilhjálmsson, B. J., Joshi, A. D., Price, A. L. & Kraft, P. Adjusting for heritable covariates can bias effect estimates in genome-wide association studies. Am. J. Hum. Genet 96, 329–339 (2015).
Fürtjes, A. E. et al. General dimensions of human brain morphometry inferred from genome-wide association data. Hum. Brain Mapp. 44, 3311–3323 (2023).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Ng, B. et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat. Neurosci. 20, 1418–1426 (2017).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Ramasamy, A. et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat. Neurosci. 17, 1418–1428 (2014).
GTEx Consortium The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Võsa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
Lee, S., Wu, M. C. & Lin, X. Optimal tests for rare variant effects in sequencing association studies. Biostatistics 13, 762–775 (2012).
Viechtbauer, W. Conducting meta-analyses in R with the metafor package. J. Stat. Softw. 36, 1–48 (2010).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524–537 (2018).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Stahl, E. A. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat. Genet. 51, 793–803 (2019).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Davies, G. et al. Study of 300,486 individuals identifies 148 independent genetic loci influencing general cognitive function. Nat. Commun. 9, 2098 (2018).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Karlsson Linnér, R. et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat. Genet. 51, 245–257 (2019).
de Kovel, C. G. F. & Francks, C. The molecular genetics of hand preference revisited. Sci. Rep. 9, 5986 (2019).
Grasby, K. L. et al. The genetic architecture of the human cerebral cortex. Science 367, eaay6690 (2020).
Traylor, M. et al. Genetic variation in PLEKHG1 is associated with white matter hyperintensities (n = 11,226). Neurology 92, e749–e757 (2019).
Hibar, D. P. et al. Common genetic variants influence human subcortical brain structures. Nature 520, 224–229 (2015).
Burgess, S., Thompson, S. G. & CRP CHD Genetics Collaboration Avoiding bias from weak instruments in Mendelian randomization studies. Int J. Epidemiol. 40, 755–764 (2011).
Andrews, S. J., Fulton-Howard, B., O’Reilly, P., Marcora, E. & Goate, A. M. Causal associations between modifiable risk factors and the Alzheimer’s phenome. Ann. Neurol. 89, 54–65 (2021).
Mavromatis, L. A. et al. Association between brain structure and alcohol use behaviors in adults: a Mendelian randomization and multiomics study. JAMA Psychiatry 79, 869–878 (2022).
Davies, N. M., Holmes, M. V. & Davey Smith, G. Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. Br. Med. J. 362, k601 (2018).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Thompson, D. J. et al. UK Biobank release and systematic evaluation of optimised polygenic risk scores for 53 diseases and quantitative traits. Preprint at medRxiv https://doi.org/10.1101/2022.06.16.22276246 (2022).
Petersen, R. C. et al. Alzheimer’s Disease Neuroimaging Initiative (ADNI): clinical characterization. Neurology 74, 201–209 (2010).
Acknowledgements
We thank all participants and cooperating institutions. The UKB analyses were conducted using the UKB Resource under application no. 19542. This study was supported by grants from the Science and Technology Innovation 2030 Major Projects (grant no. 2022ZD0211600 to J.-T.Y.), the National Natural Science Foundation of China (grant nos 92249305 and 82071201 to J.-T.Y., 82071997 to W.C. and 81971032 to L.T.), the Shanghai Municipal Science and Technology Major Project (grant no. 2018SHZDZX01 to J.-F.F.), the Research Start-up Fund of Huashan Hospital (grant no. 2022QD002 to J.-T.Y.), the Shanghai Rising-Star Program (grant no. 21QA1408700 to W.C.), the 111 Project (grant no. B18015 to J.-F.F.), and the ZHANGJIANG LAB, the Tianqiao and Chrissy Chen Institute, and the State Key Laboratory of Neurobiology and Frontiers Center for Brain Science of Ministry of Education, Fudan University. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Consortia
Contributions
J.-T.Y. designed the study. Y.-J.G. and B.-S.W. organized the data, carried out the statistical analysis and participated in writing the first draft of the manuscript. Y.-J.G., B.-S.W. and Y.Z. designed and drew the figures. S.-D.C., J.-J.K., Y.-T.D., X.-Y.H., Y.-L.Z. and Q.M. organized and analysed the data. Y.Z., S.-D.C., Y.-R.Z., Y.-T.D., Y.-N.O., X.-Y.H., K.K., H.L., T.P., J.-F.F., Q.D., L.T., G.S., W.C. and J.-T.Y. critically revised the manuscript. IMAGEN provided data used for this study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
T.B. served in an advisory or consultancy role for eye level, Infectopharm, Lundbeck, Medice, Neurim Pharmaceuticals, Oberberg GmbH, Roche and Takeda. He received conference support or a speaker’s fee from Janssen, Medice and Takeda. He received royalties from Hogrefe, Kohlhammer, CIP Medien and Oxford University Press; the present work is unrelated to these relationships. G.J.B. has received honoraria from General Electric Healthcare for teaching on scanner programming courses. L.P. served in an advisory or consultancy role for Roche and Viforpharm and received a speaker’s fee from Shire. She received royalties from Hogrefe, Kohlhammer and Schattauer. The present work is unrelated to the above grants and relationships. The other authors report no competing interests.
Peer review
Peer review information
Nature Human Behaviour thanks Andre Altmann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic diagram of the study design.
First, we aim to elucidate the genetic basis of six ventricular phenotypes, including volumes of the lateral, inferior lateral, third, and fourth ventricles, as well as asymmetries of the lateral and inferior lateral ventricles. Second, we aim to leverage genetic data to estimate the value of ventricular traits in monitoring common neuropsychiatric disorders and promoting brain health across the lifespan. The brain and DNA images were produced using Servier Medical Art (http://smart.servier.com/) licensed under CC BY 3.0.
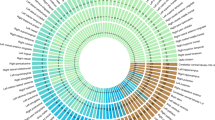
Extended Data Fig. 2 Top hits in FUMA gene set enrichment analysis.
Bar plots show the top enriched GO terms (a), pathways (b), and phenotypes in the GWAS Catalog (c) through the summary statistics of multivariate ventricular GWAS using hypergeometric tests in FUMA. The length of the bars indicates the -log10 P-value of the enrichment analysis. The color of bars shows the number of genes overlapping between cerebral ventricles and other traits. This figure displays raw P-values without FDR correction and full results could be found in Supplementary Table 17.
Extended Data Fig. 3 Associations between age and volumetric metrics in cohorts with different age groups.
Individual-level data from ABCD, IMAGEN, HCP, and UKB were used to construct the density plots with no covariates included. This plot only includes data from cross-sectional genetic analyses (N = 38,441).
Extended Data Fig. 4 The volume of the inferior lateral ventricles is a consequence of prodromal AD and could predict AD risk.
a-b, Bidirectional MR shows high genetic liability to AD is potentially causal to large inferior lateral ventricles, but not vice versa. The dot refers to each variant’s mean value of association estimate with exposure and outcome, while the cross symbol represents SE. The slope of the line corresponds to the total estimated causal effect. The number of instruments is 31 for inferior lateral ventricular volume (a) and 31 for AD (b), respectively. c-f, Multivariate COX analyses indicate the volume of the inferior lateral ventricles could predict the risk of indicant dementia (c-d) and AD (e-f). Analyses were performed without restriction (c and e, sample size = 15,198) and restricted in participants with at least three years of follow-up (d and f, sample size = 4,930). Compared to volumes of the hippocampus, the volume of the inferior lateral ventricles has even stronger significance and larger effect size.
Supplementary information
Supplementary Information
Supplementary Methods, acknowledgments and references.
Supplementary Data 1
Supplementary Tables 1–41.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ge, YJ., Wu, BS., Zhang, Y. et al. Genetic architectures of cerebral ventricles and their overlap with neuropsychiatric traits. Nat Hum Behav 8, 164–180 (2024). https://doi.org/10.1038/s41562-023-01722-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41562-023-01722-6